Abstract
OBJECTIVES:
Cervical cancer (CC) is a common malignant tumor in the female reproductive system that is characterized by a high metastatic potential. LncRNA ANRIL has been found to be a cancer oncogene in multiple tumors. In our study, we altered the expression of ANRIL in CC cells and evaluated its ability on influencing proliferation, migration and invasion of CC cells and associated mechanism.
METHODS:
Differentially expressed lncRNAs in CC were identified by microarray and TCGA analyses. CC tissues and adjacent tissues were collected in order to extract CC cells. The expression of ANRIL was determined by RT-qPCR. The CC cells were transfected with siRNA or si-NC against ANRIL to find out whether ANRIL can influence the expression of Cyclin D1, CDK4, CDK6, E-cadherin, vimentin and N-cadherin, as well as affect cell proliferation, cell apoptosis, cell migration and cell invasion of CC cells.
RESULTS:
Based on TCGA and microarray analyses, ANRIL was predicted to be highly expressed in CC and CC with migration. Then further verification was obtained by means of RT-qPCR that ANRIL was highly expressed in CC tissues. In addition, high expression of ANRIL was related to increased E-cadherin expression, high migration of CC as well as decreased cell apoptosis rate. On the other hand, inhibition of ANRIL expression led to decreased expressions of Cyclin D1, CDK4, CDK6, N-cadherin and Vimentin, along with attenuated cell proliferation, migration and invasion of CC cells.
CONCLUSION:
The key findings of our study demonstrated that the inhibition of lncRNA ANRIL reduces the proliferation, migration and invasion capabilities of CC cells. Down-regulation of ANRIL may serve as a potential therapeutic target in the treatment of CC.
Introduction
Cervical cancer (CC) is the second leading cancer occurring in women. Its incidence is high in developing countries with approximately 500,000 patients diagnosed and 280,000 deaths each year. It is mainly linked to a human papillomavirus (HPV) infection whereby the fusion of its DNA with the host genome has been linked to the occurrence of CC [1, 2]. There are many causes of CC, this includes: genetic susceptibility, viral infections such as HPV, smoking, oral contraceptives and environmental factors [3, 4, 5, 6]. Up until now, the mechanism of how HPV infections can contribute to the progression of CC is not fully understood. Metastases of CC outside or the progression to a late stage with poor responses to treatments generally result in unfavorable prognosis of patients with CC [7]. Despite treatment approaches such as surgery, chemotherapy and radiotherapy in treating patients with CC, the 5-year survival rate of patients with advanced CC is still very low [8].
Long non-coding RNAs (lncRNAs) are generally defined as genomic transcriptions longer than 200 nucleotides (nt) in length that are not translated into protein. Their lack of function in protein coding is due to a lack of an open significant reading frame length [9]. A previous study demonstrated that lncRNAs played a crucial role in a variety of cellular processes by interacting with some key component proteins during gene regulation. The changes in tissue-specific and cell-specific expression as well as primary or secondary structures were thought to influence cell proliferation, metastasis and invasion [10]. In addition, there has been growing evidence which revealed that dysregulated expressions of lncRNAs were implicated in various types of cancers including gastric cancer, breast cancer, lung cancer and CC [11, 12, 13, 14]. LncRNA ANRIL has been found to encode a kind of 3834-nt RNA which contains 19 exons located in the antisense direction of the INK4B-ARF-INK4A gene cluster that is usually upregulated in nasopharyngeal carcinoma [15]. Previous evidence has also proved that lncRNA ANRIL expression was increased in GC which was correlated with advanced TNM stage and tumor size. In addition, further experiments have been done to reveal that inhibition of ANRIL expression significantly suppressed cancer cell proliferation [16]. Thus, it can be seen that a lot of data support lncRNA ANRIL’s key role in cancer progression. However, the expression of levels of lncRNA ANRIL in CC and the underlying mechanism have yet to be discovered. Therefore, this study aims to investigate the effects of lncRNA ANRIL expression levels on the proliferation, migration and invasion of CC cells.
Materials and methods
Microarray and statistical analysis
CC microarray profiles (GSE26511) and probe annotation from the Gene Expression Omnibus (GEO) database from the National Center for Biotechnology Information (NCBI) (
Ethics statement
The study was approved by the Ethics Committee of The First Affiliated Hospital of Henan university of Chinese Medicine. All participating patients have understood experimental procedures and signed the informed consents.
