Abstract
INTRODUCTION:
Ferroptosis is a recently discovered type of programmed cell death that plays a crucial role in tumor occurrence and progression. However, no prognostic model has been established yet for clear cell renal cell carcinoma (ccRCC) using ferroptosis-related long non-coding RNAs (lncRNAs).
METHODS:
In the present study, lncRNA expression profiles, sex, age, TMN stage, and other clinical data of ccRCC samples were extracted from The Cancer Genome Atlas database. In addition, ferroptosis-related lncRNAs were identified using co-expression analysis, and the risk model was established using Cox regression and least absolute shrinkage and selection operator regression analyses. Log-rank test and Kaplan-Meier analysis were performed to evaluate the predictive accuracy of the risk model for the overall survival (OS) of patients with ccRCC. Moreover, the functional enrichment of ferroptosis-related lncRNAs was performed and visualized using Gene Ontology and Kyoto Encyclopedia of Genes and Genomes.
RESULTS:
Eight prognostic ferroptosis-related lncRNAs were identified, such as LINC01615, AC026401.3, LINC00944, AL590094.1, DLGAP1-AS2, AC016773.1, AC147651.1, and AP000439.2, making up the ferroptosis-related lncRNA risk model. The risk model effectively divided patients with ccRCC into high- and low-risk groups, and their survival time was calculated. The high-risk group showed significantly shorter OS compared to the low-risk group. The nomogram to predict the survival rate of the patients revealed that the risk score was the most critical factor affecting OS in patients with ccRCC. The ferroptosis-related lncRNA risk model was an independent predictor of prognostic risk assessment in patients with ccRCC.
