Abstract
BACKGROUND:
Intercellular adhesion molecules (ICAMs) in the tumor microenvironment are closely related to immunity and affect the prognosis of cancer patients.
OBJECTIVE:
The aim of our study is to explore the correlation between ICAM expression, mutation, methylation and immunity and their prognostic value in breast cancer (BC) is not clear.
METHODS:
Online databases and tools such as UALCAN, COSMIC, cBioPortal, MethSurv, PrognoScan, Kaplan-Meier Plotter, GSCA and TIMER were utilized in this study.
RESULTS:
We found that the mRNA and protein expression levels of ICAM1 were upregulated in triple-negative breast cancer (TNBC) compared with normal tissues, and TNBC patients with high expression of ICAM1 had better overall survival (OS) and recurrence-free survival (RFS). The main types of ICAM1 gene variants were missense mutation and amplification, and ICAM1 showed a lower level of methylation in TNBC cancer tissues than in normal tissues, which was contrary to the high expression levels of ICAM1 mRNA and protein. Next, the function of ICAM1 was mainly related to the activation of apoptosis, epithelial-mesenchymal transition (EMT) and inhibition of the androgen receptor (AR) and estrogen receptor (ER) pathways. Meanwhile, functional pathway enrichment results showed that ICAM1 was also involved in the immune regulation process of BC. Furthermore, the expression of ICAM1 was positively associated with 6 types of tumor-infiltrating immune cells (CD8
CONCLUSIONS:
Our research indicated that ICAM1 was likely to be a potential therapeutic target in TNBC.
Introduction
Breast cancer (BC) is the most common malignant tumor among women worldwide [1]. BC is a heterogeneous disease with different treatment strategies according to different molecular subtypes [2]. Recently, immunotherapy for cancer has developed rapidly and has been proven successful in the treatment of lymphoma, non-small cell lung cancer, melanoma, renal cell carcinoma and other tumor types [3]. Traditionally, BC is considered to have weak immunogenicity, which varies among different molecular subtypes. Triple-negative breast cancer (TNBC) and human epidermal growth factor receptor 2-positive (HER2
In the past few years, growing evidence has shown that tumor-infiltrating immune cell infiltration exhibits a positive impact on the neoadjuvant chemotherapy response and prognosis of BC, and tumor-infiltrating immune cells can reflect the intensity of the tumor bed immune response [6]. Several immune-based therapies have been evaluated in BC, including cancer vaccines, immune checkpoint inhibitors, and cytokine adoptive T-cell therapy. The drugs or antibodies that block the immune checkpoint include lymphocyte activating 3 (LAG3) [7], cytotoxic T lymphocyte-associated antigen-4 (CTLA4) [8], programmed cell death-1 (PD-1) [9] and programmed cell death-ligand 1 (PD-L1) [10], which have been used in clinical trials for BC treatment. However, although immunotherapy for BC has been extensively studied, only a small number of patients showed a response, and little is known about the underlying mechanisms. Therefore, it is necessary to identify more potential biomarkers of antitumor immunity, which may help the development of individualized immunotherapy for BC patients.
Intercellular adhesion molecules (ICAMs) are type I transmembrane glycoproteins containing 2–9 immunoglobulin-like C2 domains and binding to leukocyte adhesion lymphocyte function-associated antigen-1 (LFA-1) proteins [11], with a total of 5 members, namely, ICAM1, ICAM2, ICAM3, ICAM4 and ICAM5. ICAMs are closely related to immunity, such as mediating the antigen-specific immune response, lymphocyte circulation, NK-cell-mediated clearance and other adhesion interactions [12]. To date, ICAM1/3 have been confirmed to play an important role in the development of BC. For example, ICAM1/3 expression was upregulated in endothelial cells of BC, especially in ER-negative tumors, and tumors with high ICAM1/3 expression were proven to have more invasive characteristics and worse prognosis [13]. ICAM1 was also demonstrated to be a prognostic molecular target of TNBC [14]. Some studies have proven that ICAM2 plays a key role in immunotherapy of pancreatic cancer [15] and in preventing the spread of gastric cancer [16]. Overexpression of ICAM2 induced sensitivity of oral squamous cell carcinoma cells to radiotherapy [17]. ICAM4 variants are associated with glioma [18], and ICAM-5 may play a role in tumorigenesis and perineural invasion, most likely through the P13K/Akt signaling pathway [19]. However, ICAM2/4/5 has not been reported in BC.
The immune response of BC was initiated by neoantigens expressed by tumor cells that encode altered genes, such as gene mutations and abnormal methylation [5]. In summary, the literature mainly reports the expression and prognosis of ICAM1/3 in BC and its involvement in EMT and inflammation-related functional pathways. Mutations and methylation patterns of ICAMs in BC and their relationship with immune response and immunotherapy have not been reported. Therefore, the purpose of this study was to investigate the expression, mutation and methylation levels of ICAM family members in breast cancer and the correlation of ICAMs expression with survival, tumor immune cell invasion levels, and PD-1/PD-L1/CTLA4 expression levels.
