Abstract
BACKGROUND:
Chronic HCV infection progresses to fibrosis, cirrhosis and hepatocellular carcinoma (HCC). The latter represents the third most common cause for cancer mortality. Currently, there is no reliable non-invasive biomarker for diagnosis of HCV mediated disorders.
OBJECTIVE:
Profiling expression signature for circulatory miRNAs in the plasma of 167 Egyptian patients (40 healthy, 48 HCV fibrotic, 39 HCV cirrhotic and 40 HCV-HCC cases).
METHODS:
QRTPCR was used to quantify expression signature for circulatory miRNAs.
RESULTS:
MiR-676 and miR-650 were powerful in discriminating cirrhotic and late fibrosis from HCC. MiR-650 could distinguish mild (f0-f1) and advanced (f2-f3) fibrosis from HCC cases. MiR-650 and miR-147b could distinguish early fibrosis from healthy controls meanwhile miR-676 and miR-147b could effectively distinguish between mild chronic and (f1-f3) cases from healthy individuals. All studied miRNAs, except miR-512, can differentiate between (f0-f3) cases and healthy controls. Multivariate logistic regression revealed three potential miRNA panels for effective differentiation of HCC, cirrhotic and chronic liver cases. MiR-676-3p and miR-512-5p were significantly correlated in (f1-f3) fibrosis meanwhile miR-676 and miR-512 could differentiate between cirrhosis and (f0-f3) cases. Both miR-650 and miR-512-5p were positively correlated in the cirrhotic group and in (f0-f4) group. Putative targets for investigated miRNAs were also determined.
CONCLUSIONS:
Investigated miRNAs could assist in staging and diagnosis of HCV associated disorders.
Keywords
Introduction
Hepatitis C virus (HCV) infection is a contagious disorder with an estimated 185 million infected cases globally [1]. The highest prevalence rates are found in Eastern Mediterranean region followed by European and the African regions [2]. Egypt has the highest HCV prevalence rate worldwide where 14.7% of its population is infected with HCV genotype 4 [3]. Approximately 70% of individuals infected with HCV develop chronic HCV (CHC) which could end up with more devastating disorders such as fibrosis, irreversible cirrhosis and hepatocellular carcinoma (HCC) [2].
Liver fibrosis is a wound-healing response to chronic liver injury [4]. Persistence of fibrosis could lead to hepatic scar formation and irreversible cirrhosis that might ultimately lead to HCC progression [5, 6]. HCC is the fifth most common cancer and the world’s third leading cause of cancer death [7]. HCV associated HCC is the leading cause of death for about one million cases annually [8]. Nearly 31% of HCC cases arise from HCV infection [9]. The incidence of HCC is expected to increase in the next two decades, primarily due to HCV infection and secondary owing to development of cirrhotic stages [10].
Despite recent development of successful strategies against HCV, death rate arising from HCV associated disorders is still high [11]. Therefore, early detection of HCV mediated disease progression is very crucial. Although liver biopsy is the gold standard approach for this purpose, it is not widely implemented as it is expensive, invasive with the possibility for sampling error in addition to inter- and intra-observer variation in pathology reporting [12]. Few non-invasive hepatic markers were developed such as fibrosis-4 FIB-4, AST/platelet ratio index (APRI) and Göteborg University Cirrhosis Index (GUCI) [13]. Alpha feto-protein (AFP) and des-gamma-carboxy prothombin (DCP) are the main non-invasive biomarkers used for HCC diagnosis. However, all those biomarkers can not accurately distinguish between various stages of HCV mediated liver disorders and lack sensitivity [5, 14, 15]. There is urgent need for investigating other reliable non-invasive biomarkers for early staging, diagnosis and/or prognosis.
MicroRNAs (miRNAs) are small noncoding RNAs that regulate gene silencing by targeting mRNA [10, 16]. Aberrant expression of miRNAs has been linked to variety of cancer diseases [17, 18]. Circulating miRNAs leak out of cells into body fluids such as plasma and urine [19, 20]. They were reported to be stable against endogenous RNases and to display consistent profiles among healthy individuals [21, 22]. These characteristics established their potential value as biomarkers during pathological changes [23]. For instance, miR-25 and miR-223 were shown to be reliable serum biomarkers for lung cancer [24] while miR-122 was reported as a promising biomarker in HCC cases [23]. Moreover, miR-184 and miR-92a were reported for squamous cell carcinoma and leukemia detection respectively [25, 26]. Furthermore, miR-141 and miR-375 were the most promising biomarkers correlated with prostate tumor progression [27].
The main objective of the current study is to investigate expression profiles for 5 plasma miRNAs namely; miR-650, miR-512-5p, miR-552-3p, miR-147b and miR-676-3p in various stages of HCV associated hepatic disorders among Egyptian patients in an attempt to identify potential non-invasive biomarkers for early diagnosis and staging purposes.
