Abstract
BACKGROUD:
Hepatocellular carcinoma (HCC) is characterized by occult onset, rapid progression and poor prognosis. CXC chemokines play an important role in tumor microenvironment and development.
OBJECTIVE:
The potential mechanistic values of CXC chemokines as clinical biomarkers and therapeutic targets in HCC have not been fully clarified.
METHODS:
ONCOMINE, UALCAN, GEPIA, cBioPortal, SurvExpress, MethSurv, SurvivalMeth, String, GeneMANIA, DAVID, Metascape, TRRUST, LinkedOmics, and Timer were applied in this study.
RESULTS:
The transcriptional levels of CXCL9/16/17 in HCC tissues were significantly elevated while CXCL1/2/5/6/7/12/14 were significantly reduced. Significant correlation was found between the expression of CXC3/5 and the pathological stage of HCC patients. High level of CXCL4 was associated with a longer disease-free survival. For overall survival, lower expressions of CXCL1/3/5/8 and higher expressions of CXCL2 were associated with a better outcome. In addition, the prognostic values of CXC chemokines signature in HCC were explored in four independent cohorts, the high-risk group displayed unfavorable survival outcome compared with the low-risk group. And for the prognostic value of the DNA methylation of CXC chemokines, we identified the CpGs which were significantly associated with prognosis in HCC patients. DNA methylation signature analysis also showed a statistically significant association between the high- and low-risk groups. For potential mechanism, the neighbor gene networks, interaction analyses, functional enrichment analyses of CC chemokine receptors in HCC were performed, the transcription factor targets, kinase targets, and miRNA targets of CXC chemokines were also identified in HCC. We also found significant correlations among CXC chemokines expression and the infiltration of immune cells, the tumor infiltration levels among HCC with different somatic copy number alterations of these chemokine receptors were also assessed. Moreover, the Cox proportional hazard model showed that CCR2/6/8/12, B cell, macrophage and dendritic cell were significantly related to the clinical outcome of HCC patients.
CONCLUSION:
CXC chemokines might serve as therapeutic targets and prognostic biomarkers in HCC.
Introduction
Inflammation is considered to be the seventh major feature of tumor [1]. Tumor-associated inflammation is closely related to the occurrence and development of tumors. Hepatocellular carcinoma (HCC) is one of the malignant tumors with high incidence rate and mortality rate [2, 3]. According to the Surveillance, Epidemiology, and End Results (SEER) Program provided information on cancer statistics, the rate of new cases of liver and intrahepatic bile duct cancer was 9.0 per 100,000 men and women per year. The death rate was 6.6 per 100,000 men and women per year [4]. There is an estimated 42,230 new cases, and 30,230 deaths from U.S. mortality in 2021. Tumor invasion and metastasis is a series of complex, multi-step and multi-factor interaction between tumor cells, host and tumor microenvironment [5]. The interaction between tumor cells and different immune/inflammatory cells and stroma in tumor microenvironment has an important influence on the metastasis and recurrence of HCC [6]. Further study of the inflammatory microenvironment is of great value for the diagnosis and treatment of HCC.
As a small secretory protein, chemokines can regulate the migration and localization of immune cells in tissues. Recent studies have shown that chemokines play an important role in tumor progression and metastasis. For us humans, a total of 16 CXC chemokines (CXCL1-17, CXCL15 is specific to mice) were found to play key roles in physiological and pathological processes [7, 8]. Previously, we have investigated the effects of macrophages on HCC proliferation and metastasis through CXCL8, and found that cancer cells and macrophages interaction promoted cancer cell proliferation and metastasis through the up-regulation of CXCL8/miR-17 cluster [9]. Recently, Yan et al. revealed that the interferon regulatory factor 1 contributes to the anti-tumor microenvironment in HCC through CXCL10/CXCR3 axis [10]. Nie et al. have demonstrated that CXCL5 is a potential prognostic marker of HCC and provided clues about immune infiltration [11]. Liu et al.’s research indicated that cancer associated fibroblast derived CXCL11 regulates HCC cell migration and metastasis through the circUBAP2/miR-4756/IFIT1/3 axis [12]. Wang et al. provided new evidence that CXCL17 enhanced malignant invasion and inhibited autophagy through the LKB1-AMPK pathway in HCC [13]. Thus in HCC microenvironment, a variety of chemokines and their receptors shape a unique cross-talk mechanism between immune and tumor cells.
