Abstract
Pancreatic adenocarcinoma (PDAC) is one of the deadliest cancers, with five-year survival rates of 9%. We hypothesized that secreted frizzled-related protein 2 (SFRP2) may influence stromal growth in pancreatic cancer, since it increases fibrosis and collagen production in non-neoplastic pathologies. We assessed SFRP2 value as a biomarker and assessed its function in PDAC. SFRP2 gene expression in patients with PDAC was analyzed using TCGA data. Disease free survival (DFS) was analyzed using Kaplan Meier test. The effect of KRAS inhibition on SFRP2 expression in PDAC cells was assessed. The associations of stromal content with SFPR2 mRNA and protein with fibrosis were analyzed. The role of SFRP2 in mesenchymal transformation was assessed by western blot in fibroblasts. Of all cancers in TCGA, SFRP2 levels were highest in PDAC, and higher in PDAC than normal tissues (
Introduction
Pancreatic adenocarcinoma (PDAC) is the fourth leading cause of cancer deaths in the United States [1], and KRAS is mutated in 90% of patients with PDAC [2]. Even with improved chemotherapy regimens, the 9% five-year survival remains low [3]. One of the barriers to pancreatic cancer treatment is its desmoplastic stroma. Pancreatic cancer has the highest level of stroma (
Secreted frizzled-related protein 2 (SFRP2) is a glycoprotein upregulated in multiple cancer types that contributes to tumor growth, migration, anti-apoptosis [7, 8, 9, 10, 11, 12, 13, 14], and angiogenesis [7, 9, 13]. SFRP2 upregulates markers of immune dysfunction (PD-1 and CD38) in T-cells and contributes to their exhaustion [15]. We previously developed a humanized monoclonal antibody to SFRP2 (hSFRP2 mAb) that inhibits tumor growth, angiogenesis, induces tumor apoptosis [15, 16], and overcomes resistance to PD-1 inhibition [15]. We hypothesized that SFRP2 may influence stromal growth in cancer, since it increases fibrosis in non-neoplastic pathologies [17, 18]. A high correlation between SFRP2 mRNA expression and stromal mRNA expression in bladder, breast, colorectal, glioblastoma, head and neck, kidney, lung, ovarian, and endometrial cancers has also been found [19]. In correlation network analyses, genes involved in EMT process and ECM structure were associated with SFRP2 expression [19]. Because of the high stromal content in pancreatic cancer, we hypothesized that SFRP2 may play a larger role in this tumor type, not only through reducing apoptosis, increasing angiogenesis, and affecting immune cell activation, but also through activation of fibroblasts in the tumor stroma and development of a myofibroblast phenotype.
We sought to explore the relationship between SFRP2 and pancreatic cancer to determine whether SFRP2 expression is associated with survival and tumor fibrosis, and whether it is regulated by KRAS, a major driver of PDAC. Additionally, we investigated the ability of SFRP2 to promote the mesenchymal transformation of fibroblasts and stimulation of fibrinogen production.
Methods
Cell lines, cell isolation, and tissue culture
Pancreatic cancer cell lines AsPC-1, MIA PaCa-2, and Panc02-luc were donated by Dr. Ernest Ramsay Camp (Medical University of South Carolina). Panc02-luc were originally obtained from the NCI DCTD Tumor Repository and transfected as previously described [20]; AsPC-1 and MIA PaCA-2 were from American Type Culture Collection (ATCC CRL-1420). PANC-1R15 and PSN-1R15 cells were generated as previously described [21]. Primary human skin fibroblasts [22] were donated by Dr. Carol Feghali-Bostwick (Medical University of South Carolina). The mouse angiosarcoma cell line, SVR [23], was purchased from ATCC (#CRL-2280).