Rationale for selecting the five miRNAs in this study and their role in liver pathogenesis:
It was reported that miR-512 promoted oncogenic effects by targeting two members of the downstream tumor suppressors of JNK axis namely MAP3K2 and MAP2K4. The latter were believed to be significantly down-regulated in hepatocellular carcinoma (HCC) [28]. Concerning miR-650, it was postulated to promote migration, invasion, and Epithelial-Mesenchymal Transition (EMT) of HCC cells by targeting the large tumor suppressor kinase 2 gene (LATS2) [29]. It is also believed that it inhibits expression of the tumor suppressor gene namely inhibitor of growth protein 4 (ING4) and causes the progression of HCC [30]. ING4 is an emerged multimodal tumor suppressor gene of ING family where its expression is downregulated or even completely lost in various human cancers including HCC. Moreover, low expression of ING4 in liver cancerous tissues for patients with HCC is significantly correlated with rapid cancer progression and patient poor prognosis. Based on these facts, ING4 is being considered as a potential candidate in the gene-based cancer therapy approach [31]. In vitro studies indicated that miR-552 promotes HCC progression by blocking WIF1mediated GSK3
The expression levels for miR-650, miR-512-5p, miR-552-3p, miR-147b and miR-676-3p were not profiled previously in the plasma of HCV mediated cirrhosis, fibrosis and HCC patients across the globe in general and among Egyptians in particular. Therefore, this study aimed at investigating actual expression signature for those 5 miRNAs in the plasma of Egyptian patients affected with various stages of HCV associated hepatic disorders including fibrosis, cirrhosis and HCC. Our study also provided further insights and deep analysis for the possible mechanistic pathways and targets triggered by those 5 miRNAs.
Materials and methods
Subjects