Materials and methods
UALCAN
UALCAN (
Catalog of somatic mutations in cancer (COSMIC)
COSMIC (
cBioPortal
cBioPortal (
MethSurv databases
MethSurv (
Kaplan-Meier Plotter
The Kaplan-Meier Plotter (
PrognoScan databases
The flow-process diagram of this study.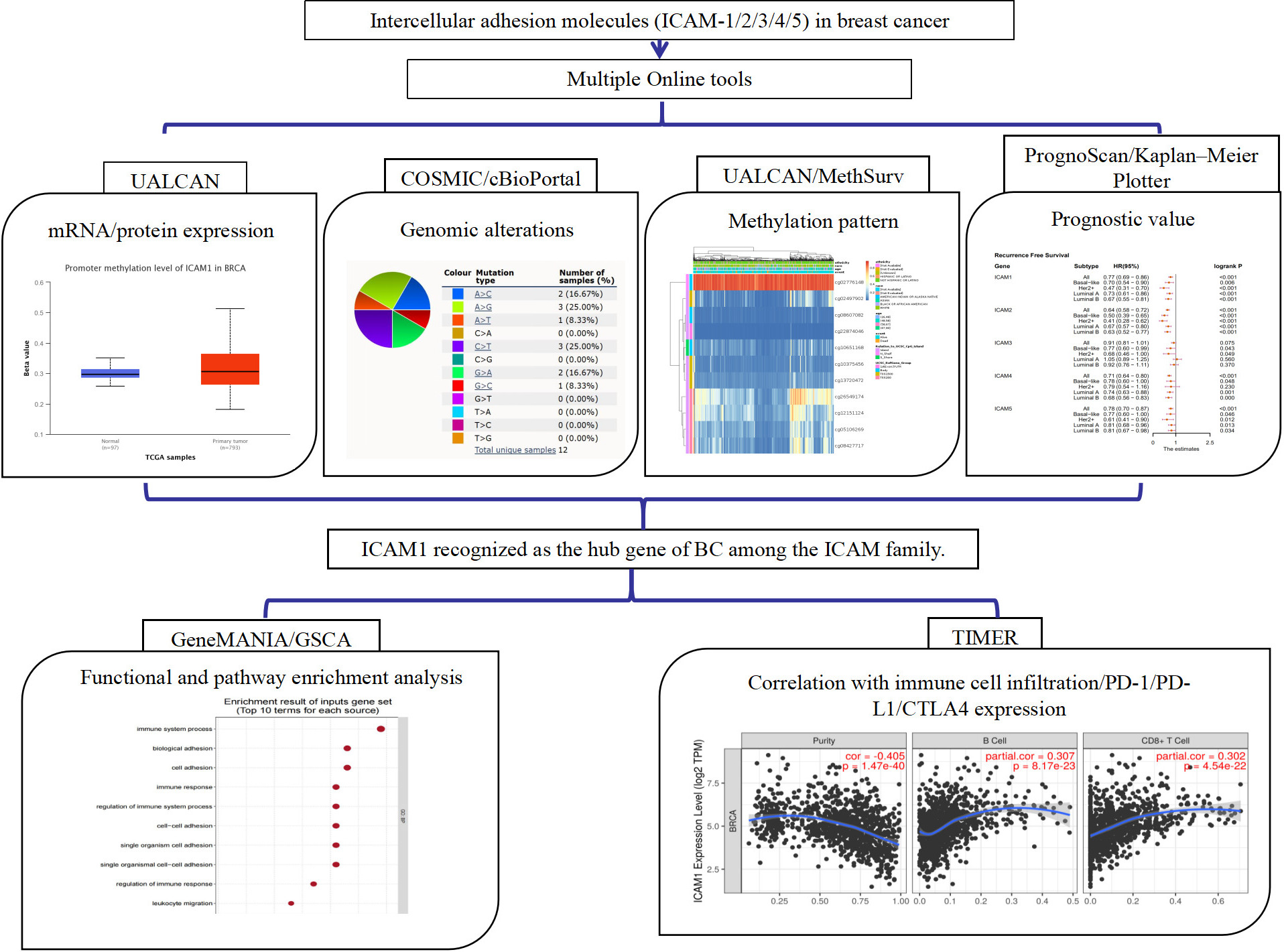
PrognoScan (
GeneMANIA (
Genome set cancer analysis (GSCA)
GSCA (
The mRNA expression levels of the ICAM family in BC (UALCAN). (A) Differences in the mRNA expression levels of the ICAM family in breast cancer tissues and normal tissues. (B) Differences in the mRNA expression levels of the ICAM family in different molecular subtypes of BC. 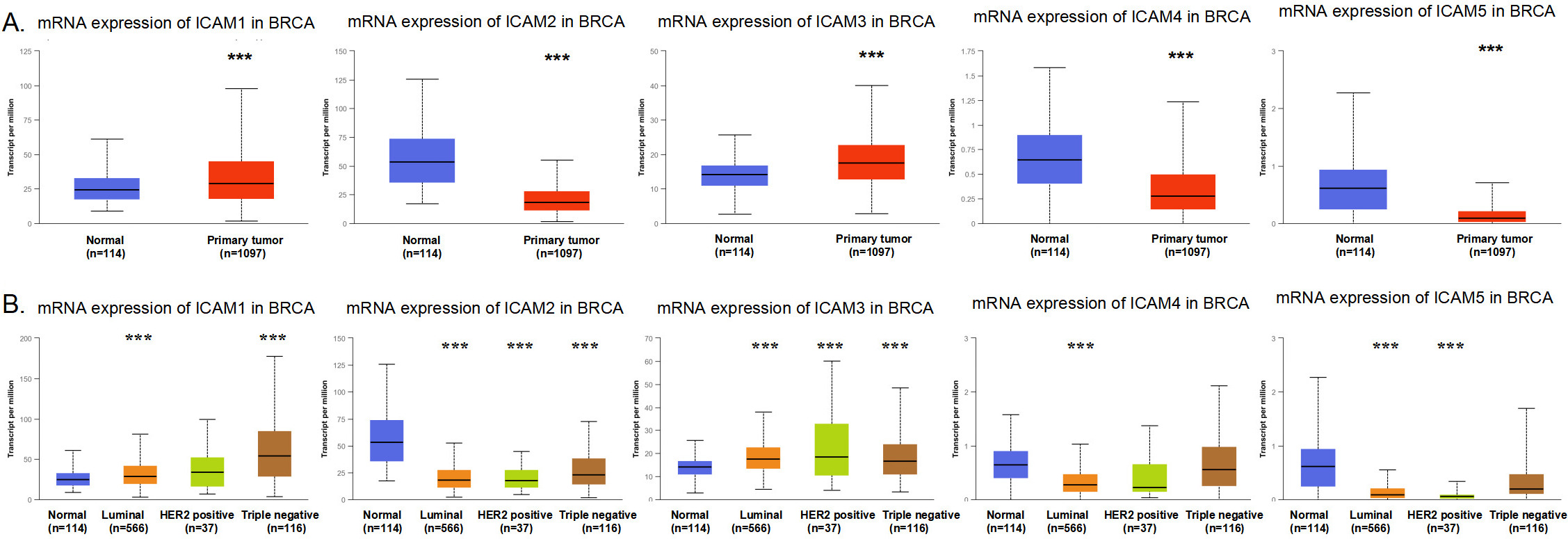
The protein expression levels of the ICAM family in BC (UALCAN) (UALCAN). (A) Differences in the protein expression levels of the ICAM family in breast cancer tissues and normal tissues. (B) Differences in the protein expression levels of the ICAM family in different molecular subtypes of BC. 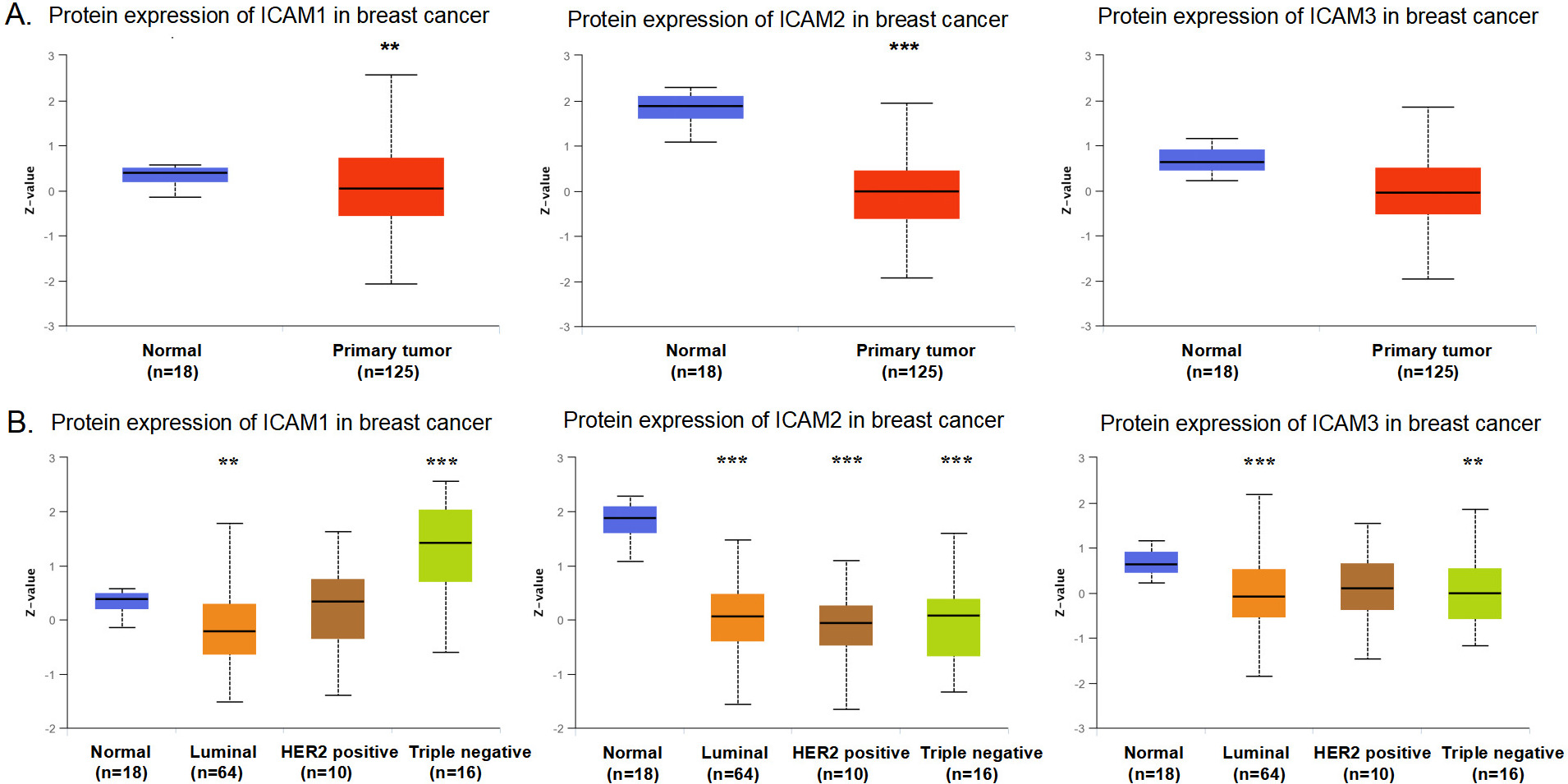
TIMER (
Finally, we provided a flowchart figure (Fig. 1), which shows our research step by step.
The genetic alteration characteristics of the ICAM family in BC. (A–B) Pie chart showing the percentage of the different mutation types of ICAMs in BC (COSMIC). Mutation types mainly include missense mutations, nonsense mutations, and synonymous mutations. (B) Pie chart showing the percentage of the different base mutation types of ICAMs in BC (COSMIC). (C) ICAMs mutation level in BC (cBioPortal); (D) Distribution of ICAMs mutations in different molecular subtypes of BC (cBioPortal). (E) Relationship between ICAMs mutations and common mutations of BC (cBioPortal).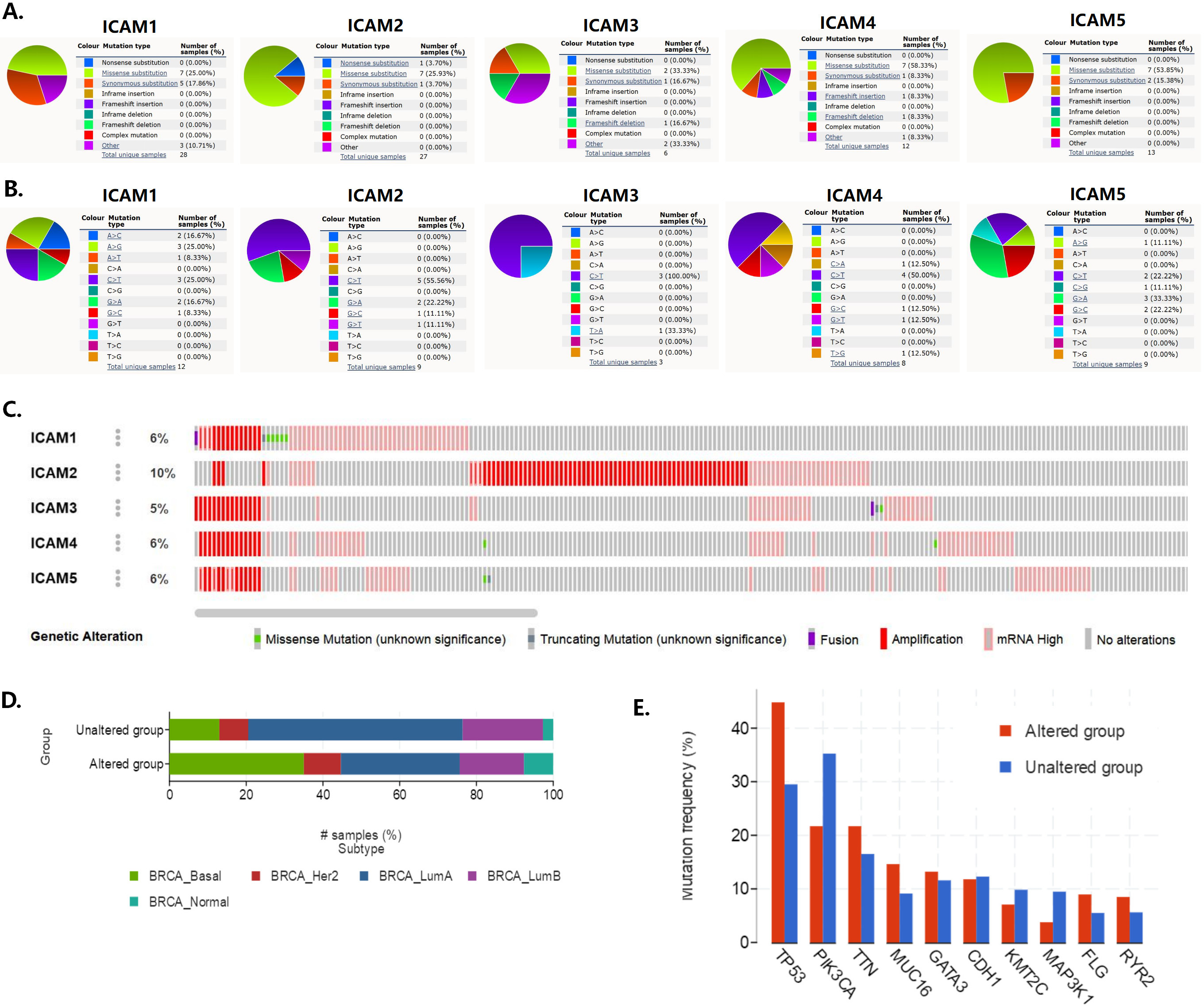
