Abstract
BACKGROUND:
Molecular and clinical diversity of neuroblastomas is notorious. The activating TERT rearrangements have been associated with dismal prognosis. Suppression of miR-128-3p may complement and enhance the adverse effects of TERT overexpression.
OBJECTIVE:
The study aimed at evaluation of prognostic significance of the miR-128-3p/TERT expression in patients with primary neuroblastoma.
METHODS:
RNA samples isolated from fresh-frozen tumor specimens (
RESULTS:
Both TERT overexpression and miR-128-3p downregulation were independently associated with superior rates of adverse events (
CONCLUSIONS:
Combined expression levels of miR-128-3p and TERT represent a novel prognostic biomarker for neuroblastoma.
Introduction
Neuroblastoma is the most common extracranial solid malignancy of childhood. Its clinical behavior varies widely from the relatively benign forms prone to spontaneous regression or maturation to the highly aggressive tumors with high relapse rates, treatment resistance and dismal outcomes. Diverse genetic characteristics of neuroblastoma cells determine the clinical heterogeneity and can be utilized as prognostic markers. Common aberrations used for the prognosis and clinical stratification of neuroblastoma include MYCN amplification (MNA) and chromosome 1p and 11q deletions [1, 2]. The list of prognostically significant copy number alterations and gene expression signatures is being constantly updated with the use of high-throughput molecular approaches [3, 4].
Activating rearrangements in TERT promoter have been recently associated with aggressive neuroblastoma [5]. These aberrations cause telomerase overexpression leading to the abnormal telomere maintenance; a similar effect is conferred by MYCN amplification [5]. When coupled with pathogenic mutations in the components of RAS and TP53 pathways, TERT aberrations determine extremely high risks; in the absence of complementing mutations, the abnormal telomere maintenance is still an unfavorable prognostic factor. Strong association of TERT rearrangements with a number of unfavorable genomic and transcriptomic signatures has been demonstrated [6].
The protein-encoding gene expression profiling of neuroblastoma revealed distinct mRNA signatures associated with favorable and adverse courses of the disease [7, 8, 9]. Significance of the matching microRNA expression profiles remains ambiguous. Preclinical findings indicate that microRNAs may promote (miR-332-3p, miR-3934-5p, miR-146b) as well as suppress (miR-15a-5p, miR-15b-5p, miR-16-5p, miR-34a, miR-145, miR-149, miR-429, miR-490-5p) the viability of neuroblastoma cells with corresponding influence on the survival [10, 11, 12, 13, 14, 15, 16, 17, 18].
Small non-coding RNA hsa-miR-128-3p (miR-128-3p, formerly known as hsa-miR-128a) is a candidate tumor suppressor microRNA. In normal conditions its expression is restricted to nervous tissue [19] but miR-128-3p is abundantly expressed in various malignancies including head and neck squamous carcinoma, hepatocellular carcinoma, lung cancer, gastric cancer and glioblastoma [20, 21, 22, 23, 24]. The mature isoform of miR-128-3p specifically targets mRNA molecules of cancer-related genes involved in cell cycle and apoptosis control (E2F3, E2F6, E2F7, BMI1, SPTAN1, BAX, BCL2), JAK-STAT and receptor tyrosine kinase pathways (JAK1, PDGFRA, PDGFRB, NTRK2, NTRK3, RPS6KB2) and angiogenesis (VEGFB, VEGFC) [20, 22, 23, 25, 26, 27].
A number of findings suggest that miR-128-3p may act as a tumor suppressor in neuroblastoma cells. For instance, miR-128-3p inhibits cellular motility and invasiveness and induces proliferation arrest and apoptosis by targeting RELN, DCX and HOXA7 genes [28, 29]. Bao et al. identified SNHG16 (long non-coding RNA associated with neuroblastoma aggressiveness) as a direct inhibitor of miR-128-3p. Expression of miR-128-3p is significantly reduced in neuroblastoma specimens (compared to the normal tissue of adrenal medulla or sympathetic ganglia) while being upregulated during the retinoid acid-induced differentiation of neuroblastoma cells [29]. According to Roy et al., miR-128-3p can block WNT signaling by targeting its key components (including WNT5B, DVL and LEF1) [30].
