Abstract
BACKGROUND:
Detection of circulating cell-free DNA (ccfDNA) methylated in BCAT1 and IKZF1 is sensitive for detection of colorectal cancer (CRC), but it is not known if these biomarkers are present in other common adenocarcinomas.
OBJECTIVE:
Compare methylation levels of BCAT1 and IKZF1 in tissue and plasma from breast, prostate, and colorectal cancer patients.
METHODS:
Blood was collected from 290 CRC, 32 breast and 101 prostate cancer patients, and 606 cancer-free controls. Tumor and matched normal tissues were collected at surgery: 26 breast, 9 prostate and 15 CRC. DNA methylation in BCAT1 and IKZF1 was measured in blood and tissues.
RESULTS:
Either biomarker was detected in blood from 175/290 (60.3%) of CRC patients. The detection rate was higher than that measured in controls (48/606 (8.1%), OR
CONCLUSIONS:
Detection of circulating DNA methylated in BCAT1 and IKZF1 is sensitive and specific for CRC but not breast or prostate cancer.
Introduction
Colorectal cancer (CRC) is a common disease with high mortality rates. Estimates from 2020 showed that CRC was the fourth most commonly diagnosed non-cutaneous cancer, with almost 2 million newly diagnosed cases globally and almost 1 million deaths [1]. Survival rates for CRC can be improved to over 90% if the cancer is detected at an early stage [2]. Formation of pre-cancerous lesions, such as adenomas and sessile serrated lesions, can take over 10 years to progress to malignancy after accumulation of genetic and epigenetic abnormalities [3]. Such an extended period between benign and malignant disease enables opportunity to detect and remove pre-invasive lesions before they turn cancerous. Strategies that can accurately detect CRC, particularly early-stage disease or recurrence when monitoring cases after treatment, would benefit outcomes and improve treatment.
Detection of circulating tumor DNA (ctDNA) in the blood for identifying tumor-specific changes (such as epigenetic changes and mutations) is a promising strategy to detect CRC as well as other cancer types [4, 5, 6]. We have established a sensitive DNA methylation assay that can detect circulating hypermethylated BCAT1 (branched chain amino acid transaminase 1) and IKZF1 (IKAROS family zinc finger 1) in CRC cases with a sensitivity of up to 77% [7]. When combined with the current CRC screening fecal immunochemical test (FIT), the sensitivity for detecting cancer rises to 89% [8]. We have also shown that levels of methylated BCAT1 and IKZF1 DNA in the blood increase with tumor stage and that over 95% of CRC tissues have aberrant methylation of these two genes [7, 9] providing clinical utility for the detection and monitoring of recurrence in CRC patients. Whether these biomarkers are also present in patients with other common adenocarcinomas such as breast and prostate cancer is not known.
Breast and prostate cancer are hormone dependent cancers and are common in both women and men, respectively [1], and as such may be present but unsuspected in a CRC screening population. Although originating in different organs, breast and prostate cancer share many genetic and epigenetic characteristics [10]. There are also many genetic commonalities between adenocarcinomas of the breast, prostate and colorectum, including somatic mutations in oncogenes and/or tumor suppressor genes (e.g. TP53 and BRAF) [11], but also common sites of DNA hypermethylation in key genes controlling carcinogenesis, such as the Wnt, Hippo and retinoblastoma tumor suppressor pathway [12]. Previous studies have pointed to a role of BCAT1 and IKZF1 in breast and prostate cancer progression [13, 14, 15, 16, 17, 18]. However, less is understood about the extent to which these specific epigenetic changes characteristic of CRC tissues occur in breast and prostate cancers.
To support the clinical utility and specificity of the methylated BCAT1 and IKZF1 assay for detection and management of CRC, it is important to determine whether these other common cancers also harbor these aberrantly methylated genes and release them into the circulation. Thus, we measured levels of BCAT1 and IKZF1 DNA methylation in blood and tissues from patients diagnosed with adenocarcinomas of the breast and prostate and compared these to levels observed in patients with CRC and individuals with no evidence of cancer.
