Abstract
Pancreatic cancer is an aggressive and lethal cancer with the highest mortality rate. Hence, the development of new targeting and innovative treatment strategies is needed. Recent studies reported that the histone chaperone anti-silencing function 1B (ASF1B) can be used as a diagnosis and prognosis cancer biomarker. However, functional studies of ASF1B in pancreatic cancer have not been performed. This study compared expression levels of ASF1B in pancreatic cancer specimens with those of normal tissues using publicly available online databases. We found that ASF1B was commonly overexpressed in pancreatic cancer specimens, which is associated with poor prognosis. ASF1B downregulation in pancreatic cancer cells reduced their colony formation, proliferation, migration, and invasion abilities, and inhibited MMP9 activity. Furthermore, ASF1B expression downregulation increased cell cycle S-phase arrest and DNA damage though activation of the checkpoint kinases Chk1 and Chk2 pathways. Additionally, increased caspase (caspases-3 and -9) activation and PARP cleavage led to enhanced caspase-dependent apoptosis and improved cisplatin sensitivity. Collectively, our results indicate that ASF1B may serve as a potential biomarker of pancreatic cancer and a novel therapeutic target.

Introduction
Pancreatic cancer is the seventh leading cause of cancer-associated death in both men and women worldwide. Pancreatic cancer is an extremely malignant tumor with a poor prognosis and almost equal incidence and mortality [1]. A recent European study predicted pancreatic cancer to be the third-leading cause of cancer-related deaths by 2025 [2]. Despite ongoing research into cancer screening and treatment, the five-year survival estimates of pancreatic cancer patients is still less than 10% [3]. Therefore, discovering biomarkers for the prediction of pancreatic cancer progression and potential targets for treatment is urgent.
Nucleosomes are sections of DNA wrapped around histone proteins and are the basic organizational unit of the genome. They act as barriers to DNA-related processes such as gene transcription, DNA repair, and DNA replication, which require the disassembly and assembly of nucleosomes [4, 5]. It has been reported that histone alterations are related to tumor progression and development [6]. Histone chaperones are an heterogeneous group of proteins that perform a variety of functions, such as storing, transporting, modifying, or depositing histones in DNA [6, 7]. ASF1 (anti-silencing function 1) is a highly conserved chaperone that participates in the assembly or disassembly of H3 and H4 heterodimers [8, 9, 10, 11]. Higher eukaryotes possess two isoforms of ASF1, ASF1A and ASF1B, which only differ in the C-terminal domain (30% of the sequence) [12, 13]. Both of these isoforms have been reported to be altered in various cancers. ASF1A is upregulated in human malignancies such as liver, lung, prostate or stomach cancer, since it is required for the indefinite proliferation of cancer cells [14, 15]. Silencing or removal of ASF1A causes DNA damage and cellular senescence, and higher sensitivity to doxorubicin [16]. ASF1B is overexpressed in breast and lung cancer patients, where it acts as a marker of poor prognosis [15, 17]. On the contrary, ASF1A is not dysregulated in lung cancer or associated with the survival of patients [17]. In cervical cancer, ASF1B inhibition induces cell cycle arrest and apoptosis, whereas ASF1B overexpression enhances cancer cell proliferation [18]. These findings suggest the potential use of ASF1A and ASF1B as biomarkers and attractive biological targets for innovative anticancer therapies.
Herein, we explored whether high expression of ASF1B is associated with poor prognosis for overall survival. It was found that downregulation of ASF1B significantly reduced colony formation, proliferation, migration, and invasion of pancreatic cancer cells and increased the sensitivity to cisplatin, indicating the potential value of ASF1B as a prognostic indicator and treatment target in pancreatic cancer.