CONCLUSION:
The ferroptosis-related lncRNAs risk model and genomic clinicopathological nomogram have the potential to accurately predict the prognosis of patients with ccRCC and could serve as potential therapeutic targets in the future.
Introduction
Renal cell carcinoma (RCC) is a prevalent malignancy of the urinary system, comprising approximately 3% of the total incidence of malignant tumors in adults [1]. It has several histological subtypes, with clear cell RCC (ccRCC) being the most common, accounting for 75%–80% of the total cases [2]. Despite recent advances in targeted drugs and immunotherapy for patient prognosis, clinical efficacy is limited due to tumor heterogeneity, drug side effects, and adaptive drug resistance, resulting in over 100,000 annual deaths due to RCC progression [3, 4]. Therefore, new therapeutic targets or strategies are needed to improve patient prognosis. Research on the role of ferroptosis in ccRCC is currently underway as the cytoplasm of ccRCC cells is rich in lipids [5].
In comparison to traditional apoptosis or necrosis, ferroptosis is a type of regulated cell death (RCD) triggered by iron-dependent lipid peroxidation and controlled by oxidation and antioxidant systems in the body [6, 7]. Previous research showed that ferroptosis differs from RCD regarding morphological changes, biochemical pathways, and genetic levels. Recent studies have identified various proteins, including glutathione peroxidase 4, heat shock protein