The ICAM family promoter methylation profile based on sample types, major subclasses and TP53 mutation status in BC (UALCAN). (A) The ICAM family promoter methylation profile based on sample types. (B) The ICAM family promoter methylation profile based on major subclasses. (C) The ICAM family promoter methylation profile based on TP53 mutation status. The beta value indicates the level of DNA methylation ranging from 0 (unmethylated) to 1 (fully methylated). Different beta value cut-offs have been considered to indicate hypermethylation [beta value: 0.7–0.5] or hypomethylation [beta-value: 0.3–0.25]. 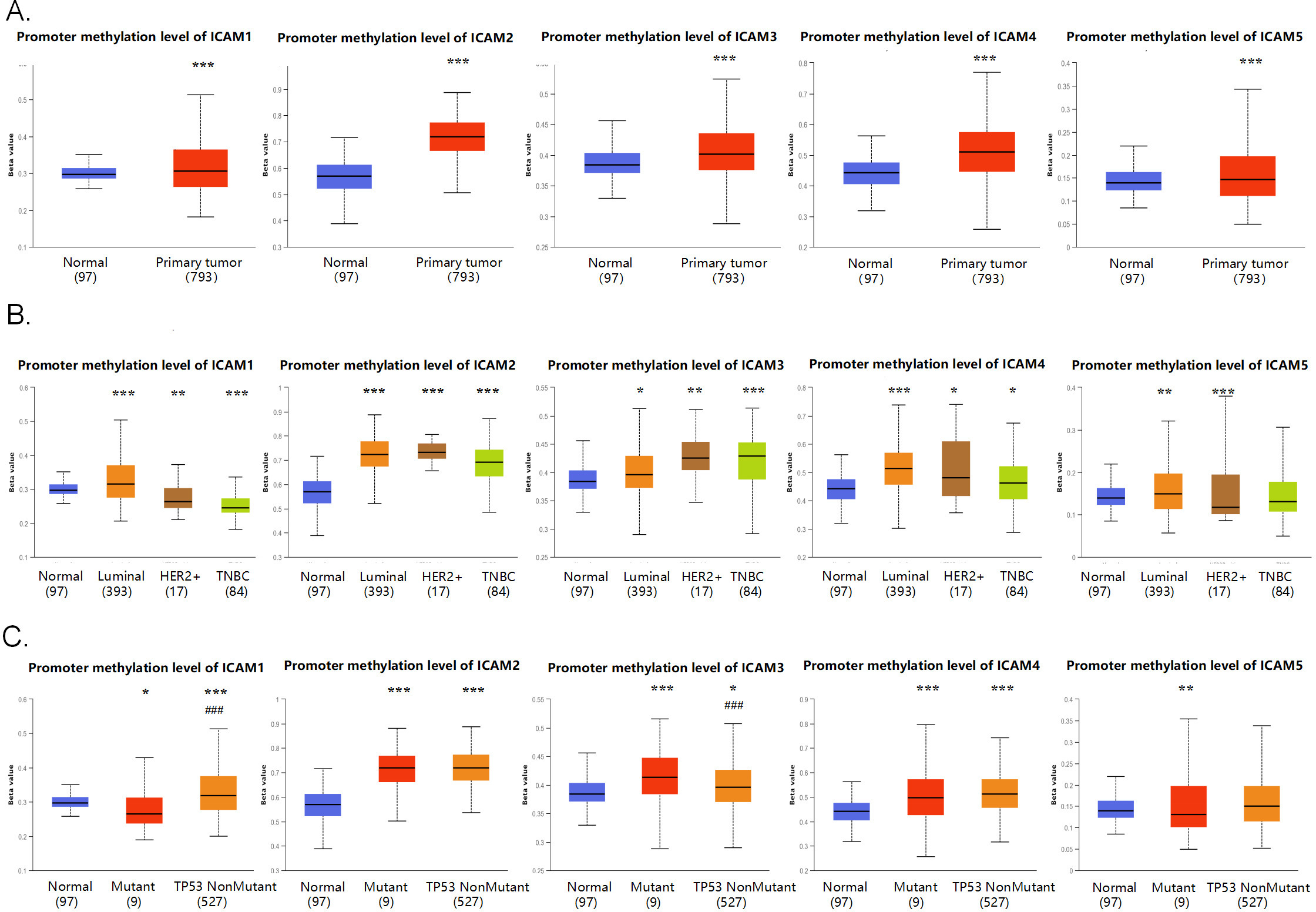
The ICAM family mRNA and protein expression in BC
First, we used the TCGA analysis and CPTAC analysis modules of the UALCAN platform to explore the differences in the mRNA and protein expression levels of ICAMs in normal breast tissue and breast cancer tissue, as well as in different molecular subtypes. The results of TCGA analysis showed that the mRNA expression levels of ICAM1 and ICAM3 in breast primary tumors were higher than those in breast normal tissues, whereas the expression patterns of ICAM2, ICAM4 and ICAM5 were reversed (Fig. 2A). From the perspective of different molecular subtypes of BC, the mRNA expression levels of ICAM1 in triple-negative tumors and ICAM3 in HER2
The prognostic value of the ICAM family for OS and RFS in BC patients. (A–B) The OS and RFS of the ICAM family mRNA expression in BC (Kaplan-Meier Plotter). (C–D) The OS and RFS of the ICAM family mRNA expression in BC (PrognoScan). A 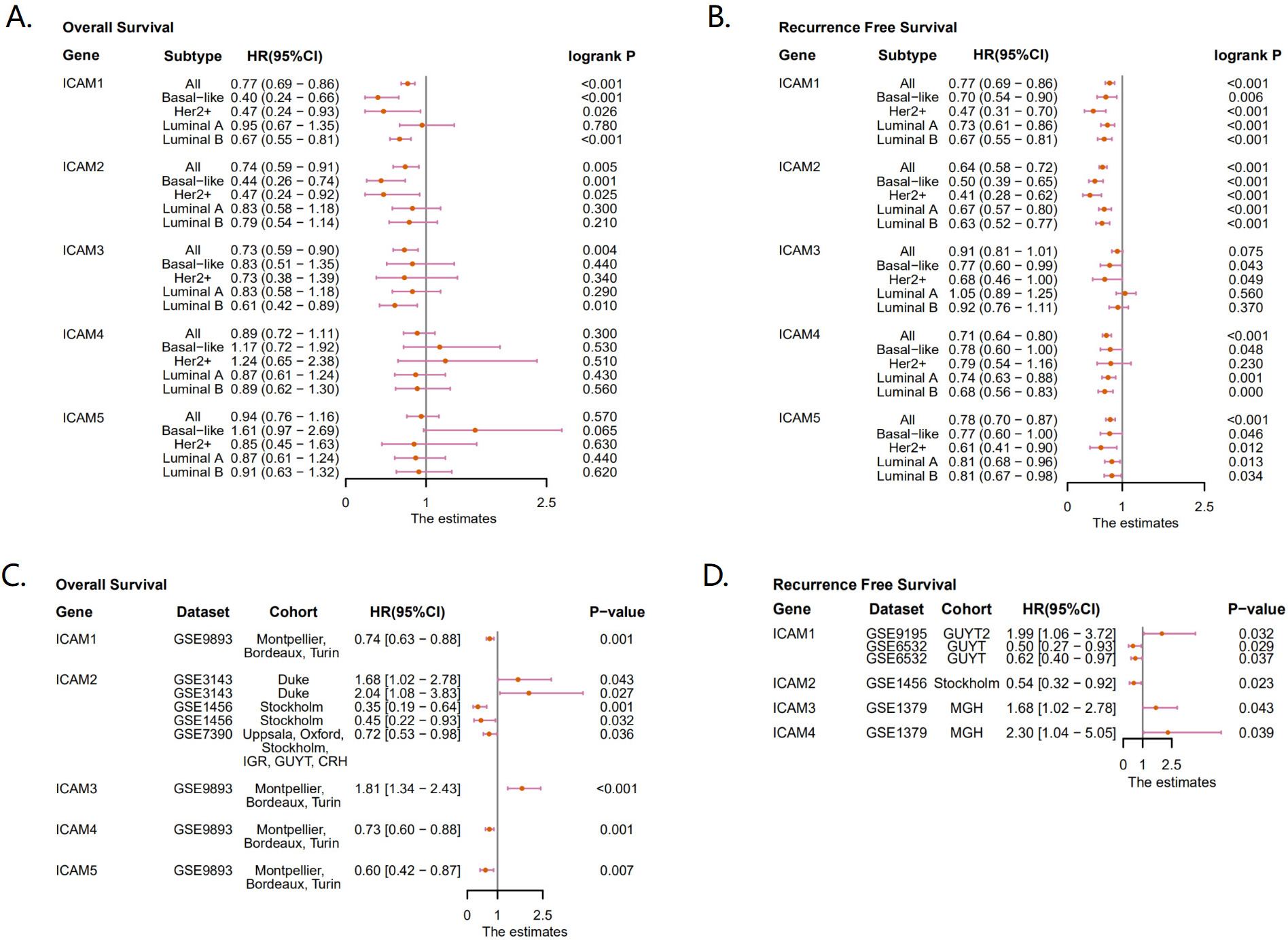