Results
Elevated expression of ASF1B in pancreatic cancer correlates with poor survival outcome
We first identified clinical pancreatic cancer and normal specimens from The Cancer Genome Atlas (TCGA) (
ASF1A and ASF1B expression patterns and association with survival outcome in pancreatic cancer patients. A, Expression levels of ASF1A and ASF1B mRNA in tumor (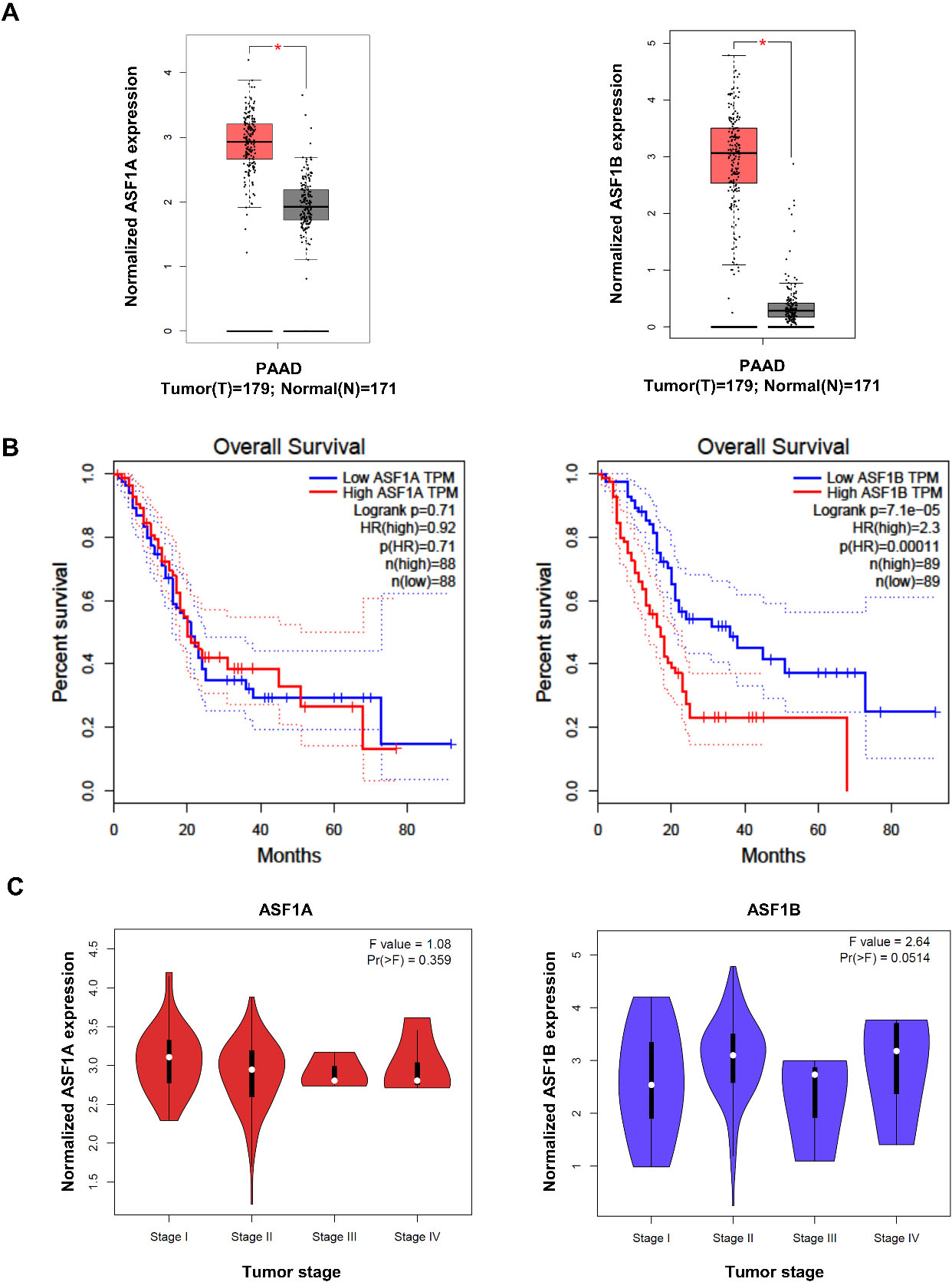
ASF1B-downregulated pancreatic cancer cells show reduced colony formation and cell proliferation capacity. A, Analysis of ASF1B protein expression levels in HPDE and pancreatic cancer cell lines (AsPC-1, PANC-1, MIA PaCa-2, and Capan-1). The ASF1B/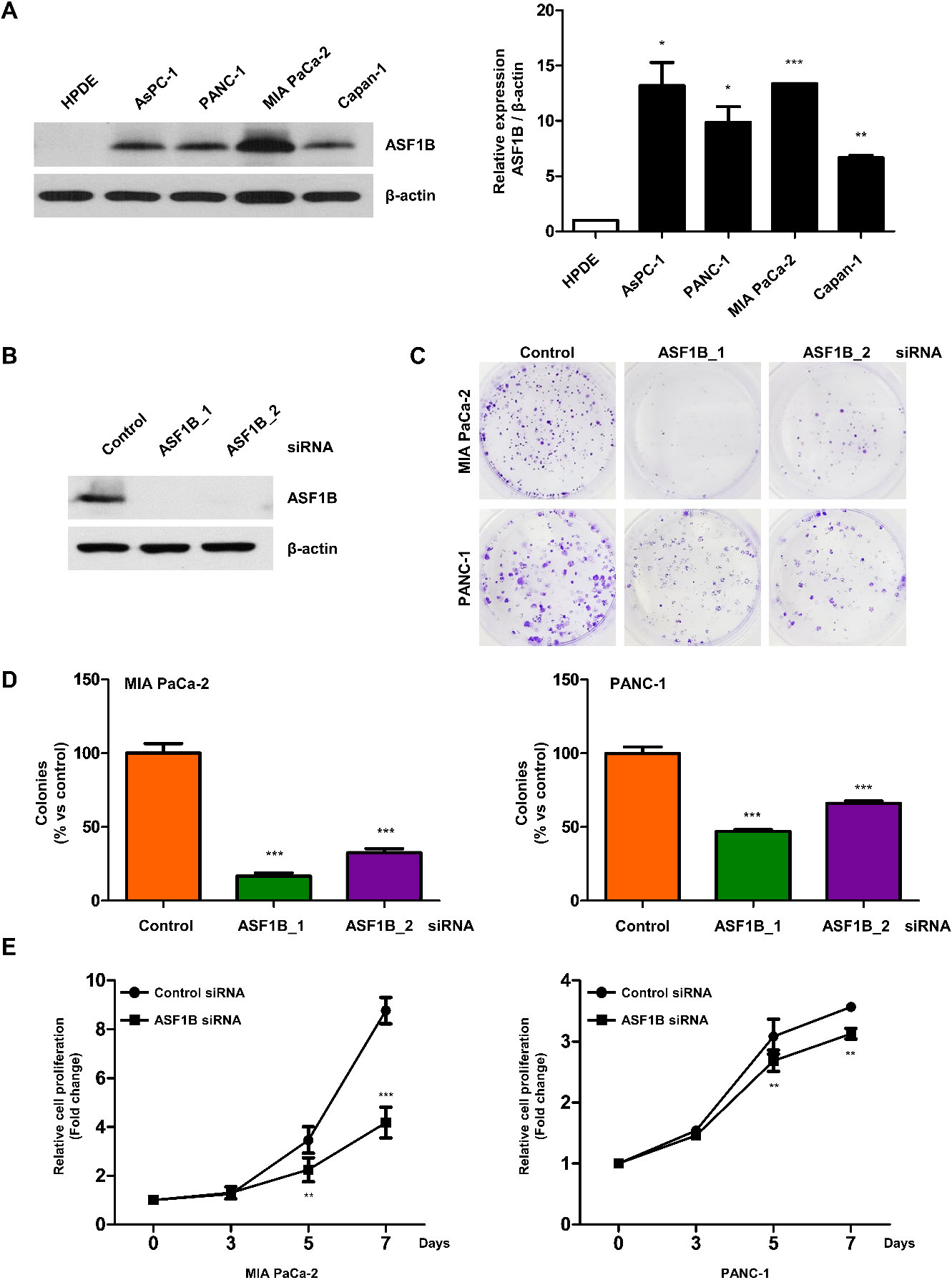
Downregulation of ASF1B in pancreatic cancer cells is observed to reduce migration, invasion, and wound-healing abilities. A–B, Migration and invasion assay of MIA PaCa-2 cells transfected with control siRNA or ASF1B siRNA, and treated with rMMP9 (recombinant MMP9, 50 ng/ml) for 48 hours. Values represent means 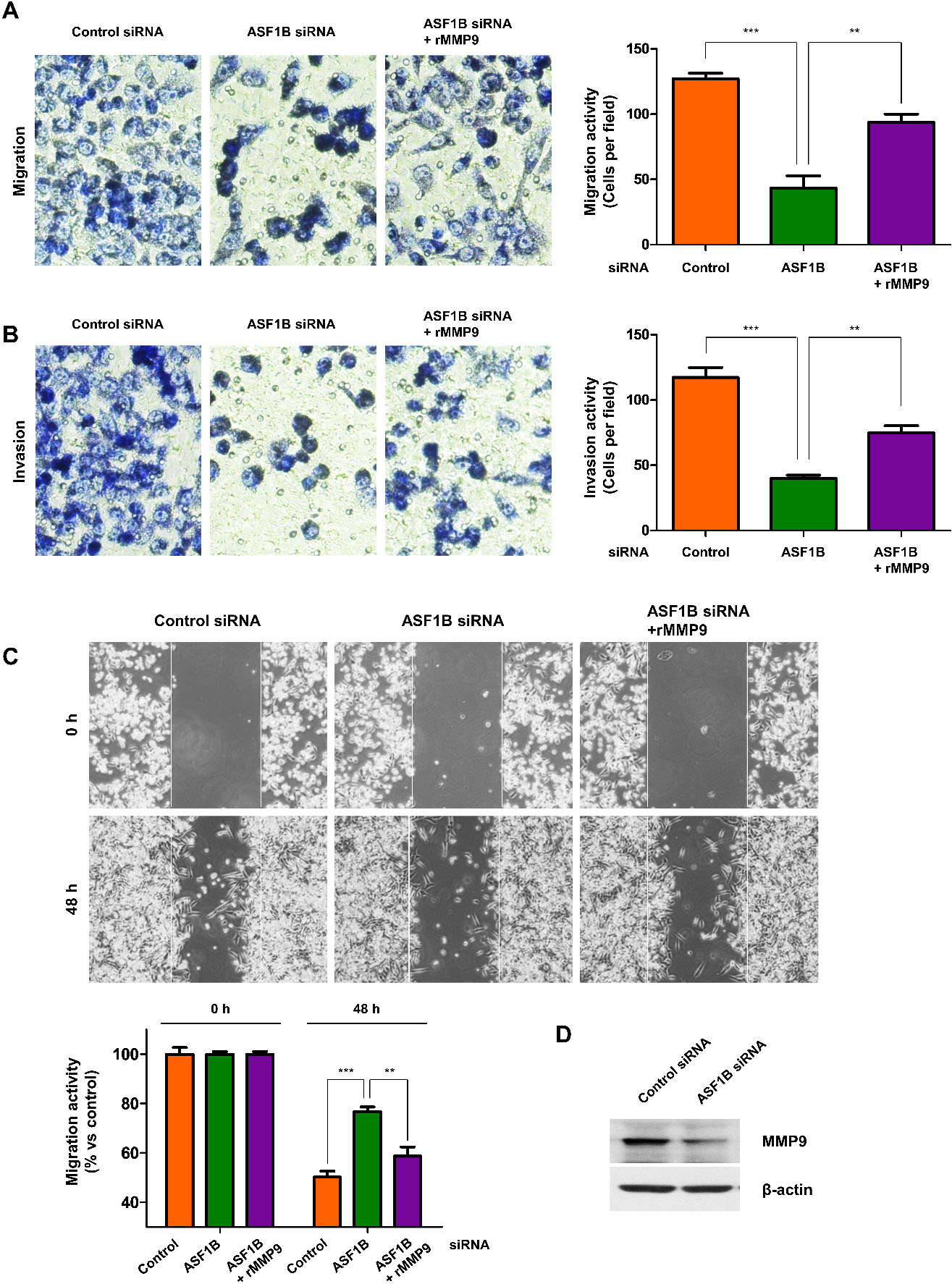
In addition, 50 genes with co-expression pattern similar to ASF1B mRNA in 177 pancreatic cancer patients were selected from the co-expression network analysis from the publicly accessible TCGA database at the cBioPortal portal (Supplementary Fig. 1A). To further understand the functions and pathways of these genes, including ASF1B, we performed GO and KEGG analysis using the web-based DAVID tools (2021 Update) [19, 20]. As a result, GO terms classified into three types: cell components (CC), biological process (BP), and molecular function (MF), and analyzed by arranging them into the 33 categories with the highest number of genes (Supplementary Fig. 1B). The KEGG pathway analysis resulted in six categories (Supplementary Fig. 1C).