There is growing evidence that ferroptosis can suppress tumor growth and progression and may hold potential as cancer therapy [8, 9, 10]. However, the expression of ferroptosis-related genes (FRGs) in ccRCC development has not been fully explored [11].
Long non-coding RNA (lncRNA) is a type of RNA that does not code for proteins and has a length exceeding 200 nucleotides. It regulates gene expression in a cell type-specific, developmental stage-specific, and cell function-specific manner [12, 13]. Abnormal lncRNA expression can disrupt the regulation of cell growth, proliferation, development, cell cycle, or function, such as ferroptosis, glycolysis, cell differentiation, and DNA repair, leading to various diseases [14, 15, 16]. Therefore, investigating abnormal lncRNA expression in ccRCC may identify potential biomarkers and novel therapeutic targets for treating patients with ccRCC.
The present study aimed to develop a novel prognostic risk model for ccRCC using The Cancer Genome Atlas (TCGA) database, focusing on ferroptosis-related lncRNAs as potential predictors of patient prognosis.
Materials and methods
Selection of ferroptosis-related lncRNA using the Lasso model. (A) Lasso coefficient values of 16 ferroptosis-related lncRNAs in ccRCC. The vertical dashed lines are at the optimal log (lambda) value. (B) Profiles of the Lasso coefficients. ccRCC, clear cell renal cell carcinoma; lncRNAs, long non-coding RNAs.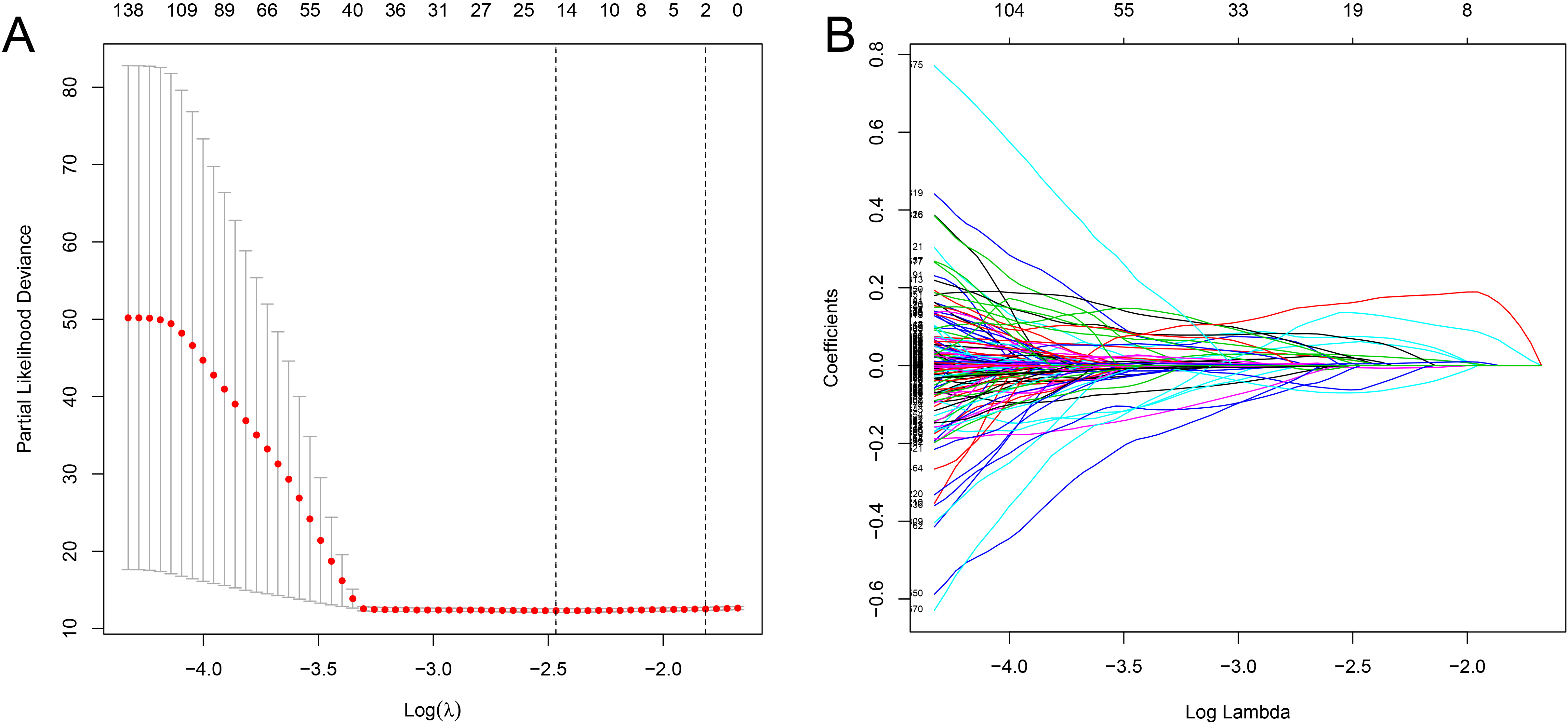
Identification of ferroptosis-related lncRNAs