COSMIC provides information on ICAM mutations in different cancers, resulting in a pie chart. Only ICAM2 was found to have a 3.7% nonsense mutation rate in BC, while missense mutations in ICAM1 (25%), ICAM2 (25.93%), ICAM3 (33.33%), ICAM4 (58.33%) and ICAM5 (53.85%) were found in BC. In addition, synonymous mutations were observed in ICAM1 (17.86%), ICAM2 (3.7%), ICAM3 (16.67%), ICAM4 (8.33%) and ICAM5 (15.38%) (Fig. 4A). C
Then, we used cBioPortal to determine the type and frequency of genetic alterations in the ICAMs. Alterations in ICAM1, ICAM2, ICAM3, ICAM4 and ICAM5 were 6%, 10%, 5%, 6% and 6% in 994 BC patients, respectively (Fig. 4C). Most of the alterations were amplifications. From the perspective of molecular subtypes of BC, the mutation rate of ICAMs in the basal-like subtype was the highest, followed by the mutation rate in the luminal B subtype (Fig. 4D). In addition, the mutation rate of ICAMs in the TP53 mutant group was significantly higher than that in the TP53 wild-type group, while the mutation rate of ICAMs in the phosphatidylinositol-4,5-bisphosphate 3-kinase catalytic subunit alpha (PIK3CA) mutant group was lower than that in the wild-type group in BC (Fig. 4E).
Functional pathway enrichment analysis of ICAM1 and its interacting genes (GSCA). (A) The biological processes involved in ICAM1. (B) ICAM1 is involved in cell composition. (C) Molecular functions in which ICAM1 was involved. (D) ICAM1 participates in the KEGG pathway. BP, biological process; CC, cell composition; MF, molecular function; KEGG, Kyoto Encyclopedia of Genes and Genomes; FDR, false discovery rate.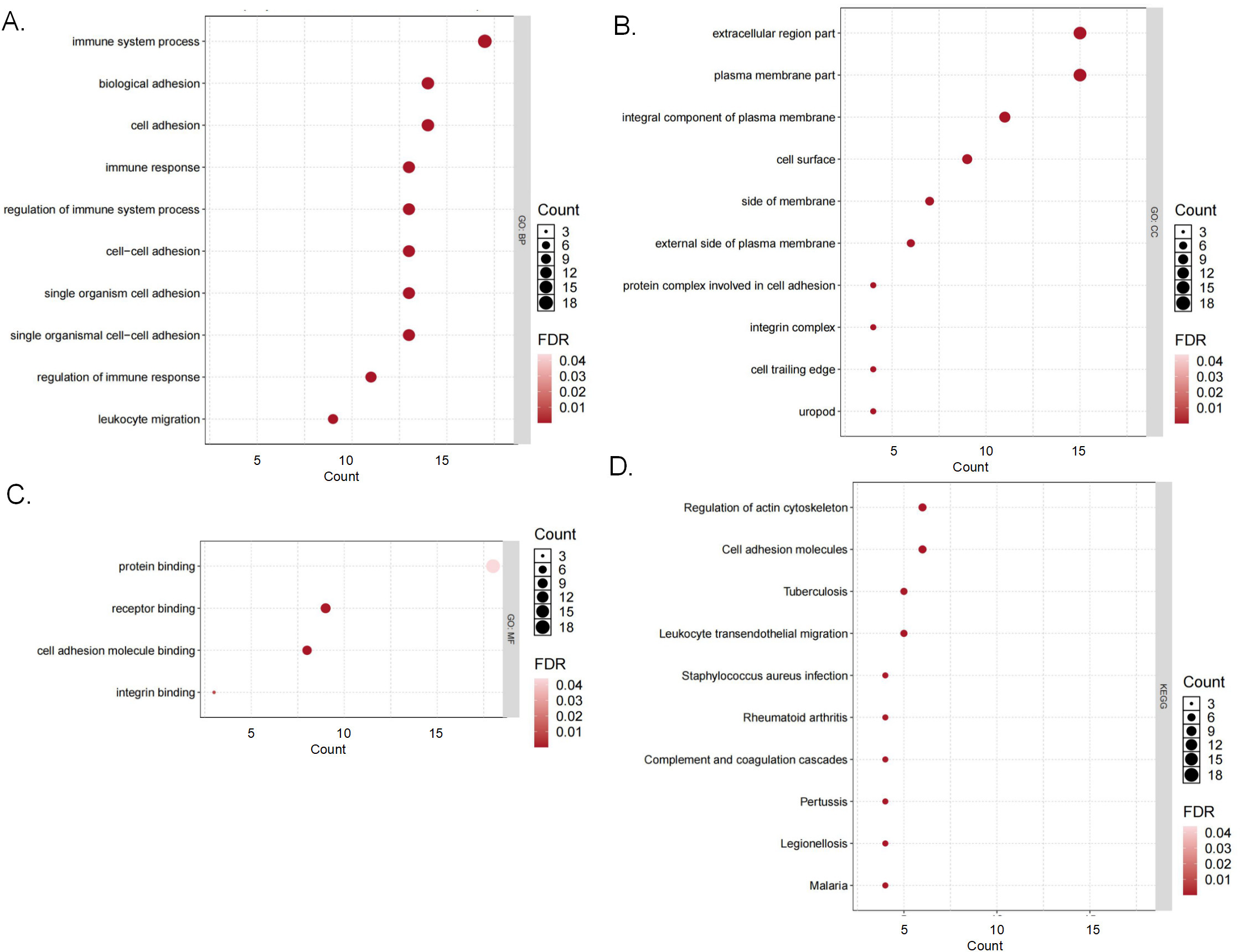