We also considered the mRNA expression levels of ASF1B in various tumor stages of pancreatic cancer using the GEPIA data set. The expression levels of ASF1B varied at different tumor stages and were associated with the tumor stage. However, this was not the case for ASF1A (Fig. 1C). Based on these results, we hypothesized that ASF1B, but not ASF1A, was related to the growth of pancreatic cancer cells.
We compared protein expression levels of ASF1B in normal human pancreatic duct epithelial (HPDE) cells and various pancreatic cancer cell lines, AsPC-1, PANC-1, MIA PaCa-2, and Capan-1 cells using Western blotting. AF1B protein levels were higher in all pancreatic cancer cell lines compared to those in HPDE cells (Fig. 2A). To further investigate the role of ASF1B in pancreatic cancer, we downregulated it and evaluated effects on pancreatic cancer progression using several cellular assays. To achieve the silencing of ASF1B, we used two distinct siRNAs, both of which inhibited the expression of the ASF1B protein and found that both ASF1B siRNAs inhibited the protein expression of ASF1B (Fig. 2B). Subsequent colony formation revealed that downregulation of ASF1B significantly reduced clonogenic survival in two pancreatic cancer cell lines (both MIA PaCa-2 and PANC-1 cells). In particular, ASF1B_1 siRNA was more effective in suppressing clonogenic survival than ASF1B_2 siRNA (Fig. 2C and D). Therefore, we chose to use ASF1B_1 siRNA for subsequent experiments. Furthermore, after 5–7 days of ASF1B