Abstract
BACKGROUND:
Hepatocellular carcinoma (HCC) is a highly heterogeneous disease, which makes prognostic prediction challenging.We aimed to investigate association of TNFRSF4 expression with the immune infiltration and gene mutation in HCC.
METHODS:
In this study, the expression profiles and corresponding clinical data of HCC patients were downloaded from the Cancer Genome Atlas (TCGA) database.
Kaplan-Meier and Cox regression were used to evaluate the clinical value of TNFRSF4. ESTIMATE and CIBERSORT algorithms were applied to investigate the infiltration ratio of 22 immune cells. The WGCNA and LASSO COX algorithms were performed, establishing a prognostic risk model that was then validated by HCC samples from GEO. Finally, the effects on gene mutation occurring in HCC patients of TNFRSF4 expression and risk score were appraised.
RESULTS:
In HCC tissues, it was found the TNFRSF4 expression profile was significantly different with age, gender, tumor grade, disease stage, prominently affecting the survival outcome and prognosis of patients. Univariate and multivariate COX regression analysis suggested that TNFRSF4 was an independent prognostic marker. Samples of high/low expression of TNFRSF4 were screened for differential genes, and then the WGCNA and LASSO COX constructed a 13-gene signature, excellently dividing samples into hign/low risk groups. Compared with the low-risk group, the overall survival (OS) of high-risk group was markedly lower, with
CONCLUSIONS:
As an independent prognostic marker of HCC, TNFRSF4 was found simultaneously to affect the immune infiltration of cells and the frequency of gene mutations.
Abbreviations
AUC, area under curve;
GEO, gene expression omnibus;
GM, gene mutation;
HCC, hepatocellular carcinoma;
IRGs, immune-related genes;
KM, Kaplan-Meier;
LASSO COX, least absolute shrinkage and selection operator;
ROC, receiver operating characteristic;
RS, risk score;
TCGA, cancer genome atlas;
WGCNA, weighted gene co-expression network analy- sis.
Introduction
HCC is the most common type of liver cancer globally, is expected to significantly increase over the next 10 years [1, 2].
Although there are many ways to treat HCC, less than 9% of HCC patients can survive over five years [3]. The high recurrence and metastasis lead to the poor overall survival [4]. Therefore, the determination of reliable prognostic markers is crucial to improve the prognosis of HCC patients.
The tumor immune microenvironment is the environment where tumor cells interact with the immune system. In addition, the mutual interactions between tumor cells and the surrounding tumor microenvironment will eventually promote tumor development and metastasis [5]. Immunotherapy is the most significant way to treat cancer [6], wherein PD-1/PD-L1 pathway is regared as the best target for immunotherapy. Although many studies have reported the impact of PD-1 pathway on prognosis, fewer patients benefit from PD-1/PD-L1 immunotherapy [7, 8]. Thus, the immune microenvironment urgently awaits to be better understood to improve prognosis prediction.