Study subject selection
From November 12, 2011 to February 15, 2016, 277 CC patients (aged between 18 to 75 years, median age of 46 years) diagnosed with cervical cancer in The First Affiliated Hospital of Henan university of Chinese Medicine were selected for this study. Patients with CC were divided into T
Cell extraction
Resected CC tissues were cut into small pieces and then detached with 1% trypsin (Sinopharm Chemical Reagent Co., Ltd., Shanghai, China). After incubation for 30 minutes, serum was added to terminate the reaction. A suction pipette was used to resuspend CC tissues. A Cytoscreener with an aperture of 70
Cell transfection and culture
CC cells were initially inoculated in 100 mL/L 1640 medium containing fetal bovine serum (FBS). Cells were then incubated at 37
RNA extraction and reverse-transcription quantitative polymerase chain reaction (RT-qPCR)
RNA extraction was performed on ice. RNA extraction from tissue was as follows: 15 random samples of CC and adjacent tissues were selected and cut into small pieces and grounded into fine powder in liquid nitrogen. In the next step, we used spoon pre-cooled by liquid nitrogen to add 50 to 100 mg of tissue powder to a 1 mL Trizol eppendorf (EP) tube. Cell RNA pretreatment was performed as follows: the medium in 6-well plates was aspirated and removed, then washed with PBS twice. Next, 1 mL of Trizol (Invitrogen Inc., Carlsbad, CA, USA) was then added into each well. Cells were scraped repeatedly using a pipette tip and transferred into 1.5 mL EP tubes.
Total RNA of cells or tissues was exacted using Trizol. When extracting, DNase I (TransGen Biotech Co., Ltd., Beijing, China) was added in order to remove genomic contamination. A reverse transcription kit (Takara Biotechnology Ltd., Dalian, China) was used to reverse transcribe RNA into cDNA, which was used as a template for RT-qPCR amplification. The reaction was performed on ordinary PCR amplification instrument (Bio-Rad Laboratories, Inc., CA, USA). The PCR instrument collected fluorescence signal by preheating at 95
Primer sequences for reverse-transcription quantitative polymerase chain reaction
Primer sequences for reverse-transcription quantitative polymerase chain reaction
Notes: F, forward; R, reverse; GAPDH: glyceraldehyde-3-phosphate dehydrogenase.
The total protein from CC cells was lysed using RIPA cell lysate (BB-3209, Bestbio Biological, Shanghai, China), followed by electrophoretic separation with sodium dodecyl sulfate-polyacrylamide gel electrophoresis (SDS-PAGE). After separation, the samples were transferred onto a polyvinylidene fluoride (PVDF) membrane using a voltage of 80 V. The membrane was sealed for 1 hour and incubated overnight at 4
Cell Counting Kit-8 (CCK-8) assay
