Abstract
Background:
Glioma is the central nervous system tumor with the highest incidence rate and the molecular detection of gliomas has been the focus of research. This study aimed to investigate the guiding effect of cluster of differentiation 276 (CD276) expression on the clinical prognosis of glioma.
Methods:
The TCGA and CGGA databases were used to study whether CD 276 can be used as an independent prognostic factor for gliomas. Immunohistochemistry was used to detect the expression of CD276, isocitrate dehydrogenase-1 (IDH1), matrix metallopeptidase 9 (MMP9), p53, and Ki-67, and 1p/19q co-deletion was detected by fluorescence in situ hybridization (FISH). The effects of CD276 RNA interference (RNAi) on cell invasion, cell cycle and the expression of β-catenin, tumor necrosis factor receptor 1 (TNFR1), and MMP9 were observed. Furthermore, the biological effects of CD276 gene knockout on intracranial transplanted tumors in nude mice were studied.
Results:
CD276 expression was positively correlated with the extracellular matrix, collagen decomposition, and cell adhesion molecules. Immunohistochemistry and FISH showed that CD276 expression positively correlated with the glioma grade, p53 mutation, Ki-67 proliferation, and MMP9 expression; however, it negatively correlated with IDH1 mutation, 1p/19q co-deletion, and the survival rate. CD276 RNAi in U87 cells inhibited cell proliferation, migration, and invasion, but had no effect on the cell cycle. CD276 inhibited the expression of β-catenin, TNFR1, and MMP9 in U87 cells at the mRNA and protein levels. In vivo experiments showed that the tumor formation and invasion of the CD276 small interfering RNA glioma cell line in nude mice were reduced and the survival time was prolonged.
Conclusions:
The present study demonstrated that high expression of CD276 in gliomas indicates a poor prognosis.
Keywords
Introduction
Glioma is the central nervous system tumor with the highest incidence rate, accounting for 30%–40% of all the primary intracranial tumors and approximately 80% of the adult intracranial malignant tumors. 1 In the recent years, the incidence of glioma has been increasing annually, 1 and the molecular detection of gliomas has been the focus of research. In the fourth edition of the World Health Organization’s Classification of Central Nervous System Tumors in 2016, oligodendrogliomas were classified according to isocitrate dehydrogenase (IDH) mutation and 1p/19q co-deletion. 2
Gliomas encompass a range of tumors and are classified into grade I to IV according to World Health Organization (WHO). The most common type of grade I, II, III, and IV are pilocytic astrocytoma, diffuse astrocytoma and oligodendroglioma, anaplastic astrocytoma and anaplastic oligodendroglioma, and glioblastoma (GBM), respectively. The median onset age of pilocytic astrocytoma is 12 years old. 3 The median age of diagnosis of diffuse astrocytoma, oligodendroglioma, anaplastic astrocytoma, anaplastic oligodendroglioma, and GBM 47, 43, 57, 50, and 65 years old respectively. 3 The overall 5-year survival rates of pilocytic astrocytoma, diffuse astrocytoma, oligodendroglioma, anaplastic astrocytoma, anaplastic oligodendroglioma, and GBM are 94.4%, 51.6%, 82.7%, 30.3%, 60.2%, and 6.8% respectively. 3 IDH mutation and 1p/19q co-deletion are good prognostic factors for gliomas4,5; however, most GBMs with the worst prognosis lack the IDH mutation and 1p/19q co-deletion. Therefore, finding new molecular changes in gliomas, especially potential therapeutic targets, is of great significance for its diagnosis.
At present, tumor immunotherapy and its related therapeutic targets are one of the research hotspots, especially the B7 immune checkpoint family. Programmed death-ligand 1 (PD-L1) (B7-H1, cluster of differentiation 274 [CD274]) is one such member of the B7 family. Some neurooncologists have used PD-L1 in the treatment of gliomas, but its therapeutic effect is unsatisfactory. Studies have shown that the therapeutic effect of PD-L1 combined with temozolomide is better than that of temozolomide alone, 6 however, some other studies have demonstrated the unsatisfactory treatment effect of PD-L1 antibody on glioma. 7 At present, the method of treating glioma using PD-L1 antibody is unclear; nevertheless, other members of the B7 family are being constantly considered cluster of differentiation 276 (CD276, B7-H3), is overexpressed in many tumors, including glioma, indicating that it has comparative advantages in the targeted therapy of glioma. 8 To date, CD276 targeting has not taken as a form of therapy.
As a co-stimulatory/co-inhibitory molecule, CD276 was first reported by Chapoval et al. 9 It is located on the human chromosome 15, encoding a transmembrane glycoprotein of 316 amino acids, with a molecular weight of 45–66 kDa. CD276 can co-stimulate CD 4+ and CD 8+ cells, thereby inducing cellular immunity. 9 The protein includes an intracellular domain, a transmembrane region, and an extracellular domain, and its extracellular domain includes two immunoglobulin constant regions and two variable regions. 10 CD276 is widely expressed in many human tissues and organs. Overexpression of CD276 in cancers indicates that CD276 plays an important role in the occurrence and development of tumors. Many studies have confirmed that the expression of CD276 is positively correlated with tumor grade and negatively correlated with tumor prognosis11-13; however, some studies have shown that the high expression of CD276 is associated with a good prognosis of some tumors. 14 It has been demonstrated that the expression of CD276 is increased in glioma and that CD276 redirected CAR-T cells can effectively control tumor growth, which may become a new target for glioma treatment. 15 At present, the role of CD276 in glioma-related biological characteristics is not completely clear.