A total of 14,142 lncRNAs were identified from the TCGA Kidney Renal Clear Cell Carcinoma transcriptome dataset, and 174 FRGs were extracted from the FerrDb database. Hence, 1628 ferroptosis-related lncRNAs were finally obtained through correlation analysis.
Results of the multivariate Cox analysis of long non-coding RNAs based on Cancer Genome Atlas Kidney Renal Clear Cell Carcinoma data
Results of the multivariate Cox analysis of long non-coding RNAs based on Cancer Genome Atlas Kidney Renal Clear Cell Carcinoma data
Co-expression network and Sankey diagram of prognostic ferroptosis-related lncRNAs. (A) Co-expression network between the prognostic lncRNAs and ferroptosis-related genes in ccRCC. Grass-green diamond nodes represent prognostic lncRNAs, and red nodes represent ferroptosis-related genes. (B) The Sankey diagram shows the association among prognostic ferroptosis-related lncRNAs, ferroptosis-related genes, and risk types. ccRCC, clear cell renal cell carcinoma; lncRNAs, long non-coding RNAs.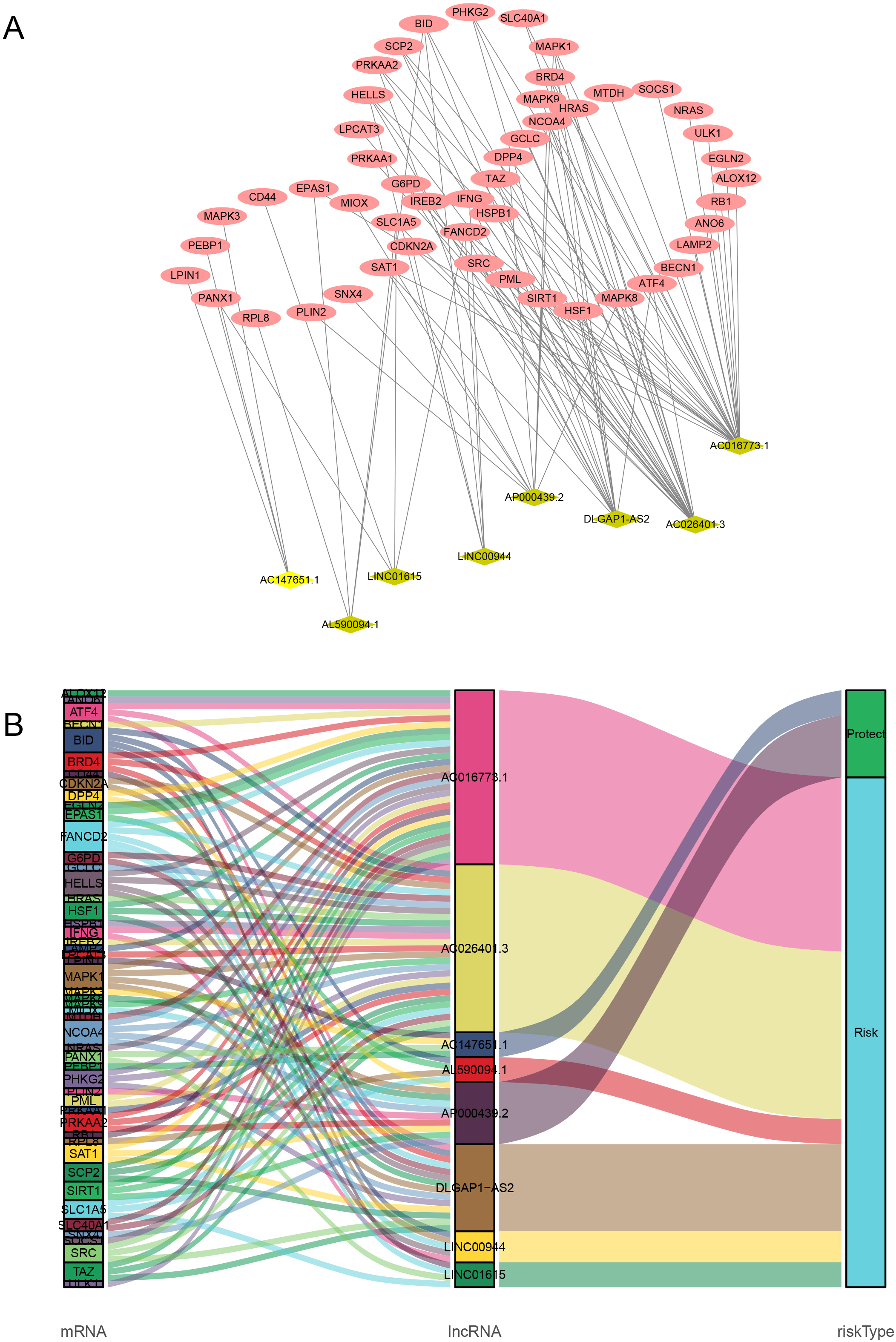
Kaplan-Meier survival curves of the eight prognostic ferroptosis-related lncRNAs. Six of them (LINC01615, AC026401.3, LINC00944, AL590094.1, DLGAP1-AS2, and AC016773.1) were unfavorable factors, and two (AC147651.1 and AP000439.2) were protective factors for ccRCC. ccRCC, clear cell renal cell carcinoma; lncRNAs, long non-coding RNAs.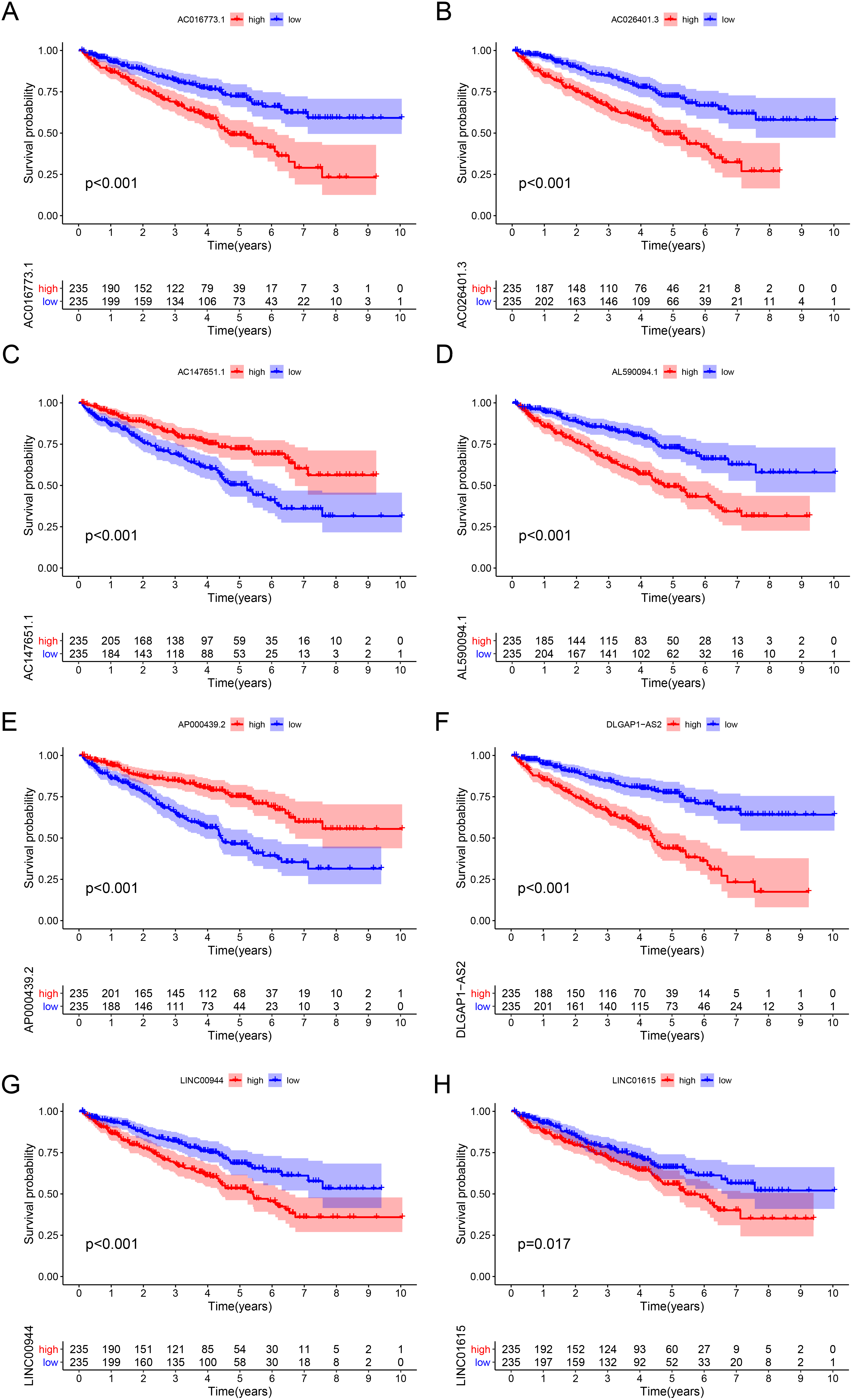
Kaplan-Meier (KM) survival curve of the risk score based on the eight ferroptosis-related lncRNAs. (A) KM survival curve based on TCGA data. (B) KM survival curve based on ICGC data.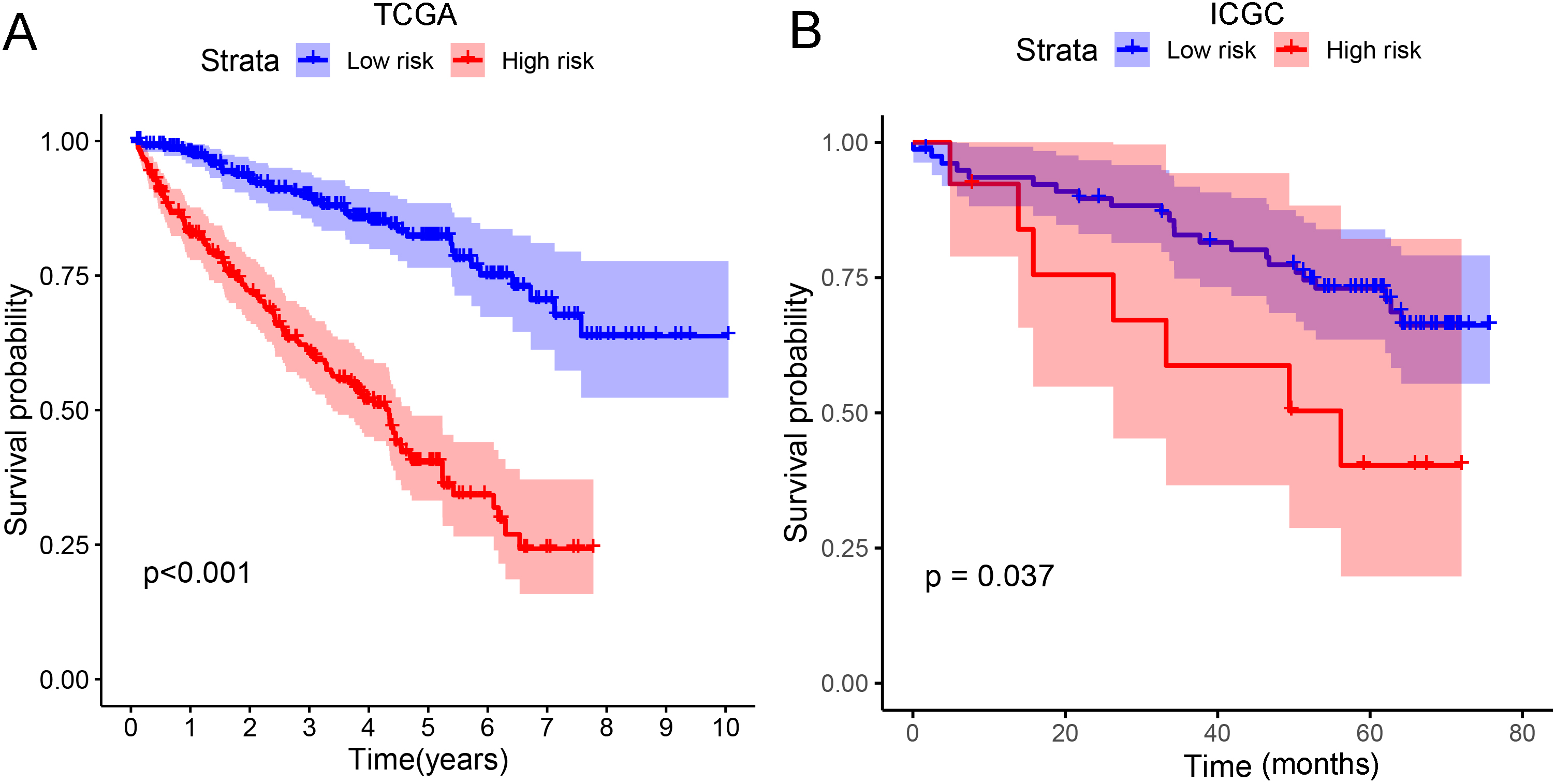