A male C57/BL6NCrl mouse purchased from Charles River Laboratory (Wilmington, MA) was used to isolate pancreatic fibroblasts. The pancreas was resected after euthanasia and was washed with ice-cold modified Krebs-Hensleit buffer supplemented with 2.1 g/L sodium bicarbonate (pH 7.4). Next, 1 mL of ice-cold digestion mix (34.4 mL Hanks buffered saline solution (HBSS; #55037C-1000 mL, Sigma-Aldrich, St. Louis, MO, USA), 1 mL Liberase mix (5 mg in 2 mL of HBSS; # 541020001, Roche, Basel, Switzerland), 200 mL 0.25% Trypsin (# 25-050-CL, Corning, Corning, NY, USA), 360 mL 1M HEPES buffer (#H4034-25G, Sigma-Aldrich) was added, and tissue was macerated using a razor blade. The macerated tissue and digestion mix was placed into a 15 mL falcon tube, and an additional 3 mL of digestion mix at 37∘C was added. The mix was incubated on a tube rotator at 37∘C for 8 min. 10 mL of modified Krebs-Hensleit buffer containing 2.1 g/L sodium bicarbonate (#K3753, Sigma-Aldrich) was added to quench the digestion, and the mixture was briefly rested to allow the tissue chunks to collect at the bottom. The supernatant containing the cellular suspension was removed and placed on ice. Digestion mix (3 mL at 37∘C) was added and another round of incubation on the tube rotator was performed for 8 min. Once again, the supernatant was placed on ice, and the digestion was repeated until no visible chunks of tissue remained in the tube. The supernatants were then combined in a 50 mL falcon tube, centrifuged for 5 min at 1200 rpm, and washed with 10 mL Krebs-Hensleit buffer and centrifuged again at 1200 rpm for 5 min. Each pellet was finally resuspended in Human Cardiac Fibroblast growth media (#316-500, Cell Applications, San Diego, CA, USA) containing 1% penicillin/streptomycin (#15140122, Thermofisher, Waltham, MA, USA), placed into a 10 cm Petri dish, and grown to confluence.
SVR angiosarcoma, pancreatic cancer MIA PaCA-2 cell line, human skin fibroblasts, and mouse pancreatic fibroblasts were cultured in Dulbecco’s Modified Eagle Medium (DMEM, #10-013-CV, Corning) with 10% fetal bovine serum (FBS) (#BT 201-500-D, BioFluid, Fleming Island, FL, USA). AsPC-1 and Panc02-luc pancreatic cancer cell lines were cultured in Roswell Park Memorial Institute (RPMI) media with 10% FBS. All cell lines were cultured at 37∘C, 5% CO2, and 95% humidity.
In vivo pancreatic cancer GEMM mouse model
Mice experiment protocols were in accordance with the National Institutes of Health guidelines for the care and use of laboratory animals and approved by the Animal Care and Use Committee (IACUC) at the Medical University of South Carolina. KPC male and female mice (Krastm4TyjTrp53tm1BrnTg(Pdx1-cre/Esr1*)#Dam/J) (
Antibodies and proteins
Primary antibodies used included rabbit anti-actin (#A2103, Sigma-Aldrich), mouse anti-fibronectin (#610077, BD Biosciences, Franklin Lakes, NJ, USA), rabbit anti-frizzled-5 (H00007855-PW1, Abnova, Taipei, Taiwan), rabbit anti-GAPDH (glyceraldehyde-3-phosphate dehydrogenase, #ab9485, Abcam, Cambridge, MA, USA), rabbit anti-N-cadherin (#ab1221, Abcam), mouse anti-Vinculin (SC #73614, Santa Cruz Biotechnology, Dallas, TX, USA). Rabbit anti-Vinculin (E1E9V) (#13901, Cell Signaling Technologies, Danvers, Massachusetts, USA), Phospho-p44/42 MAPK (Erk1/2) (Thr202/Tyr204) Antibody (#9101, Cell Signaling Technologies, Danvers, Massachusetts, USA), p44/42 MAPK (Erk1/2) Antibody (#9102, Cell Signaling Technologies, Danvers, Massachusetts, USA). Rabbit anti-SFRP2 (#ab86379, Abcam), rabbit anti-SFRP2 (#ab137560 Abcam), and rabbit anti-SFRP2 (#LS-C61791, Lifespan Biosciences, Seattle, Washington, USA) were purchased based on manufacturers’ availability. Secondary antibodies used were biotinylated-conjugated anti-rabbit (#BA-1000, Vector Laboratories, Burlingame, CA, USA), horse-radish peroxidase (HRP)-conjugated anti-mouse (#7076, Cell Signaling, Danvers, MA, USA), HRP-conjugated anti-rabbit (#403005, Southern Biotech, Birmingham, AL, USA), and Goat Anti-Rabbit IgG-HRP (#4030-50, Southern Biotech, Birmingham, AL, USA). Recombinant human SFRP2 protein (SFRP2) was prepared as previously described [25], and provided by the Protein Expression and Purification Core facility from the University of North Carolina, Chapel Hill.