TNFRSF4 (tumor necrosis factor receptor 4), a costimulatory receptor of the TNF family, plays a significant role in the immune microenvironment [9]. TNFRSF4 expression could be reduced by regulatory T (Treg) cell, thereby contributing to developments of various malignant tumors [10]. However, the clinical and prognostic role of TNFRSF4 in HCC development is perplexing. Therefore, the study aims to investigate the prognostic performance of TNFRSF4 in HCC.
General flow chart.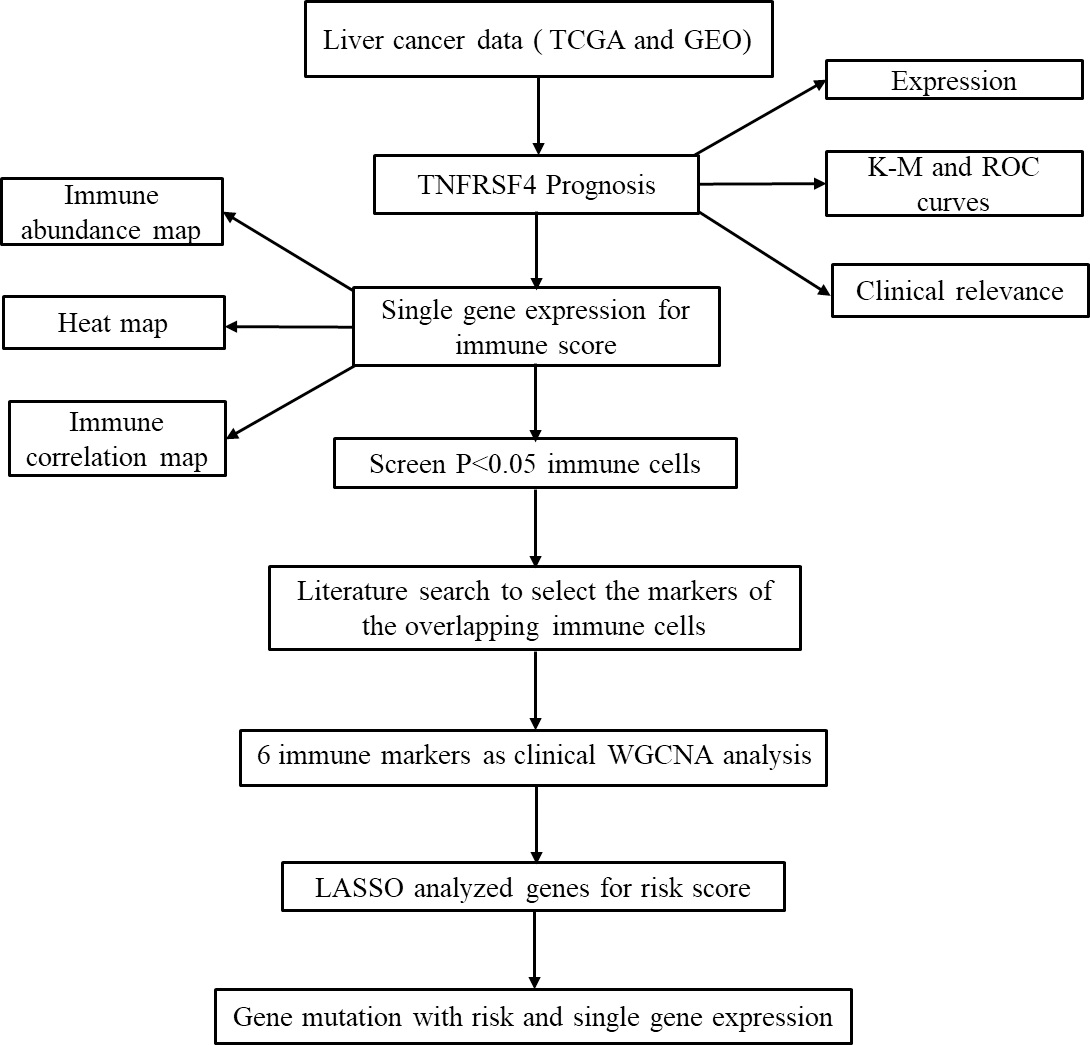
By our previous researches, several prognostic biomarkers and potential inflammation-modulating therapeutic targets were identified through experimental and bioinformatics method [11, 12, 13]. In this study, univariate and multivariate COX were utilized to confirm the potentiality of the TNFRSF4 for improving the survival prediction for HCC. CIBERSORT algorithm was used to score immune cells including 22 kinds. Prognostic immune signatures were constructed based on WGCNA and LASSO COX algorithms. Finally, the effects of TNFRSF4 expression and risk score were evaluated on gene mutation frequency.
TCGA provided patient gene expression and clinical characteristics data (
Prognostic analysis
Univariate Cox analysis was implemented to appraise the association of the overall survival outcome with TNFRSF4 expression level as well as clinical characteristics as age, gender, stage (T, N, M). Following, Multivariate Cox analysis were performed, indicating TNFRSF4 was an independent prognostic marker of HCC (referring to Appendix 1, 2).
Immune cell infiltration
For all the 363 HCC samples (referring to Appendix 3), ESTIMATE algorithm was first used to calculate immune score, and then Wilcoxon test adopted to analyze differential expression of TNFRSF4, and the end, the CIBERSORT software [14] implemented to calculate the expression ratio of TNFRSF4 in 22 immune cells.
Differentially expressed genes (DEG)
The DESeq2 method was taken to yield differential expression of TNFRSF4 using the statistical test. Fold-change (FC) of gene expression was calculated, taking a threshold of logFC
WGCNA analysis
The WGNCA (R package) was performed to construct the co-expression network [15]. In the network, paired Pearson coefficient was employed to assess the co-expression correlation among all genes in the adjacency matrix. Genes with a high degree of full correlation were clustered into the same module. Then, the topological overlap of internal modules and adjacent modules was served as the functional modules. Finally, the correlation between gene expression modules and traits were constructed.
In Eq. (1),
Finally, we conduct module mining to identify the relationship between gene expression modules and immune markers.
Under the univariate COX regression and LASSO regression analysis, 13 genes were incorporated into the model. The risk score (RS) of each sample was calculated using Eq. (4) below:
Almonst,
The ‘maftools’ package in R [16] was executed to observe the effects of TNFRSF4 expression and RS grouping on gene mutations occuring in HCC samples (in this step, excluding samples occurring no mutations, 353 samples were left).