This study included 127 CHC Egyptian patients (26 mild fibrosis and 22 advanced fibrosis, 39 liver cirrhotic and 40 HCC subjects) with HCV viral load as measured by qRT-PCR (Step-One Plus, Applied Biosystems, USA). A written informed consent was obtained from all participants and the study protocol was approved by ethics review committee at Cairo University (Serial: BC2116) and was conducted following Helsinki declaration. Inclusion criteria set for this study were; age
Routine laboratory tests, RNA extraction and cDNA synthesis
Blood samples were collected and the following biochemical tests were performed; RBCs count, CBC, Hb, total leukocyte count, platelet count, prothrombin time, MCV, INR, AFP, ALP, albumin, ALT, AST and total bilirubin as reported previously [40, 41, 42]. Total RNA was extracted using Direct-zol™RNA MiniPrep Kit (Zymo Research, USA) according to manufacturer’s protocol. Prior to the ethanol precipitation step, cel-miR-39-3p (Qiagen, Germany) was used as a spike in control for all samples. Extracted RNA samples were quantified using Nanodrop Q5000 UV-Vis (Quawell, USA) [43, 44, 45] and then 100 ng RNA sample was poly-adenylated using E. coli Poly(A) Polymerase (New England Biolabs, UK). This was followed by reverse transcription using the GoScript Reverse Transcription kit (Promega, USA) according to manufacturer’s manual.
Quantitative Real-time PCR (qRT-PCR)
QRT-PCR was carried out using the GoTaq
Primers sequences used for amplification in qRT-PCR
Primers sequences used for amplification in qRT-PCR
The Data analysis was performed using SPSS software version-15 and GraphPad Prism-5.0). Data are represented as mean
Results
Demographic and clinical features of healthy control, CLD (F0-F3), cirrhotic and HCC patients
Healthy controls had negative HCV viral load while all other participating groups showed positive HCV viral load with comparable viral titer Supplementary Table 1. Hepatic enzymes (AST, ALT and ALP) and bilirubin levels were significantly upregulated among all diseased groups (
Differential expression of the investigated miRNAs among the studied groups