In this study, the TCGA and CGGA databases were used to analyze the clinical information related to CD276, enrich the related proteins, and speculate their possible molecular biological functions. Subsequently, the clinical and molecular changes in the glioma wax block were verified. Finally, the CD276 silencing cell line was established in vitro and in vivo to verify the proliferation, invasion, and cell cycle changes of glioma tumor cells after CD276 silencing in order to clarify its influence on the biological functions of glioma.
Materials and methods
Screening of differentially expressed genes
Gene expression and clinical follow-up data of glioma were downloaded and extracted from the TCGA (https://cancergenome.nih.gov/) and CGGA (http://www.cgga.org.cn/) databases. Bioinformatics analysis was carried out in the R language (https://www.r-project.org/). Survival analysis filtering was performed using the packages survival and survimner for R with the Kaplan-Meier method (KM). A p value for KM < 0.001 and Coxp < 0.001 and the difference in the 5-year survival rate between high and low expression groups (> 0.2) were used for gene filtering. Independent prognostic analysis filtering was then conducted with p < 0.001. Survival receiver operating characteristic (ROC) packages were used to construct the ROC, and the area under the curve (AUC) was calculated using AUC > 0.7. The Beeswarm package was used for clinical correlation, the Wilcoxon test was performed between two groups, and the Kruskal test was conducted among different groups. TCGA is an open database; therefore, the use of TCGA data required no permission. Since the database does not involve patient privacy, ethical approval was not required.
Enrichment analysis
Enrichment analysis of the CGGA data was performed using the Gene Set Enrichment Analysis (GSEA) software. Protein coding genes (PCGs) related to CD276 in the TCGA database were screened using the limma package. The screening criteria were: a Pearson correlation coefficient of >0.4 and p < 0.01. Gene ontology enrichment analysis was performed using clusterprofiler and org.Hs.eg.db packages, with p < 0.05. A dot map was created using the dotplot package.
Immunohistochemistry
Paraffin-embedded glioma tissue blocks from the Department of Pathology of the Fifth Affiliated Hospital and the Second Affiliated Hospital of Zhengzhou University were used for immunohistochemistry (IH). The study protocol of IH was approved by the Institutional Ethical Review Committee of The Fifth Affiliated Hospital of Zhengzhou University (No.2018033). Informed consent for the use of tissues in this study was obtained from all the patients. The surviving patients were 1 point, while those of the dead patients were 2. The expression of CD276 and the survival status of the patients were statistically analyzed. A total of 147 glioma cases were studied, including 26 cases of WHO grade II (diffuse astrocytoma and oligodendroglioma), 32 cases of WHO grade III (anaplastic astrocytoma and anaplastic oligodendroglioma) and 89 cases of WHO grade IV (GMB). There were 81 males and 66 females. None of the patients received radiotherapy or chemotherapy before surgery. In addition, IH analysis was performed on mouse tissue sections. Avidin–biotin peroxidase complex technology was used for IH. Briefly, the blocks were cut into 4 μm slides, deparaffinized with xylene baths, and rehydrated using graded concentrations of alcohol. After steaming with 10 mmol/L citrate buffer solution, the slides were immersed in methanol containing 3% hydrogen peroxide and incubated in 10% horse serum. The slides were then incubated with specific primary antibodies against IDH1, CD276, p53, matrix metalloproteinase 9 (MMP9), and Ki-67. Immu-nostaining was performed using the iVIEW DAB Detection Kit (Ventana Medical Systems Inc., AZ, USA). Different scoring standards for the different markers are presented in Supplemental Table S1.
Fluorescence in situ hybridization
Fluorescence in situ hybridization (FISH) detection was carried out on each sample with two slides, and deletions of chromosome 1p and 19q were detected. In brief, these sections were cut into 10–20 μm sections. The slides were incubated at 65℃ overnight, and then subjected to a series of elution with xylene, 100%, 90%, and 70% gradient ethanol, and distilled water at room temperature. Subsequently, the slides were digested with pepsin and washed with 70%, 90%, and 100% gradient ethanol. Hybridization was performed after adding 10 μL 1p and 19q probes for each sample with denaturation at 85℃ for 5 minutes and hybridization at 37℃ for 10–18 hours. After washing with 2 × SSC, 0.1%NP-40/2 × SSC, and 70% ethanol, the slides were stained and observed under a fluorescence microscope. The laser beam was focused on different areas of the surface with a ×100 objective lens. Tumor nuclei were observed by DAPI (4’,6-diamidino-2-phenylindole) light, red signals were excited by green light, green signals were excited by blue light, and red and green signals of cell nuclei were observed by green and blue lights. The number of red and green signals for 100 cells were counted in each section and the ratio of red to green signals was calculated. A red signal/green signal of <0.5 was regarded as a 1p or 19q deletion. Patients with simultaneous deletions of 1p and 19q had 1p/19q co-deletion.
Cell culture
U87, U251, and U118 cell lines, the commonly used experimental cell models of GBM were purchased from Shanghai Research Biotechnology Co., Ltd. and stored in our laboratory. After screening, the cell line with higher expression abundance of CD276 mRNA was used for subsequent interference experiments. Additionally, 293T cell lines were used for the cell study. The cells were maintained in Dulbecco’s Modified Eagle’s medium (DMEM), supplemented with 5% fetal bovine serum (FBS), 1% Gln, and 1% penicillin/streptomycin at 37℃ under 5% CO2 and 95% air.