Screening of single gene TNFRSF4.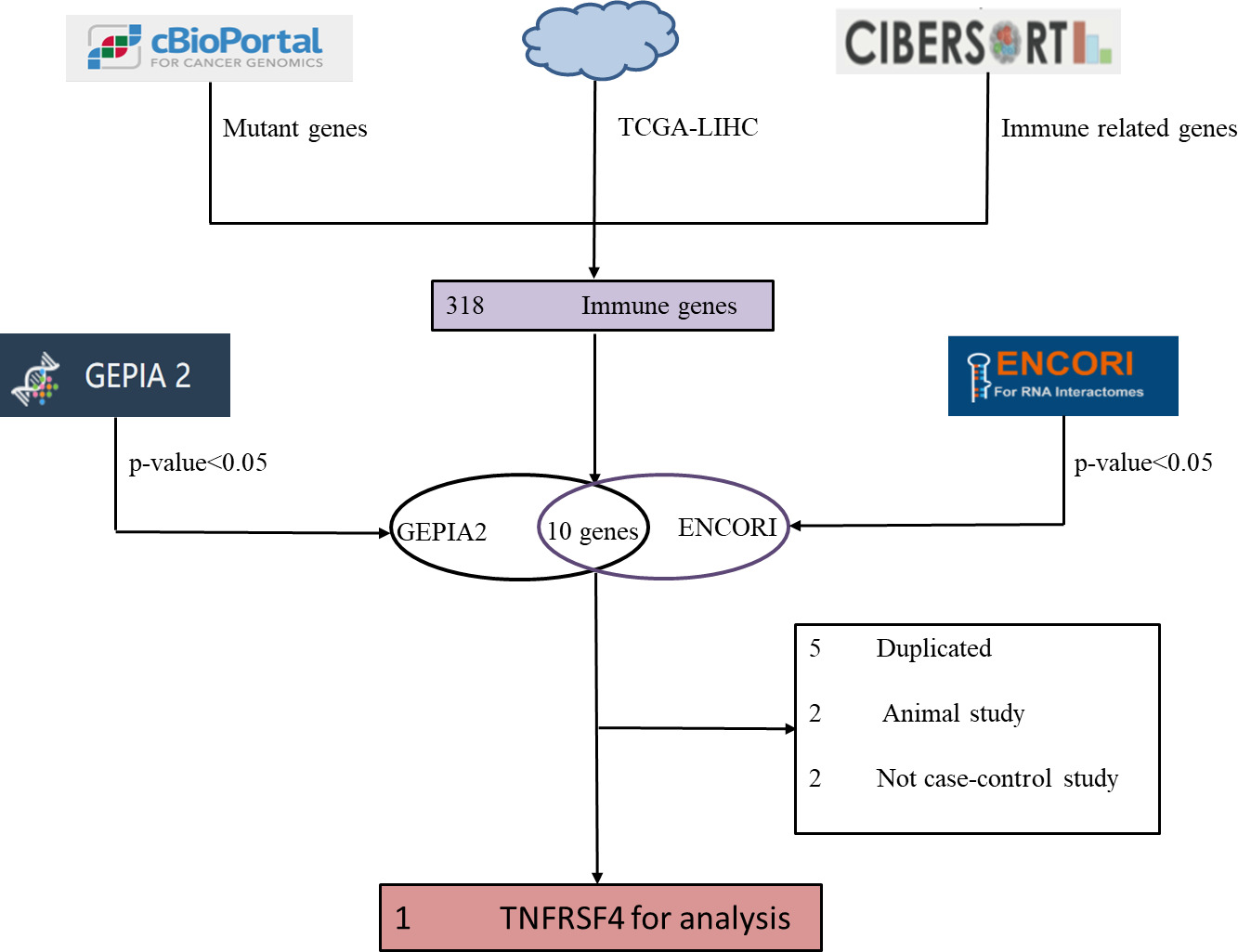
Clinical analysis of TNFRSF4. (A) Violin chart shows the relative expression of TNFRSF4 in normal and tumor samples; (B) KM curve of TCGA cohort high (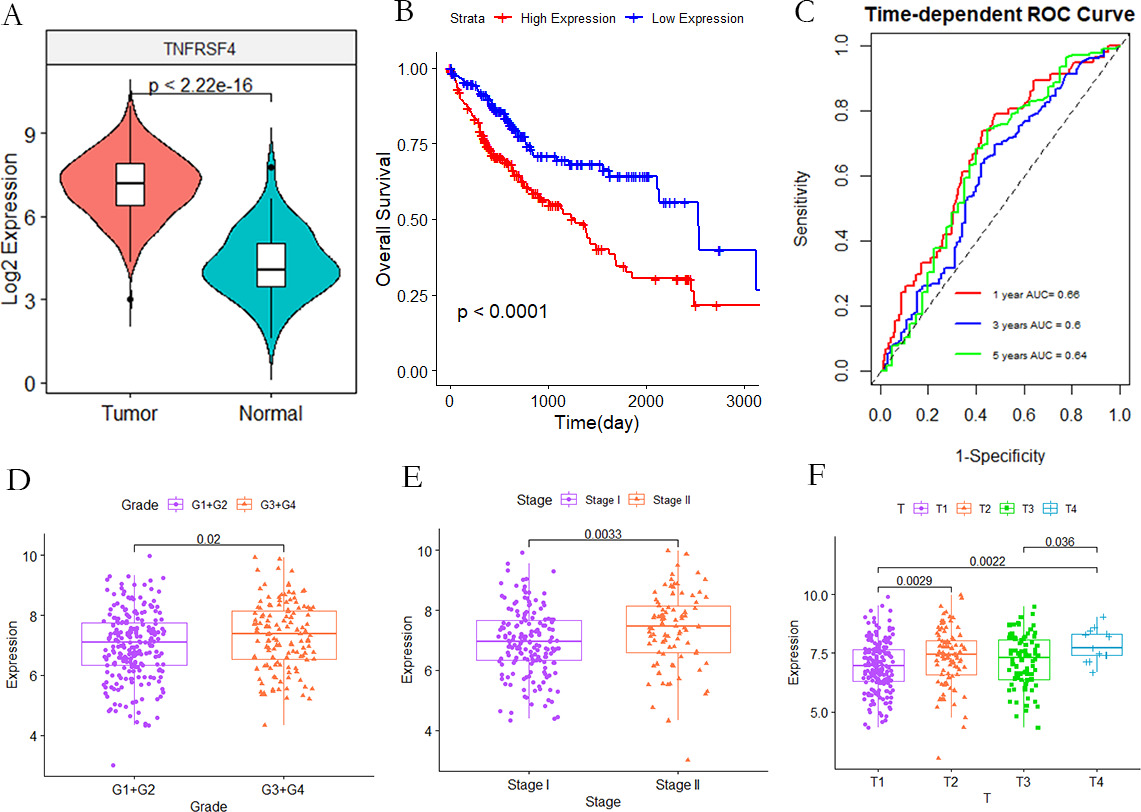
All the statistical analysis in this manuscript were performed by R software (
TNFRSF4 identification in HCC and its prognosis evaluation
318 genes were resulted from the intersection of HCC whole genome, immune-related genes, and the mutation genes, then subject to GEPIA 2 and ENCORI databases to analyze the survival outcome and differential expression, yielding 10 genes with the
The original microarray data was downloaded from TCGA, and the TNFRSF4 expression in HCC samples were analyzed. In Fig. 3A, it is indicated TNFRSF4 expression in