downregulation in MIA-PaCa-2 and PANC-1 cells, we monitored cell proliferation and viability using the MTT assay. The cell proliferation of ASF1B-downregulated cells was greatly reduced in both pancreatic cancer cell lines (Fig. 2E). These results indicate that ASF1B plays a central role in the ability to form colonies and proliferate of pancreatic cancer cells.
Downregulation of ASF1B expression inhibits migration and invasiveness of pancreatic cancer cells
To investigate the inhibitory influence of ASF1B on the cell phenotype, we transiently transfected pancreatic cancer cells with ASF1B siRNA and control siRNA and analyzed the effect of downregulation of ASF1B in pancreatic cancer cells on the migratory capacity of cells. For this purpose, cell growth was repressed by culturing in low fetal bovine serum (FBS) conditions, and the migrating ability of cells was measured using a transwell assay. Downregulation of ASF1B greatly decreased the number of migratory cells compared to those observed in the control group (Fig. 3A). We evaluated the influence of ASF1B downregulation on cell invasion ability, where the transwell inserts were Matrigel-coated wells. Consistent with the results of the cell migration assay, the invasion capacity of ASF1B-downregulated pancreatic cancer cells was significantly lower than that in the control cells (Fig. 3B). Next, downregulation of ASF1B in pancreatic cancer cells showed to delay wound-healing capacity, effect that was especially prominent after 48 hours (Fig. 3C). Also, ASF1B-downregulated pancreatic cancer cells presented decreased expression levels of matrix metalloproteinase9 (MMP9) (Fig. 3D).