After transfection for 48 hours, cells were then seeded in a 96-well plate at a density of 2
Flow cytometry
After transfection for 48 hours, CC cells were incubated, centrifuged at 800 r/min for 5 minutes (centrifugal radius was 15 cm) and washed twice with ice-cold PBS. According to the instructions provided by the Annexin-V-FITC cell apoptosis detection kit (K201-100, Biovision, Moutain View, CA, USA), cells were resuspended after the addition of 200
Scratch test
The cells in each group were seeded in 6-well plates and transfected when the cells reached 50% confluence. Forty-eight hours after transfection, culture solution was replaced and cells were cultured for another two days until cells reached adequate confluence. A vertical scratch was made in the cell culture surface using a 100
Transwell assay
Serum free medium (300
Statistical analysis
All data was analyzed statistically using an SPSS 20.0 software (IBM Corp. Armonk, NY, USA). Results are expressed as a mean
Based on TCGA and microarray analysis, ANRIL is predicted to be highly expressed in CC and CC with metastasis. Panel A, LncRNA ANRIL is highly expressed in CC metastasis by using a GSE26511 chip data; Panel B, ANRIL is highly expressed in CC according to the TCGA database. The abscissa refers to the sample number, and the ordinate refers to names of DEGs. The upper right histogram refers to color shades. The color change from the top to the bottom corresponds to a change in expression from high to low. Each block represents the level of gene expression in one sample, and each column represents the expression of all genes in samples. The left dendrogram refers to the cluster analysis results of different genes in different samples. The transverse line at the top represents the types of samples, and the upper right square represents the sample color reference: the blue refers to the metastasis group and the red refers to the non-metastasis group.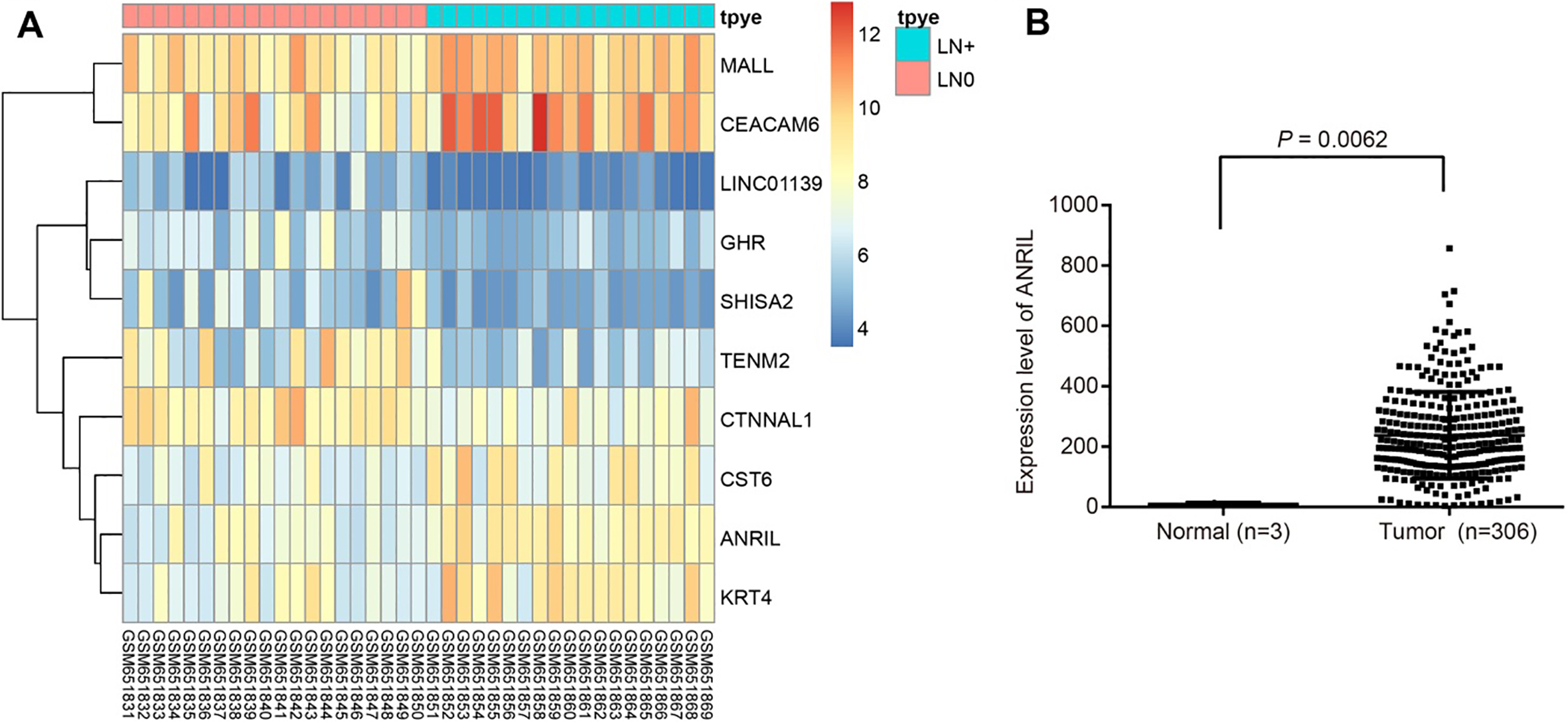
ANRIL is predicted to be highly expressed in CC
In recent years, lncRNAs have been shown to play important roles in tumor biological function in CC. In order to identify differentially expressed lncRNAs in CC, we performed a comprehensive lncRNA profiling analysis that was made available from the GEO database of the NCBI. Through CC microarray analysis (GSE26511), the expression of lncRNA ANRIL was found to be elevated in CC with cell migratory features (Fig. 1A); based on the TCGA database, ANRIL was highly expressed in CC (Fig. 1B).
Relationship between clinicopathological characteristics and CC metastasis
Relationship between clinicopathological characteristics and CC metastasis
Note: CC, cervical cancer; TNM, tumor-node-metastasis.