Materials and methods
Experimental subjects
Individuals with confirmed diagnosis of prostate cancer, breast cancer and CRC, and who attended clinic appointments at one of three South Australian public hospitals (Repatriation General Hospital, Flin- ders Medical Centre, and Noarlunga Health Service) between 2011 and 2020 were assessed for presence of methylated BCAT1 and IKZF1 in blood. Where possible, and providing that neoadjuvant therapy had not been given prior to surgery, tumor tissue and adjacent normal tissue samples were collected at the time of cancer resection or retrieved from archival samples obtained through the Flinders Tissue Biobank (Bedford Park, South Australia) for assessment of hypermethylation in these two genes. Inclusion criteria comprised patients aged 18 years and older with known diagnosis of primary invasive adenocarcinoma (i.e. not stage 0) of any pathological stage, of the breast, prostate, or colorectum. Cases were excluded from the study if they had already commenced any type of treatment, if an insufficient blood volume was collected for assay, or if they had a known active cancer in any other organ. Controls recruited from the same hospital sites between 2011 and 2014 were males and females aged over 18 years who had a blood sample collected prior to colonoscopy, with no known history of cancer diagnosis and who were negative for precancerous lesions (adenoma or sessile serrated lesions) and CRC at colonoscopy. Informed written consent was obtained prior to sample collection.
Experiments were undertaken with the understanding and written consent of each subject, and that the study conforms with The Code of Ethics of the World Medical Association (Declaration of Helsinki), printed in the British Medical Journal (18 July 1964). Approval was granted by the Southern Adelaide Clinical Human Research Ethics Committee (human ethics committee approvals 162.16, 134.045). The study was prospectively registered through the Australian New Zealand Clinical Trials Registry (registration numbers: ANZCTR 12616001138471 and 12611000318987). All authors had access to the study data and had reviewed and approved the final manuscript.