HCC samples was up-regulated compared with the case of non-tumor samples. Kaplan-Meier analysis showed that the overall survival outcome of high TNFRSF4 expression group was poor (
TNFRSF4 prognostic value. Univariate and multivariate regression analysis of different clinicopathological variables in TCGA (A–B) and GEO (C–D).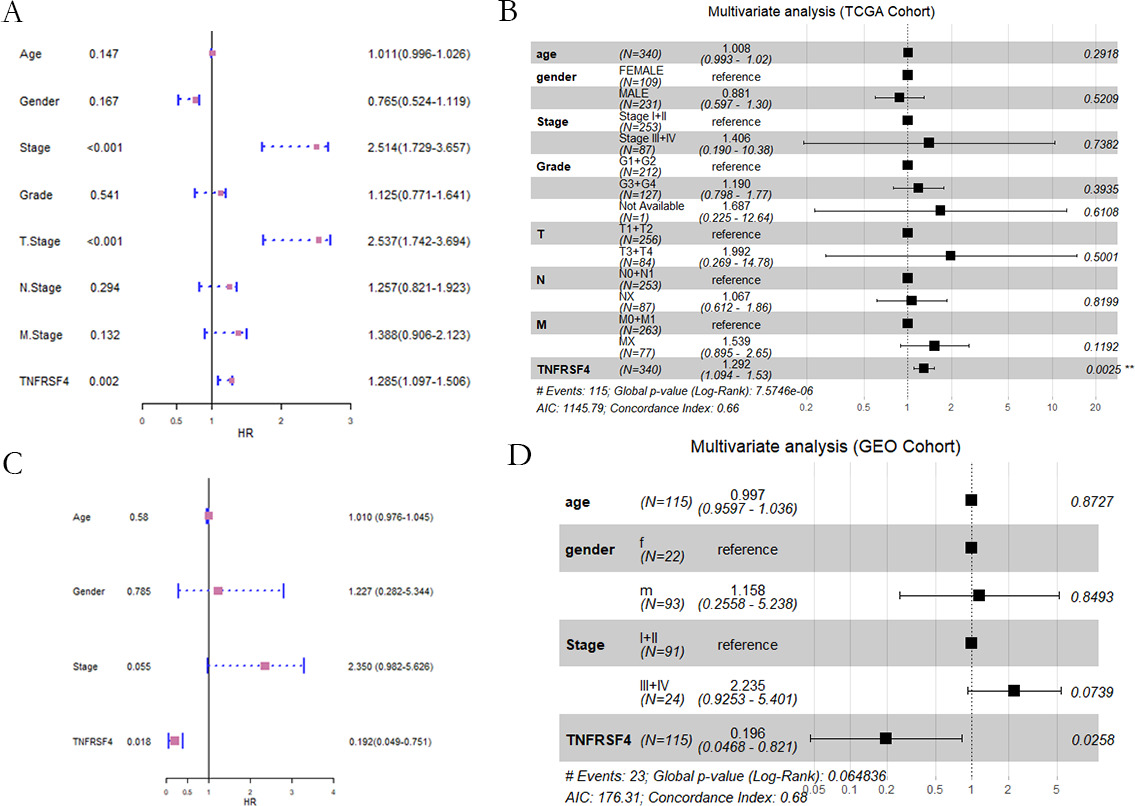
Subsequently, Univariate Cox regression analysis was implemented to evaluate the prognostic value of TNFRSF4 expression, age, gender, tumor grade and disease stage (Fig. 4A). Then subject to multivariate Cox regression, it was found TNFRSF4 expression was an independent prognostic biomarker (Fig. 4B). The conclusion was further confirmed by the GEO dataset (Fig. 4C and D), demonstrating the TNFRSF4 could be an independent biomarker for HCC.