SFRP2 mRNA expression
SFRP2 mRNA expression levels were determined using the Cancer Genome Atlas (TCGA) database that was accessed on cBioPortal [26, 27]. On cBioPortal, the SFRP2 gene was queried for all cancer types in the TCGA database. SFRP2 mRNA expression levels were collected for each patient and each cancer type, and descriptive statistics were performed. Gene Expression Profiling Interactive Analysis (GEPIA) [28] is an online tool that includes RNA sequencing data from several databases and can be used for interactive analysis of differential gene expression, profile plotting, survival analysis, and correlation analysis. GEPIA was used to analyze the differential RNA expression of SFRP2 in pancreatic cancer in the TCGA compared to normal pancreatic tissue. Survival analysis with the Kaplan-Meier method was also conducted through GEPIA to determine the association of disease-free survival with high or low RNA expression of SFRP2 in TCGA samples. We looked at publicly available datasets to explore percentiles at which there was a statistically significant decrement in survival. Based on these mRNA data, the cutoffs that we can use 20the percentile or less for low group and 50th percentile or greater for the high group provided a statistical separation of survival.
SFRP2 mRNA expression correlation to stroma in pancreatic cancer
The level of SFRP2 mRNA expression in pancreatic cancer was obtained in units of fragments per kilobase of transcript per million (FPKM) from the Human Protein Atlas (
Protein extraction from pancreatic tumor tissues
Human pancreatic tumor samples and their matched normal adjacent tissues from pancreatic resections were obtained from the Medical University of South Carolina Tissue Biorepository. These de-identified tissues were obtained from institutional surgical specimens under an IRB-approved protocol and were stored at
Immunohistochemistry
Pancreatic tumors were resected from KPC mice, fixed in formalin, paraffin-embedded, and 8
Evaluation of epithelial mesenchymal transformation in fibroblasts
Mouse pancreatic fibroblasts and human skin fibroblasts were plated at 60% confluence in DMEM with 10% FBS and treated with either TGF
Effect of KRAS inhibition on SFRP2 expression in PDAC cells
PANC-1R15 and PSN-1R15 were treated with doxycycline (DOX) to induce expression of R15 Monobody, which selectively inhibits a subset of oncogenic KRAS proteins including KRASG12D and KRASG12R [21]. Following DOX treatment (1–2
To confirm if KRAS inhibition alters SFRP2 expression, we inhibited KRAS activity by a second method using a KRASG12D inhibitor, MRTX1133 (Selleckchem, E1051, Houston, TX) [31], on PANC-1 pancreatic cancer cells and performing western blot probing for SFRP2. In this experiment, PANC-1 cells were treated with MRTX1133 (20
Western blot analysis
The protein content was equalized across samples using a Bradford solution, Bio-Rad Protein Assay (#500-0205, Bio-Rad Laboratories). Samples containing equal amounts of total protein were run on 10% bis-tris SDS-PAGE gels (#NW00100BOX Invitrogen) for 1 hour at 140 V. Proteins were transferred onto a PVDF membrane (#LC2005 Life Technologies; Carlsbad, CA, USA) at 30 V for 70 min. Membranes were then incubated for 30 min at RT in blocking buffer (PBS, Twin 0.1%, 5% milk), and then at 4∘C overnight with the primary antibody resuspended in blocking buffer. The following day, membranes were incubated with a HRP-conjugated secondary antibody (1:5000) for 1 hour at RT. The ECL Prime Western Blotting System was used to visualize protein bands (#RPN2232; GE Healthcare). Protein levels from test samples were compared to the loading controls (actin, GAPDH, or vinculin). To do this, densities were calculated by multiplying the average intensity by the surface of each band, and the percentage of control was calculated using the following formula: marker of interest total density x 100/loading control total density. For each experimental sample, values were then normalized to a control which was given the value of 1.00.