Initially, relations of CC and age, TNM stage, differentiation degree, pathological classification, initial sexual behavior time, myometrial invasion depth, vessel invasion and ANRIL expression were assessed (Table 2). The results indicated that there was no significant difference in age between the metastatic and non-metastatic groups (
RT-qPCR shows that there is high expression of ANRIL in CC tissues. ANRIL mRNA expression in T1 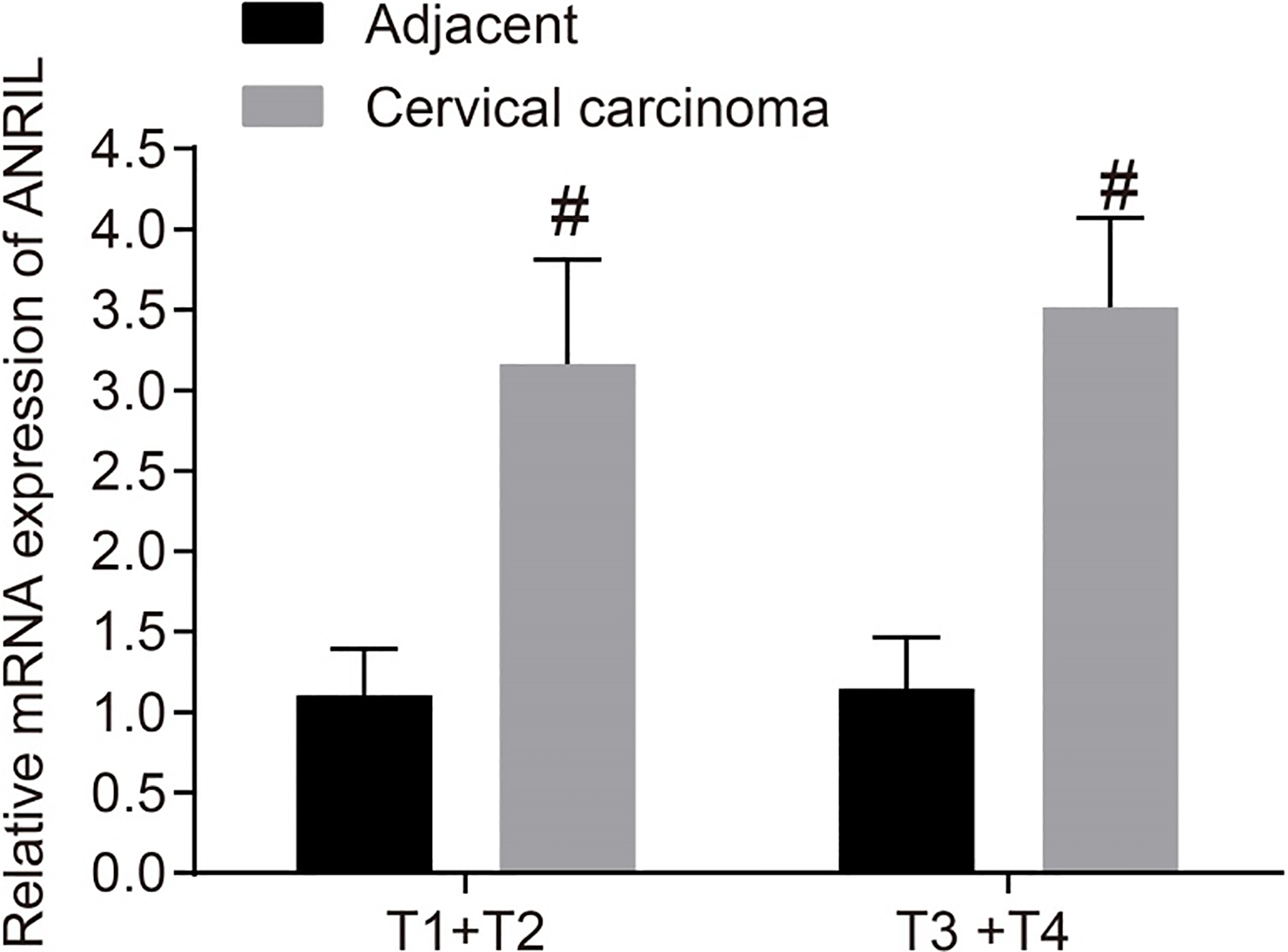
The expression of ANRIL in CC tissues and adjacent tissues at different TNM stages was measured by RT-qPCR. The results are shown in Fig. 2. The ANRIL expression in CC tissues was significantly higher in both T1
siRNA lowers ANRIL expression and the inhibition of ANRIL hinders CC cell proliferation. Panel A, after ANRIL silencing for 48 hours, ANRIL mRNA expression was detected by RT-qPCR; Panel B, cell proliferation in each time point at 48 hours after transfection was measured by CCK-8 assay; Panel C, protein bands of Cyclin D1, CDK4 and CDK6 at 48 hours after transfection; Panel D, protein levels of Cyclin D1, CDK4 and CDK6 48 hours after transfection; Statistical values are expressed as mean 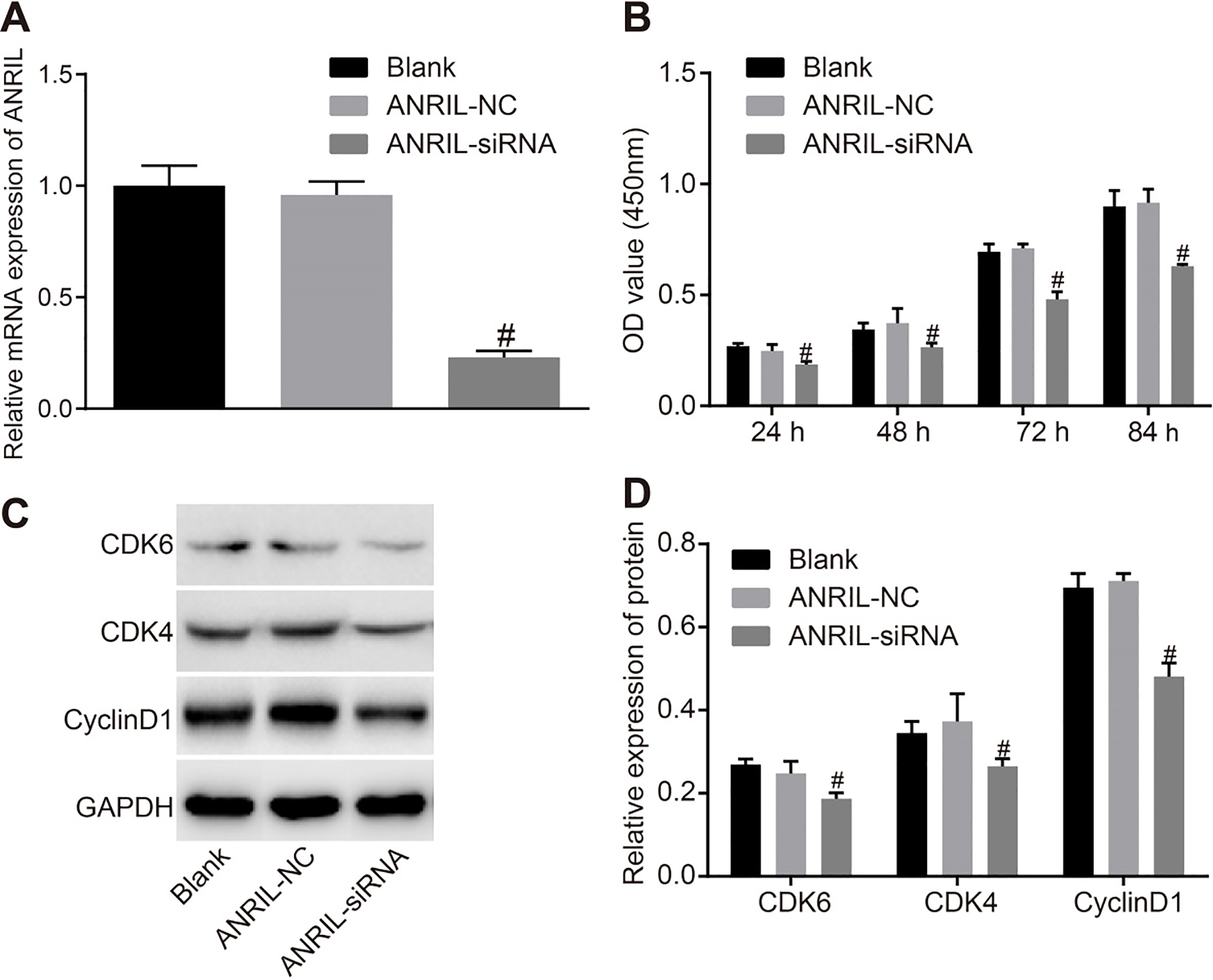