There is a lot of evidence that chemokines play an anti-tumor role in guiding immune cells to gather into tumor microenvironment. At the same time, there are many conclusions that chemokines can also induce immune cells to reshape the tumor microenvironment and promote the progress of cancer [14, 15]. In this study, based on several large public bioinformatics databases, an in-depth and comprehensive analysis of the potential values of CXC chemokines as clinical markers and immunotherapeutic targets in HCC was performed, thus to provide additional data to help clinicians choose appropriate treatment drugs and predict the long-term prognosis of patients with HCC more accurately.
Materials and methods
HCC samples of all online platform databases are: GTEx normal (110 cases), TCGA adjacent to cancer (50 cases) and TCGA tumor (371 cases).
GEPIA
GEPIA (Gene Expression Profiling Interactive Analysis), a web server for cancer and normal gene expression profiling and interactive analyses developed at Peking University (
Oncomine
The Oncomine
UALCAN
UALCAN (
cBioPortal
cBioPortal (www.cbioportal.org) integrates the data of 126 tumor genome studies, including large-scale tumor research projects such as TCGA and ICGC, covering 28,000 samples [19]. cBioPortal is widely used to explore, visualize and analyze multidimensional cancer genome data. 184 PDAC samples (TGCA, PanCancer Atlas), genetic alteration, and co-expression module of CXC chemokine were achieved and analyzed. mRNA expression
GeneMANIA
GeneMANIA (
STRING
STRING (
SurvExpress
SurvExpress (
MethSurv
MethSurv (
SurvivalMeth
SurvivalMeth (
LinkedOmics
LinkedOmics (
TRRUST
Transcriptional Regulatory Relationships Unraveled by Sentence-based Text mining (TRRUST) database (
mRNA levels of CXC chemokines in HCC (ONCOMINE). The figure shows the numbers of datasets with statistically significant mRNA over-expression (red) or downregulated expression (blue) of CXC chemokines.
The transcription of CXC chemokines in HCC (UALCAN). The transcriptional levels of (A) CXCL1, (B) CXCL2, (C) CXCL3, (D) CXCL4, (E) CXCL5, (F) CXCL6, (G) CXCL7, (H) CXCL8, (I) CXCL9, (J) CXCL10, (K) CXCL11, (L) CXCL12, (M) CXCL13, (N) CXCL14, (O) CXCL16, (P) CXCL17 displayed. (Q) Heatmap of the expression pattern of the input genes in HCC were displayed. The 
The relative level of CXC chemokines in HCC.
Timer (Tumor Immune Estimation Resource,
Results
CXC chemokine expression in HCC
There are 16 chemokines of CXC ligand family in human, including CXCL1-17 (CXCL15 is specific to mice). The Oncomine
Furthermore, the transcription of CXC chemokine in HCC was explored by UALCAN. Figure 2: As expected, the transcriptional levels of CXCL9 (
Correlation between different expressed CXC chemokines and the pathological stage of HCC patients (GEPIA). 
The prognostic value of different expressed CXC chemokines in HCC patients in the disease-free survival curve (GEPIA). The disease-free survival curve of (A) CXCL1, (B) CXCL2, (C) CXCL3, (D) CXCL4, (E) CXCL5, (F) CXCL6, (G) CXCL7, (H) CXCL8, (I) CXCL9, (J) CXCL10, (K) CXCL11, (L) CXCL12, (M) CXCL13, (N) CXCL14, (O) CXCL16, (P) CXCL17 in HCC.
The correlation of differentially expressed CXC chemokines and clinical outcomes were explored by GEPIA. Figure 5, a notably longer disease-free survival was demonstrated with the high transcriptional level of CXCL4 (
The prognostic value of different expressed CXC chemokines in HCC patients in the overall survival curve (GEPIA). The disease-free survival curve of (A) CXCL1, (B) CXCL2, (C) CXCL3, (D) CXCL4, (E) CXCL5, (F) CXCL6, (G) CXCL7, (H) CXCL8, (I) CXCL9, (J) CXCL10, (K) CXCL11, (L) CXCL12, (M) CXCL13, (N) CXCL14, (O) CXCL16, (P) CXCL17 in HCC.