Reverse transcription-polymerase chain reaction
Total RNA was extracted from fresh cells or paraffin sections using TRIzol (INvitrogen), according to the manufacturer’s protocol. Reverse transcription was performed using the extracted RNA. The primers used were as follows: CD276: forward- TGAATGCGCTTTTGCAGGC, reverse- TGCCCCTGCCTCCCAG; β-catenin: forward- CCGCATGGAAATAGTTGAAG, reverse- CAATTCGGTTGTGAACATCCC; tumor necrosis factor receptor 1 (TNFR1): forward- AACACCGTGTGTAACTGCCA, reverse- CTCTTTGACAGGCACGGGAT; MMP9: forward-GCTGGCAGAGGAATACCTGTAC, reverse-CAGGGACAGTTGGCTTCTGGA; β-actin: forward- CCTGGCACCCAGCACAAT, reverse- GGGCCGGACTCGTCATAC. The program was as follows: 95℃ for 5 minutes, followed by 40 cycles of 95℃ for 10 seconds, 60℃ for 30 seconds, and 75℃ for 30 seconds. β-actin was used as an internal reference gene. The relative quantitation of CD276 gene in different cell lines was detected by △△CT.
Lentiviral interference
The CD276 sequence was analyzed and the interference targets were designed (Supplemental Table S2). Single-stranded oligonucleotides were synthesized by Sangon Biotech (Shanghai, China), linked to the pcDNA6.2-GW vector with the BsaI site, and transformed into the competent DH5α cells. The plasmids were extracted, sequenced, and amplified, with primers containing NHeI and Asc1 restriction sites (forward: Lenti-NHeI-F, 5’-GTCAGCTAGCGCCACCATGGTGAGCAAGGGCGAGGA; reverse: Lenti-Asc1-R, 5’-TCAGTGGCGCGCCTGCGGCCAGATCTGGGC). Subsequently, the amplified products were ligated to into pLenti6.3-MCS/V5 DEST and transformed into the competent DH5α cells. Plasmids were extracted from positive colons and sequenced. The 293 T cells were infected with the constructed anti-virus interference vector and packaging plasmid, and then the virus was packaged. The virus stock solution was collected and concentrated by ultracentrifugation, and the virus was stored at −80℃. U87 and CD276 knockout cells (U87-CD276-KD) were inoculated into a six-well culture plate, and 1 ml of target virus solution and control virus solution were added, following which a polyaromatic hydrocarbon solution was added to promote infection. After 48 hours, 1 m mL of TRIzol reagent was added to lyse the cells and extract the RNA. The expression of CD276 was detected by reverse transcription-polymerase chain reaction (RT-PCR).
Cell counting kit-8 detection
U87 cells were inoculated into six-well plates, with 3 × 105 cells per well. Then, 1 mL of the target virus, negative control virus and a certain volume of polyaromatic hydrocarbon were added. After 16 hours of culture, the cells were incubated for another 48 hours in a lentivirus-free medium. The cells were then inoculated into a 96-well plate, with 5 × 103 cells per well. Subsequently, 10 µL CCK-8 solution (Cell Counting Kit-8, Dojindo, Kumamoto Prefecture, Japan) was added to each well and incubated for 2 hours. Absorbance was measured at 450 nm. The control group was regarded as 100% absorbance.
Flow cytometry
U87-CD276-KD and U87-NC tumor cell suspensions were inoculated into six-well plates at a concentration of 3 × 105 cells/well. After culturing for 24, 48, and 72 hours, the cells were photographed, recorded, and collected. The cells were digested with trypsin and centrifuged at 1,000 rpm for 5 minutes. After washing twice with phosphate buffer saline (PBS), the cells were fixed with 70% ethanol. Then, the cells were washed with PBS, mixed with propidium iodide (PI, with RNAase), and incubated in the dark at 4℃ for 30 minutes. Flow cytometric analysis was performed to measure cell cycle distribution.
Transwell invasion and migration assays
U87 cells were infected with Lenti-CD276-EGFP-mir lentivirus and Lenti-EGFP negative control lentivirus, and the cells were inoculated into Transwell chambers (Costar 3422, Corning, NY, USA) to observe cell invasion and migration. The Matrigel and DMEM culture mediums were diluted to 1 mg/mL suspension in a ratio of 1:30, and 50 μL suspension was added to the inner surface of the Transwell chamber and then left at room temperature for 1 hour for later use. The cells were resuspended at a density of 1 × 105 cells/mL in DMEM without FBS. Subsequently, 0.1 mL of the cell suspension and 0.2 mL of serum-free culture medium were added to each upper chamber of the 24-well plate, and 0.6 mL of DMEM containing 10% FBS was added to the lower chambers. After culturing for 20 hours, the cells in the upper chambers were thoroughly cleaned. The cells were fixed with 4% paraformaldehyde at room temperature for 15 minutes. The lower surface of the upper chambers was stained with 0.25% crystal violet for 20 minutes at room temperature. Cell migration was evaluated using Transwell inserts without Matrigel. The ratio of migrated U87 cells was analyzed by imaging the cells from three independent tests in randomly selected fields of view, and then quantifying them with ImageJ software (NIH Image).