At the same time, the possible role of miR-128-3p in neuroblastoma progression is ambiguous. Other reported targets of miR-128-3p include the truncated isoform of NTRK3, which participates in membrane remodeling and reorganization of the cytoskeleton but cannot support the transduction of proliferative signals like the full-length isoform [25]. Moreover, overexpression of miR-128-3p in SH-SY5Y neuroblastoma cell cultures leads to upregulation of BCL2, ETS1, ETV4 and CCND2 resulting in the immature phenotype (rounded cell bodies, shorter neurites and smaller cell size) and apoptosis inhibition thus acting as an oncogenic event [25].
Clinical significance of miR-128-3p expression in neuroblastoma was evaluated just once by Evangelisti et al. on the limited cohort of patients. The study revealed the association of high miR-128-3p levels with favorable indicators such as early age at diagnosis, favorable Shimada category, localized and 4S stages and the absence of MYCN amplification [28].
In this study, we evaluate clinical significance of hsa-miR-128-3p expression in primary neuroblastomas with diverse molecular contexts including known prognostic factors, including overexpression of TERT.
Nucleotide sequences of primers and probes used for reverse transcription reaction and qPCR
Nucleotide sequences of primers and probes used for reverse transcription reaction and qPCR
Experimental subjects
The study was approved by the Local Ethics Committee of Regional Children’s Hospital, Yekaterinburg, Russian Federation. The written informed consent was obtained from parents or legal representatives of all patients. Totally, 103 patients admitted to the Pediatric Oncology and Hematology Center during the period 02/2005–04/2014 were enrolled in the study. The inclusion criteria were histopathologically verified neuroblastoma and the availability of fresh-frozen tumor samples. Of the studied cohort, 68 (66%) patients had localized tumors, 26 (25%) patients were diagnosed with stage 4 disease and 9 (9%) had stage 4S disease as defined by INSS criteria. In terms of molecular characterization, 16 (15%) patients harbored MYCN amplification and 22 (21%) had chromosome 1p deletions. Forty-six (45%) patients were under 12 months of age at the time of diagnosis. All patients were managed according to the NB2004 protocol of German Society of Pediatric Oncology and Hematology, (GPOH), with 61 (59%) patients assigned to the observation group, 11 (11%) to the intermediate risk group and 31 (30%) to the high-risk group.
Molecular genetic techniques
Total RNA and genomic DNA were extracted from fresh-frozen tumor tissues with TRI-reagent (Molecular Research Center, USA) and Wizard Genomic DNA Purification Kit (Promega, USA), respectively. In a pilot series of experiments, the RNA samples were enriched with a pool of short fragments extracted from 10 tissue specimens using mirVana miRNA Isolation Kit (Ambion, USA). As the enrichment showed no significant influence on the miR-128-3p quantification, all subsequent experimental workflow was conducted on the total RNA. The MMLV reverse transcriptase-mediated first strand cDNA synthesis was carried out using 1
To validate the results of miR-128-3p quantification, independent analysis was implemented using molecular “barcodes” (NanoString) technology with nCounter
MNA and 1p36 deletions were assessed by fluorescence in situ hybridization (FISH) with N-MYC, 2p24.3, Red and LAF4, 2q11.2, Green probe combination (Cytocell, UK); and SRD (CHD5), 1p36.31, Red and 1qter, 1q44, Green combination (Cytocell).
Statistics
Statistical analysis was performed in XLstat software (Addinsoft, USA). The quantitative data distribution fitting was evaluated by Shapiro-Wilk, Anderson-Darling and Lilliefors tests. The measured values of TERT and miR-128-3p expression did not follow a normal distribution (all
Results
The normalized expression of TERT ranged from
The normalized expression of miR-128-3p ranged from
As patients with MNA tumors have the worst prognosis, to avoid the bias in subsequent Cox regression analysis, the patients with MNA were sorted out into a separate group (
patients with TERT expression below patients with TERT expression above patients with TERT expression below patients with TERT expression above
Group 1 consisted predominantly of low-risk patients with localized tumors, whereas groups 2–4 consisted of patients with diverse risk status harboring localized or metastatic tumors. 1p deletion was distributed randomly across the groups showing no association with TERT and miR-128-3p expression levels. In general, expression of the studied markers and the presence of MNA were major determinants of survival rates for the entire cohort (Table 2).