Kruskal-Wallis multiple comparisons revealed significant differentiation for all investigated miRNAs among all the participating groups (Fig. 1). Comparing HCC to healthy controls using Mann-Whitney showed that both miR-676-3p and miR-512-5p were significantly upregulated in HCC group (
Differential expression of plasma miRNA levels in Healthy Control, CLD, cirrhotic and HCC patients. The box represents the 25%–75% percentiles; the line inside the box represents the median and the upper and lower lines represent the 10%–90% percentiles of fold change in expression levels of the studied miRNAs. Data were analyzed by Mann-Whitney U-test. Groups with (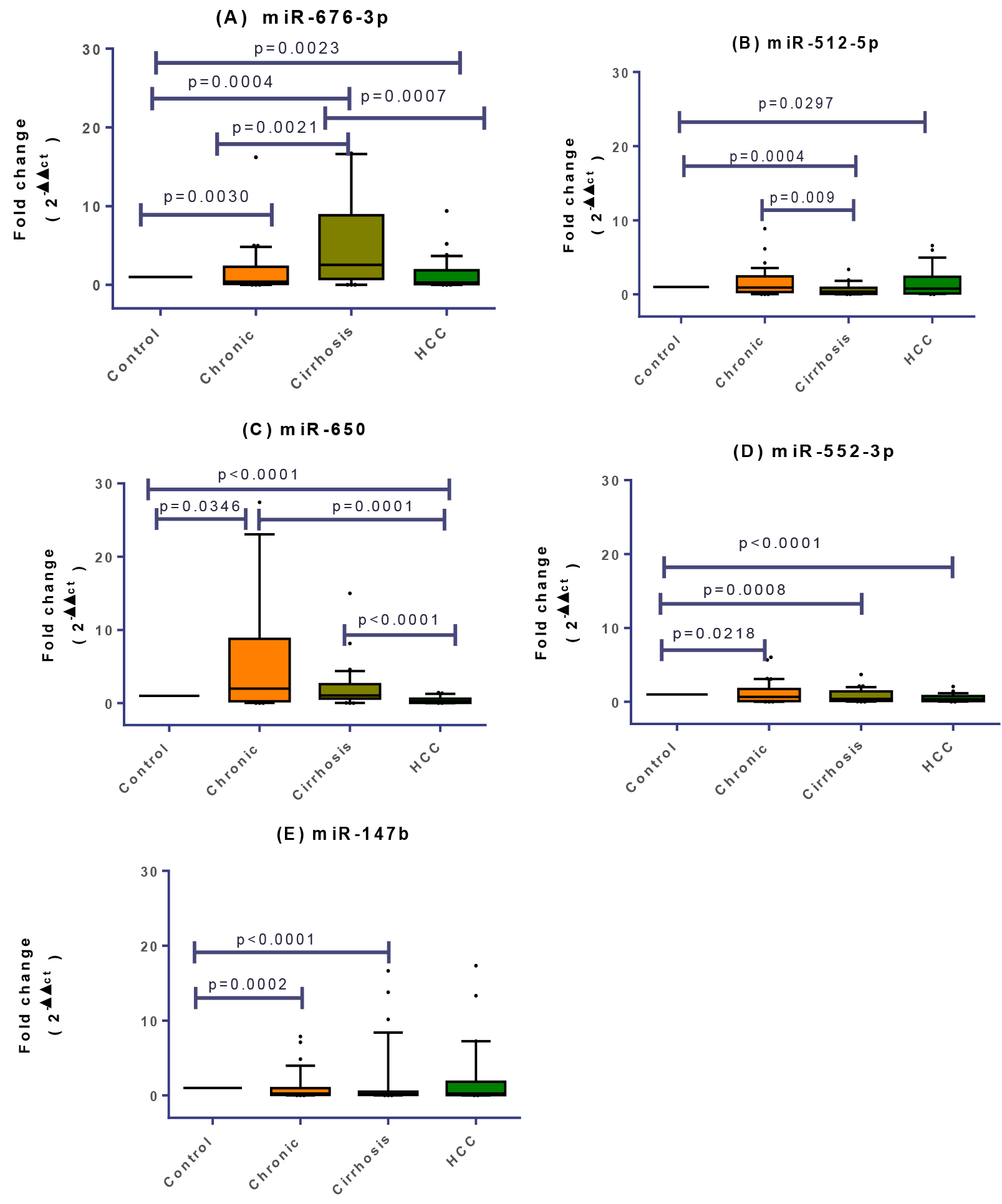
Individual miRNA expression signature in every fibrotic stage (f1 to f4) was compared in order to monitor disease progression. Expression profile for miR-650 exhibited significant upregulation in early fibrotic (f1-f2) group as compared to healthy controls meanwhile miR-147b exhibited significant downregulation (
Signature of plasma miRNAs relative expression profiles in healthy, f0, f1-f2, f3-f4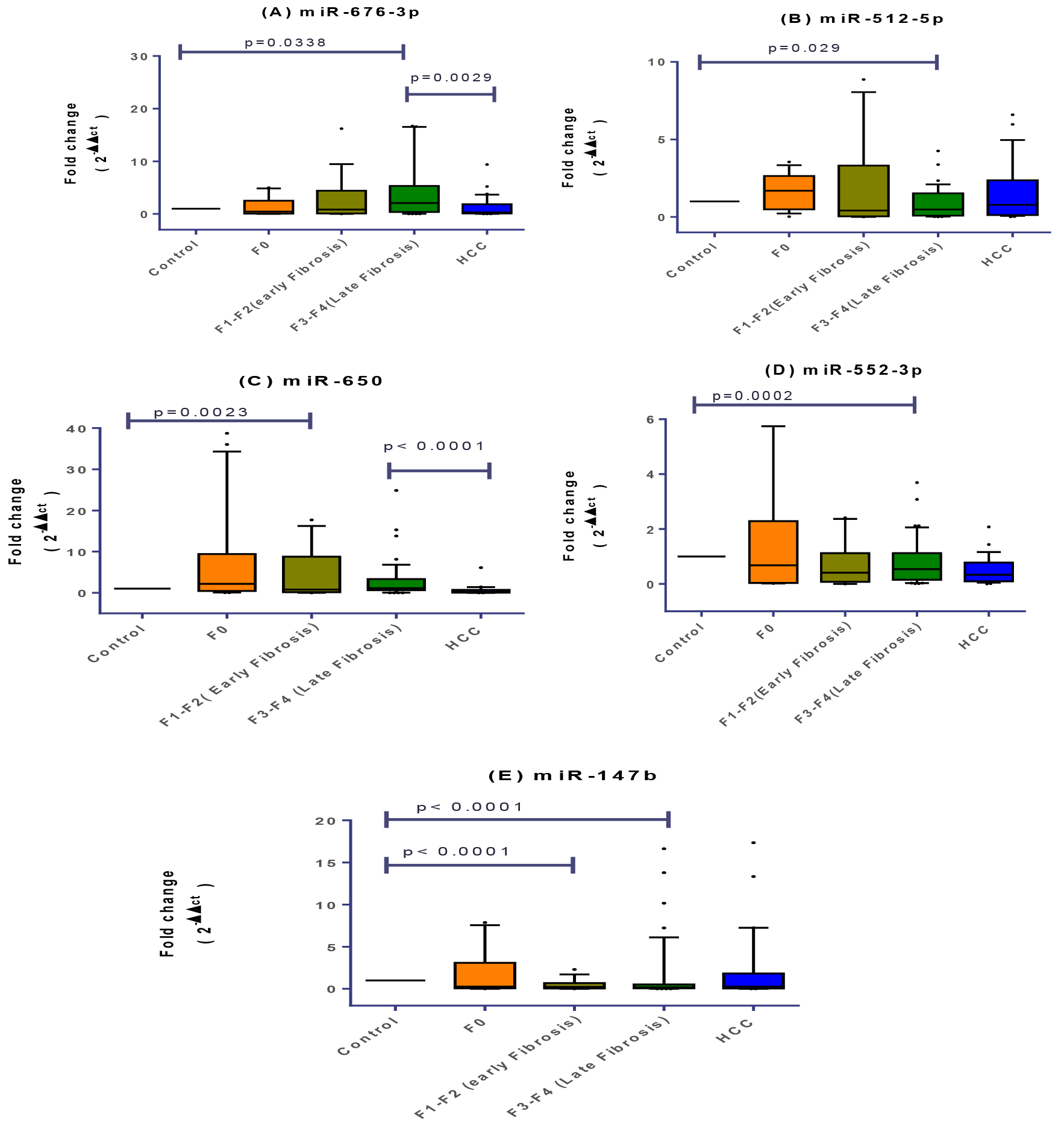
Signature of plasma miRNAs relative expression profiles in healthy, f0, f1-f3, cirrhosis and HCC groups. Data were analyzed by Mann-Whitney U-test. Groups with (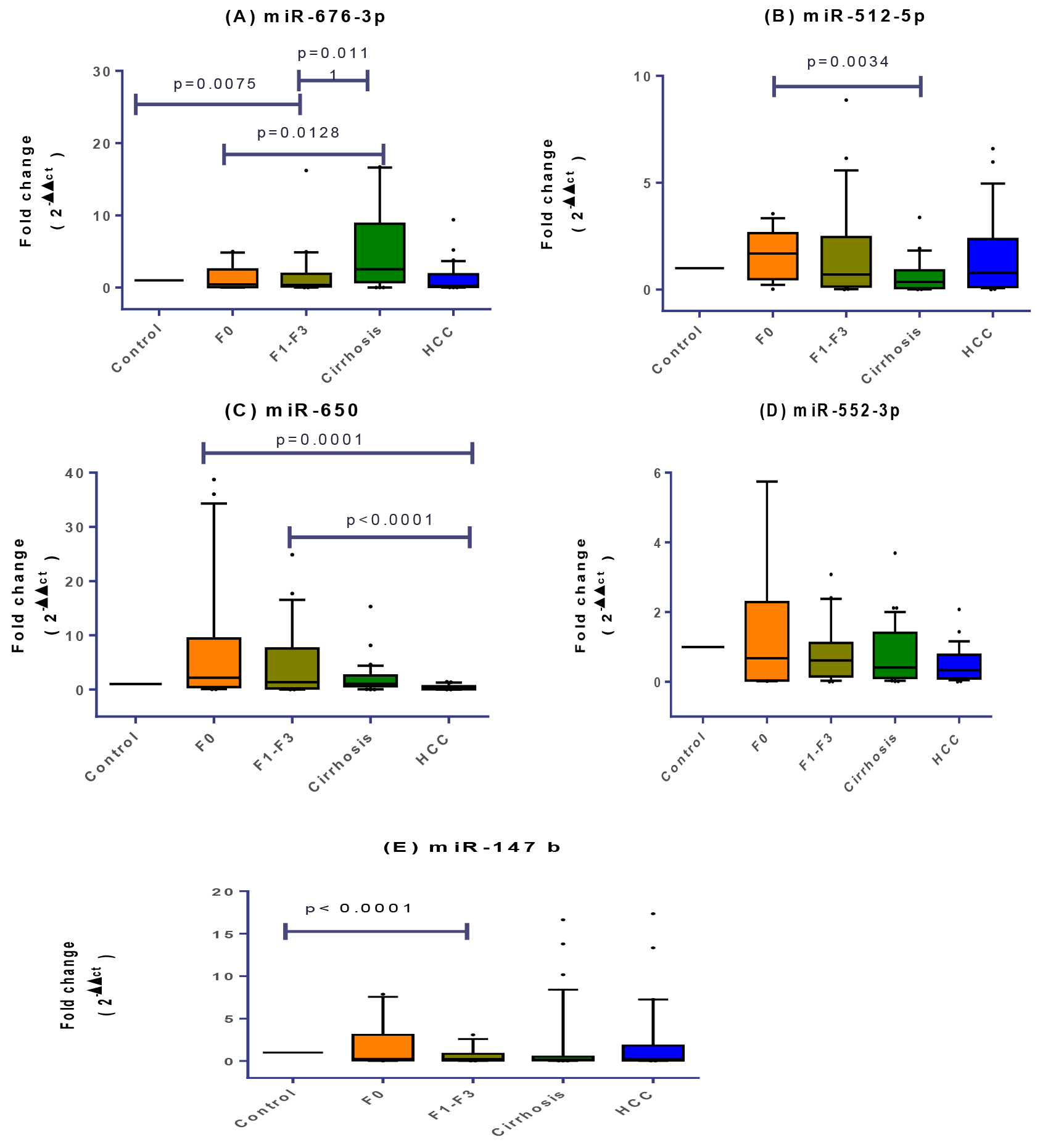
Signature of plasma miRNAs relative expression profiles in Non HCC and HCC groups. The box represents the 25%–75% percentiles; the line inside the box represents the median and the upper and lower lines represent the 10%–90% percentiles of fold change in expression levels of the studied miRNAs. Data were analyzed by Mann-Whitney U-test. Groups with (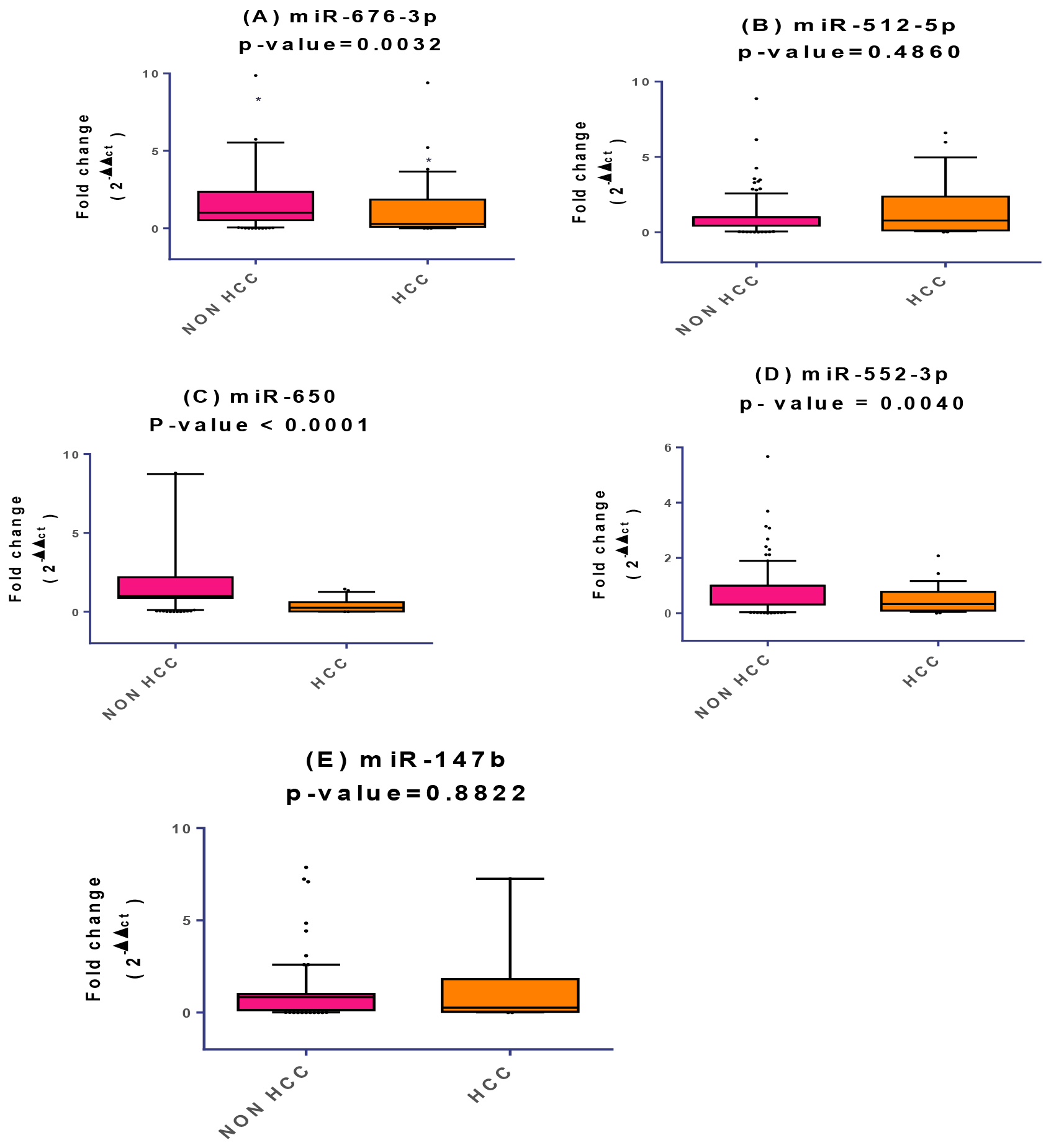
Fold change of miRNA expression levels in non-cirrhotic and cirrhotic group. The box represents the 25%–75% percentiles; the line inside the box represents the median and the upper and lower lines represent the 10%–90% percentiles of fold change in expression levels of the studied miRNAs. Data were analyzed by Mann-Whitney U-test. Groups with (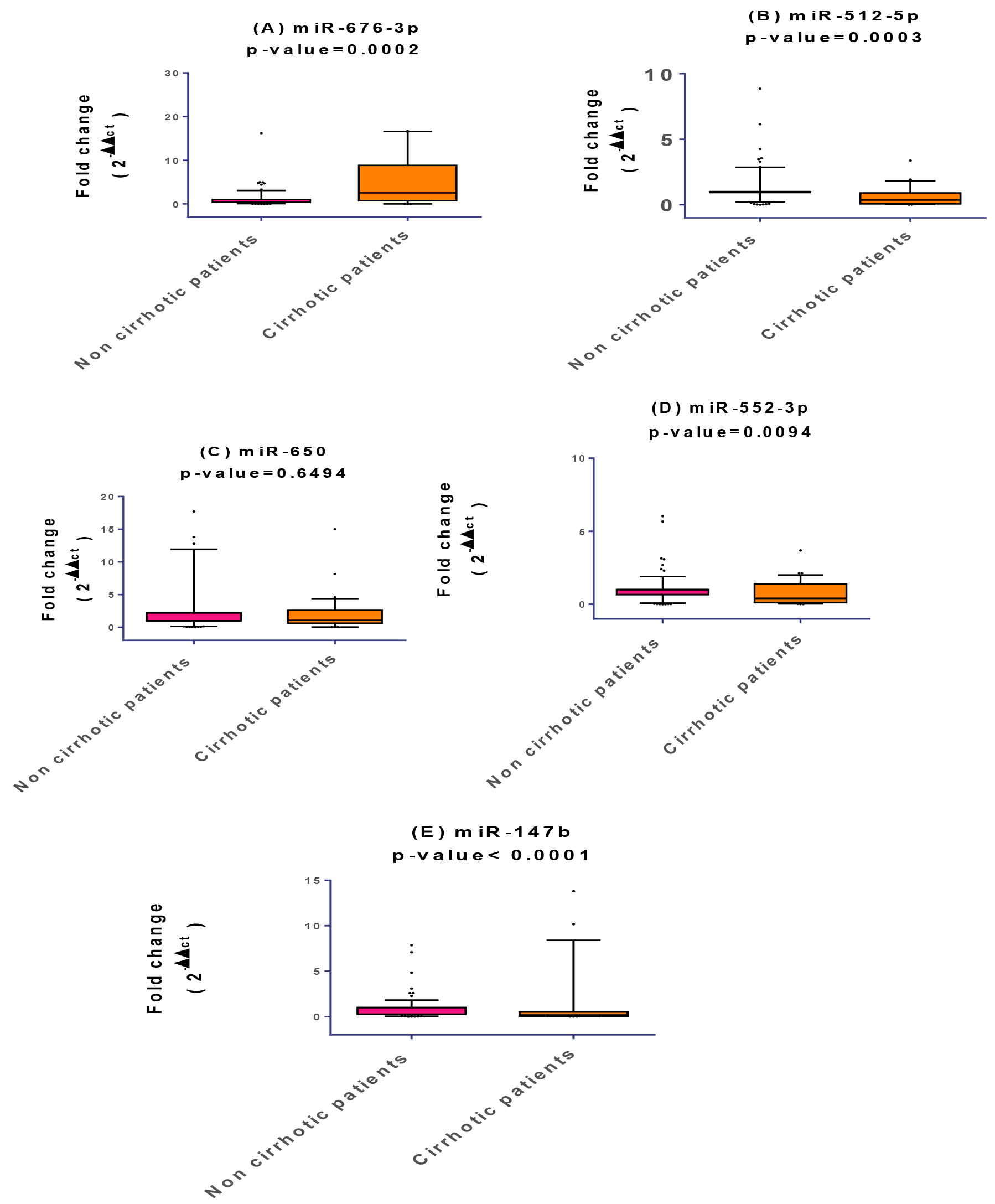