The methylation level of the oncogene promoter region is closely related to the occurrence and development of cancer, including BC. The UALCAN database was used to explore the levels of promoter region methylation in ICAMs in BC from multiple perspectives. In general, the promoter methylation level of the ICAM family in BC tissues was higher than that in breast normal tissues (Fig. 5A). From the perspective of the molecular subtype of BC, ICAM1 and ICAM5 showed similar methylation patterns: the methylation level of the luminal subtype was higher than that of normal tissues, while the methylation level of the HER2
MethSurv is also a well-known DNA methylation research database, whose data are mainly derived from TCGA. We used it to further investigate CpG methylation sites in the ICAMs in BC. The results showed that ICAM1 methylation sites were concentrated in CpG island, N-shelf and S-Shore regions, ICAM2 was only located in the Open Sea, and ICAM3 had four concentrated methylation site areas, including CpG Island, S-shelf, S-Shore and Open Sea regions (Fig. S1A–C). Additionally, ICAM4 and ICAM5 have the same methylation site regions, including island and N-shore regions (Fig. S1D–E).
Prognostic value of the ICAM family in BC
According to Kaplan-Meier Plotter’s research results, ICAM1/2/3 were favorable prognostic factors for OS in BC patients. Based on specific molecular subtypes, it was found that ICAM1 and ICAM2 were favorable prognostic factors for OS in BC patients with HER2