Risk indicators (EFS, CIRP) with molecular and clinical characterization of tumors for the cohort stratified by MYCN amplification (MNA) and TERT/miR-128-3p expression in non-MNA cases
Risk indicators (EFS, CIRP) with molecular and clinical characterization of tumors for the cohort stratified by MYCN amplification (MNA) and TERT/miR-128-3p expression in non-MNA cases
The patients with unaltered levels of TERT mRNA and miR-128-3p (group 1) had favorable outcomes, with only two patients of this group experiencing disease progression (EFS 0.91
Comparisons of survival probability between the analyzed groups (
Prognostic significance of TERT/miR-128-3p expression levels (groups 1–4, lines 1–4) and MYCN amplification (lines 5) assessed by EFS (A) and CIRP (B).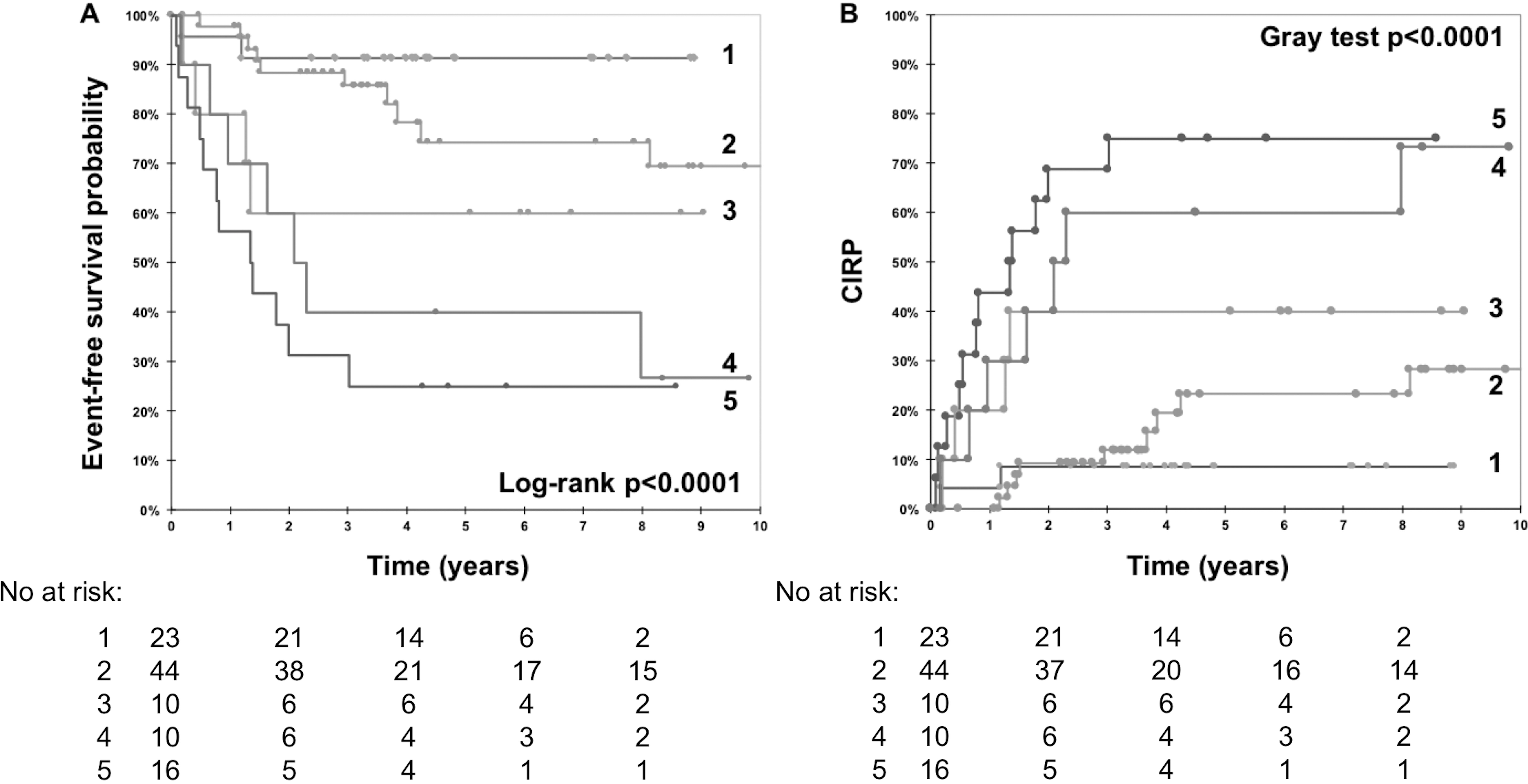
Cox regression analysis of survival probability for the entire cohort
Cox regression analysis of survival probability for the non-MNA group (single-copy MYCN patients)
Bland-Altman plot showing agreement of the miR-128-3p expression obtained by qPCR and NanoString assay. Horizontal axis represents mean of miR-128-3p normalized expression assessed by two techniques, vertical axis shows difference between the methods; line A represents the mean difference (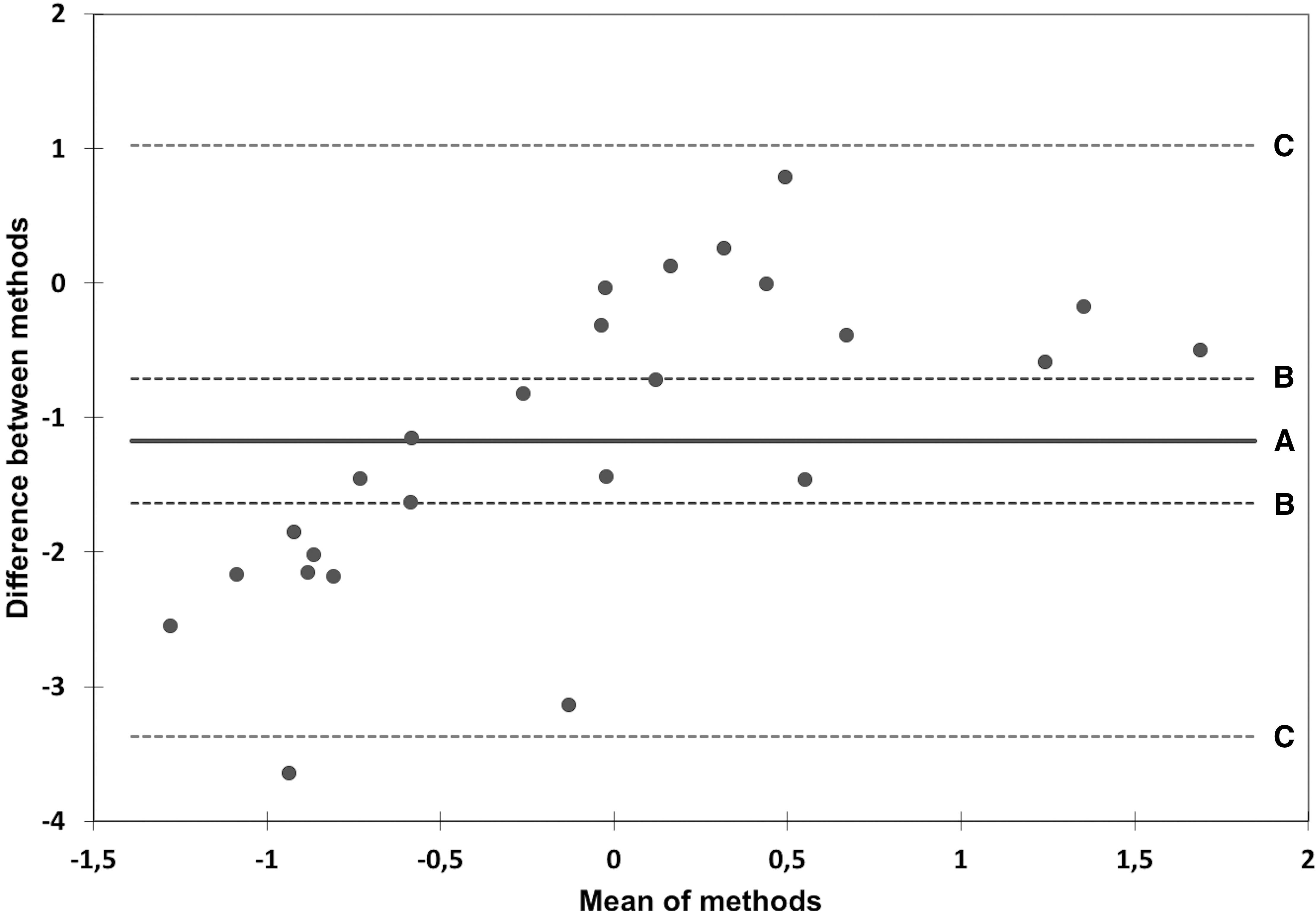
The univariate analysis of EFS in the entire cohort confirmed the adverse prognostic significance for a number of risk factors, both clinical (stage 4 of the disease, age at the diagnosis over 18 months and the assignment to high-risk group according to the NB2004 stratification scheme) and molecular (MNA, TERT/miR-128-3p profiles). Noteworthy, the loss of miR-128-3p expression was the only significant risk factor identified by the multivariate analysis, HR 2.35,
According to the recent mechanistic classification of clinical phenotypes in neuroblastoma introduced by Ackermann et al., abnormal telomere maintenance via TERT activation or alternative pathway is strongly associated with the tumor aggressiveness. When complemented by pathogenic mutations in RAS and/or TP53 pathway components, the abnormal telomere lengthening determines the most adverse clinical course. Moreover, it has been linked to other known risk factors including segmental chromosomal aberrations and unfavorable gene expression signatures [6].