Diagnostic performances of the studied plasma miRNAs to discriminate HCC, CLD, cirrhotic patients and healthy controls from each other’s. Significance is indicated by (
Comparing
Combining (f0-f4) categories revealed also significant upregulation in the expression profiles of miR-512-5p and miR-552-3p (
ROC analysis revealed that miR-650 was the best among all other studied miRNAs in discriminating HCC from healthy controls with an AUC
Correlations between the studied plasma miRNAs in various HCV mediated liver diseases
There were various positive and significant correlations between the studied miRNAs in different stages of HCV mediated liver diseases. For instance, miR-676-3p was correlated with marginal significance not only with miR-512-5p (Spearman
Logistic regression analysis of miRNAs
Univariate and multivariate logistic regression analyses revealed that miR-650 and miR-552 were significant predictors associated with the chances of HCC diagnosis with an AUC value of 0.6320. The same miRNA panel could also discriminate HCC from healthy controls (AUC
Target prediction analysis
Based on the TargetScan and miRDB prediction programs, miR-676-3p, miR-512-5p, miR-650, miR-552-3p and miR-147b were found to strongly target genes related to fibrosis, cirrhosis and HCC development and progression. We closely reviewed the putative targets for the chosen miRNAs in order to explain the significance of the dysregulation of their expression in the context of HCV related hepatic diseases. The putative targets for miR-676 are PTP1B (protein tyrosine phosphatase, receptor type-B); SMURF2 (SMAD specific E3 ubiquitin protein ligase-2); ANP32B (acidic nuclear phosphoprotein 32 family member-B); TAB2 (TGF-beta activated kinase-1 (MAP3K7) binding protein-2); SCN8A (sodium voltage-gated channel alpha subunit-8) and B3GALNT2 (beta-1,3-N-acetylgalactosaminyltransferase-2). The predicted targets for miR-512 are; HLTF (helicase like transcription factor); PTMA (prothymosin alpha); DDX6 (DEAD-box helicase 6) and ATRX (ATRX, chromatin remodeler). The putative targets for miR-650 are; AXL (AXL receptor tyrosine kinase); EBF3 (EBF transcription factor 3); TRAF4 (TNF receptor associated factor 4); Rac1 (Ras-related C3 botulinum toxin substrate 1); NCOA1 (nuclear receptor coactivator 1) and KLF6 (Kruppel like factor 6). The possible targets for miR-552-3p are; TAOK1 (TAO kinase1); NLK (nemo like kinase); SAMD9L (sterile alpha motif domain containing 9 like); DTWD1 (DTW domain containing 1); TSC22D1 (TSC22 domain family member 1). TSPYL5 (TSPY like 5) and NLK (nemo like kinase).The putative targets for miR-147b are NDRG4 (NDRG family member 4) and BDNF (brain derived neurotrophic factor). Target prediction analysis for all investigated miRNAs is summarized in (Table 3).
Discussion
Recently circulating miRNAs have attracted great
Targets prediction analysis using TargetScan and miRDB prediction programs for the five investigated miRNAs
Targets prediction analysis using TargetScan and miRDB prediction programs for the five investigated miRNAs
attention as stable, promising and reliable biomarkers in several diseases with remarkable changes in their expression levels in response to physiological and pathological conditions [21]. Their expression profiles have been quantified mainly using either RNA-Seq, microarrays or RT-PCR platform [17, 18, 24, 25, 26, 27, 51, 52, 53, 54, 55]. In this study and for the first time, we reported that miR-676 was upregulated in CLD, mild fibrosis (f0-f1), (f1-f3), late fibrosis (f3-f4), cirrhotic and HCC groups. To the best of our knowledge, no study was conducted previously to evaluate the expression profile for miR-676 in liver related disorders. Previous studies reported that it was down-regulated in the tissues of aggressive neuroblastoma tumor phenotype [56], in signet-ring cell carcinomas [38] and in breast cancer [57]. On the contrary, it was upregulated in the plasma of colorectal cancer patients [58]. Based on the TargetScan and miRDB prediction programs, The putative targets for miR-676 