A total of 682 ferroptosis-related lncRNAs with prognostic values for ccRCC were identified using univariate Cox analysis (
Risk score
Prognostic influence of the established risk model
The survival probability of the low-risk group was significantly higher than that of the high-risk group (
Clinical characteristics and risk scores of clear cell renal cell carcinoma using multivariate Cox regression
Clinical characteristics and risk scores of clear cell renal cell carcinoma using multivariate Cox regression
Analysis of ferroptosis-related lncRNA signature for patients with ccRCC. (A) Risk scores between the high and the low-risk groups. (B) Survival time of the patients. (C) Heat map of eight ferroptosis-related lncRNA expressions. The color from green to red shows a rising tendency from low to high levels. ccRCC, clear cell renal cell carcinoma; lncRNAs, long non-coding RNAs.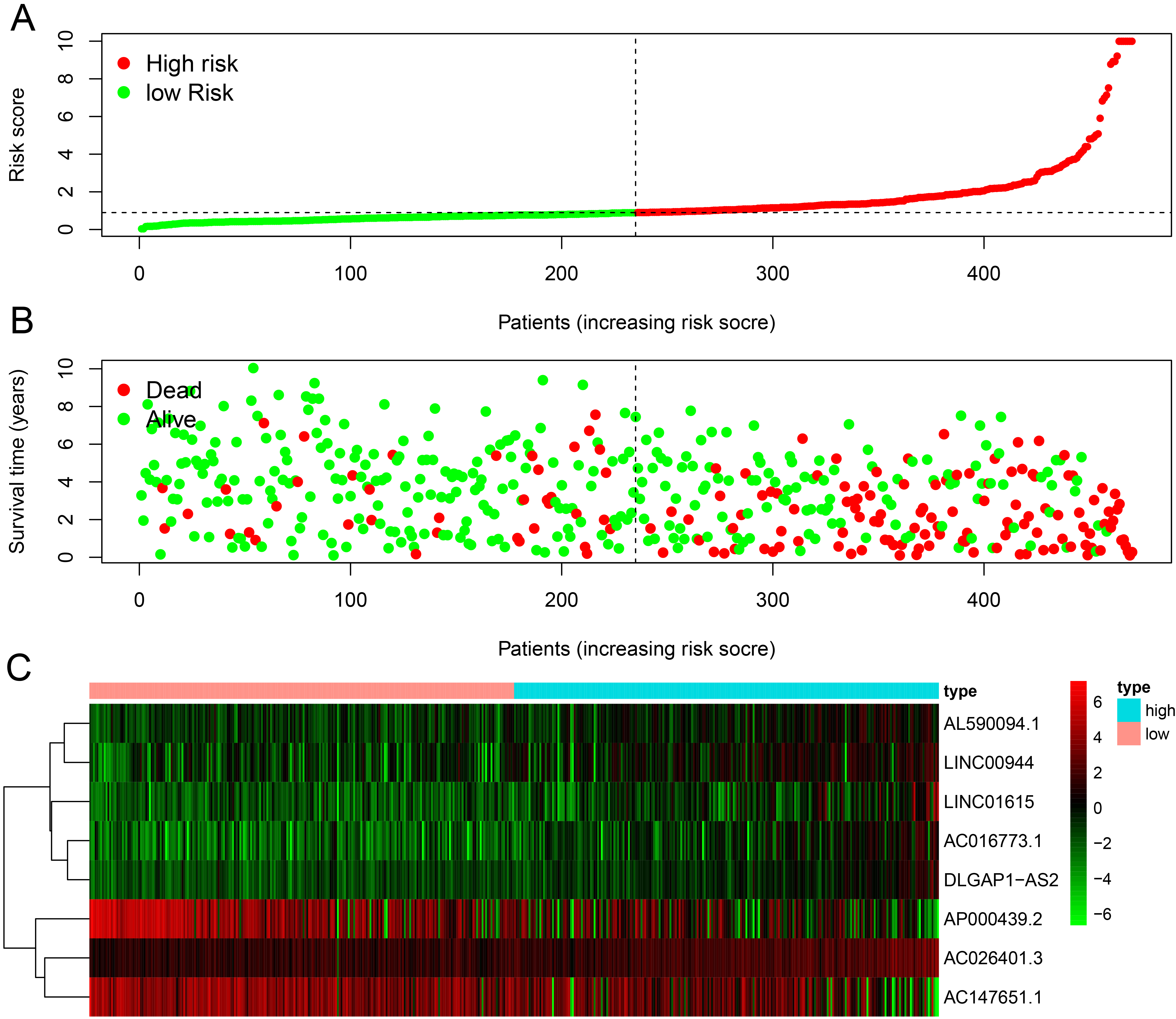
Prognostic indicators based on ferroptosis-related lncRNAs showed significant predictive performance. The forest plots for the univariate (A) and multivariate (B) Cox regression analyses in ccRCC. ccRCC, clear cell renal cell carcinoma; lncRNAs, long non-coding RNAs.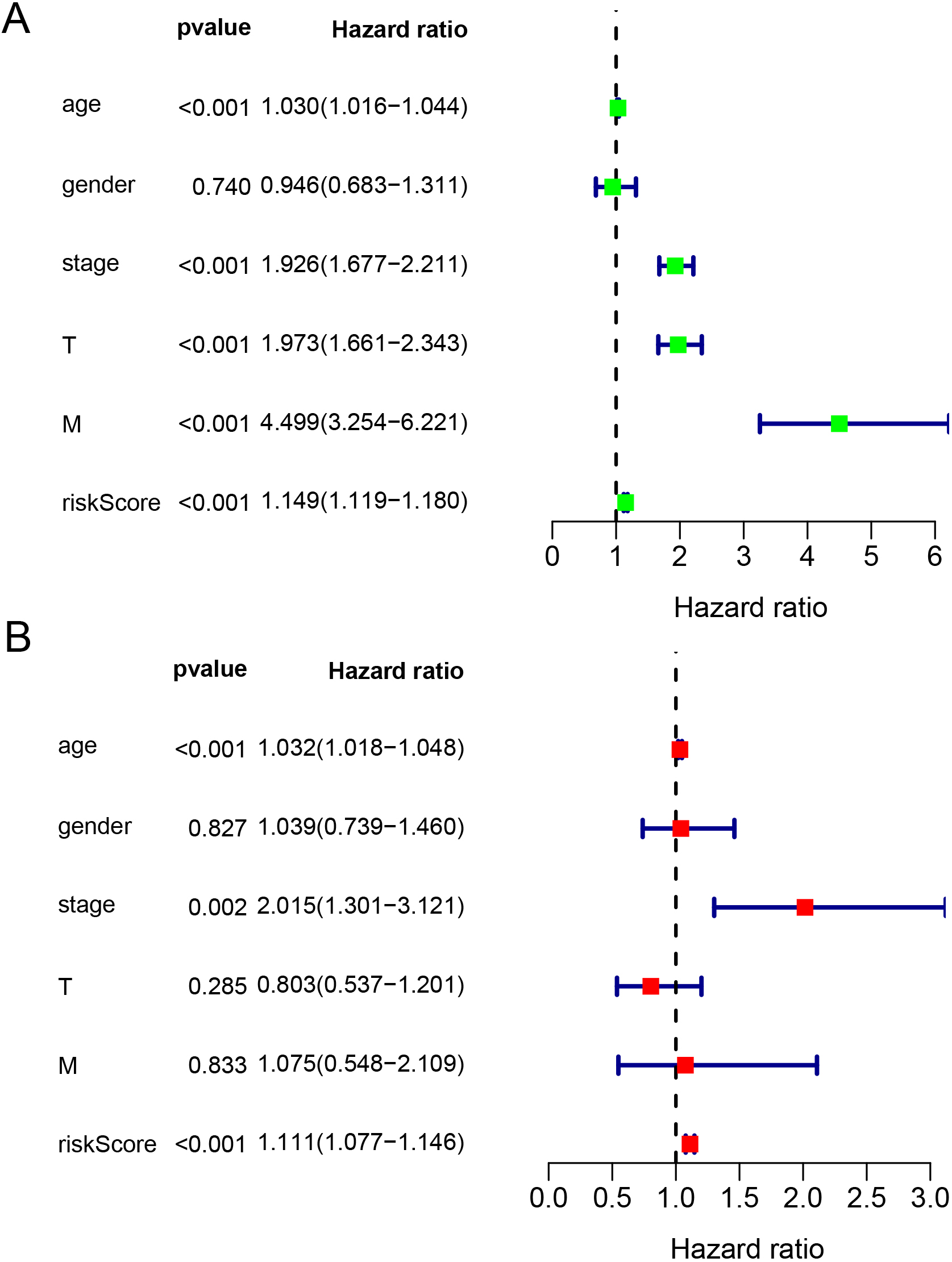
Areas under the receiver operating characteristic curve at 1, 3, and 5 years.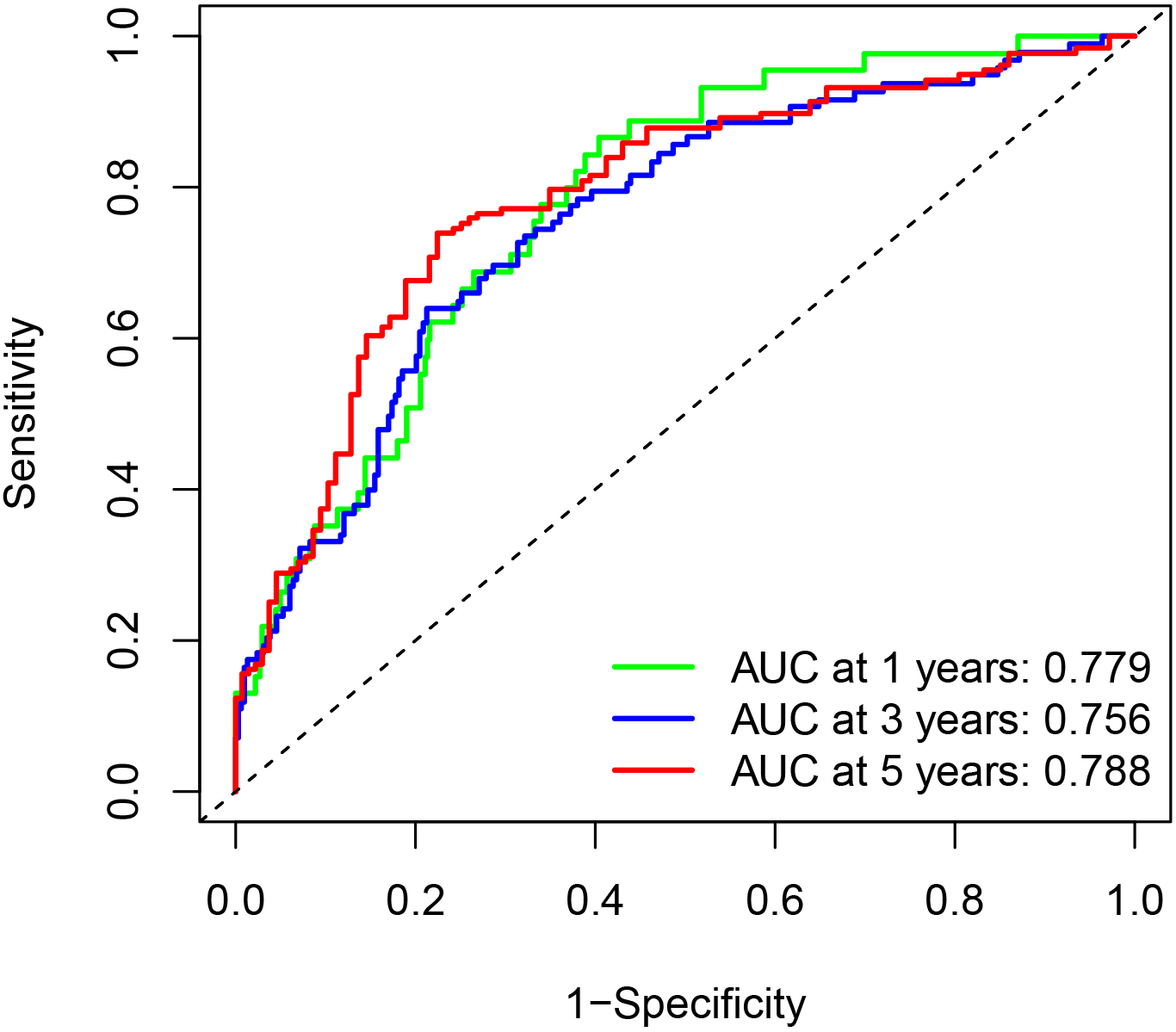