Finally, to determine whether MMP9 is responsible for cell migration, invasion, and wound healing ability, ASF1B-downregulated pancreatic cancer cells were treated with recombinant MMP9 protein (rMMP9). As shown in Fig. 3A–C, rMMP9 rescued the inhibitory effects caused by downregulation of ASF1B on migration, invasion, and wound healing ability in pancreatic cancer cells (Fig. 3A–C). Altogether, these results suggest that ASF1B plays an important role in migration and invasiveness by MMP9 activation in pancreatic cancer cells.
Downregulation of ASFB1 in pancreatic cancer cells induces S-phase cell cycle arrest and increases DNA damage
To determine the influence of ASF1B on cell cycle progression of the pancreatic cancer cells, we evaluated the cellular population percentage at different stages of the cell cycle (G1, S and G2/M) using flow cytometry analysis. Downregulation of ASF1B in pancreatic cancer cells resulted in no significant differences in cell cycle at 48 hours post-downregulation. However, at 72 hours, ASF1B-downregulated pancreatic cancer cells showed an increased fraction of cells in S-phase compared to the control group cells, and the G1 population decreased in a concentration-dependent manner in ASF1B-downregulated pancreatic cancer cells (Fig. 4A and B). To decipher the molecular mechanism associated with the observed cell cycle dysregulation, we measured expression levels of cyclin A2, an S phase regulator, using Western blotting. The level of cyclin A2 was significantly increased in ASF1B-downregulated pancreatic cancer cells 24 hours post-transfection (Fig. 4C).
In addition, to measure the effect of ASF1B downregulation on DNA damage, we analyzed expression levels of phosphorylated H2AX (
S-phase arrest and increased DNA damage were observed in ASF1B-downregulated pancreatic cancer cells. A, Flow cytometry analysis of MIA PaCa-2 cells transfection with control or ASF1B siRNA after 48 and 72 hours. The flow cytometry plots and data are representative of at least three independent experiments. B, Percentages of cells in G1 (blue), S (orange), and G2/M (green) phases. C, Western blotting to determine the levels of ASF1B, cyclin A2, 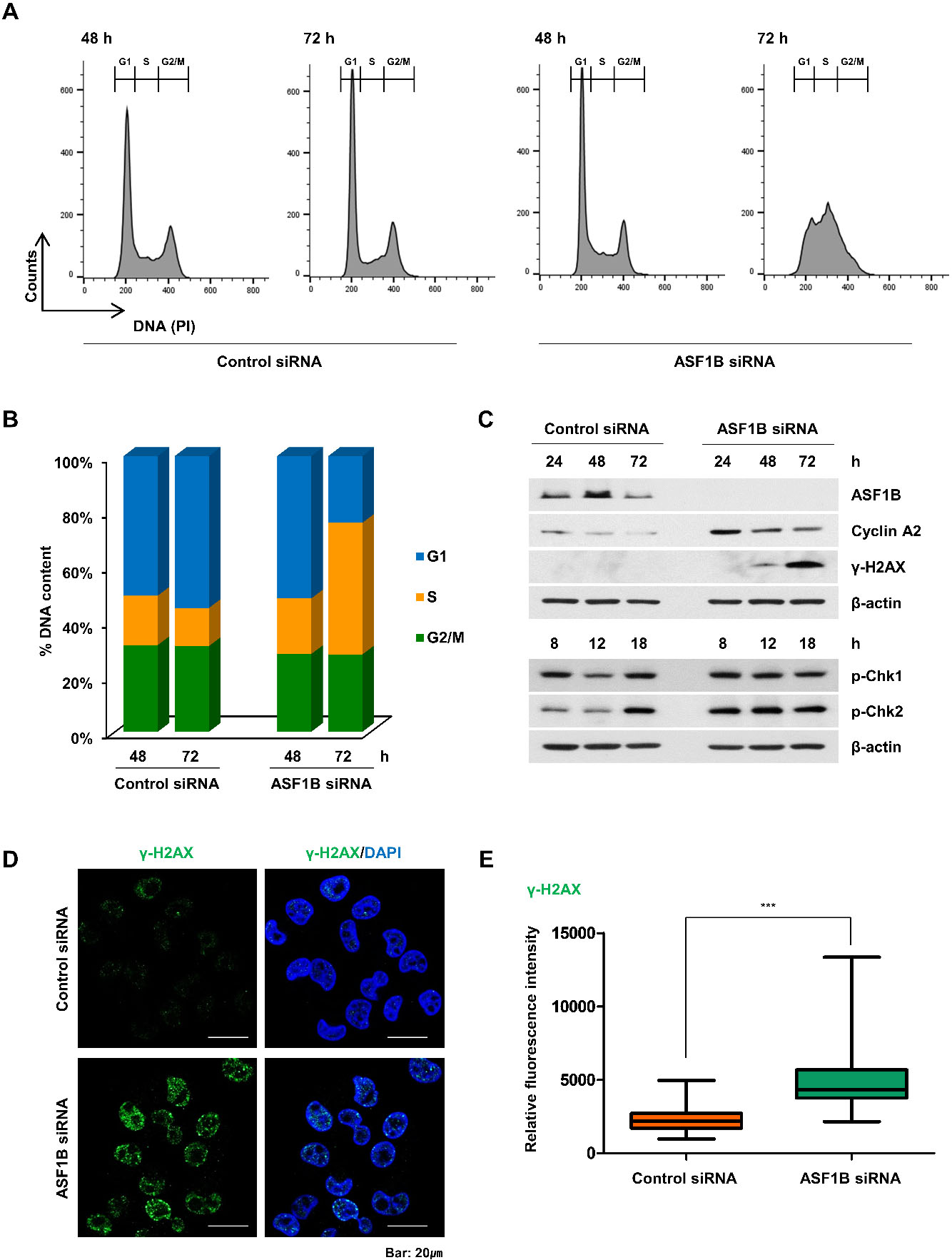
ASF1B-downregulated pancreatic cancer cells had increased sensitivity to cisplatin. A, Flow cytometry analysis of MIA PaCa-2 cells transfection with control or ASF1B siRNA, treated with cisplatin (1 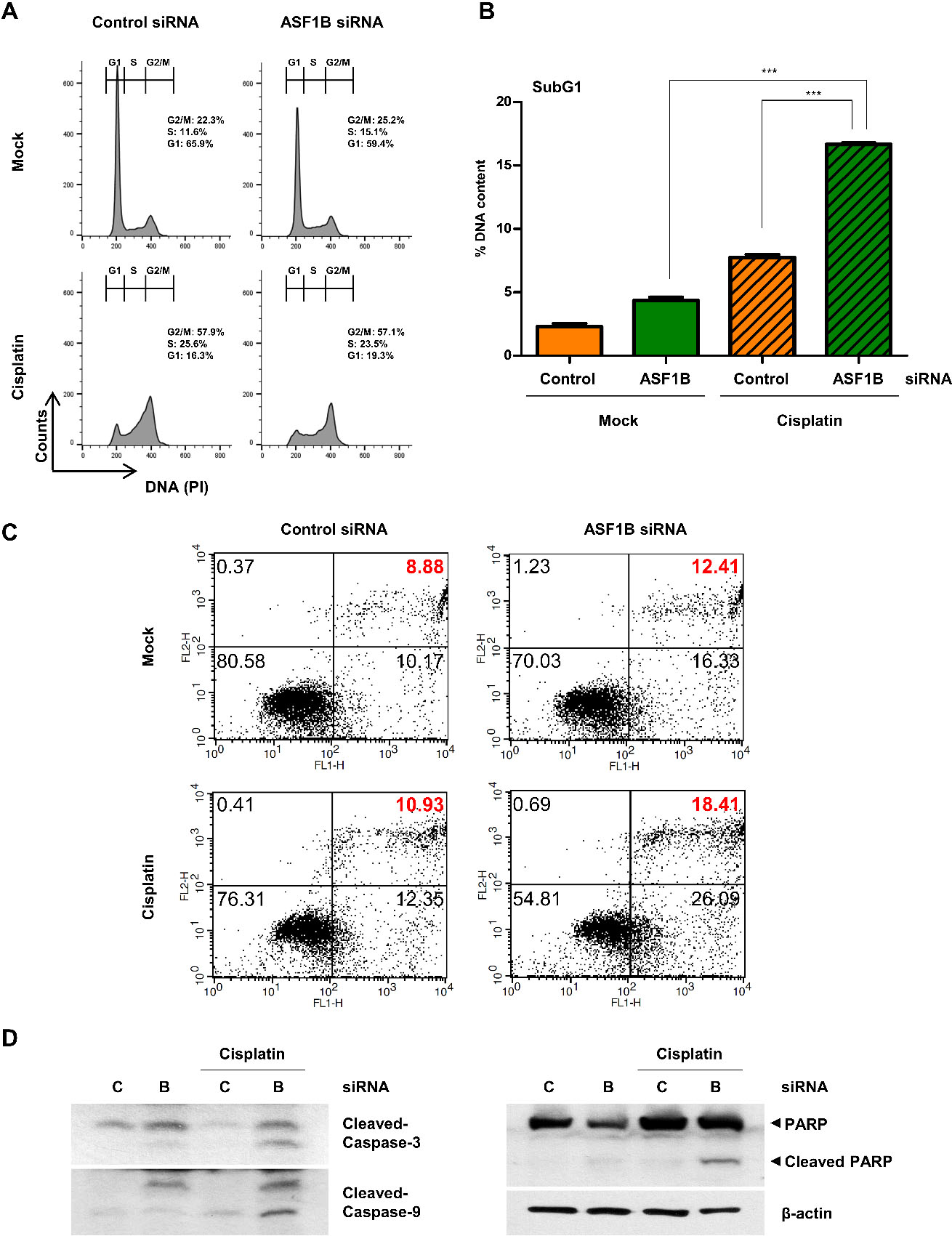