Statistical analysis
Paired
Results
SFRP2 RNA levels are the highest in pancreatic cancer and are associated with poor survival
SFRP2 protein is present in multiple cancers including angiosarcoma, hepatocellular carcinoma, prostate, colon, renal, lung, ovarian, and pancreatic cancers [7]. Therefore, we first assessed which cancer had the highest SFRP2 expression levels. To evaluate SFRP2 mRNA levels, we used the RNAseq data from the TCGA. We collected RNAseq data for SFRP2 expression from all 32 cancer types in the database (Fig. 1A). The median number of samples for each cancer type was 294 (range 36-1,082). Pancreatic cancer was found to have the highest levels of SFRP2 mRNA compared to all other tumor types, with a median expression of 13,155 RNAseq by expectation maximization (RSEM) (5.11-120,416 RSEM). In the vast majority of tumor samples among the 32 types of cancer, the SFRP2 gene was not mutated. Next, we compared the mRNA expression levels between normal pancreas and pancreatic adenocarcinoma using the GEPIA online tool. Pancreatic adenocarcinoma mRNA expression data were obtained from the TCGA RNAseq database (
SFRP2 mRNA expression is increased in pancreatic cancer. (A) SFRP2 mRNA levels were compared between 32 cancer types in the TCGA database. Pancreatic cancer had the highest SFRP2 median expression levels. (B) The online interactive tool GEPIA was used to compare SFRP2 mRNA levels between pancreatic cancer samples (
The GEPIA online tool was used to evaluate the association between SFRP2 mRNA expression and disease-free survival for pancreatic cancer patients. High SFRP2 mRNA levels were defined as SFRP2 mRNA expression in the 50th percentile or higher, and low SFRP2 mRNA levels were defined as SFRP2 mRNA expression in the 20th percentile or lower. Disease-free survival was compared between high SFRP2 mRNA levels (
Inhibition of KRAS reduces SFRP2 expression in PDAC. A, B, C, D) Band densities (average intensity x band surface) were normalized to the loading control, Vinculin, to eliminate inter-sample variability, to quantitatively obtain the Intensity Ratio. For each cell type, results were normalized to the untreated control: -Dox (A), and 0 h (B). (A) Western blot analysis of PDAC expressing a doxycycline (DOX)-inducible CFP-R15 monobody. DOX-induced R15 (+) reduced SRFP2 levels in both PDAC lines and a PANC-1 tumor sample. DOX: doxycycline. (B) Western blot analysis of PANC-1 cells treated with DMSO or MRTX1133 (10 or 20 nM), a noncovalent selective KRASG12D inhibitor, for various periods of time shows a reduction of p-ERK and SFRP2 at 4 hours compared to control. (C) Treatment of MIA PaCa-2 cells with trametinib (MEK inhibitor) at 100 nm for 4 hours shows a reduction in p-ERK and SFRP2. (D) Treatment of MIA PaCa-2 cells with buparlisib (PI3Kinase inhibitor) at 1 uM for 4 hours shows a reduction in p-AKT and SFRP2 compared to control.
We previously showed that frizzled-5 (FZD5) is the receptor for SFRP2 in the non-canonical WNT-calcineurin pathway in endothelial, angiosarcoma, and breast cancer cells [25]. Therefore, we measured SFRP2 and FZD5 protein levels in pancreatic cell lines. Both KRAS and p53 mutation-bearing cell lines were selected, since they are the most common mutations in pancreatic cancer, with a rate of 90% and 75%, respectively for each marker [32]. SFRP2 and FZD5 protein levels were assessed by western blot in two human pancreatic cancer cell lines, AsPC-1 and MIA-PaCa2, and one murine cell line, Panc02-luc. SFRP2 and FZD5 protein levels were further compared to the levels in SVR angiosarcoma, used as a positive control [25]. PBS and media were used as negative controls. All three pancreatic cell lines showed high SFRP2 and FZD5 protein levels compared to the positive control, with no protein in the negative controls (Fig. 1D).
KRAS regulates SFRP2 protein in PDAC cells
To determine whether mutant KRAS influences SRFP2 expression, we utilized the recently described R15 monobody to directly inhibit oncogenic RAS activity in PDAC cells (Fig. 2A) [21]. DOX-induced expression of R15 in either PANC-1R15 or PSN-1R15 led to a reduction in SRFP2 protein levels. In addition, PANC-1R15 xenograft tumors treated with the monobody contained lower SFRP2 levels than untreated tumors (Fig. 2A), further supporting regulation of SFRP2 expression by mutant KRAS signaling in PDAC. To confirm this result with a different KRAS inhibitor, PANC-1 cells were treated with MRTX1133, a non-covalent and selective inhibitor of KRASG12D, for periods of time ranging from 0 to 48 hours. As shown in Fig. 2B, MRTX1133 reversibly inhibited the activity of ERK, a known target of KRASG12D, and reduced the expression SFRP2, with a maximum effect at 4 hours. To further evaluate downstream mediators of the KRAS pathway, MIA PaCa-2 cells were treated with dabrafenib (B-Raf inhibitor) and trametinib (MEK inhibitor), which reduced phospho-ERK and SFRP2 protein levels (Fig. 2C). Likewise, the inhibition of PI3Kinase by buparlisib, another downstream target of KRAS, reduced phospho-AKT and SFRP2 protein levels (Fig. 2D). This demonstrates that in pancreatic cancer, SFRP2 is regulated by both KRAS/RAF/MEK/ERK and KRAS/ PI3Kinase pathways.