Western blot
U87-CD276-KD cells, U87 negative control (U87-NC) cells, and mouse brain tissue were subjected to Western blotting (WB). The cells were collected by pipetting, washed with PBS, and homogenized. Mouse brain tissue wax block was sliced into 5 μm sections and homogenized with radioimmunoprecipitation assay lysis buffer. After heating at 100 ℃ for 10 minutes, the lysate was centrifuged at 12,000 rpm for 1 minute. Equal amounts of total protein were separated by 12% sodium dodecyl sulphate–polyacrylamide gel electrophoresis and transferred to a polyvinylidene difluoride membrane. After blocking with skimmed milk in TBST, the membranes were incubated with primary antibodies overnight at different dilution ratios according to the manufacturer’s instructions. The membranes were then washed three times with TBST and incubated with secondary antibodies at room temperature for 1 hour. Finally, the membranes were washed and developed using an enhanced chemiluminescence system. Relative gray scale = target protein gray scale/β-actin gray scale.
Tumor formation in nude mice
U87-NC and U87-CD276-KD cells growing in an exponential phase were collected, washed twice with serum-free medium, digested with trypsin, and centrifuged at 1,000 rpm for 5 minute. The cells were adjusted to 1 × 107/mL with PBS, and placed on ice for further study. Thirty-one BALB/C nude mice, aged 6–8 weeks, weighing 20 ± 3 g, were purchased from Beijing Weitong Lihua Experimental Animal Science and Technology Co., Ltd. and raised in SPF class experimental animal room of the Zhengzhou University Experimental Animal Center. The mice were raised at room temperature (26–28 °C) and humidity (45%–55%).
The mice were randomly divided into three groups: sham operation group (n = 5), U87-NC group (n = 13) and U87-CD276-KD group (n = 13). The mice were anesthetized by intraperitoneal injection of pentobarbital sodium (60 mg/kg) and fixed in prone position on the stereotactic apparatus. After wiping and disinfecting the skin in the middle of the skull with 75% alcohol, the skin was cut off by approximately 1 cm. The skull was drilled 0.15 mm outside the posterior part of the anterior chamber and 2 mm to the right of the sagittal suture. Then, 10 μL of PBS, U87-NC tumor cells (105 cells), and U87-CD276-KD tumor cells (105 cells) were dripped into the puncture site at a speed of 2.5 μL/min. Two minutes after injection, the needle was pulled out and the needle hole was sealed with bone wax to suture the skin of the head. The general conditions of nude mice were observed daily for approximately 35 days. All mice were sacrificed by cervical dislocation under anesthesia (60 mg/kg intraperitoneal pentobarbital sodium) at the end of the study. All dead nude mice were dissected, the brain tissue was exposed, and the brain and cerebellum tissues were removed. It was observed whether there was tumor formation, and the maximum diameter of the tumor size in the brain was measured with a Vernier caliper. The criteria for successful modeling are unstable walking or hemiplegia before death, tumor found by anatomy, and glioma observed under a microscope after hematoxylin and eosin stain (HE) staining. If no tumor was found after dissection or no tumor cells were detected by HE staining, the nude mice were ruled out. The animal study protocol was approved by the Institutional Ethical Review Committee of The Fifth Affiliated Hospital of Zhengzhou University (No.2018069).
Hematoxylin-eosin staining
Brain tissues were fixed in 4% paraformaldehyde and embedded in paraffin. The tissues were then sliced into 4 μm thick sections. The slices were stained with HE for 10 minutes. Histological changes were observed under an optical microscope, and images were obtained.
Statistical analysis
SPSS software version 21.0 (SPSS Inc., Chicago, Illinois, USA) was used for statistical analyses, and GraphPad Prism 5 was used to produce statistical charts. The results were expressed as mean ± standard deviation. The Spearman correlation test was used to check the correlation between the two samples. The
Results
Differential expression of CD276 in normal and tumor samples
TCGA data of glioma and normal brain tissue were downloaded and compared to reveal the differences in CD276 expression. The logical value was greater than 2, and the false discovery rate (FDR) was less than 0.05, which indicated that there was a significant difference between the two sample types. The relationship between the expression of CD276 and the survival time and status of patients in the TCGA and CGGA databases was analyzed. Patients with high expression of CD276 had a poor prognosis (p < 0.05, Figure 1A and B). Univariate factors in the CGGA database (CD276 gene expression, postoperative-recurrence survival (PRS) type, histopathologic types, tumor grade, gender, age, radiotherapy, chemotherapy, IDH mutation, 1p/19q co-deletion) were used to identify the factors that are related to the survival rate. Visualization of hazard ratio (HR) values was performed using a forest map (Figure 1C). The results showed that CD276 expression was negatively correlated with the survival rate (p < 0.001, HR > 1), indicating that CD276 is a high-risk survival factor. In order to obtain more accurate results, multivariate prognostic analysis was conducted on the clinical factors related to CD276 in the CGGA database. The p value of the CD276 gene was less than 0.001, indicating that CD276 is related to survival. At the same time, HR > 1 proved that CD276 is a high-risk factor for survival; that is, the higher the expression of CD276, the worse the survival (Figure 1D). The AUC areas of all the three curves of CD276, calculated using the CGGA database, were greater than 0.7, which indicates that CD276 is highly reliable as a predictor (Figure 1E).