Comparisons of 22 important immune fractions between high/low TNFRSF4 groups. (A) Differences in the immune invasion of 22 cases with high and low TNFRSF4 expression; (B) In TCGA-LIHC, Violin chart showing the difference in the distribution of 22 immune cells in the high TNFRSF4 expression (red) and the low TNFRSF4 expression (blue). (C) In GEO-LIHC, Violin chart showing the difference in the distribution of 22 immune cells in the high TNFRSF4 expression (red) and the low TNFRSF4 expression (blue).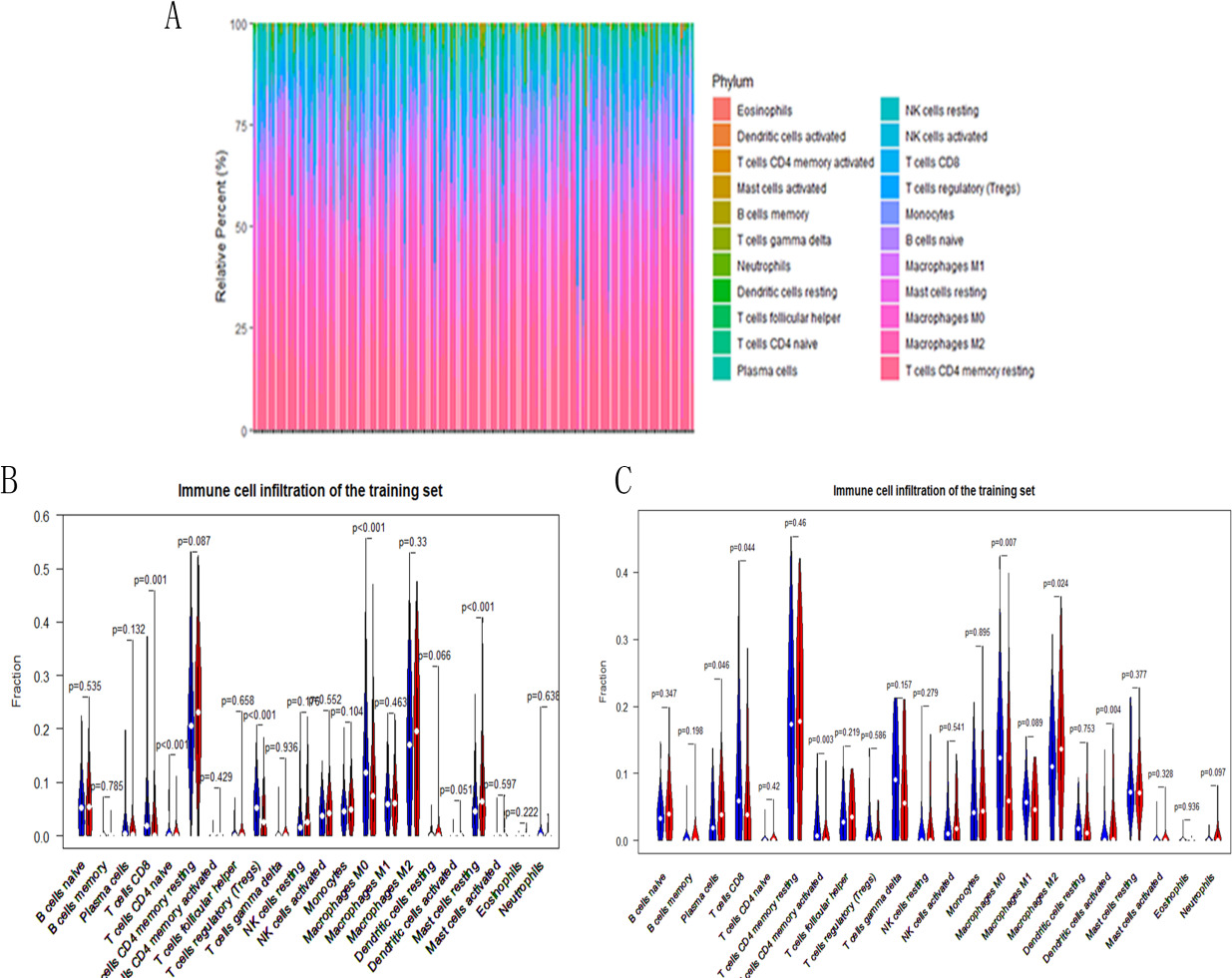
We used the CIBERSORT algorithm to investigate the immune infiltration level involved in 22 resistant cell types for HCC patients. There was a significant difference in the immune invasion of 22 cases with high and low TNFRSF4 expression (Fig. 5A). Also, we assess the correlation of 22 anti-tumor cell groups. We compare the effects of TNFRSF4 expression in the distribution of immune infiltration in both the TCGA and GEO datasets after removing samples with
Screening the highly relevant modules based on WGCNA. (A) The volcano map of differential genes in TNFRSF4 expression group; (B) Unscaled fitting index of soft threshold power (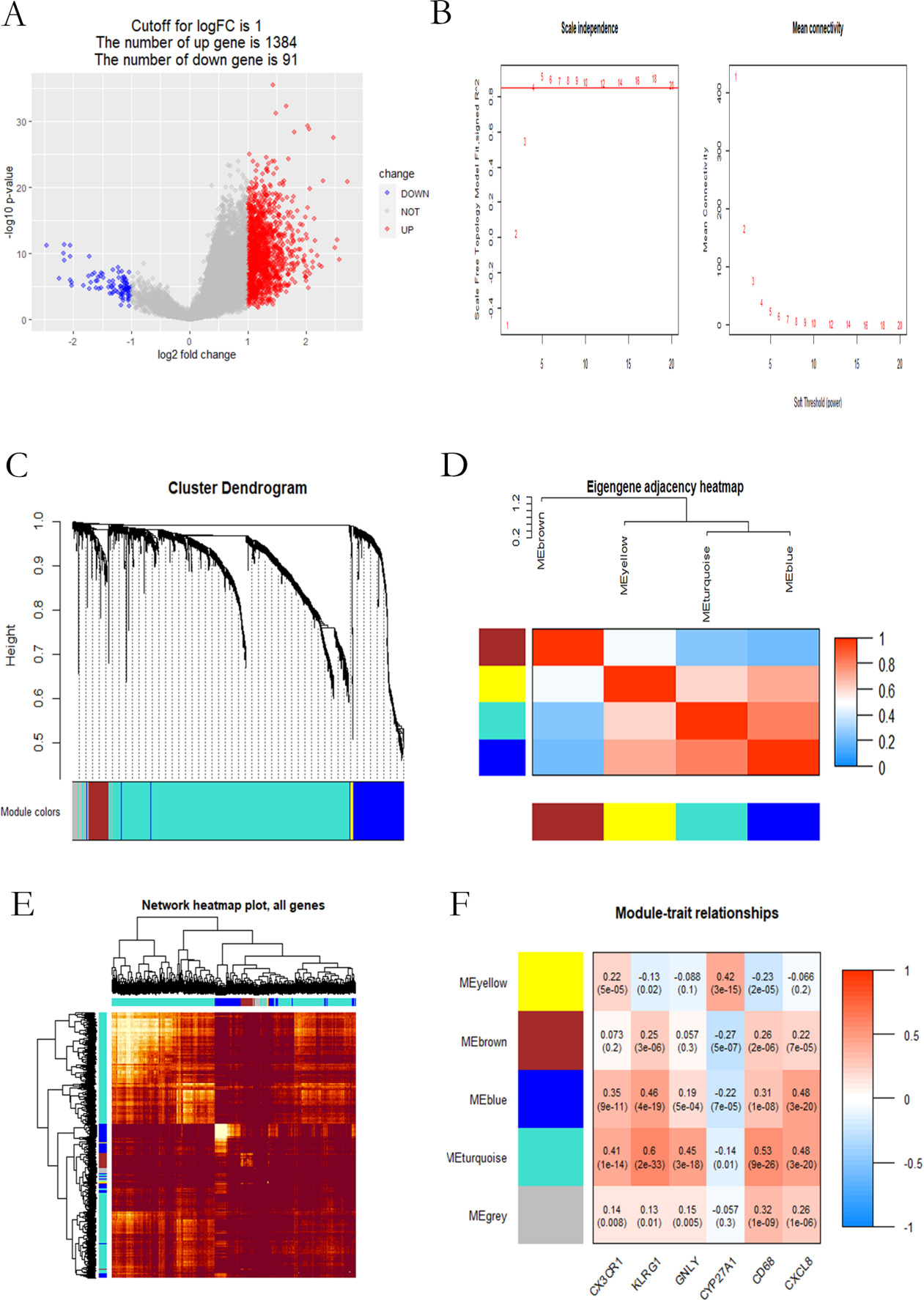