SFRP2 protein expression is increased in human pancreatic cancer compared to adjacent normal pancreatic tissue
To compare SFRP2 protein expression levels in human pancreatic cancer to matched normal adjacent pancreatic tissue, we measured the levels of SFRP2 protein by western blot in 23 pairs of samples. Table 1 shows the clinical characteristics of these patients. Forty-two percent were male, and the median age was 67.5 (range 47–87). Seventy five percent of the patients were Caucasian while the remaining 25% were non-Hispanic Blacks. Seventy-five percent of patients had Stage IIB disease (ranged from IA to IIB). The densitometry of SFRP2 protein bands was measured in tumor and normal tissues and normalized to the loading control, GAPDH, for each tumor and matched normal tissue assessed. Results showed significantly higher expression of SFRP2 in tumor tissues with an average percent GAPDH of 57.0
Clinical Characteristics of Patients Analyzed
Clinical Characteristics of Patients Analyzed
SFRP2 protein expression is increased in pancreatic cancer. (A) Blot plot of the densitometry values from representative western blot comparing the levels of SFRP2 between tumor tissue from patients in Table 1 and their normal adjacent pancreatic tissue. (B) Representative western blot of tumor tissues (T) and their normal adjacent tissues (N). Densitometry was calculated by measuring the total intensity of each SFRP2 band in tumor and normal samples, and comparing them to the loading control, GAPDH.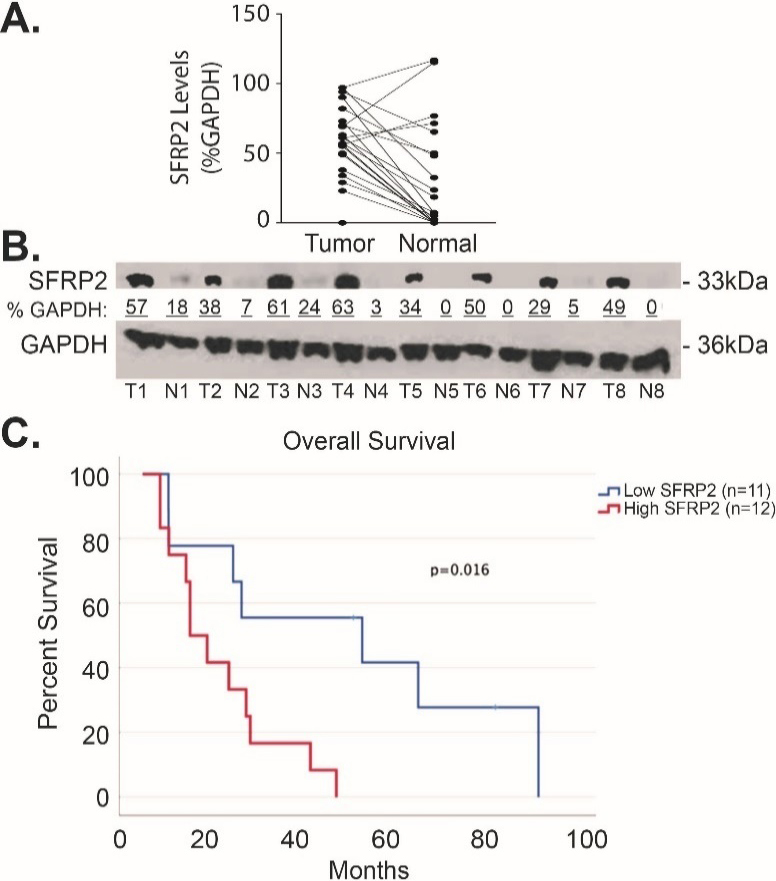
Pancreatic cancer has the highest stroma content of any solid organ tumor, which is a major barrier to chemotherapy tumor biodistribution [6]. Since SFRP2 is associated with fibrosis in other pathologies [18], including cancers [19], we hypothesized that SFRP2 expression levels would be associated with the degree of fibrosis in pancreatic cancer. SFRP2 mRNA expression from pancreatic cancer patients in the TCGA (
SFRP2 RNA expression is associated with fibrosis in pancreatic cancer. mRNA expression of SFRP2 in pancreatic cancer patient samples from the TCGA (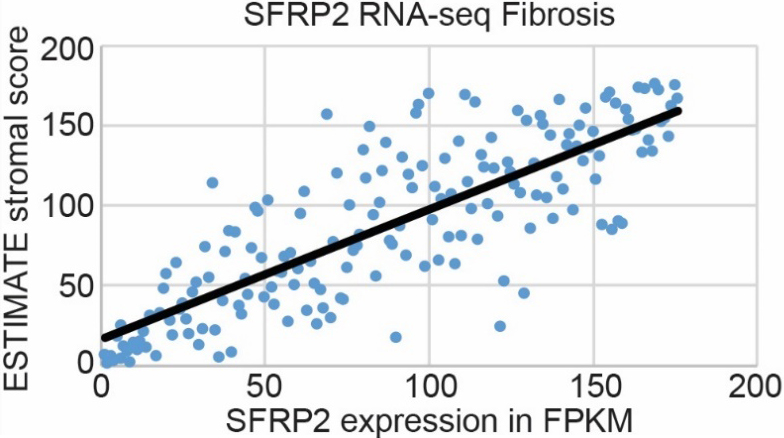
Next, we tested the correlation between fibrosis in pancreatic tumors and SFRP2 protein expression in tumors in vivo. KPC mice (