Differential expression of CD276 in normal and tumor samples. Analysis with: (A) TCGA database, (B) CGGA database showed that high expression of CD276 was associated with poor prognosis, (C) single factor prognostic analysis of CGGA database demonstrated that CD276 high expression, postoperative-recurrence survival (PRS) type, histopathologic types, grade, age, and chemotherapy were associated with poor prognosis while IDH mutation and 1p/19q co-deletion were correlated with good prognosis. No association was noted in gender and radiation therapy with prognosis, (D) multivariate prognostic analysis of CGGA database also showed that CD276 high expression, PRS type, and grade were associated with poor prognosis while IDH mutation and 1p/19q co-deletion were correlated with good prognosis. No association was noted in histopathologic types, gender, age, and radiation therapy with prognosis, and (E) the ROC curve showed that CD276 could be used as an independent prognostic factor.
Predictive effect of CD276 in glioma
We analyzed the correlation between CD276 and the different tumor subtypes using the CGGA data. CD276 expression was positively correlated with glioma grade. The higher the grade of glioma, the higher was the expression of CD276 (Figure 2A). Age is an important prognostic factor for glioma and is negatively correlated with prognosis, that is, higher the age of diagnosis the worse the prognosis. GBMs, the most common and malignant form of gliomas, are rare in individuals aged <40 years. 3 Many studied included in the TCGA database have used 40 years old as a cut-off for studying glioma grouping,16,17 and as a result, 40 years old was set as the cut-off. Our result showed that CD276 was positively correlated with age, with 40 years as the boundary. The expression rate of CD276 in patients over 41 years of age was higher than that in patients below 40 years of age (Figure 2B). CD276 was positively correlated with chemotherapy status, and the expression of CD276 in chemotherapy patients was higher than that in non-chemotherapy patients (Figure 2C). CD276 was positively correlated with tumor recurrence, and the level of CD276 in recurrent patients was higher than that in primary patients (Figure 2D). The expression of CD276 was negatively correlated with IDH mutation and 1p/19q co-deletion (Figure 2E and F).

Predictive effect of CD276 in glioma: (A–D) CD276 was positively correlated with glioma grade, age, chemotherapy status, tumor recurrence and (E and F) the expression of CD276 was negatively correlated with IDH mutation and 1p/19q co-deletion.
Enrichment of CGGA and TCGA databases
To unravel the functional changes that CD276 expression may affect, enrichment was performed according to the data formats in the CGGA and TCGA databases. CD276 gene related functionally enriched genes in the CGGA database were analyzed using GSEA. The functions of six positively correlated and two negatively correlated expressed protein genes were enriched, which were mainly related to CD276 (Supplemental Figure 1). The PCGs related to CD276 in the TCGA database were enriched. The results showed that the proteins related to CD276 were rich in cell adhesion molecule binding, actin binding, carbohydrate binding, cadherin binding, and cytokine receptor binding (Figure 3). Together, our results indicate that CD276 may be involved in the occurrence and development of glioma.

Enrichment results using CGGA and TCGA databases. These enriched genes were correlated with adhesion molecule binding, actin binding, carbohydrate binding, cadherin binding, and cytokine receptor binding.
Relationship between CD276 and common prognostic factors in glioma revealed by IH
IDH mutation and 1p/19q co-deletion are important benign prognostic factors of glioma, 18 which is the basis of the WHO classification in 2016. p53 gene mutation is an important molecular change in glioma and a characteristic molecular change in diffuse astrocytoma (>60%), 19 which plays an important role in the development of glioma. 20 Ki-67 is an important factor that reflects the proliferation activity of glioma cells and plays an important auxiliary role in the histological classification of gliomas. 21 Therefore, we carried out IH to verify the relationship between the expression of CD276 and these factors and glioma grade. Statistical analysis showed that the higher the grade of glioma, the higher the expression of CD276 (Figure 4A). The expression of CD276 was positively correlated with glioma grade and the expression of p53, Ki-67, and MMP9. In addition, it was negatively correlated with IDH1 mutation (Figure 4B). FISH showed that CD276 was negatively correlated with 1p/19q co-deletion, with a correlation coefficient of −0.275 (p < 0.01, Figure 4C and D). Survival statistical analysis showed that high expression of CD276 was positively correlated with death, with a correlation coefficient of 0.38 (p < 0.001, Figure 4E)

Immunohistochemical detection and FISH results: (A and B) immunohistochemical detection showed that CD276 expression was positively correlated with glioma grade, p53 mutation, Ki-67 proliferation and MMP9 expression, but negatively correlated with IDH1 mutation and (C–E) FISH and correlation analysis demonstrated that CD276 expression was negatively associated with 1p/19q co-deletion and survival rate.