The CXC chemokines signature was input for prognostic analysis in SurvExpress. For overall survival, in the training cohort 1 (GSE10143, 162 patients), the mRNA expression of CXCL1, CXCL14 was higher in the low-risk group than that in the high-risk group, while CXCL4, CXCL5, CXCL6, and CXCL11 were lower expressed in the low-risk group than that in the high-risk group (Fig. 7A and B), the high-risk group displayed an unfavorable overall survival compared with the low-risk group (Fig. 7C). For relapse-free survival, in the training cohort 2 (GSE17856, 95 patients), the mRNA expression of CXCL9, CXCL10 and CXCL16 was higher in the low-risk group than that in the high-risk group. However, a comparable relapse-free survival was observed of the low- and high-risk groups (Fig. 7D–F). Similarly, these results were also displayed in the training cohort 3 (GSE10186, 118 patients), the high-risk group had a worse OS compared with the low-risk group (Fig. 7G–I). And the prognostic value of CXC chemokines signature in HCC was also confirmed in the validation cohort (TCGA, 361 patients) (Fig. 7J–L).
The prognostic values of CXC chemokines signature for survival analysis in HCC cases from the training cohort 1 (GSE10143, 162 patients), training cohort 2 (GSE17856, 95 patients), training cohort 3 (GSE10186, 118 patients), and validation cohort (TCGA, 361 patients) via SurvExpress platform. Notes: (A, D, G and J) the heat maps of mRNA expression of CXC chemokines; (B, E, H and K) the mRNA expressions of CXC chemokines between high-and low-risk groups; (C, F, I and L). Survival curves of CXC chemokines between high- and low-risk groups.
The heat map of DNA methylation clustered expression level of CXC chemokines (MethSurv). Notes: cg10350689 of CXCL1, cg26013975 of CXCL2, cg13468041 of CXCL3, cg14882398 of CXCL4, cg15478045 of CXCL5, cg22670329 of CXCL6, cg26523478 of CXCL7, cg125799590 of CXCL10, cg08046471 of CXCL11, cg06671614 of CXCL12, cg12020230 of CXCL13, cg23510026 of CXCL14, cg08313880 of CXCL16, cg22276896 of CXCL17 showed the highest DNA methylation level in HCC.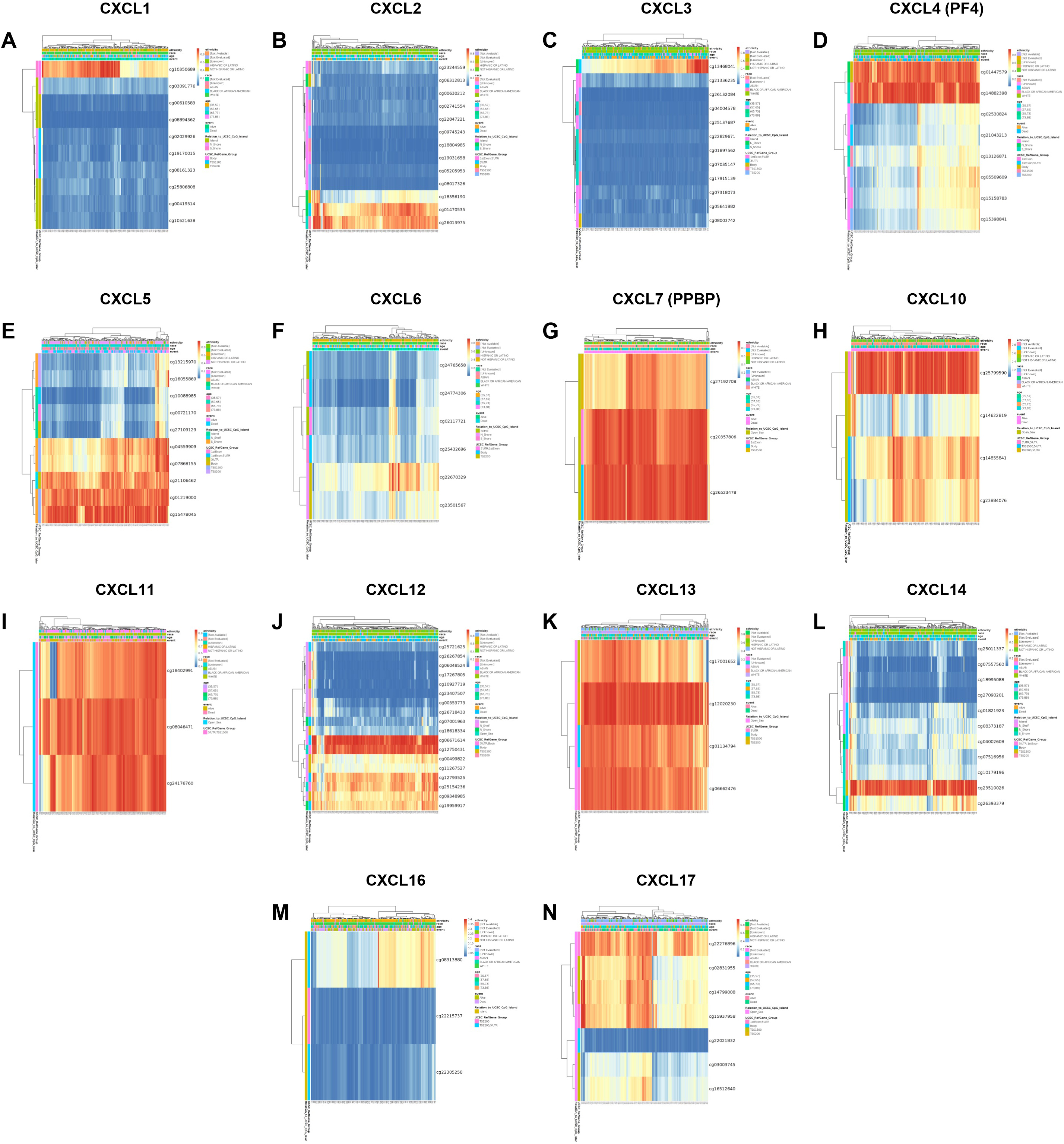