In the next step, we used RNA interference technology to conduct ANRIL silencing. After cellular transfection of ANRIL-siRNA and ANRIL-NC, ANRIL mRNA expression in each group was detected using RT-qPCR. As shown in Fig. 3A, ANRIL mRNA expression was significantly lower in the ANRIL-siRNA group compared with the control group (
CCK8 assay was used to determine the effect of ANRIL on proliferation of CC cells. The results are shown in Fig. 3B. As time progressed, all three groups showed increased trend in proliferation of CC cells. At 24 hours, the ANRIL-siRNA group showed a lower OD value compared to the control and ANRIL-NC groups (
Inhibition of ANRIL increases cell apoptosis
We used flow cytometry to determine the effects of ANRIL silencing on CC cell apoptosis. Results showed that the cell apoptotic rate was higher in the ANRIL-siRNA group compared with that in the control and ANRIL-NC groups (
Flow cytometry analysis shows that the cell apoptotic rate increased as a response to the inhibition of ANRIL. Panel A, apoptosis of CC cells after transfection for 48 hours; Panel B, apoptotic rate of CC cells after transfection for 48 hours; Statistical values are expressed as mean 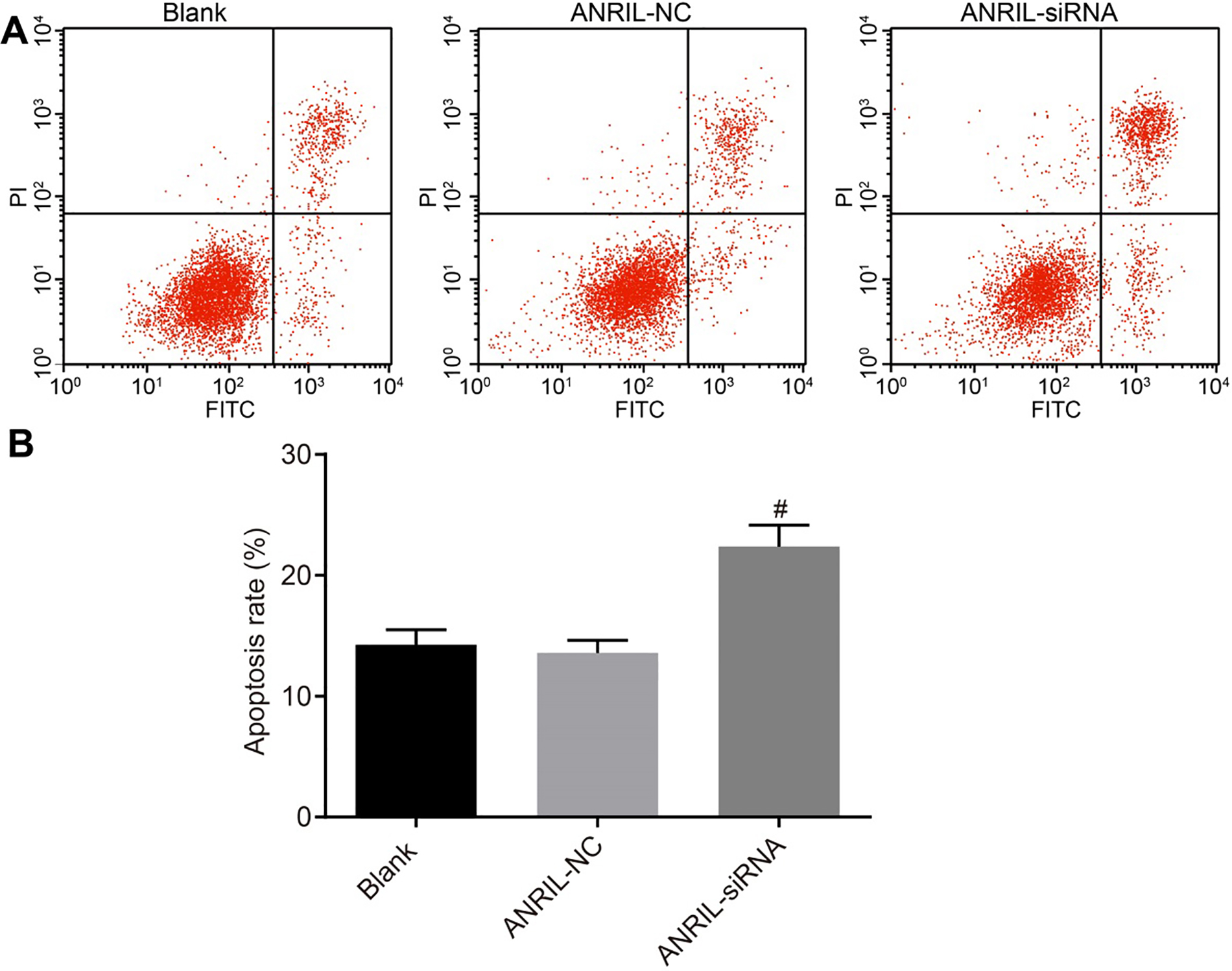
Detection by Scratch test and western blot analysis demonstrates that cell migration of CC cells was suppressed after ANRIL inhibition. Panel A, Scratch test results in each group after transfection for 48 hours; Panel B, comparisons of scratch distance in each group after transfection for 48 hours; Panel C and D, protein levels of E-cadherin, vimentin and N-cadherin were detected by western blot analysis after transfection for 48 hours; statistical values are expressed as mean 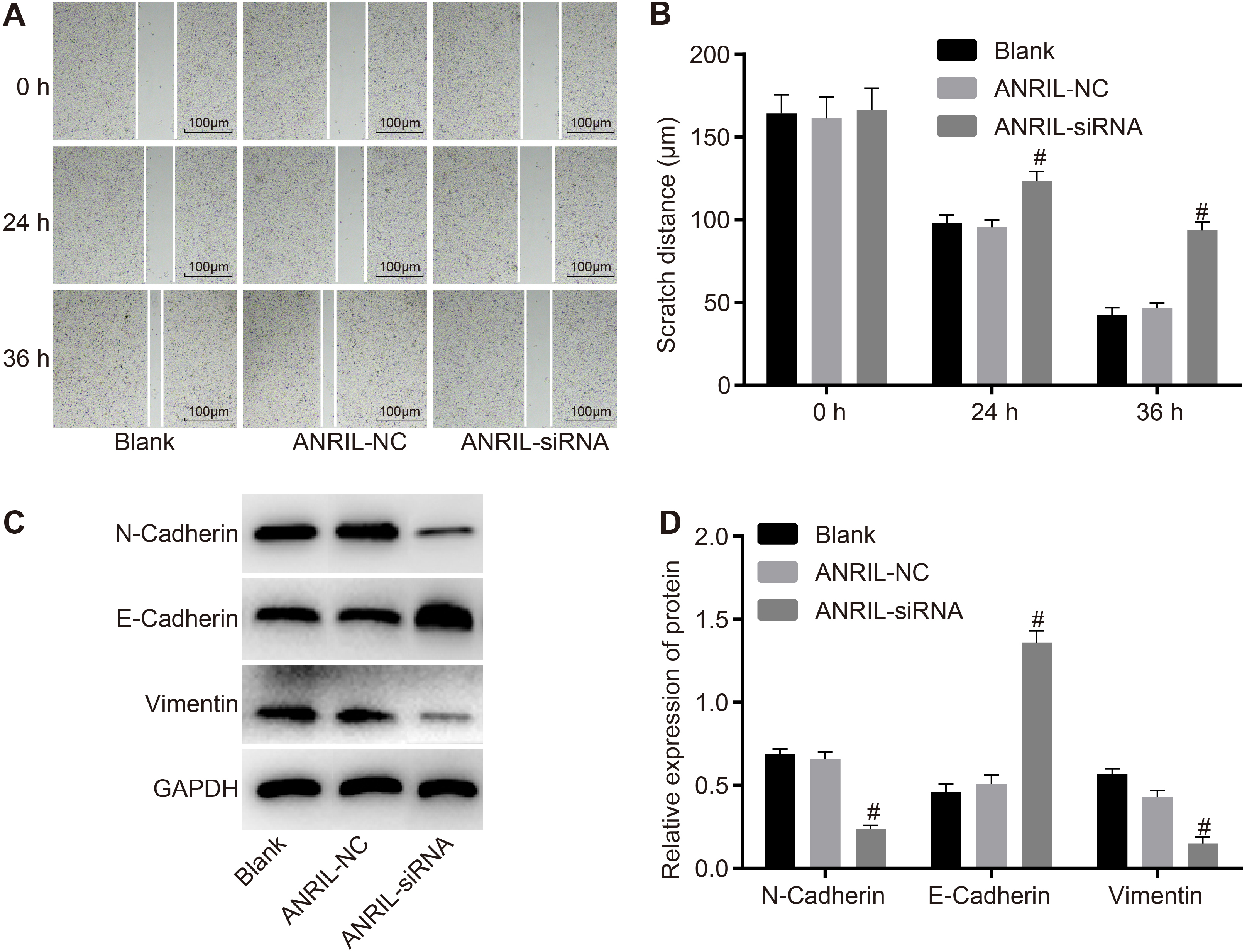
Inhibition of ANRIL attenuates cell invasion in CC cells, based on the Transwell assay. #, 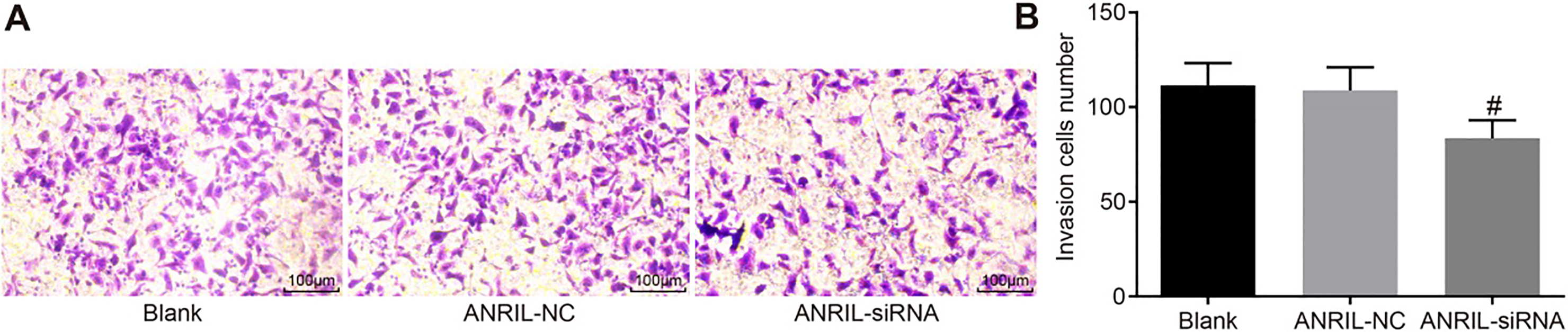
To determine the effect of ANRIL on the migration of CC cells, a scratch test was performed. The results are shown in Fig. 5A and B. There was no significant difference in the cell scratch distance among the three groups (
Inhibition of ANRIL dampens invasion of CC cells
Finally, the Transwell assay was used to evaluate the extent of cell invasion of CC cells via inhibition of ANRIL. The results are shown in Fig. 6. Compared with the ANRIL-NC and control groups, the number of cells passing through the micropore membrane covered by Matrigel was significantly less in the ANRIL-siRNA group (
Discussion
Over the past few years, there have been an increasing amount of studies that investigate the different types of LncRNAs. These include PVT1, MEG3, CCAT2 and many others which play indispensable roles in inhibiting CC. Due to their role in tumor suppressing, LncRNAs have been widely studied and if properly developed, they may be used as a therapeutic target and biomarkers for treating CC [22, 23, 24]. LncRNA ANRIL was reported to contribute to tumor progression in various cancers, however, its function and expression in CC still remain unclear [25]. Therefore, we initially aimed to further investigate the effects of lncRNA ANRIL on the proliferation, migration and invasion of CC cells in this study. Our results demonstrated that the inhibition of lncRNA ANRIL could effectively suppress the proliferation, migration and invasion of CC cells.