were; i) PTP1B which was elucidated for its tumor-suppressing characteristics and is likely found to be upregulated in fibrosis and down regulated in HCC [59, 60, 61]; ii) SMURF2 which was upregulated in fibrosis but down regulated in cirrhosis [62, 63]; iii) ANP32B which was reported to be down regulated in fibrosis and HCC [64]; iv) TAB2 that regulates the activity of TAK1 kinase which is a key modulator in activating NF-
Concerning miR-512-5p, it was down regulated in cirrhotic group, CLD, (f0), advanced fibrosis and also in non-cirrhotic group. On the other hand, it was upregulated in HCC, (f0-f4) and late fibrosis. Previous studies did not focus on miR-512-5p expression profile during various stages of HCV mediated liver disorders. For instance, its levels were reported to be upregulated only in cells of HCC [69], colorectal and liver metastasis [70, 71]. This aligns well with our findings for HCC category. On the contrary, it was reported that miR-512-5p was down regulated in 4 different HCC cell lines [72]. Moreover, miR-512-5p was found to be down regulated in other cancer types such as gastric cancer [73], head and neck squamous cell carcinoma (HNSCC) [74]. TargetScan and miRDB prediction programs revealed the following putative targets for miR-512-5p; i) HLTF which plays an important role in carcinogenesis as it was suggested to be a tumor suppressor gene since it was found to be down regulated in HCC [1]; ii) PTMA which was found to be upregulated in HCC [75]; iii) DDX6 was found to be upregulated in HCV related diseases and in HCC [76]; iv) ATRX was down regulated in HCC [77]. Accordingly, we hypothesized that over expression or down expression of the above mentioned genes might be regulated through miR-512 dysregulation which was upregulated in our HCC patients and down regulated in cirrhosis and CLD groups. Therefore, miR-512 seems to be a possible oncogenic regulator and could serve also as a promising biomarker candidate for HCC, cirrhosis and CLD associated with HCV chronic infection (Table 3).