Correlation between ICAM family expression and tumor-infiltrating immune cells in BC (TIMER). The horizontal axis is the infiltration level of tumor-infiltrating immune cells, and the vertical axis is the expression level of the ICAM family. A 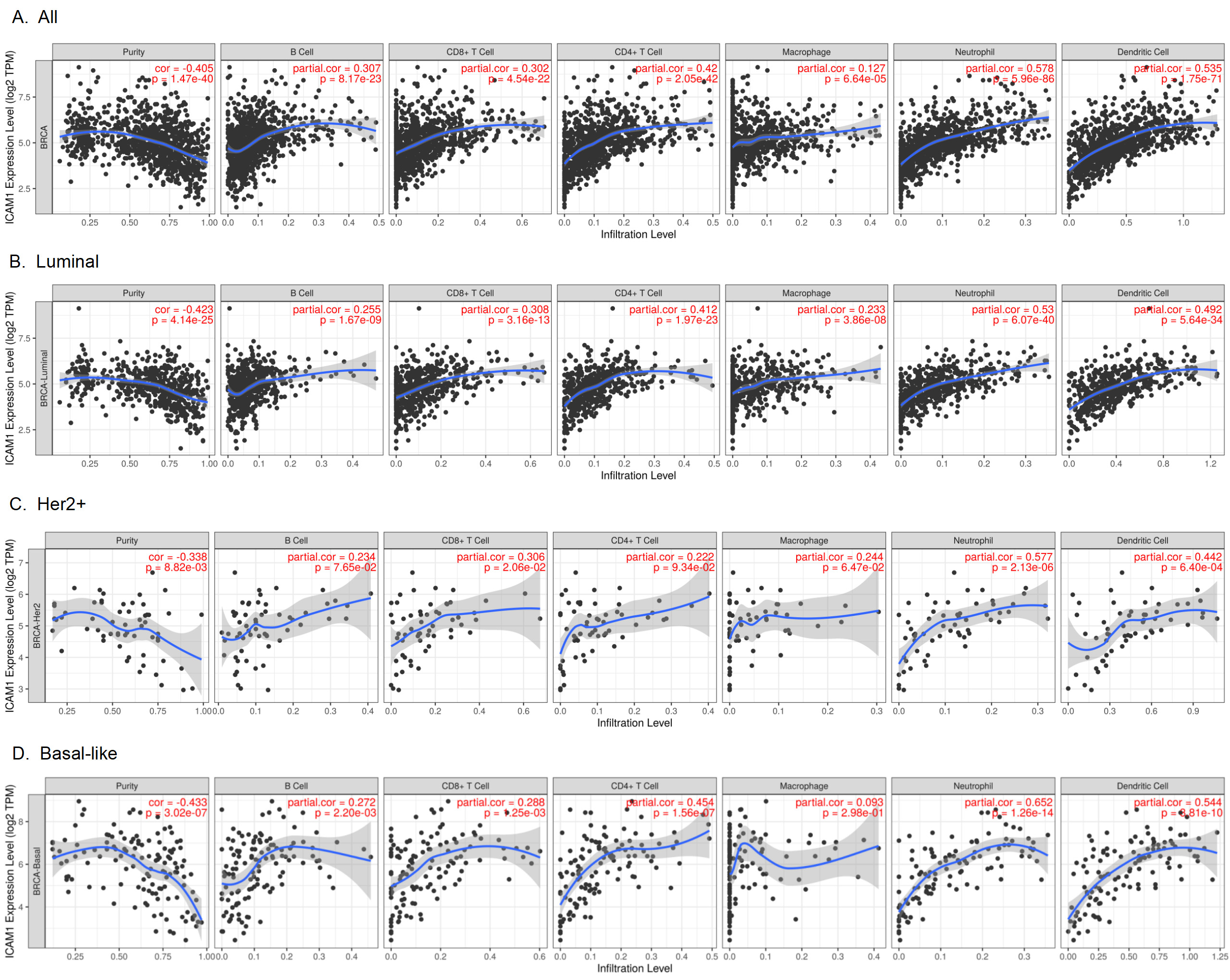
Since ICAM1 showed favorable prognostic value in all the datasets analysed, it is recognized as the hub gene of BC among the ICAM family. Then, a gene interaction network was constructed via the GeneMANIA database. The results suggested that among the top 20 genes associated with ICAMs, integrin subunit alpha L (ITGAL) was the most significant (Fig. S2A). Next, we used GSCA to explore the activity of ICAM1 in 10 cancer-related pathways in BC. The results suggested that ICAM1 was potentially involved in the activation of apoptosis and epithelial-mesenchymal transition (EMT) pathways and the inhibition of the hormone androgen receptor (AR) and hormone ER pathways (Fig. S2B). Additionally, we utilized the GSCA website to conduct KEGG and GO analyses on the 20 genes that were most significantly associated with ICAM1 obtained from GeneMANIA. The results of GO analysis showed that ICAM1 was mainly involved in biological processes such as immune response regulation and cell adhesion (Fig. 7A). It was mainly involved in the formation of the integral components of the extracellular region or plasma membrane (Fig. 7B) and the function of binding integrin (Fig. 7C). The results of KEGG pathway analysis showed that ICAM1 was mainly enriched in the pathways regulating the actin cytoskeleton and cell adhesion molecules (Fig. 7D).