The prognostic value of DNA methylation of CXC chemokines in HCC was analyzed by MethSurv. The heat maps of DNA methylation of the CXC chemokines are displayed in Fig. 8. Among them, cg10350689 of CXCL1, cg26013975 of CXCL2, cg13468041 of CXCL3, cg14882398 of CXCL4, cg15478045 of CXCL5, cg22670329 of CXCL6, cg26523478 of CXCL7, cg125799590 of CXCL10, cg08046471 of CXCL11, cg06671614 of CXCL12, cg12020230 of CXCL13, cg23510026 of CXCL14, cg08313880 of CXCL16, cg22276896 of CXCL17 showed the highest DNA methylation level. And overall, we found that 7 CpGs of CXCL1, 5 CpGs of CXCL2, 4 CpGs of CXCL3, 5 CpGs of CXCL4, 10 CpGs of CXCL5, 1 CpG of CXCL6, 1 CpG of CXCL7, 2 CpGs of CXCL10, 3 CpGs of CXCL12, 3 CpGs of CXCL14, and 5 CpGs of CXCL17 were significantly associated with prognosis in HCC patients (Supplementary Table 2).
The prognostic value of the DNA methylation of CXC chemokine signature in HCC via SurvivalMeth. Notes: (A) The methylation level of CpGs in high- and low-risk group, (B) The survival curve of Kaplan-Meier plot, (C) The heatmap of CpG methylation level.
The gene symbols of CXC chemokines were input for prognostic analysis in SurvivalMeth. As shown in Figs 9A–C, the significant expression patterns were found in CXCL4/5/7/8/9/10/12/13 between low- and high-risk groups. A significantly better survival outcome was observed for the low-risk group as compared to the high-risk group (
Genetic alteration, neighbor gene network, and interaction analyses of different expressed CXC chemokines in HCC patients. (A) Summary of alterations in different expressed CXC chemokines in HCC. (B) Correlation heat map of CXC chemokines in HCC. (C and D) Protein-protein interaction network of different expressed CXC chemokines.