CD276 small interfering RNA interference
To screen out a cell line suitable for the interference of CD276 small interfering RNA (siRNA), we first carried out RT-PCR. The results showed that the △Ct values of CD276 in U87, U251, and U118 cell lines were 11.08, 13.73, and 12.99, respectively. The expression abundance of CD276 mRNA in the U87 cell line was higher than that in the other two cell lines, which could be used for subsequent interference experiments (Figure 5A). The quantitative RT-PCR results showed that the silencing efficiency of the CD276 gene reached more than 90% (Figure 5B). The relative expression of CD276 in cultured U87-NC and U87-CD276-KD stable transformants was detected by RT-PCR, which indicated that the construction of CD276 RNA interference (RNAi) stable transformants was successful. To investigate the effect of CD276 knockout on cell proliferation, the CCK-8 assay was performed. The 450 nm OD values of the U87-NC and U87-CD276-KD groups were detected for 1 to 5 days. The results showed that the expression of CD276 was downregulated and the proliferation of U87 cells was inhibited (Figure 5C). The effect of CD276 gene knockout on the cell cycle was detected by flow cytometry. The cell cycle of U87-CD276-KD and U87-NC tumor cells showed no significant changes (Figure 5D and E). Cell migration and invasion detected by the Transwell method showed that compared with U87-NC tumor cells, U87-CD276-KD tumor cells had weaker migration and invasion abilities (p < 0.05, Figure 5F and G). To test the expression of the corresponding genes related to CD276 at different levels, RT-PCR and WB were performed. As shown in Figure 5H and I, the mRNA and protein expressions of CD276, β-catenin, MMP9, and NFR1 in U87-CD276-KD tumor cells were significantly decreased compared with those in U87-NC tumor cells.
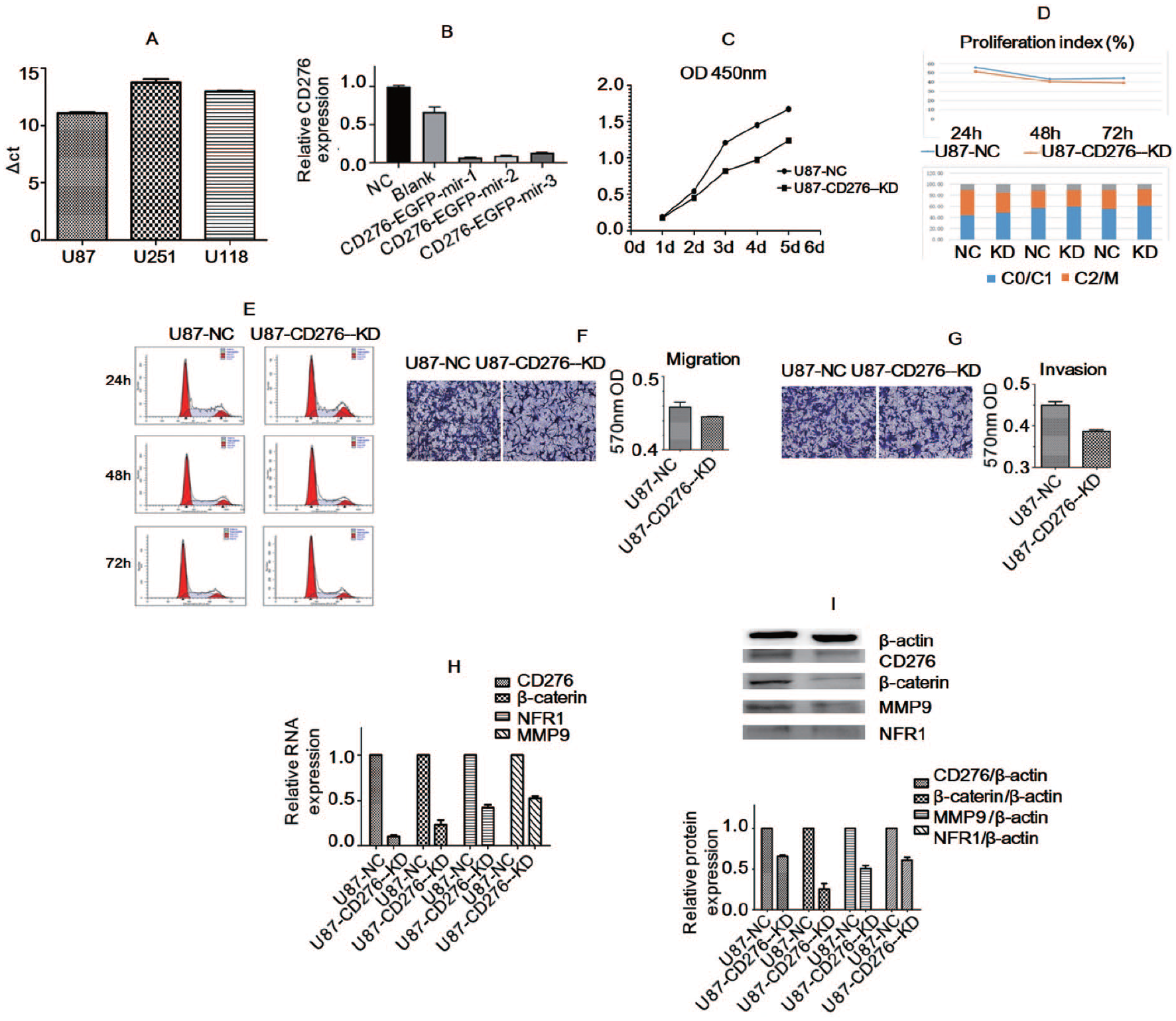
Detection results of CD276 expression in glioma U87 cell line: (A) compared with U251 and U118 cell lines, the abundance of CD276 genes in U87 cell lines was higher, (B) the silencing efficiency of CD276 was satisfactory, (C) the proliferation of U87 cells was inhibited, (D and E) CD276 gene knockout had no significant effect on cell cycle of U87-CD276-KD tumor cells and U87-NC tumor cells, (F and G) cell migration and invasion reduced in CD276 gene knockout cells. The expressions of CD276, β-catenin, MMP9 and NFR1 in U87-CD276-KD tumor cells were significantly reduced at (H) mRNA and (I) protein level compared with the U87-NC tumor cells.