To determine whether downregulation of ASF1B expression could reverse the resistance to cisplatin, a platinum-based anticancer drug in pancreatic cancer cells, we used MIA PaCa-2 that was reported to be resistant to platinum drugs [21]. After downregulation of ASF1B, cells were treated with cisplatin, and consequences in the cell cycle distribution and apoptotic cells were assessed via flow cytometry analysis. Cisplatin-treated ASF1B-downregulated cells presented an increase in subG1 (apoptotic fraction) compared to that of cells without cisplatin treatment, whereas there was no significant difference in cell cycle phase distribution (Fig. 5A and B). To further determine whether ASF1B-downregulated pancreatic cancer cells died through apoptosis or necrosis, cells were dual-stained with annexin V and propidium iodide (PI) and phenotypic changes in dying cells were assessed. The proportion of late-apoptotic cells (annexin V-positive and annexin V/PI-positive) was greatly higher in cisplatin-treated ASF1B-downregulated pancreatic cancer cells compared to cells not treated with cisplatin (Fig. 5C, upper right quadrant). Finally, Western blotting revealed that the cleaved forms of caspase-3, caspase-9, and poly ADP ribose polymerase (PARP) were highly expressed in cisplatin-treated ASF1B-downregulated pancreatic cancer cells compared to cells not treated with cisplatin (Fig. 5D). These results suggested that ASF1B confers cellular resistance to platinum-based drugs, including cisplatin.
Discussion
It was found that ASF1B is upregulated in pancreatic cancer tissues and cell lines, and is associated with poor prognosis. Functional studies using several pancreatic cancer cell lines showed that downregulation of ASF1B reduced cell proliferation, migration, and invasiveness of pancreatic cancer cells. Moreover, it induced cell cycle arrest and DNA damage through the Chk1 and Chk2 pathway, and enhanced the sensitivity to cisplatin of cancer cells. These results imply that ASF1B contributes to pancreatic cell progression, and support its use as a prognosis biomarker.
Our results are consistent with previous studies that have evaluated the importance of ASF1B in cancer. Our current study evaluated the expression level of ASF1B in pancreatic cancer tissues and cells and found that high expression of ASF1B was associated with poor prognosis. We subsequently investigated its function in the growth, metastasis, and migration of pancreatic cancer by inducing downregulation of ASF1B expression. We observed that downregulation of ASF1B expression reduced the cell proliferation and colony