SFRP2 expression is associated with fibrosis in vivo. (A) Correlation of SFRP2 staining and degree of fibrosis in tumors from KPC mice (
Because we consistently observed a positive correlation between SFRP2 levels and fibrosis, and since SFRP2 is connected to epithelial-mesenchymal transition (EMT) in network analysis [18], we hypothesized that SFRP2 induces a mesenchymal transformation in fibroblasts. To test this hypothesis, we used two cell types: normal human skin fibroblasts and normal mouse pancreatic fibroblasts. Cells were treated for 48 hours either with TFG
SFRP2 induces epithelial mesenchymal transformation and collagen production in fibroblasts. (A) Human skin and mouse pancreatic fibroblasts were either untreated (control) or treated for 48 hours with SFRP2 (30 nM) or TGF
The challenges surrounding pancreatic cancer treatment lie in its tremendous resistance to chemotherapeutics. The dense stroma of pancreatic cancer is a key contributor to this resistance, establishing an environment with low vascularity that impedes drug delivery and immune cell infiltration [4]. In this study, we found SFRP2 to be highly expressed in pancreatic cancer compared to normal adjacent pancreatic tissue, and high SFRP2 expression was predictive of worse survival using a TCGA data set. Based on these findings, a future prospective clinical study to confirm the high and low cutoffs will need to be obtained in future prospective clinical trials. SFRP2 as a reliable biomarker is warranted. SFRP2 also correlated with pancreatic cancer fibrosis, and fibroblasts treated with SFRP2 protein expressed higher levels of mesenchymal markers involved in EMT, a phenotype that correlates with cancer invasion and metastasis.
With the TCGA RNAseq data, we found that SFRP2 expression levels are the highest in pancreatic cancer. Conversely, both SFRP2 mRNA and protein levels appeared to be significantly lower in normal pancreatic tissue, compared to adjacent tumor tissue. This suggests that the neoplastic transformation of the pancreatic tissue correlates with an upregulation of SFRP2. KRAS mutation is the most frequent mutation in PDAC, occurring in more than 90% of tumors [2], and our data show that SFRP2 protein levels in PDAC are regulated by KRAS activity. SFRP2 has the potential to serve as a prognostic biomarker and is a potential therapeutic target, which both warrant further investigation.