Effect of CD276RNAi on intracranial in situ tumor formation in nude mice
To further investigate the effect of CD276 in vivo, tumor formation was performed in nude mice (BALB/C). U87-NC and U87-CD276-KD stably transformed cell lines were implanted into the skull of nude mice to form tumors in situ, and the effects of CD276 on the invasion and proliferation of glioma model in vivo were studied. Compared with the U87-NC group, the U87-CD276-KD group significantly prolonged the survival time of experimental animals, and the difference was statistically significant (p < 0.05, Figure 6A). The brain was removed, and the diameter was measured. Compared with the U87-NC group, the tumor diameter in the U87-CD276-KD group was evidently reduced (Figure 6B and C). HE staining was then performed to validate the histopathological changes and the results showed that the tissue was malignant (Figure 6D). IH was carried out to test the expression of tumor CD276 protein, and the results showed that CD276 protein in the U87-CD276-KD stable transformation group was clearly lower than that in the U87-NC group (p < 0.05, Figure 6E). RT-PCR and WB were conducted to test the expression of the related genes at the mRNA and protein levels. Compared with the U87-NC group, RT-PCR and WB showed that the gene and protein levels of CD276, β-catenin, MMP9T, and TNFR1 in the U87-CD276-KD group were significantly decreased (p < 0.05, Figure 6F and G).

Biological effects of CD276 knockout of U87 cells on intracranial orthotopic transplanted tumor in nude mice. CD276 knockout significantly prolonged the: (A) survival time of experimental animals, (B and C) the tumor diameter was evidently reduced in the knockout group, (D) HE staining showed that the tissue was malignant, (E) CD276 expression was obviously lower in the knockout group than the U87-NC group. RT-PCR and WB showed that the expressions of CD276, β-catenin, MMP9 and NFR1 in U87-CD276-KD tumor cells were significantly decreased at the (F) mRNA and (G) protein levels compared with the U87-NC group.
Discussion
By analyzing the data from the TCGA and CGGA databases, we found that the expression of CD276 was negatively correlated with the prognosis of glioma and can be regarded as an independent prognostic factor for malignant glioma. Gene enrichment analysis showed that CD276-related genes are enriched in collagen and have functions of decomposition, extracellular matrix, and cell adhesion, which suggests that CD276 is closely related to glioma invasion. The functions of proteins enriched by the two statistical methods in the TCGA and CGGA databases were similar, which can validate each other. Astrocytoma, such as WHO grade I hairy cell astrocytoma, mostly occurs in children with a good prognosis, and its molecular mechanism is different from diffuse astrocytoma, anaplastic astrocytoma, oligodendroglioma, anaplastic oligodendroglioma and GBM. 1 Therefore, WHO grade I gliomas were not included in this study. In previous studies, gliomas were divided into GBM (WHO grade IV) and low-grade gliomas (LGG, WHO grade II/III) and related biological behavior and clinical research was conducted. 22 LGGs account for 10%–20% of all the primary brain tumors, with slow growth and median survival time of 4.7–9.8 years. 23 However, high-grade gliomas, especially GBMs, are more aggressive and have poor prognosis. 24 CD276 is co-expressed with stemness genes in GBM stem cells, suggesting that the CD276 gene can promote the occurrence and development of glioma. 25 This study shows that the grade of glioma is positively correlated with CD276, which is an important prognostic factor of glioma, further proving that the expression of CD276 is associated with a poor prognosis of glioma. Consistent with our results, Wang et al 26 found that gliomas with high expression of CD276 have a poor prognosis, and IDH1 mutation may have an important impact on the expression of CD276.
IDH mutation is an important benign prognostic factor and classification basis of glioma, and 90% of glioma IDH mutations are IDH1 mutations. 27 In this study, the mutation of IDH1 and the expression of CD276 in gliomas were detected by IH. The results showed that the expression of IDH1 was negatively correlated with the expression of CD276 in gliomas. 1p/19q co-deletion is a benign prognostic factor of glioma, and it is also an important factor in molecular classification. 28 FISH showed that 1p/19q co-deletion was negatively correlated with the expression of CD276 in gliomas. Owing to the low incidence of 1p/19q, the correlation between 1p/19q co-deletion and CD276 was not significant; the correlation coefficient was -0.275, but the p value was <0.01, which still had a statistical difference. According to the TCGA data, 78% of GBM mutates in the signaling pathway of p53. 29 The p53 pathway plays an important role in many different cell reactions, such as cell cycle regulation, apoptosis, cell differentiation, and DNA damage. Our results showed that the mutation of p53 in glioma was positively correlated with CD276, which suggests that CD276 may be related to the cell cycle and DNA damage in glioma. Ki-67 protein exists in all active stages of the cell cycle, such as G1, S, G2 and M stages, but does not exist in the static G0 stage, which makes it an excellent indicator of the growth fraction. 30 The expression of CD276 was positively correlated with Ki-67, suggesting that CD276 might affect cell proliferation. Functional enrichment analysis showed that CD276 was related to the extracellular matrix and collagen. MMP9 is an important member of the matrix metalloproteinase family, which plays an important role in the hydrolysis and remodeling of the extracellular matrix. 31 Numerous studies have shown that MMP9 is closely related to many biological functions of tumors, especially tumor invasion, metastasis, and angiogenesis. Our results showed that the expression of CD 276 in glioma was positively correlated with MMP9, which further indicated that the expression of CD 276 in glioma had an important relationship with tumor invasion function.