In this study, miR-650 levels were down regulated in HCC as compared to remaining groups. On the other hand, it was up regulated in CLD, (f1-f2) as compared to healthy controls. This comes in agreement with Abdalla & Haj-Ahmad study (2012) which reported down-regulation of miR-650 in urine samples of HCC-post HCV positive group [78]. This suggests that it may serve as a potential marker for early diagnosis of HCC among high-risk HCV patients. Contradictory to this, miR-650 was found to be upregulated in the tissues of different cancer types such as HCC [30, 79], colorectal cancer [80], prostate cancer [81] and gastric cancer [82].
Based on the TargetScan and miRDB prediction programs, the putative targets for miR-650 are; TRAF4, Rac1, AXL, EBF3 and NCOA1 which are most likely to play an important role in carcinogenesis as it was suggested that their over-expression could promote tumorigenesis. On the other side, KLF6 is considered to be a tumor suppressor gene which was found to be deregulated in HCC. Rac1 and AXL were also found to be upregulated in fibrosis. Accordingly, we hypothesized that the over-expression of TRAF4, Rac1, AXL and NCOA1 along with the down regulation of KLF6 could be regulated by miR-650 [83, 84, 85, 86, 87, 88, 89]. Therefore, miR-650 could be a possible tumor-suppressor regulator and could serve as a potential candidate biomarker for HCC and fibrosis (Table 3).
Concerning miR-552-3p, it was found to be upregulated in cirrhosis, CLD, (f3-f4), (f0-f4) and downregulated in HCC. Its expression in the tissues of several types of cancer was reported to increase in HCC [34], colon cancer [90, 91], ovarian and gastric cancer as well [92, 93, 94]. TargetScan and miRDB software predicted the following putative targets for miR-552-3p; DTWD1, TSC22D1, SAMD9L which play an important role in carcinogenesis as they were reported to be down regulated in tumorigenesis. On the contrary, TSPYL5 and NLK are considered to be oncogenes which were found to be upregulated in HCC [95, 96, 97, 98] meanwhile TAOK1 was found to be upregulated in fibrosis [99]. Accordingly, we hypothesized that the over-expression of TSPYL5 and NLK, and the down regulation of DTWD1, TSC22D1 and SAMD9L could be initiated as a result of down-regulating miR-552-3p [95, 96, 97, 98, 99, 100, 101]. We can therefore hypothesize that miR-552-3p could serve as a possible tumor-suppressor regulator and its measurements could assist in early screening for HCV associated HCC and fibrosis (Table 3).
Concerning miR-147b, it was found to be down regulated in cirrhotic, CLD, (f0-f1), (f0-f2), (f1-f3), (f1-f2), (f3-f4) and (f0-f4). Another study reported significant upregulation of miR-147-b in HCV DLBCL (diffuse large B-cell lymphoma) [102]. Its expression in lung and ovarian cancer was also reported to be upregulated [103, 104]. Contradicting results were reported concerning its expression in CRC [105, 106]. Based on the TargetScan and miRDB prediction programs, the putative targets for miR-147b are NDRG4 which was upregulated in fibrosis and BDNF which was down regulated in non-alcoholic fatty liver fibrosis and was also found to be down regulated in cirrhosis [107, 108, 109]. Accordingly, we hypothesized that the over-expression of NDRG4 and the down regulation of BDNF may result from down-regulation of their regulator miR-147b. MiR-147b could therefore be used as an interesting biomarker for detection of fibrosis and cirrhosis (Table 3).
Our study profiled, for the first time, expression signatures for (miR-676, 512-5p, 650, 552-3p and 147b) in the plasma of Egyptian patients affected with various stages of HCV mediated hepatic disorders. The investigated miRNAs seem to be promising in staging and diagnosis of various fibrotic, cirrhotic and HCC complications among Egyptian patients. They could be utilized as potential non-invasive and reliable biomarkers in HCV mediated hepatic disorders. Further clinical evaluation on larger populations with different HCV genotypes would be beneficial.
Author contribution
AAG, MFI and NIS conceived and designed the experiments. SME, NZ and AY provided part of the samples. HS and AAG performed experimental work and statistical analysis. AAG designed primers of this study. AAG and HS drafted the manuscript. MFI, NIS, HS and AAG critically revised the manuscript.
Informed consent statement
Informed consent was obtained from all subjects involved in the study.
Data availability
This article and its supplementary file include all data generated and analyzed during the study.
Supplementary data
The supplementary files are available to download from http://dx.doi.org/10.3233/CBM-210456.
sj-docx-1-cbm-10.3233_CBM-210456.docx - Supplemental material
Supplemental material, sj-docx-1-cbm-10.3233_CBM-210456.docx
Footnotes
Acknowledgments
Authors thank all participating individuals.
Conflict of interest
All authors declare that they have no conflict of interest.