The correlation of ICAM1 expression with PD-1, PD-L1 and CTLA4 expression in BC (TIMER). (A) The correlation of ICAM1 expression with PD-1, PD-L1 and CTLA4 expression in all BC. (B) The correlation of ICAM1 expression with PD-1, PD-L1 and CTLA4 expression in the luminal subtype. (C) The correlation of ICAM1 expression with PD-1, PD-L1 and CTLA4 expression in the HER2+ subtype. (D) The correlation of ICAM1 expression with PD-1, PD-L1 and CTLA4 expression in the basal-like subtype. A 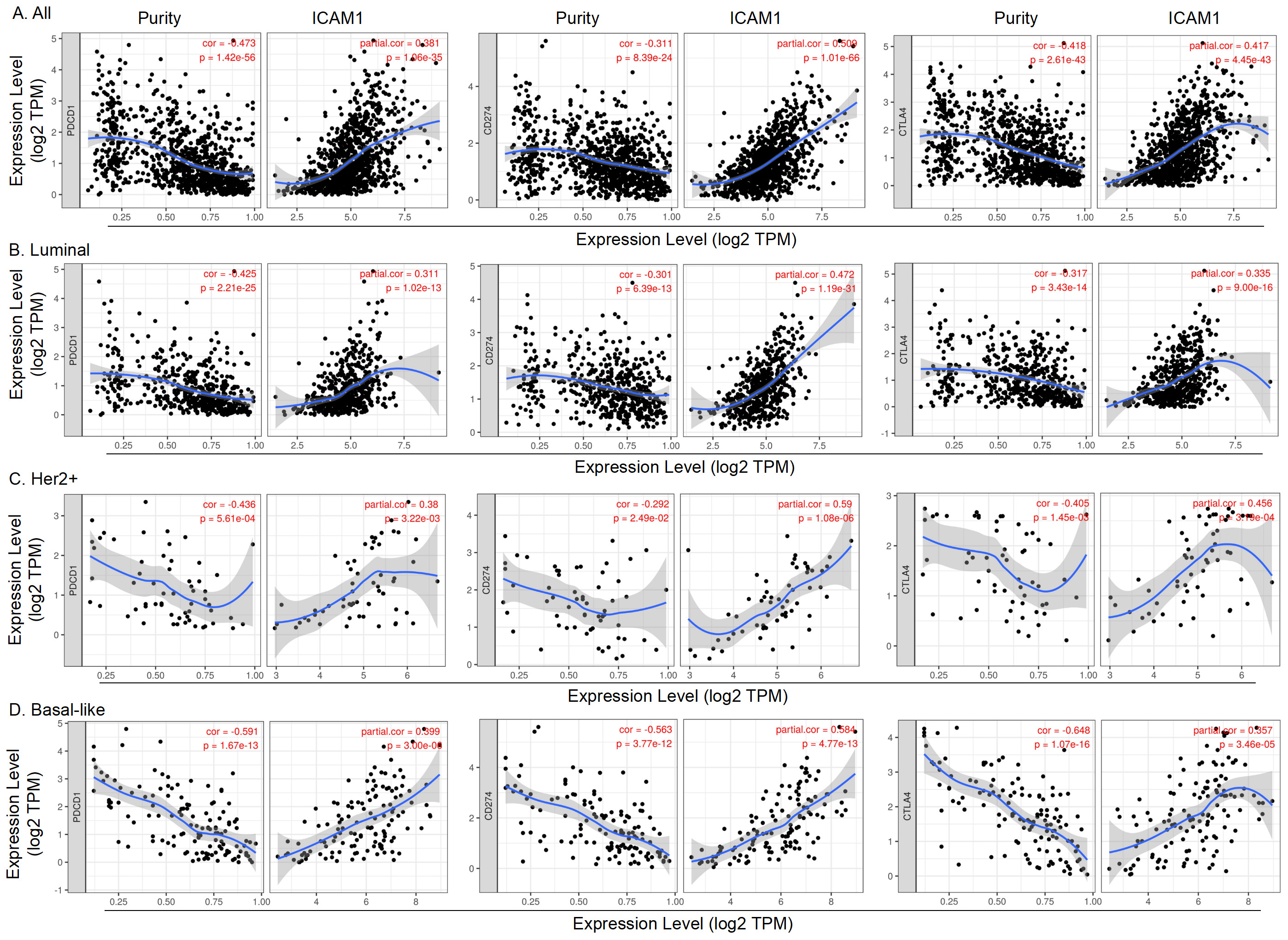
Tumor-infiltrating immune cells can affect the survival of patients, and the results of enrichment analysis of the above functional pathways supported the important role of ICAMs in immune regulation. Therefore, it is of great significance to explore the relationships between ICAM1 expression and immune infiltration/PD-1/PD-L1/CTLA4 expression in BC.
First, we used TIMER to calculate the coefficient of ICAM1 expression and immune cell invasion level in all BCs and different molecular subtypes to determine whether ICAM1 expression is related to the abundance of neoplastic infiltrating lymphocytes. The results showed that ICAM1 expression was negatively correlated with the purity of BC and was significantly positively associated with the infiltration level of B cells, CD8
Furthermore, we explored the correlation between the ICAM1 expression level in BC and the expression of PD-1, PD-L1, and CTLA4 via TIMER. The results suggested that the ICAM1 expression level in BC was significantly positively associated with the expression of PD-1, PD-L1 and CTLA4 (Fig. 9A). In different molecular subtypes, the expression level of ICAM1 was positively correlated with PD-1, PD-L1 and CTLA4 (Fig. 9B–D).