formation ability of pancreatic cancer cells and demonstrated inhibition of migration and invasion. A recent study reported that ASF1B is highly expressed in multiple cancers, including pancreatic cancer and is associated with poor prognosis in patients [22, 23]. These results were consistent with those reported in studies on cervical cancer, lung adenocarcinoma, and myeloma [17, 18, 24, 25]. In cervical cancer, ASF1B knockdown is associated with cell proliferation and clonogenic survival, and ASF1B overexpression promotes cell proliferation [18]. Furthermore, in breast cancer and lung adenocarcinoma cells, it has been reported that overexpression of ASF1B promotes proliferation and metastasis, suggesting an association with poor prognosis [15, 17]. Additionally, in several cancers such as breast cancer, lung cancer, and esophageal carcinoma, the expression of ASF1B is higher in tumors at stage IV than in tumors at stage I, suggesting that the expression of ASF1B is related to the stage of tumor progression [22]. In the present study, we show that the expression of ASF1B was higher as the stage progressed, highlighting the need of further research into this issue. However, in the case of ASF1A, the expression was higher in pancreatic cancer than that in normal tissues, but it was not related to overall survival or to tumor stage. In line with these results, a previous study also reported that ASF1A was not significantly related to progression-free survival and overall survival in lung adenocarcinoma [22]. These discrepant results suggest that ASF1A and ASF1B do not perform the same cellular functions.
Using different molecular analyses, we could observe that downregulation of ASF1B caused S-phase cell cycle delay and increased DNA damage response through the activation of Chk1 and Chk2 signaling pathway. In cervical cancer and prostate cancer, knockdown of ASF1B causes G1 or G2/S-phase cell cycle arrest and cell death owing to its action on Bcl-2, Bax and caspase-3 [18, 26]. ASF1B forms a stable complex with CDK9, a member of the cyclin-dependent kinase family, and thereby promotes cell proliferation [18]. These results suggest the potential use of ASF1B as a novel proliferation marker in pancreatic cancer, aiding diagnosis and prognosis prediction, and potentially supporting the discovery of new drugs in pancreatic cancer. However, further validation using tissue samples obtained directly from pancreatic cancer patients is still necessary to confirm the validity of the results from this study.