Clinical procedures
Pathology and clinical data from cases where blood was collected for methylation analysis were recorded to determine any associations with blood test positivity. TNM and AJCC staging [19, 20, 21] were assessed based on findings at surgery or at MRI if no surgery was performed. Staging by MRI in prostate and rectal cancer patients not undergoing surgery is a widely accepted clinical practice [22, 23], and demonstrates good accuracy for diagnosis of T stage [24, 25]. For colorectal tissue sampling, normal biopsies were collected at a minimum distance of 20 mm adjacent to the tumor (range 20–350 mm). For prostate tissue samples, 5 mm biopsy punch samples were collected from each quadrant of intact whole resected prostate glands after radical prostatectomy, with half of the biopsy sent for pathology diagnosis and the other half banked for DNA extraction. For breast tissues, whole resected breast samples were sectioned and tumor tissue identified by macroscopic examination by a pathologist. Normal breast tissue was taken from the quadrant with no visible macroscopically confirmed tumor tissue for DNA extraction. For prostate cancer, indicators of disease severity including Gleason score of the tumor and blood PSA were also collected. For breast cancer, tumor sub-type (invasive ductal carcinoma, lobular or mixed ductal/lobular), histology grade, and tumor size (at surgical resection or by imaging) were recorded, and hormone receptor status for ER, PR and HER2 were assessed by standard clinical immunohistochemical staining.
Blood DNA methylation testing
Approximately 18 mL of blood was collected via venipuncture (K3-EDTA Vacuette tubes, Greiner Bio-One, Frickenhausen, Germany), followed by plasma isolation, which was stored at