A comprehensive molecular characterization of differentially expressed CXC chemokines was performed. The genetic alteration of differentially expressed CXC chemokines was analyzed by the provisional datasets of TCGA. Fig. 10A, the analytical results showed that CXCL-17 were altered in 3, 5, 4, 3, 6, 2.6, 4, 4, 6, 5, 7, 3, 4, 5, 7, and 4% of the queried HCC samples, respectively. Gene amplification, mutation, deep deletion and multiple alterations were the main genetic alteration types in HCC. Enhanced mRNA expression was the most common change in these samples. Next, the potential co-expression of the differentially expressed CXC chemokines was explored. There was a moderate to high correlation among the expression of CXCL1, CXCL2, CXCL3, and CXCL17, as showed by the heatmap of the co-expression of the differentially expressed CXC chemokines (Fig. 10B). Figure 10C: A PPI network analysis of differentially expressed CXC chemokines with STRING was conducted to explore their possible interactions. Combined with GeneMANIA analysis (Fig. 10D), the effect of 16 CXC chemokines was mainly associated with chemokine receptor binding and the inflammatory response. Besides, the top 50 most regularly interacting neighboring genes related to 16 CXC chemokines were analyzed by STRING. These data suggested that ACKR1, ACKR3, CCL1, CCL11, CCL19, CCL2, CCL20, CCL21, CCL25, CCL3, CCL4, CCL5, CCR1, CCR2, CCR3, CCR4, CCR5, CCR7, CD4, CD80, CD86, CSF1, CSF2, CSF3, CXCR1, CXCR2, CXCR3, CXCR4, CXCR5, CXCR6, ICAM1, IL10, IL10RA, IL10RB, IL13, IL18, IL1B, IL4, IL6, ITCH, JAK1, MMP9, PTK2, PTPRC, RELA, STAT3, STAT4, TIMP1, TNF, TNFRSF1B are mainly related to the regulation and capability of CXC chemokines.
CXC chemokine functional enrichment analysis in HCC