Evaluation of prognostic models based on the eight ferroptosis-related lncRNAs. (A) Nomogram of 1-, 3- or 5-year OS based on the risk score, age, and TNM stage. (B) ROC curve analysis based on the risk score and clinicopathological parameters. (C) Calibration plots for evaluating the agreement between the predicted and actual OS for the prognosis model. The 45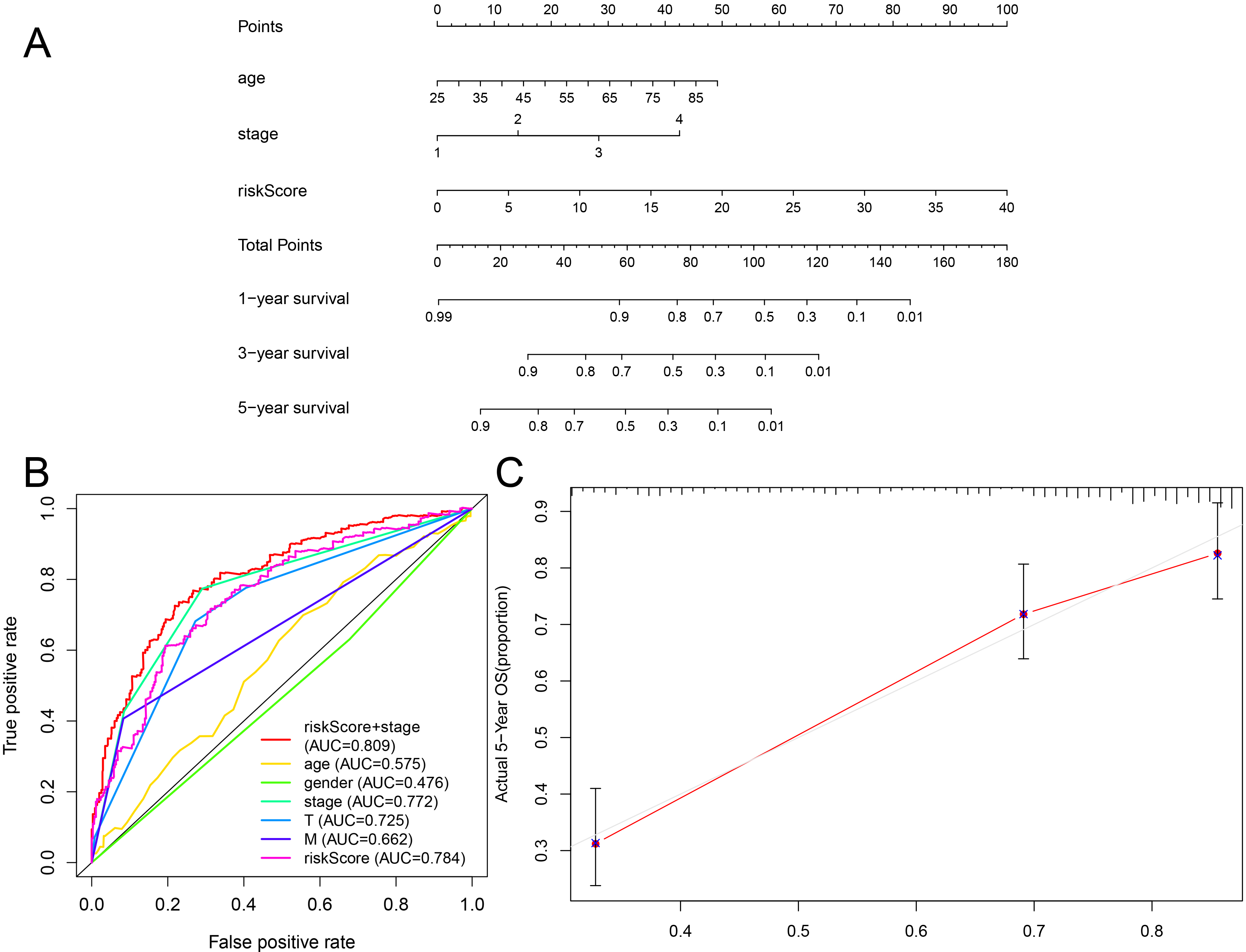
In the univariate Cox regression analysis, age, stage, risk score, T stage, and M stage were identified as indicators with predictive ability. The risk score demonstrated a hazard ratio (HR) of 1.149 (95% confidence interval [CI]: 1.119–1.180,
The AUC value of the risk score in the ROC curve for the 5-year survival rate was 0.784, indicating its strong prognostic potential for patients with ccRCC (Fig. 8B). The combination of the risk score and TNM stage resulted in an improved AUC value of 0.809, suggesting that the risk score could provide valuable complementary information to the TNM stage. Factors influencing the risk score were analyzed, revealing no significant correlation between age and gender and risk score (
GSEA
The GSEA software was used to analyze the functional enrichment of lncRNAs. In the GO analysis, the ferroptosis-related lncRNAs were mainly associated with biological processes such as lipid oxidation, ammonium ion metabolic, hormone-mediated signaling pathway, and endothelium development (Fig. 9A, Table S3). The KEGG analysis revealed that the ferroptosis-related lncRNAs were predominantly involved in pathways related to peroxisome, fatty acid metabolism, RCC, prostate cancer, homologous recombination, and primary immunodeficiency. These pathways may have significant associations with tumorigenesis and cancer progression (Fig. 9B, Table S4).