Cisplatin is one of the most commonly used anticancer drugs and DNA-damaging agents since it was first discovered as an inhibitor of bacterial cell growth. This platinum drug is a very effective treatment of patients with a wide variety of solid malignancies, including lung, ovarian, head and neck, and gastric cancers [27, 28, 29]. The mode of action of cisplatin relies on the activation of DNA damage responses and the creation of DNA lesions following induction of mitochondrial apoptosis [30]. However, it is known that pancreatic cancer cells develop cisplatin resistance associated with mechanisms involving inactivation/transport of drug, DNA damage response, DNA damage-specific repair, and regulation of apoptosis [31]. In this study, it was observed that ASF1B inhibition in cisplatin-resistant pancreatic cancer cells resulted in increased apoptosis through the increase of caspase-3/9 activation and PARP cleavage. This suggests that the high expression level of ASF1B in pancreatic cancer cells may have contributed to the cisplatin resistance development. Further in vivo analyses are required to demonstrate the effect of high ASF1B expression on anti-cancer drug resistance in pancreatic cancer.
In conclusion, our study established that interfering with ASF1B can significantly inhibit the growth of pancreatic cancer cells by regulating cell growth inhibition, migration, and metastasis, as well as increase the death of cancer cells and its sensitivity to cisplatin. Our results reveal a potential target for novel pancreatic cancer treatment strategies, as well as strategies to reverse the acquired cisplatin-resistance of cancer cells, a pressing topic in ongoing cancer research.
Materials and methods
Cell culture and transfection
The human pancreatic cancer cell lines MIA PaCa-2 (ACTT number: CRL-1420) and PANC-1 (ACTT number: CRL-1469) were obtained from the American Type Culture Collection (ATCC, Manassas, VA, USA). Cells were cultured in Dulbecco’s Modified Eagle Medium (DMEM) contained high glucose. AsPC-1 and Capan-1 were cultured in Roswell Park Memorial Institute (RPMI) medium. Noncancerous immortalized HPDE cells, obtained from Joo Kyung Park, MD (Samsung Medical Center, Seoul, South Korea), were grown in defined Keratinocyte-Serum Free Medium (K-SFM). Cell culture mediums were used including 100 U/ml penicillin, 100
Cell proliferation and colony formation assay
The cells were transfected with ASF1B siRNA or control siRNA, after adhesion for 24 hours. For colony formation assays, the MIA PaCa-2 and PANC-1 cells were seeded into 6-well plates at a density of 500–1000 cells/well and incubated for 14 days. The colonies were fixed in 100% methanol, stained with 10% crystal violet, and counted. For cell proliferation assays, MIA PaCa-2 and PANC-1 cells were seeded into 12-well plates at a density of 1
Cell migration, invasion, and wound-healing assays
Abilities of migration and invasion under downregulation of ASF1B expression were evaluated using Trans-well plates (Corning, Corning, NY, USA). For the cell migration assay, 5
The motility of ASF1B siRNA and control siRNA transfected cells was examined by a wound-healing assay. The cells were seeded in 6-well plates at 5
Western blot and antibodies
Whole cells were washed once with cold 1
Fluorescence-assisted cell sorting (FACS) analysis
For cell cycle and DNA content measurements, whole cells were collected, and washed in 1
Apoptosis was also analyzed by FACS using FITC (fluorescein isothiocyanate)-conjugated annexin V (BD Biosciences) and PI (Sigma-Aldrich) staining. The analysis was performed with a FACS Calibur (BD Biosciences) instrument according to the standard protocol.
Immunofluorescence
Cells were grown on Thermo Scientific Nunc Lab-Tek II Chamber Slides & trade, fixed with 4% paraform-aldehyde for 15 minutes, and permeabilized with 0.5% Triton X-100 for 5 minutes. The fixed cells were incubated for 1 hours with 3% bovine serum albumin and then incubated overnight at 4
Statistical analysis
All data will be represented central tendency or typical value obtained in form two or three independent replicate experiments. The data of the experiments were statistically analyzed using GraphPad Prism (version 5.0, GraphPad Software Inc., San Diego, CA, USA) and presented as means
Author contributions
Conception: Jae Hyeong Kim, Ji Kon Ryu, Jin-Hyeok Hwang.
Interpretation or analysis of data: Jae Hyeong Kim.
Preparation of the manuscript: Jae Hyeong Kim.
Revision for important intellectual content: Jae Hyeong Kim, Yuna Youn, Jong-chan Lee, Jaihwan Kim.
Supervision: Jin-Hyeok Hwang.
Ethics declarations
The authors declare no competing interests.
Supplementary data
The supplementary files are available to download from http://dx.doi.org/10.3233/CBM-210490.
sj-pdf-1-cbm-10.3233_CBM-210490.pdf - Supplemental material
Supplemental material, sj-pdf-1-cbm-10.3233_CBM-210490.pdf
Footnotes
Acknowledgments
This research was funded by grant nos. NRF-2021R1A2C200708811 from the National Research Foundation of Korea (NRF) and the 2021 Seoul National University Research Fund. We would like to thank Editage (www.editage.co.kr) for English language editing.