The enrichment analysis of CXC chemokines and their closely interacting neighboring genes were performed by DAVID 6.8 and Metascape. Figure 11A–C shows that among the three most highly enriched functions in the BP category, cellular response to chemokine, response to chemokine, and chemokine-mediated signaling pathway were associated with the tumorigenesis and progression of HCC. Platelet alpha granule lumen, tertiary granule lumen and external side of plasma membrane were the three most highly enriched items in the CC category. In the molecular function (MF) category, they were cytokine activity, cytokine receptor binding, and chemokine receptor binding. Figure 11D: KEGG pathway analyses were also performed. As expected, among the top three KEGG pathways, chemokine signaling pathway, cytokine-cytokine receptor interaction, and viral protein interaction with cytokine and cytokine receptor were significantly associated with the tumorigenesis and progression of HCC.
Transcription factor targets, kinase targets and miRNA targets of CXC chemokine in HCC
Key regulated factor of CXC chemokines in HCC was performed by TRRUST. Three transcription factors, RELA, NFKB1 and SP1 were found associated with CXC chemokine regulation (Table 1). Table 2, the top two kinase targets of CXC chemokines were identified from LinkedOmics (
Key regulated factor of CXC chemokines in HCC (TRRUST)
Key regulated factor of CXC chemokines in HCC (TRRUST)
The Kinase target networks of CXC chemokines in HCC dataset (LinkedOmics)
The enrichment analysis of different expressed CXC chemokines and 50 most frequently altered neighboring genes in HCC (David 6.8). GO enrichment in cellular component terms, biological process terms, molecular function terms and KEGG enriched terms.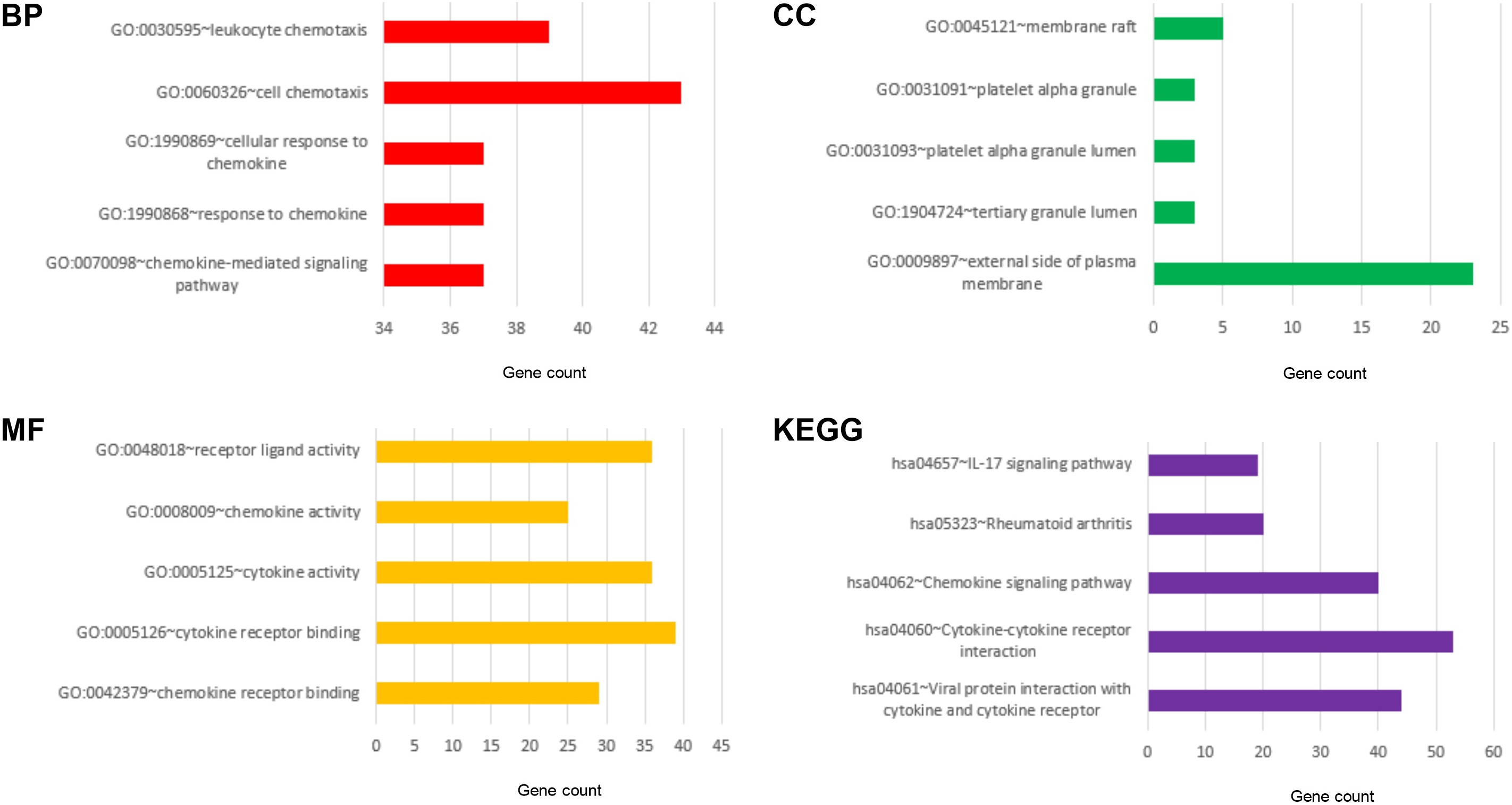
The correlation between different expressed CXC chemokines and immune cell infiltration (TIMER). The correlation between the abundance of immune cell and the expression of (A) CXCL1, (B) CXCL2, (C) CXCL3, (D) CXCL4, (E) CXCL5, (F) CXCL6, (G) CXCL7, (H) CXCL8.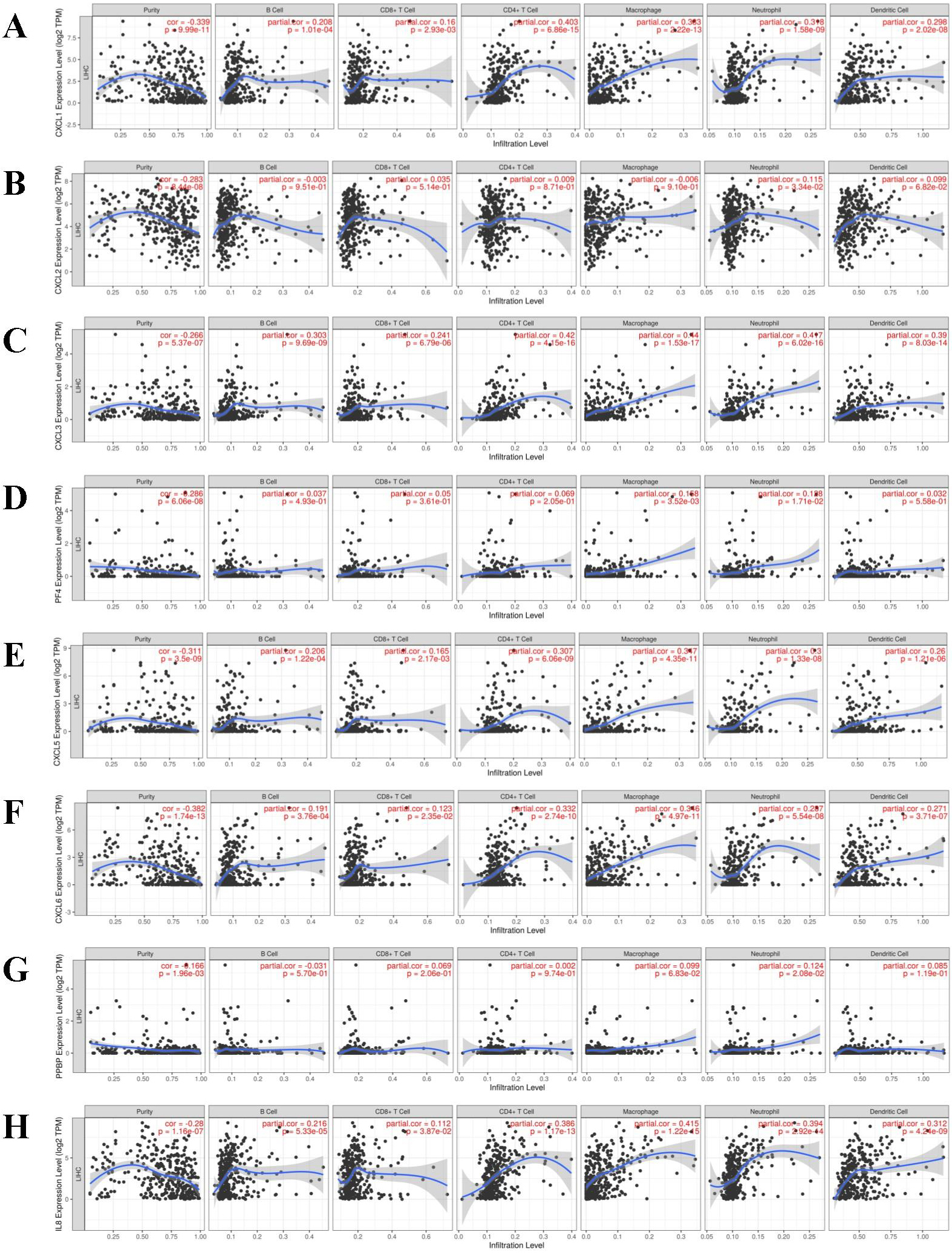