A number of preclinical findings suggest that miR-128-3p acts as a tumor suppressor by switching off the expression of cancer-related genes in various malignancies [20, 21, 22, 23, 24]. In neuroblastoma models, miR-128-3p specifically reduced the expression of oncogenic transcription factors, regulators of cell migration and components of RAS and WNT signaling, thus inhibiting the viability and metastatic potential of tumor cells [28, 29, 30]. However, the opposing line of evidence points to oncogenic effects of miR-128-3p overexpression, for example, the immature phenotypes, increased proliferation rates and apoptosis inhibition in neuroblastoma cell lines [25]. The only reported evaluation of clinical significance of miR-128-3p expression in neuroblastoma supports the association of its high levels with favorable course of the disease [28].
Our data demonstrate synergistic impact of TERT overexpression and miR-128-3p downregulation on the disease-specific survival of patients with neuroblastoma. The patients manifesting both alterations showed low EFS and high disease progression rates resembling the effects of MYCN amplification. By contrast, preserved miR-128-3p expression in the absence of TERT activation was associated with favorable outcomes. Disparate manifestations of either TERT upregulation or the loss of miR-128-3p expression were associated with intermediate EFS and CIRP values. The conventional risk factors showed no extra-significance for survival to the TERT/miR-128-3p expression modes. In the multivariate analysis, these factors were outperformed by the level of miR-128-3p expression as a covariate. The obtained qPCR measurements of miR-128-3p expression showed agreement with the results of NanoString assay, which is an excellent independent tool for the validation of qPCR data due to the absence of reverse transcription, and amplification reactions in the protocol and consequently the lack of bias associated with these steps.
Characteristic gene expression signatures identified by different research groups enable the use of transcriptomic approaches for the prediction of neuroblastoma outcomes. MicroRNAs, a subtype of small non-coding RNA molecules, target a variety of transcripts and work as pivotal negative regulators of gene expression at the post-transcriptional level. Noteworthy, 16 out of 194 transcripts included in the SVM_th10 neuroblastoma classifier developed by Oberthuer at al. [8] are predicted targets of miR-128-3p listed in the miRDB target prediction database [26]. Of those 16 transcripts, PHLDA2, AGRN, DIRAS1, TCEANC2, NCAN, NXT2, KPNB1, RLIM, MED14 and ZBTB20 have been positively associated with adverse phenotypes of neuroblastoma, including drug resistance, whereas high expression levels of BAZ2B, NCOA7, WSB1, RFX3, PGM2L1 and CNR1 correlate with favorable clinical course [31, 32, 33, 34, 35, 36, 37, 38, 39].
Considering that microRNAs naturally inhibit gene expression at the post-transcriptional level, direct binding of the preserved miR-128-3p to TERT mRNA would be a straightforward explanation for the observed inhibition of TERT. However, neither predicted nor de facto targets of miR-128-3p include TERT. At the same time, several positive regulators of telomerase activity are definitely controlled by miR-128-3p. These include BMI1 capable of inducing telomerase activity in different cell types, normal or pathologically altered. For instance, hematopoietic stem cells (HSCs) harbor high expression levels of BMI1, which decline upon HSC differentiation towards partially committed precursors [40]. Furthermore, BMI1 has been shown to promote self-renewal of neural stem cells (and HSCs as well) by tuning the balance between differentiation and the undifferentiated cell growth [41, 42]. Upregulation of TERT by BMI1 has been observed in breast and ovarian cancers [43, 44] and nasopharyngeal carcinoma [45]. Noteworthy, in these cancers, TERT is activated independently of MYC or MYCN transcription factors [46]. The BMI1-dependent activation of TERT is cell type-selective: for instance, overexpression of BMI1 in fibroblasts does not promote excessive telomerase activity and immortalization [47]. Although no immediate effect of BMI1 on TERT has been demonstrated for neuroblastoma cells, BMI1 expression is prerequisite for the malignant transformation of neuroblasts and initiation of neuroblastoma tumorigenesis in immunodeficient mice [48, 49, 50].