Our study found that a higher degree of CC metastasis was found to be associated with advanced TNM stage, poor differentiation, myometrial invasion depth at
Additionally, in our study, overexpression of lncRNA ANRIL was observed in CC cell lines and clinical specimens suggesting that ANRIL expression in CC tissues was significantly higher than those in adjacent tissues. A previous study revealed that higher expression of lncRNA ANRIL in cancer tissues was correlated with advanced TNM stage and low histologic grade [27]. Decreased expression of lncRNA LET inhibited CC, further supporting the notion that lncRNAs can function as a tumor suppressor [28]. Furthermore, higher expression of ANRIL in human cancer tissues was related to lower survival rates of the patients [29]. For these reasons, we inferred that the regulation of lncRNA ANRIL in CC cells and tissues could be a therapeutic target and a promising biomarker for the treatment of CC.
In order to study the effects of lncRNA ANRIL on CC progression, we observed the proliferation, migration and invasion capabilities of CC cells before and after inhibiting lncRNA ANRIL expression. Surprisingly, our study found that cells transfected with ANRIL-siRNA showed significantly reduced CC cell proliferation, migration and invasion compared with the cells transfected with negative controls and normal CC cells. A previous study demonstrated that the suppression of lncRNA ANRIL was carried out using siRNAs to target and inhibit lncRNA ANRIL [26]. According to a previous study, the knockdown of ANRIL inhibited proliferation, metastasis and invasion of colorectal cancer cells [29]. For these reasons, we speculate that a similar mechanism can be identified in CC in addition to colorectal whereby lncRNA ANRIL could act as a regulator. In addition, ANRIL knockdown led to an increase in E-cadherin expression, decrease N-cadherin, Vimentin, Cyclin D1, CDK4 and CDK6 expressions. E-cadherin is a tumor suppressor gene that plays a critical role in the malignant progression of epithelial tumors and inhibits epithelial to mesenchymal transition [30]. E-cadherin expression has been found to be down-regulated in CC whereby lncRNA-EBIC which is an oncogenic lncRNA, could promote tumor cell invasion in CC by inhibiting E-cadherin expression [31]. Vimentin is a type III intermediate filament that has been found to be implicated in various biological processes including maintaining cell shape and stabilizing cytoskeletal interactions. It has been designated as a methylation biomarker for early diagnosis of CC [32]. The p16 (INK4A) tumor suppressor is a critical KDM6B downstream transcriptional target and its expression is critical for CC cell survival. Oncogenic p16 activity depends on the regulation of CDK4/CDK6, suggesting that in CC cells with an inactivated retinoblastoma tumor suppressor, CDK4/CDK6 activity needs to be inhibited in order for cells to survive [33]. Furthermore, an exogenous epidermal growth factor (EGF) stimulation may enhance human papillomavirus (HPV)-related CC cell proliferation by activating Cyclin D1 that is independent of COX-2 levels, suggesting that the inhibitors of Cyclin D1 may be effective against CC cell proliferation [34].
Consequently, lncRNA ANRIL was highly express-ed in CC and participated in the regulation of migration, proliferation and invasion of CC cells. Our study provides evidence that the inhibition of lncRNA ANRIL could suppresses cell proliferation, migration and invasion of CC cells. The development of treatment strategies using ANRIL to down-regulation of lncRNAs can provide a promising and novel therapeutic target treating patients with CC. However, there are several important issues that have yet to be explored as well as the limitations of this study. Despite our findings, the specific mechanisms of ANRIL including its signal pathway and influencing factors still remain unclear. Therefore, further research needs to be carried out in order for us to proceed to discovering therapeutic strategies using lncRNAs.
Footnotes
Acknowledgments
We would like to acknowledge the reviewers for their helpful comments on this paper.
Conflict of interest
None.