In Fig. 6A, there were 1475 differential genes (referring to appendix4) of high and low TNFRSF4 expression, including 1384 up-regulated genes (colored in red) and 91 down-regulated genes (colored in blue). Then, these 1475 differential genes were subject to WGCNA analysis to explore the relation of the TNFRSF4 expression with immune markers. Genes as CX3CR1, KLRG1 and GNLY representing T CD8 cell markers, and CYP27A1, CD68 and CXLC8 denoting macrophage M0 markers, were selected out as traits for co-expression analysis [17, 18, 19, 20, 21, 22, 23]. Average linkage clustering identified 5 gene co-expression modules, with each module separately containing 42, 81, 277, 476, and 579 genes colored by gray, yellow, brown, blue, and cyan (Fig. 6C). Correlation analysis between the modules and immune markers revealed that TNFRSF4 expression level was greatly related to the turquoise module (Fig. 6F).
Identification of immune-related genes. (A–B) LASSO analysis determines the number of factors. (C) Analysis of prognostic risk scoring model. (D) Kaplan-Meier survival curve in the TCGA cohort. (E) 1, 3 and 5 year ROC curve.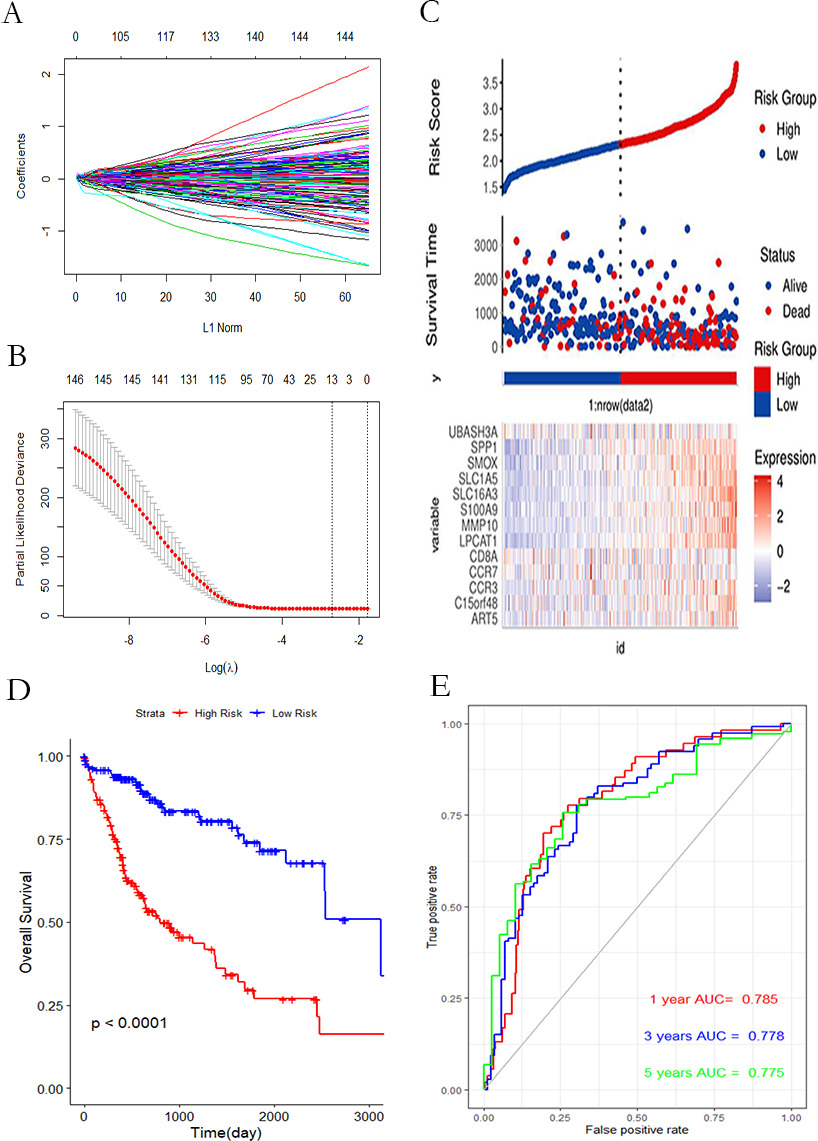
Included in the turquoise module, 579 IRGs were related to the survival outcome, and then subject to LASSO analysis, ultimately 13 IRGs were determined as the 13-gene signature, referring to ART5, C15orf48, CCR3, CCR7, CD8A, LPCAT1, MMP10, S100A9, SLC16A3, SLC1A5, SMOX, SPP1, UBASH3A (Fig. 7A and B). In Fig. 7C, HCC patients were divided into high/low risk group according to the risk scoring model. Subsequently, Kaplan-Meier