Tissue DNA methylation testing
Tumor and matched normal adjacent tissues (pro- state
Demographics and blood BCAT1 and IKZF1 positivity rates by cancer status
Demographics and blood BCAT1 and IKZF1 positivity rates by cancer status
*Individuals undergoing colonoscopy with no history of cancer diagnosis. SEM
Data are presented as percentages or as mean
For tissue methylation analysis, paired t-tests were used to compare % methylation
All data were analyzed using statistical package StataMP 16.0
Results
Experimental subjects
Overall, there were 32 breast, 101 prostate, and 290 CRC patients and 606 non-cancer controls who met inclusion criteria. Case data are outlined in Table 1. Female CRC patients and female non-cancer controls were generally younger than their male counterparts, and prostate cancer patients were the oldest cohort with a mean age of 69.3 years
Blood test positivity rates and concentration of circulating methylated BCAT1 and IKZF1. (A) Blood test positivity rates (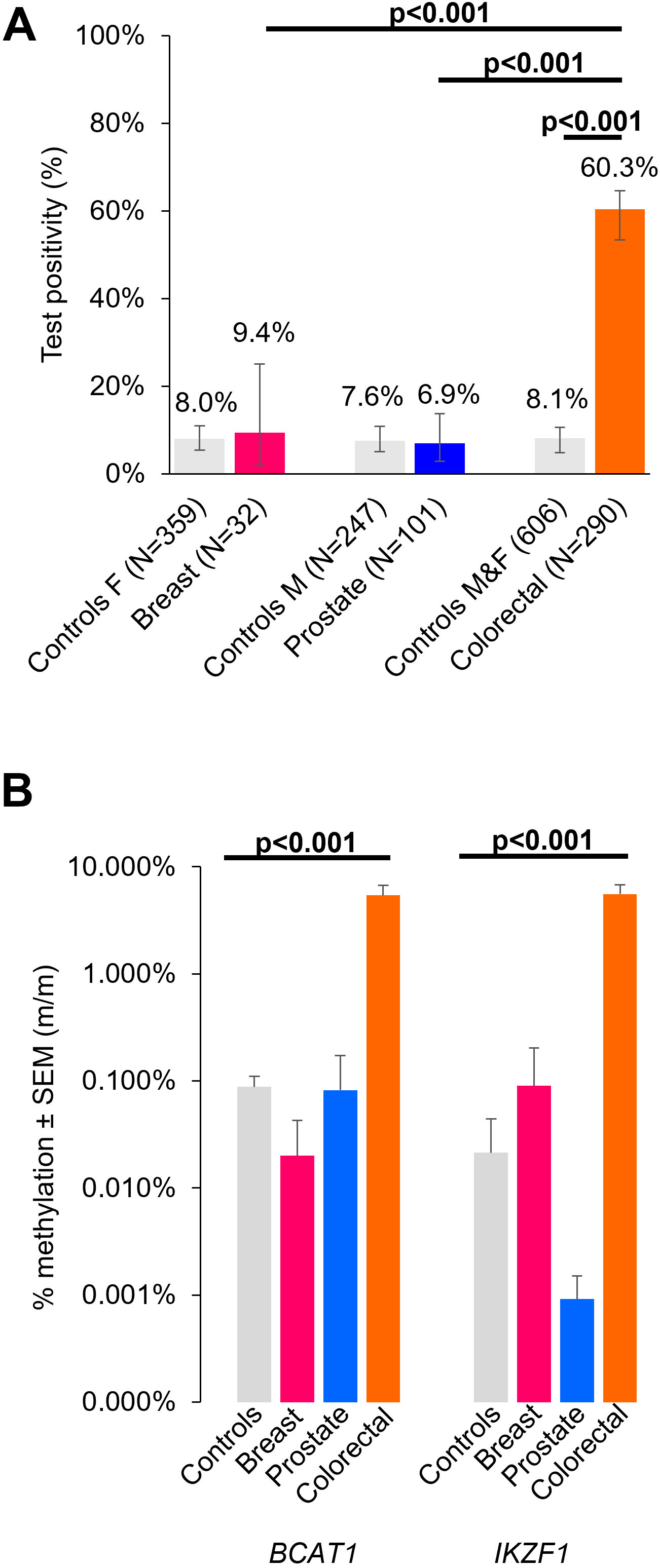
Venn diagrams showing concordance for BCAT1 and IKZF1 positive cases for non-cancer controls (A), breast and prostate cancer (B), and CRC cases (C).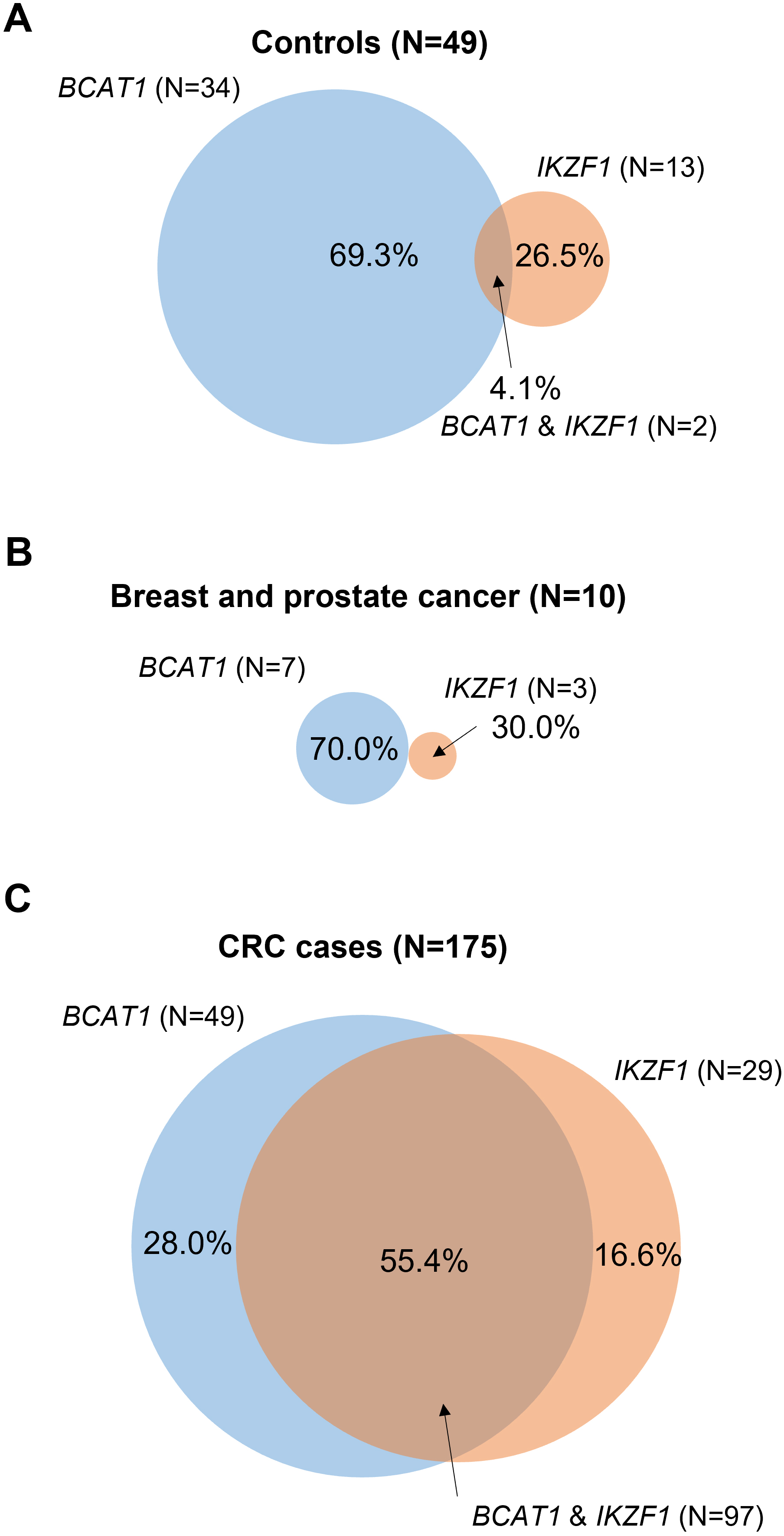
There were 9.4% (3/32) and 6.9% (7/101) breast and prostate cancer cases, respectively, who tested positive for methylated BCAT1 and/or IKZF1 DNA in blood (Table 1). The test positivity rates were no different to the test positivity rates observed for non-cancer controls at 8.0% for females (odds ratio (OR)
Average levels of % methylated BCAT1 and IKZF1 in the blood from patients testing positive were over 100-fold higher in CRC patients (BCAT1: 5.4%
Circulating methylated BCAT1/IKZF1 test result according to cancer stage
Circulating methylated BCAT1/IKZF1 test result according to cancer stage