Clinical influences of the risk score signature for Cancer Genome Atlas Kidney Renal Clear Cell Carcinoma data
Clinical influences of the risk score signature for Cancer Genome Atlas Kidney Renal Clear Cell Carcinoma data
Results of the functional analysis based on ferroptosis-related lncRNAs. (A) KEGG enrichment analysis of the low-risk group. (B) KEGG enrichment analysis of the high-risk group. (C) GO enrichment analysis of the low-risk group. (D) GO enrichment analysis of the high-risk group. GO, Gene Ontology; KEGG, Kyoto Encyclopedia of Genes and Genomes; lncRNAs, long non-coding RNAs.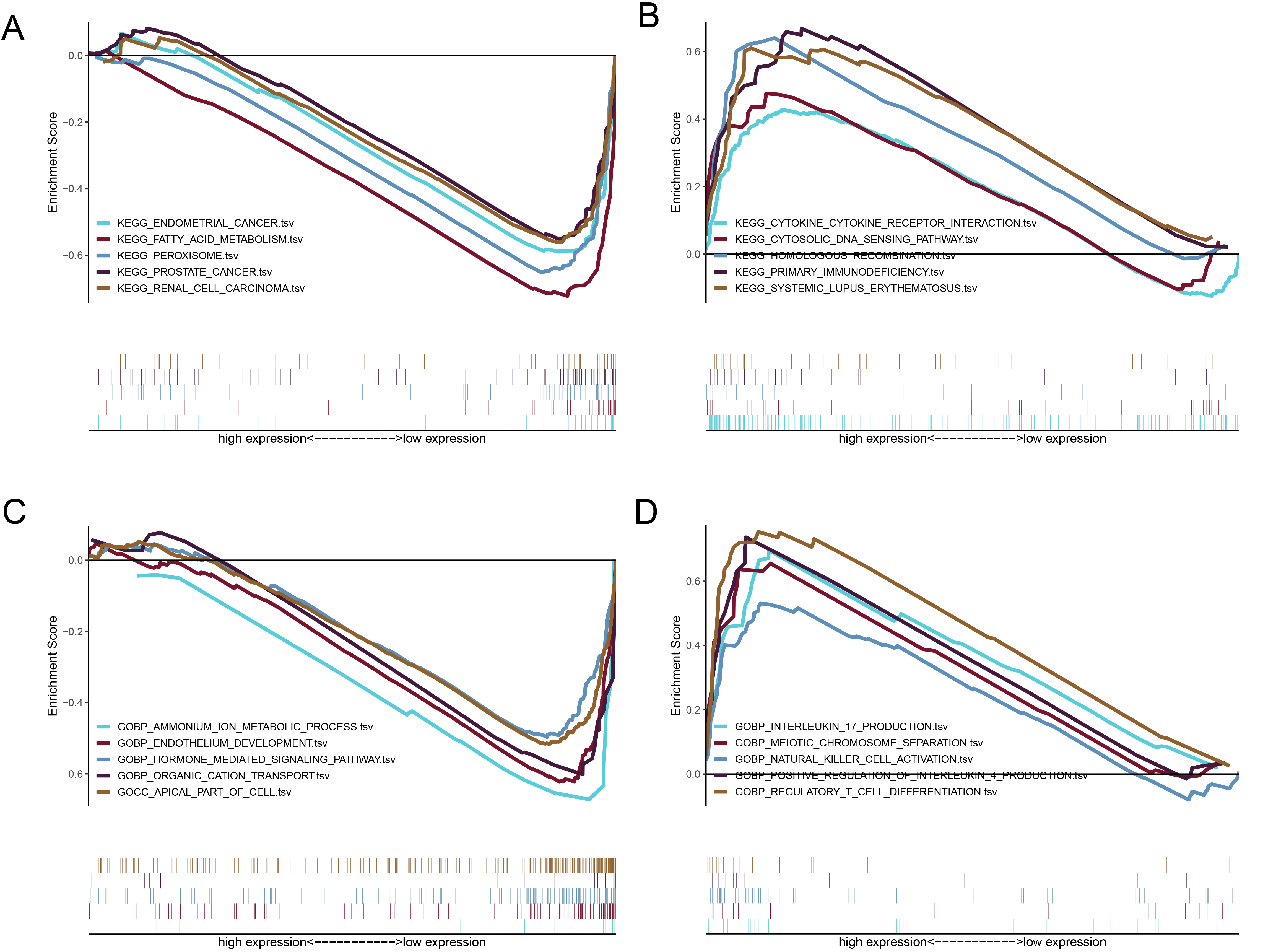
RCC is a prevalent malignancy in adults, with ccRCC being the most common subtype insensitive to radiotherapy and chemotherapy. Unfortunately, ccRCC is often unresponsive to targeted therapy and immunotherapy, and the prognosis is generally poor [21]. Current prognostic markers, such as Eastern Cooperative Oncology Group Performance Status score, tumor size, histological tumor type, and TNM stage, have limited accuracy and cannot assess genetic factors [22]. Existing prognostic indicators for RCC, including immunity and autophagy-related prognostic signatures, and for ccRCC are also insufficient, highlighting the urgent need for new molecular diagnostic and prognostic markers to guide ccRCC management [22]. To date, no prediction signature of ferroptosis-related lncRNAs has been reported.