analysis was implemented in Fig. 7D, showing that the overall survival rate of the high-risk group was lower than that of the low-risk group. In the time-dependent ROC (tROC) curves, it was demonstrated that the classifier possessed a strong predictive ability, with the AUCs of 1, 3, and 5 years of 0.785, 0.778, and 0.775, respectively (Fig. 7E).
Gene mutation waterfall chart. (A–B) The waterfall chart of gene mutation between high-risk and low-risk groups of signature. (C–D) Cascade of gene mutation between high and low TNFRSF4 expression groups.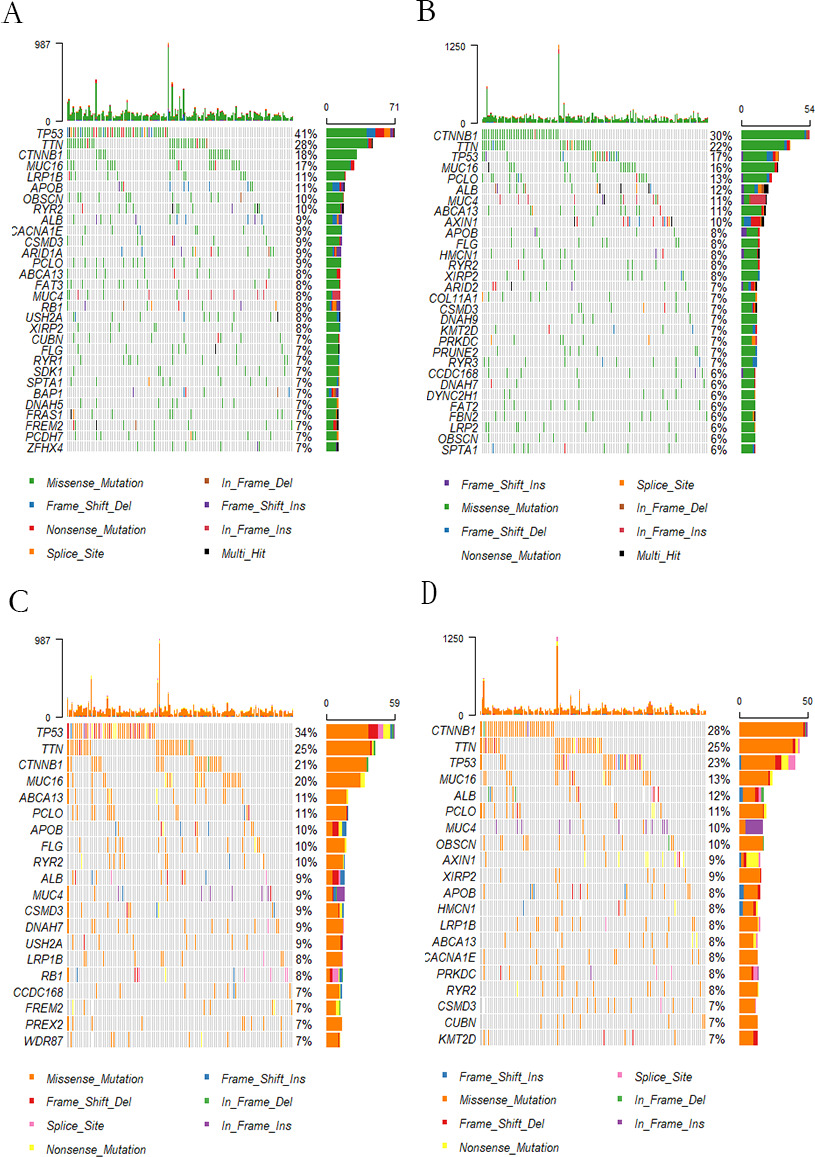
In this step, mutation landscapes were depicted for 353 samples, including 174 samples of high TNFRSF4 expression and high risk score (referring to Fig. 8A and C), and 179 samples of low TNFRSF4 expression and low risk score (referring to Fig. 8B and D). By exploring the correlation of TNFRSF4 expression and gene mutations for the TCGA dataset, it was found that the gene mutation frequency in TNFRSF4 high expression group was higher than that of the low. Moreover, the situation was the same in high and low-risk groups. Amonst, the gene mutation frequency of TP53, TIN and CTNNB1 were the highest, indicating that the gene mutation frequency was influenced by high/low TNFRSF4 expression and highlow risk score. Conclusively, TNFRSF4 could be served as a biomarker for HCC classification.