Continued. (I) CXCL9, (J) CXCL10, (K) CXCL11, (L) CXCL12, (M) CXCL13, (N) CXCL14, (O) CXCL16, (P) CXCL17 in HCC.
TIMER database was applied to display a comprehensive exploration of the correlation between differentially expressed CXC chemokines and immune cell infiltration (Fig. 12). Moreover, the Cox proportional hazard model was applied for CXC chemokines and six tumor-infiltrating immune cells in HCC (Supplementary Fig. 2). As shown in Table 3, CXCL2 (
The cox proportional hazard model of CXC chemokines and six tumor-infiltrating immune cells in HCC (TIMER)
The cox proportional hazard model of CXC chemokines and six tumor-infiltrating immune cells in HCC (TIMER)
TIMER in LIHC (362 patients with 127 dying).
In HCC, immunosuppressive cells including MDSCs, TAMs, TANs, Tregs, CAFs and TILs are important components of tumor microenvironment. They interact with each other to produce growth factors, cytokines and chemokines to participate in immunosuppression, thus promoting the progress of HCC [30]. The differentiation, maturation and function of these immune cells are also regulated by cytokines and other chemokines. By regulating a variety of signaling pathways, chemokines and their receptors directly or indirectly shape the microenvironment of tumor cells and regulate the biological behavior of tumor, and therefore, involved in inflammatory response, tumor immune response, proliferation, invasion and metastasis. Recent studies have shown that chemokines in HCC microenvironment play an important role in the occurrence and development of tumor. Through the recruitment of inflammatory cells, inhibition of anti-tumor immune response, promotion of angiogenesis/tumor resistance, it eventually leads to the occurrence of HCC, and leads to invasion and metastasis [31, 32].