Discussion
This study was the first to comprehensively analyse the correlation of the expression, mutation and methylation of ICAM family members in BC via multiple online databases. It is noteworthy that ICAM1 mRNA was upregulated and ICAM1 protein was downregulated in breast cancer tissues. There are two explanations for this phenomenon. First, breast cancer is a highly heterogeneous tumor [31], so it is common for the same molecule to have different expression levels in different breast cancer subtypes. Second, the biological process from mRNA to protein involves much posttranscriptional regulation [32], which is also one of the reasons for the inconsistency of mRNA and protein expression levels. Additionally, our study found that both the mRNA and protein levels of ICAM1 were elevated in TNBC but opposite in luminal and HER2
Next, this study further explored the variation and methylation status of the ICAM family. ICAMs in BC had a relatively high frequency of missense mutations, among which the most common base replacement was C
Furthermore, based on the analysis results of the Kaplan-Meier Plotter database, only ICAM1 and ICAM2 were found to be favorable prognostic factors for OS and RFS in patients with BLBC in this study. However, previous literature has reported that ICAM1 overexpression in breast cancer cells (especially TNBC cells) and downregulation of ICAM1 expression can inhibit the invasion and metastasis of tumor cells [33, 37, 38], indicating that ICAM1 is a potential therapeutic target for TNBC. Next, in the process of exploring its functional pathway, ICAM1-related genes were mainly involved in activating apoptosis and inhibiting the hormone AR and ER pathways. Current studies have shown that the hormone AR and ER pathways are involved in the occurrence, invasion and metastasis of TNBC, and their inhibition can lead to a better prognosis for these patients [39]. Additionally, our study found that ICAM1 was highly expressed in TNBC, and its high expression was positively correlated with the immune cell infiltration level. Collectively, we found that ICAM1 was highly expressed in TNBC but was correlated with better OS and RFS, and the most likely reason is that ICAM1 expression was positively correlated with the level of immune cell infiltration. Therefore, we believe that ICAM1 is closely related to the occurrence, development and immunotherapy effect of TNBC.
A large number of studies on the precision treatment of immune-related cancers are ongoing. Many studies have found that the methylation levels of oncogenes and TP53 mutations are closely related to the immune microenvironment and immune response of tumors. Tumor-specific antigens, or neoantigens, produced by somatic mutations can influence patients’ response to immunotherapy and promote tumor shrinkage [40]. Wild-type TP53 genes are involved in cell cycle regulation, DNA damage repair and damage-induced apoptosis. Previous studies have shown that TP53 mutation is closely related to the immune microenvironment and immunotherapy response of liver cancer and head-neck cancer [41, 42]. In addition, TP53 mutation has also been found to be related to the immunotherapy response of anti-PD-1 and anti-PD-L1 in ovarian cancer [43]. Our study found that in the altered group of ICAMs, the proportion of patients with TNBC or TP53 mutation rates was the highest. Therefore, from the perspective of mutation, ICAMs may play a regulatory role in the immune regulation and immunotherapy of BC, especially in TNBC patients.
It is well known that the first epigenetic change described in human cancer is the loss of DNA methylation. In BC, general hypomethylation of cancer genes can lead to gene activation, oncogene upregulation and chromosome instability, or local abnormal hypermethylation can lead to DNA repair gene silencing, further leading to gene suppression and genetic instability [44]. In addition, some studies have shown that DNA methylation plays an important role in the immune response of tumors or inflammation [45, 46, 47], and evidence has shown that the function of DNA methylation can be used to evaluate the immune response of BC and can improve the diagnosis and prognosis of BC. Overall, high or low levels of DNA methylation reflect, to some extent, a better predictor of the host’s response to immunotherapy. Our results indicated that the methylation level of ICAM1 in TNBC was lower than that in the normal group, the methylation level of ICAM1 in BC was negatively correlated with the immune cell infiltration level, and high expression of ICAM1 was associated with better OS and RFS, which was consistent with the research results of Jeschke J and Zhou et al. [48, 49]. In summary, from the perspective of epigenetics, ICAM1 may become the latest predictive biomarker of immunotherapy in TNBC.
In BC, the immune biomarkers of the TNBC subtype and the expression levels of PD-1, PD-L1 and CTLA4 are relatively high [50]. Through TCGA RNA sequencing, Mittendorf et al. found that the expression levels of PD-L1 in the TNBC group (
Our study has some limitations. We analysed the expression, mutation and methylation of the ICAM family in BC mainly at the transcriptional level and further explored their correlation with immune infiltration and immunotherapy. However, we did not further investigate these relationships at the protein level, mainly because there are few publicly available databases on proteins. In addition, in vivo or in vitro basic studies are needed to confirm our results.
Conclusions
In summary, ICAM1 was closely related to the occurrence, development and immunotherapy effect of TNBC. Additionally, we proposed that ICAM1 can be further explored from the perspective of mutation and methylation in immunoregulation and immunotherapy of TNBC. Therefore, our study may provide new insights into the selection of immune checkpoint therapeutic targets and prognostic biomarkers for BC.
Abbreviations
BC, breast cancer; CTLA4, cytotoxic T-lymphocyte associated protein 4; COSMIC, Catalog of Somatic Mutations in Cancer; ER, estrogen receptor; EMT, epithelial-mesenchymal transition; EGA, European Genome-phenome Archive; GEO, Gene Expression Omnibus; GSCA, Gene Set Cancer Analysis; HER2, human epidermal growth factor receptor 2; ITGAL, integrin subunit alpha L; ICAMs, intercellular adhesion molecules; LPBC, lymphocyte-dominated breast cancer; TNBC, triple negative breast cancer; PR, progesterone receptor; ICGC, International Cancer Genome Consortium; LAG3, lymphocyte activating 3; LncMAP, LncRNA Modulator Atlas in Pancancer; PD-1, programmed cell death 1; PD-L1, programmed cell death-ligand 1; PIK3CA, phosphatidylinositol-4,5-bisphosphate 3-kinase catalytic subunit alpha; TP53, tumor protein P53; TNBC, triple negative breast cancer; TNF, tumor necrosis factor; TF, transcription factor; TCGA, The Cancer Genome Atlas; TIMER, Tumor IMmune Estimation Resource; TISDB, Tumor-Immune System Interaction Database; VCAM-1, vascular cell adhesion molecule 1.
Author contributions
Conception: Heyan Chen, Huimin Zhang and Jianjun He.
Interpretation or analysis of data: Heyan Chen, Huimin Zhang, Shengyu Pu and Nan Mei.
Preparation of the manuscript: Heyan Chen, Huimin Zhang.
Revision for important intellectual content: Heyan Chen and Xiaoxu Liu.
Supervision: Heyan Chen and Huimin Zhang.
Data availability statement
The data used to support the findings of this study are available from publicly available online databases. The access sites for each of the open databases used in this study were specifically described in the methodology.
Supplementary data
The supplementary files are available to download from http://dx.doi.org/10.3233/CBM-220073.
sj-pdf-1-cbm-10.3233_CBM-220073.pdf - Supplemental material
Supplemental material, sj-pdf-1-cbm-10.3233_CBM-220073.pdf
Footnotes
Acknowledgments
We thank The Cancer Genome Atlas, Gene Expression Omnibus and European Genome-phenome Archive for sharing the large amount of data. The authors also thank the Institutional Foundation of The First Affiliated Hospital of Xi’an Jiaotong University (No. 2022YQPY08) and Shaanxi Provincial Key Research and Development Project (No. 2020SF-294) for funding to do this work.
Conflict of interest
The authors declare no conflicts of interest.