TNFRSF4 uses T cells to conduct costimulatory signals and enhance cell proliferation, survival, and migration [24]. The prognostic role of the TNFRSF4 remains to be explored in liver cancer. The study directed an in-depth analysis of the expression, mutation, and prognostic value of TNFRSF4 in HCC. It was believed the TNFRSF4 expression in HCC was higher than that in normal liver tissue, consistent with the conclusion of previous studies [25]. Upregulated in clinical HCC samples, TNFRSF4 may serve as a potential prognostic biomarker.
TNFRSF4, a T cell costimulatory molecule, is associated with activation of inflammatory immunity in HCC [26]. The activation of TNFRSF4 promoted the inflation of activated T cells and memory T cells to exacerbate autoimmune diseases [27] . Besides, TNFRSF4 imaging probe predicted early local immunotherapy for HCC [28]. Piconese et al. found that TNFRSF4 can induce the proliferation of regulatory T cells in HCC [29]. However, few efforts were made on the prognostic gene signature associated with TNFRSF4.The study dedicated to exploring the relationship of TNFRSF4 expression with immune infiltration and frequency of gene mutation.
The effects of TNFRSF4 on the immune process were reported to be significant [30, 31], and the interaction between them could reduce the immune suppression of regulatory T cells, thus enhancing the immune responses to definite antigens [32, 33]. Moreover, we also found the TNFRSF4 expression were in correlation with different immune markers and immune infiltration. Therefore, it was inferred the TNFRSF4 expression may have a potential impact on tumor immunity.
In this manuscript, it was found the TNFRSF4 expression was greatly different in the immune infiltration level of T CD8 cells and macrophages M0. By previous investigations, the average density of T CD8 cells in liver cancer was indicated to be significantly lower than that in normal liver tissue, thereby served as a benchmark for evaluating tumor immune checkpoints [34, 35]. T CD8 cells differentiate into an exhaustion state in HCC to form a strong barrier against powerful anti-tumor immunity [36]. Macrophage M0 was believed to be a poor prognostic factor of HCC tumor immune microenvironment [37]. Overexpression of immune-related genes C15orf48 [38, 39], CCR3 [40, 41], CCR7 [42], MMP10 [43], S100A9 [44], SLC1A5 [45] and SMOX [46] are involved in the progression of many malignancies play a key role. The expression levels of CD8A [47] and SLC16A3 [48] were positively correlated with high immune infiltration in multiple tumor types. LPCAT1 [49] and SPP1 [49] may promote HCC progression by accelerating cell growth, migration and metastasis. However, the biological roles of ART5 and UBASH3A in HCC remain unclear. Integratively, the relationship between different immune cell markers and the TNFRSF4 expression level suggested the TNFRSF4 was important to regulating the immune micro-environment of HCC.
Following, HCC patients were divided into high and low-risk groups to further explore the prognostic model, displaying TP53 was greatly up-regulated in both the high TNFRSF4 expression group and high-risk group. TP53, as a tumor suppressor, its mutation accounted for a relatively large proportion of HCC [51]. For the low TNFRSF4 expression group and the low-risk group, CTNNB1 were significantly up-regulated. CTNNB1 mutations can cause a high enrichment of DNA methylation at the CpG sites in the HCC [52]. Accordingly, TNFRSF4 expression was believed to not only affect the immune microenviroment, but also the gene mutation in liver cancer.
Conclusions
Conclusively, TNFRSF4 expression was suggested to be an independent indicator for clinical prognosis of HCC, and high expression of TNFRSF4 is associated with immune infiltration and frequency of gene mutation. However, the potentiality of TNFRSF4 in HCC still awaits to be further experimentally validated.
Ethical statement
Our study did not require an ethical board approval because it did not contain human or animal trials.
Consent for publication
Not applicable.
Availability of data and materials
All data generated or analyzed during this study are included in this published article.
Authors contributions
Conception: Huan Hu.
Interpretation or analysis of data: Huan Ding.
Preparation of the manuscript: Di Wang.
Revision for important intellectual content: Qingjia Chi.
Supervision: Feifei Tian, Han Zhao.
Footnotes
Acknowledgments
We appreciate the generosity of TCGA and GEO database for sharing hepatocellular carcinomadata.
Conflict of interest
The authors have declared that no competing interests exist.