Other known miR-128-3p-dependent positive regulators of telomerase activity are RUVBL2, NHP2, and PRKCB [19, 26, 27]. The RUVBL2 ATPase ensures interaction of DKC1/TERC complex with the TERT catalytic subunit, thus enabling the synthesis of telomeric DNA repeats. NHP2 (jointly with DKC1) participates in the H/ACA small nucleolar ribonucleoprotein complex (H/ACA snoRNP) responsible for the assembly of pivotal cellular RNPs; apart from its role in ribosome biosynthesis, the complex is involved in processing of TERC – the RNA component of TERT holoenzyme [51, 52]. The diverse functionalities of PRKCB, Protein Kinase C Beta type, include TERT activation, while its own PRKCB mRNA is targeted by miR-128-3p [19, 26]. Thus, despite the lack of direct destabilizing action of miR-128-3p on TERT mRNA, inhibition of telomerase activity under preserved expression of miR-128-3p can be achieved by inhibitory effects of this microRNA on translation of particular proteins essential for telomerase activity.
Conclusions
The study identifies the loss of miR-128-3p expression as a novel, potentially clinically relevant marker of aggressive neuroblastoma. The loss of miR-128-3p expression effectively complements TERT activation which is typical for high-risk tumors. The combination of two markers defines the adverse course of the disease indistinguishable from the effects of MYCN amplification. Downregulation of miR-128-3p with concomitant TERT upregulation is a promising prognostic criterion of adverse neuroblastoma for prospective validation in clinical trials.
Ethical statement
The study was approved by the Local Ethics Committee of Regional Children’s Hospital, Yekaterinburg, Russian Federation.
The study was supported by the Charity foundation Podari.Life.
Author contributions
Conception: Druy A.E., Tsaur G.A.
Interpretation or analysis of data: Druy A.E., Tsaur G.A., Shorikov E.V., Tytgat G.A.M., Fechina L.G.
Preparation of the manuscript: Druy A.E.
Revision for important intellectual content: Druy A.E., Tsaur G.A., Shorikov E.V., Tytgat G.A.M., Fechina L.G.
Supervision: Fechina L.G.
Footnotes
Acknowledgments
The authors thank Alena Alekseeva (BioVitrum LLC) for providing nCounter
In memoriam Alexander Globa (Syntol LLC), the developer of oligonucleotide primers and probes used in the research.
Supplementary data
Characteristics of patients randomly selected for the validation analysis of miR-128-3p expression by NanoString
Patient ID
Age at diagnosis, years
INSS stage
NB2004 risk group
1p deletion
MNA
263
2.48
4
High-risk group
No
No
283
0.91
3
Observation
No
No
293
3.76
4
High-risk group
No
No
311
1.11
4
High-risk group
Yes
No
346
7.86
1
Observation
No
No
376
1.81
4
High-risk group
No
No
486
2.88
4
High-risk group
No
No
577
0.17
4S
Observation
No
No
593
0.59
4S
Observation
No
No
685
0.90
1
Observation
No
No
719
4.10
4
High-risk group
No
No
755
0.24
3
Observation
No
No
783
0.35
1
Observation
No
No
789
0.94
3
Observation
No
No
790
4.46
1
High-risk group
Yes
Yes
800
0.61
3
Observation
No
No
821
0.05
1
Observation
No
No
833
5.12
4
High-risk group
No
Yes
834
3.73
1
Observation
No
No
842
0.15
4S
Observation
No
No
874
0.27
2
Observation
No
No
881
0.20
1
Observation
No
No
910
0.65
4
Intermediate risk group
No
No
986
0.77
3
Observation
No
No
994
2.72
4
High-risk group
Yes
No