β-catenin is a component of the classical Wnt pathway (Wnt/β-catenin pathway) and plays an important role in the dynamic balance of development, differentiation, tissue maturation and apoptosis. 32 Besides being a transcription regulator in the Wnt signaling pathway, β-catenin is a multifunctional protein, an E-cadherin-binding protein that participates in the regulation of intercellular adhesion. 33 E-cadherin and cadherin-catenin complex not only play a role in cell-cell adhesion, but also regulate Wnt signal transduction, nuclear factor kappa B (NF-κB) pathway, and epithelial-mesenchymal transition in epithelial cells, thus accelerating cell migration and invasion. 34 In this study, silencing of the CD276 gene confirmed that catenin expression in glioma cells decreased with a decrease in CD276 expression, indicating that CD276 could influence the extracellular matrix by affecting β-catenin, thereby, affecting invasion and other functions of tumor cells. TNFR1 is one of the most typical signaling pathways regulating cell fate, and TNFR1 triggers NF-κB to cause downstream functions. 35 MMP9 is closely related to tumor invasiveness, 36 and it is also an important factor involved in cell fusion. 37 The process of cell fusion requires many factors, including adhesion molecules, intracellular signal proteins, proteases, transcription factors, and tissue proteins. Research on cell fusion has shown that minocycline can inhibit tumor necrosis factor α (TNF-α)-induced cell fusion by directly targeting TNFR1/NF-κB signal node to inhibit the expression of the target gene MMP9 of NF-κB-induced fusion-related factors. 38 In addition, NF-κB signaling and TNF-α/TNFR1 autocrine/paracrine rings are the key mediators of GBM migration and invasion. 39 In view of this, it is speculated that glioma can also mediate tumor migration and invasion through TNFR1/NF-κB acting on MMP9. Our study confirmed that the downregulation of CD276 could induce a decrease in TNFR1 and MMP9, which suggested that CD276 could affect cell function through the TNFR1/MMP signaling pathway.
We implanted the stably transformed U87-NC and U87-CD 276-KD tumor cells into intracranial tumors in nude mice and observed the changes in the biological function of U87 glioma cells after treatment with CD276 RNAi. BALB/C nude mice were selected as experimental animals. Nude mice are important experimental animals in oncology research because of congenital thymus defects and congenital immunodeficiency. 40 Because the symptoms of glioma are closely related to the tumor site, the tumor implantation location was consistent. In vivo experiments proved that the RNAi of CD276 can reduce the malignant biological function of tumors. The survival time of experimental animals was prolonged, the average tumor diameter was reduced, and the expression level of CD276 in the implanted site decreased. This is consistent with the results of Zhang et al., who found that in xenograft mouse models of human GBM cell lines, CD276 gene knockdown inhibited tumor growth. 15 The animal model of glioma confirmed that the expression of CD276 decreased simultaneously with the expression of β-catenin, TNFR1, and MMP9, implying that CD276 also affected the invasion and other biological activities of glioma through the β-catenin and TNFR1/MMP9 pathways in vivo.
Conclusions
In summary, our in vitro and in vivo experimental results showed that CD276 was a factor of poor prognosis of glioma and its high expression was related to a poor prognosis of tumors; nevertheless, CD276 silencing could reduce the proliferation and invasion of tumors. Therefore, the use of targeted antibodies in glioma patients with high CD276 expression may provide new opportunities for the treatment of glioma.
Supplemental Material
sj-docx-1-onc-10.1177_11795549211032330 – Supplemental material for High expression of cluster of differentiation 276 indicates poor prognosis in glioma
Supplemental material, sj-docx-1-onc-10.1177_11795549211032330 for High expression of cluster of differentiation 276 indicates poor prognosis in glioma by Linchen Li, Min Zhang, Dengna Zhu and Xinjun Wang in Clinical Medicine Insights: Oncology
Supplemental Material
sj-tif-2-onc-10.1177_11795549211032330 – Supplemental material for High expression of cluster of differentiation 276 indicates poor prognosis in glioma
Supplemental material, sj-tif-2-onc-10.1177_11795549211032330 for High expression of cluster of differentiation 276 indicates poor prognosis in glioma by Linchen Li, Min Zhang, Dengna Zhu and Xinjun Wang in Clinical Medicine Insights: Oncology
Footnotes
Acknowledgements
We are sincerely acknowledge the contributions from TCGA protect (https://cancergenome.nih.gov/) and CCGA protect (![]() ).
).
Declaration Of Conflicting Interests:
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding:
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This study is financially supported by National Natural Science Foundation of China (No. 81972361).
Authors’ Contributions
LL and WX conceived and designed the study project. LL and ZM performed the experiments. ZD participated in part of experiments and data entry. LL and WX performed data analysis and wrote the manuscript. All authors read and approved the final manuscript.
Availability of data and materials
Availability of data and materials that the raw data will be made available on reasonable request.
Ethical Approval
All tissues were obtained with written informed consent from each patient. The present study conformed to the ethical guidelines of the 1975 Declaration of Helsinki and was approved by the Institutional Ethical Review Committee of The Fifth Affiliated Hospital of Zhengzhou University ((No.2018033, No.2018069).
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