In this study we found that fibroblasts treated with SFRP2 had increase protein levels of N-cadherin and fibronectin, similar to TGF
SFRP2 has been highly associated with fibrosis and EMT in other tumor types [19]. Sun and colleagues’ investigation into cancer resistance after chemotherapy showed that SFRP2 was upregulated in CAFs subjected to cytotoxic therapy [38]. They also showed that WNT16B signaling is also enhanced , which further contributes to drug resistance. Inhibition of SFRP2 may dually affect chemoresistance by decreasing the tumor stroma, and decreasing the signaling that confers therapeutic resistance. There has been recent interest in targeting the stromal tissue for pancreatic cancer treatment to dismantle the barricade to chemotherapy and immune response. The results of therapeutics used for these effects have been contradictory thus far. A small molecule inhibitor of the sonic hedgehog pathway showed efficacy in preclinical studies [39] but failed to demonstrate improved survival in clinical trials [40, 41]. Meanwhile, some improvements have been observed with other stromal inhibitors in phase I/II trials targeting TGF
Investigation of pancreatic CAFs has shown that this is a heterogeneous population of cells with different functions, which likely contributes to these different findings. Although Ozdemir and colleagues found that myofibroblast depletion led to more aggressive tumors and immune dysregulation, they also discovered that treating these myofibroblast-depleted mice with CTLA-4 checkpoint inhibitors led to less undifferentiated cancer cells, normalized parenchyma, and improved survival compared to non-myofibroblast depleted mice treated with anti-CTLA-4 antibody [44]. Therefore, inhibiting the stroma may need to be accompanied by immunotherapy for efficacy, and further studies will identify whether targeting the stroma overcomes resistance to immunotherapy in pancreatic cancer. We previously showed that SFRP2 affects immune dysregulation, specifically increasing transcription of CD38, which reduces resistance to the PD-1 inhibition [15]. Therefore, targeting SFRP2 may have the added benefit of providing immunotherapy, in conjunction with stromal regulation, inducing tumor apoptosis and inhibiting angiogenesis. Since SFRP2 is highly expressed in pancreatic cancer; its expression is significantly associated with worse survival, and it is regulated by KRAS; SFRP2 may serve as a novel therapeutic target for pancreatic cancer.
Conclusion
We demonstrated that SFRP2 is highly expressed in pancreatic cancer and its expression is significantly associated with worse disease-free survival and is regulated by KRAS. We confirmed that, like other cancer types, SFRP2 is associated with fibrosis in pancreatic cancer. Investigation of SFRP2’s impact on fibroblasts showed that SFRP2 stimulates EMT, which identifies an additional role for SFRP2 as a regulator for cancer invasion, angiogenesis, metastasis, and ECM construction. Targeting the stroma is a promising avenue for pancreatic cancer treatment, but the current literature demonstrates that further research on the communication of all heterogeneous cell populations, cytokines, and proteins in the tumor microenvironment is needed to understand how best to manipulate it to improve survival. SFRP2 is one such protein that plays a part in this crosstalk and merits additional study to determine its potential as a regulator for both stroma production and immune dysregulation.
Author contributions
Conception: NKD, JBS, PN, JPO; Interpretation or analysis of data: JBS, PN, KLH, LS, RM, JPO, JEM, IK; LH, MG, BR, Preparation of the manuscript: JBS, PN, NKD, RM, IK, JPO, NGD, EH, LH; Revision for important intellectual content: NKD; Supervision: NKD, JBS, PN.
Supplementary data
The supplementary files are available to download from http://dx.doi.org/10.3233/CBM-220044.
sj-docx-1-cbm-10.3233_CBM-220044.docx - Supplemental material
Supplemental material, sj-docx-1-cbm-10.3233_CBM-220044.docx
Footnotes
Acknowledgments
NKD was supported by grants from the Hirshberg Foundation for Pancreatic Cancer Research Seed Grant Award, and a donation from Dr. Robert and Mrs. Laura Ryan. Dr. Ryan is CEO of Innova Therapeutics, Inc. JPO was supported by grants from the Department of Veterans Affairs Biomedical Laboratory Research and Development Service MERIT Award (1I01BX002095) and the NCI (R01CA212608 and P30CA138313). NKD, JPO, and NGD were supported by grants from Hollings Cancer Center Team Science Award, and Tissue collection for this study was supported by the Biorepository & Tissue Analysis Shared Resource, Hollings Cancer Center, Medical University of South Carolina. The study sponsors were not involved in the study design or the collection, analysis, and interpretation of data.