In this study, eight ferroptosis-related lncRNAs were identified using Cox regression and Lasso regression analyses and were used to create a co-expression network with FRGs. Of these, six lncRNAs, such as LINC01615, AC026401.3, LINC00944, AL590094.1, DLGAP1-AS2, and AC016773.1, were found to be unfavorable factors, while two, such as AC147651.1 and AP000439.2, were protective factors. As lncRNAs are known to be closely related to tumor occurrence and progression, these findings have significant implications for the understanding and management of cancer.
DLGAP1-AS2 is a cytoplasmic antisense RNA2 that regulates cell proliferation, differentiation, migration, and apoptosis by modulating downstream YAP1. High expression of lncRNADLGAP1-AS2 is associated with poor prognosis in patients with glioma. Knockout of the DLGAP1-AS2 gene in glioma cells can significantly inhibit tumor cell proliferation and migration, induce apoptosis, and improve patient prognosis [23]. DLGAP1-AS2 reportedly also inhibits the migration and invasion of hepatocellular carcinoma cells by regulating the miR-154-5p methylation pathway [24]. In addition, it plays a crucial role in the development of Wilms’ tumor, cholangiocarcinoma, lung disease, and glycolysis in RCC [25, 26, 27].
The mechanism of LINC00944 in regulating ccRCC occurrence and development is unclear. However, it has been linked to the biological behavior of breast cancer. A study by De Santiago et al. found that LINC00944 expression is associated with immune-signaling pathway, apoptosis promotion, age at diagnosis, tumor size, estrogen receptor, and prognosis in patients with breast cancer [28].
AC016773.1 has been found to play a crucial role in the development and progression of various cancers, including laryngeal squamous cell carcinoma (LSCC). In a study by Kong et al., a competitive endogenous RNA network was constructed to compare expression profiles of mRNA, lncRNA, and miRNA in LSCC and adjacent normal tissues, showing a close connection between AC016773.1 and LSCC occurrence and development [29]. Similarly, a study on RCC also identified AC016773.1 as crucial in tumor biology [30].
Previous studies have reported that LINC01615 plays a role in the extracellular matrix, which has been linked to the progression of hepatocellular carcinoma and prognosis of patients with head and neck squamous cell carcinoma [31, 32]. However, no study has examined the prognostic potential of LINC01615 in RCC.
In recent studies, the prognostic potential of AC147651.1, AL590094.1, and AP000439.2 had not been assessed, but it has been found that these lncRNAs can predict the survival prognosis of patients with tumors using several prognostic models [33, 34]. These findings are consistent with the study’s results, which confirm that ferroptosis-related lncRNAs play a vital role in regulating tumor occurrence and progression. Hence, further studies are required to clarify its specific regulation mechanism. The risk score method was used to divide the cases into high and low-risk groups, and the risk score was confirmed as an independent factor in predicting the prognosis of patients with ccRCC. Furthermore, the accuracy of nomogram prediction was verified using discrimination and calibration plots, and the AUC value of the multi-factor ROC curve was 0.784, indicating high predictive accuracy. These results can provide a personalized assessment of the survival benefits for patients with ccRCC and guide personalized management decisions.
The GSEA revealed that the high and low-risk groups were enriched in pathways related to cell development and metabolism, including fatty acid metabolism, peroxisome, and endothelium development. In the low-risk group, pathways related to tumorigeneses, such as prostate cancer, RCC, and endometrial carcinoma, were significantly enriched. These results suggest that ferroptosis-related lncRNAs play a crucial role in ccRCC occurrence, providing potential therapeutic targets for ccRCC treatment.
In the present study, a risk model consisting of eight ferroptosis-related lncRNA was constructed that showed high prediction accuracy for the prognosis of patients with ccRCC. However, the study has some limitations. First, bioinformatics methods alone were used for data analysis, which was not verified by further experiments. Second, the lack of some clinical variables, such as lymph node metastasis data, may limit the strength of these results. Therefore, additional experimental studies are required to validate the predictive value of the risk model and explore its specific regulation mechanism.
Conclusion
In conclusion, the study demonstrated that the risk score model has an independent prognostic value for patients with ccRCC. The genomic clinicopathological nomogram can accurately predict the prognosis of these patients. Moreover, the eight ferroptosis-related lncRNAs identified in the study may serve as potential therapeutic targets for ccRCC.
Data availability
Part of the data used for the analysis is included in this paper and its supplementary information file. The rest of the data can be obtained from the author. The single-cell sequencing datasets can be found in online repositories (
Authors’ contributions
Conception: Xuebao Xiang, Yi Guo, Zhongyuan Chen, Fangxin Zhang, and Yan Qin.
Interpretation or analysis of data: Xuebao Xiang, Yi Guo, and Zhongyuan Chen.
Preparation of the manuscript: Xuebao Xiang, Fangxin Zhang, and Yan Qin.
Revision of important intellectual content: Xuebao Xiang, Yi Guo, Fangxin Zhang, and Yan Qin.
Supervision: Xuebao Xiang, Yi Guo, Zhongyuan Chen, Fangxin Zhang, and Yan Qin.
Funding
This work was supported by the Natural Science Foundation of Guangxi (No. Z20200351).
Supplementary data
The supplementary files are available to download from http://dx.doi.org/10.3233/CBM-210445.
sj-docx-1-cbm-10.3233_CBM-210445.docx - Supplemental material
Supplemental material, sj-docx-1-cbm-10.3233_CBM-210445.docx
Footnotes
Acknowledgments
We thank the staff at the Centre for Genomic and Personalized Medicine, Guangxi Medical University, for supporting our research.
Conflict of interest
There is no conflict of interest.