Previously, Wang et al. explored the mRNA transcriptional and survival analysis of CXCLs in patients with HCC from the databases involving ONCOMINE, GEPIA, and cBioPortal [43]. In this study, we first discussed the expression difference of CXC chemokines in HCC and its normal liver tissues, and showed the correlation between CXC chemokines and the staging of HCC. In addition, the differences of CXC chemokines expression and the survival and prognosis of HCC were also explored. We also revealed the role of epigenetic factors from methylation analysis of CXC chemokines related genes. After that, the molecular characteristics and functional enrichment of CXC chemokines have also been explored. Then we comprehensively analyzed and predicted CXC chemokines transcription factors, kinases and miRNA targets. Finally, we also introduced the correlation analysis of immune infiltration to demonstrate the influence of various CXC chemokines on immune cells and microenvironment in HCC.
In the present study, we focused on the CXC subfamily chemokines with the purpose providing novel insights as well as prognostic biomarkers for the immunotherapy strategy selection in HCC. In line with our work, a recently published meta-analysis explored the correlation of chemokine concentrations and HCC prognosis. They found that higher levels of CCL20, CXCL8 and CXCR4 could be served as HCC prediction biomarkers [33]. In consist with that study, CXCL8 was also found an increased level expression in HCC compared with that in normal tissues with the fold changes of 3.323, and a higher concentration of CXCL8 indicated a worse survival by our analysis. It has been reported that the AKT/mTOR/STAT3 signaling pathway was involved in the progression and metastasis of CXCL8 [34]. What’s more, CXCL8 was responsible for the Tregs infiltration in HCC microenvironment [35].
A significantly higher expression of CXCL10 was found in HCC tumor tissues as compared with the normal tissues. Of note, there was research showed that a positive correlation existed between CXCL10 and AFP, the size and number of tumor as well as the TNM staging in HCC, and CXCL10 could be regarded as an independent prognostic factor for HCC from the multivariate analysis [36]. Another CXC subfamily chemokine, CXCL12, is found notably reduced in HCC tissue (
Reviewing the history of systemic treatment of HCC, in 2008, sorafenib became the first-line therapy to achieve a significant survival benefit in advanced HCC [40]. Since then, sorafenib has remained alone in the first-line, and no other first-line therapy has been shown to be more effective than it. Ten years later, a study showed that the overall survival improvement of lenvatinib was no worse than that of sorafenib, and progression-free survival and time to progression were better than that of sorafenib [41]. In the NCCN Guidelines for Hepatobiliary Cancer of the United States of America, Version 2021, the combination therapy of atelizumab and bevacizumab is the only first-line treatment option recommended in the Guidelines [42]. This also completely elevates the status of “T
Conclusion
In conclusion, CXC chemokines in HCC could be regarded as prognostic indicators. Targeting the CXC chemokines may result in a valuable prognostic impact and assist novel immunity-based treatment identification in HCC. It can be predicted that HCC patients may benefit from the treatment strategies targeting CXC chemokines in the future. This study will also provide important explanations and innovation ideas for oncologists in the future.
Statement of ethics
This study is a bibliometric analysis of the articles related to management of hepatocellular carcinoma, and the data were obtained from the Clarivate Analytics Web of Science Core Collection database, so ethical approval and consent to participate were not necessary for this paper.
Funding
Natural Science Foundation of Guangdong (No. 2021A1515012234, Zi Yin); Science and Technology Program of Guangzhou (No. 201904010043, Zi Yin); National Key Clinical Specialty Construction Project (2021–2024, No. 2022YW030009).
Author contributions
Conception: Zi Yin.
Interpretation or analysis of data: Zi Yin, Tingrting Ma.
Preparation of the manuscript: Zi Yin, Min Yu.
Revision for important intellectual content: Sheng Chen.
Supervision: Zi Yin.
Supplementary data
The supplementary files are available to download from http://dx.doi.org/10.3233/CBM-210300.
sj-docx-1-cbm-10.3233_CBM-210300.docx - Supplemental material
Supplemental material, sj-docx-1-cbm-10.3233_CBM-210300.docx
Footnotes
Conflict of interest
The authors declare no potential conflicts of interest.