Concordance of blood test positive cases across the different patient groups for BCAT1 and IKZF1 and a combination of the two genes are shown in Fig. 2. Non-cancer controls (Fig. 2A) and breast and prostate cancer patients (Fig. 2B) were more likely to test positive for BCAT1 than for IKZF1 alone or for BCAT1 and IKZF1 combined (
Methylated BCAT1 and IKZF1 in matched normal and cancer patient tissues
DNA methylation levels for BCAT1 (A) and IKZF1 (B) in matched normal and cancer tissues. Test positivity cut-off was set at 10% methylation. Cases were defined as not detectable where there was no signal from three replicates in 50 PCR amplification cycles.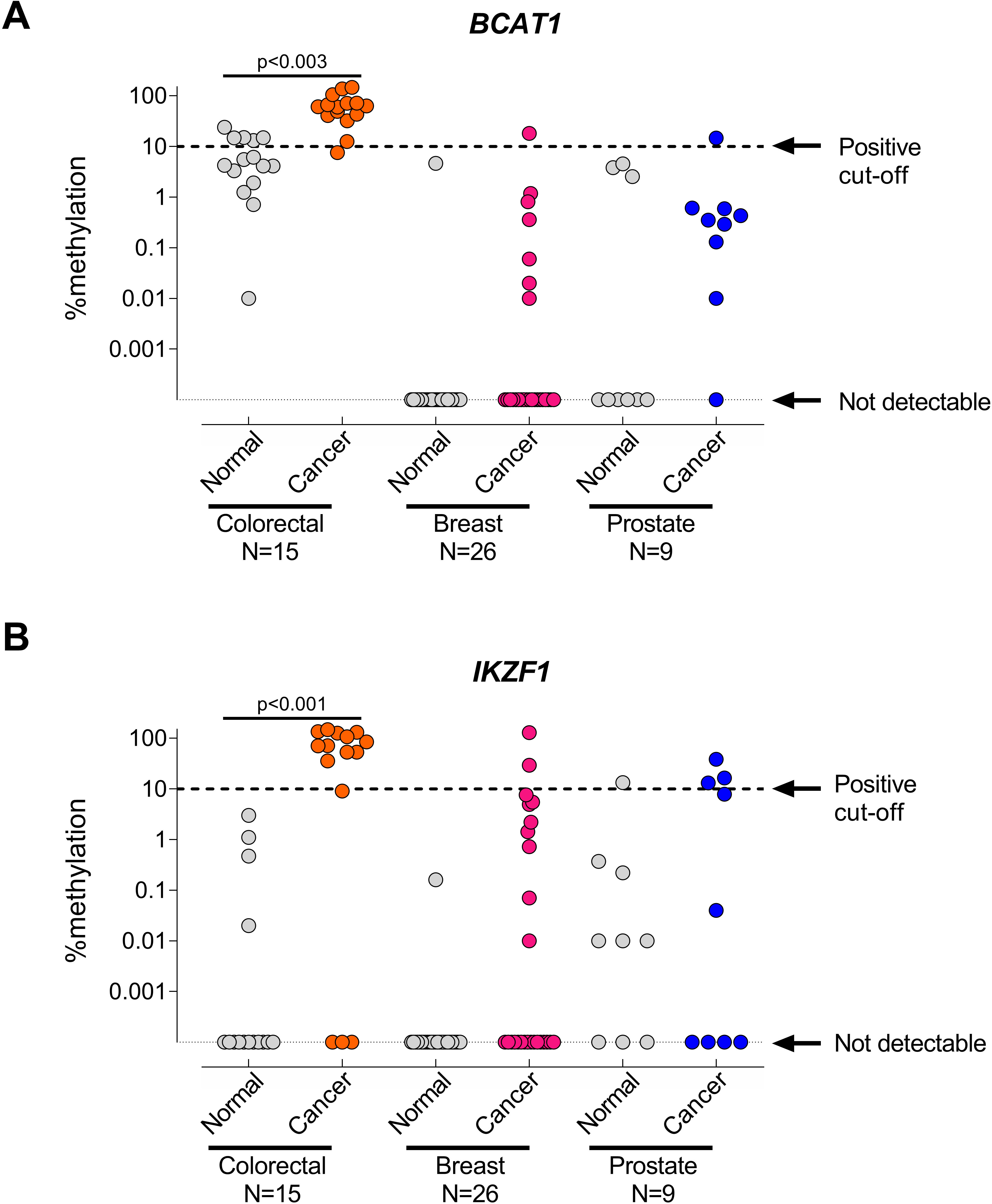
Blood test positivity for each cancer type and for each primary tumor, lymph node and metastasis (TNM) stage and AJCC (American Joint Committee on Cancer) stage is presented in Table 2. For CRC patients, there were strong associations between detection of methylated BCAT1/IKZF1 DNA in blood and advancing tumor stage (T1 4.3% vs T4 80.9%; OR
Blood samples were taken at a median time of 12 days after biopsy for breast cancer patients (range 1–39 days) and 36 days after biopsy for prostate cancer patients (range 13–796 days), where 1 day is sufficient time for clearance of
All BCAT1/IKZF1 positive breast cancer cases were diagnosed as invasive ductal carcinoma (IDC), two positive cases were estrogen receptor-
BCAT1 and IKZF1 methylation in tissues
Normal and matched cancer tissues were collected from 26 breast, 9 prostate and 15 CRC patients. Percent DNA methylation of BCAT1 and IKZF1 (Fig. 3A and B, respectively) were significantly higher in CRC tissue samples compared to their matched normal samples (
Percent DNA methylation was reported as a binary outcome using a 10% methylation cut-off value to assess hypermethylation of cancer versus matched normal tissues for breast, prostate and CRC (Fig. 3 and Table 3). Breast and prostate cancers were not significantly hypermethylated for BCAT1 and/or IKZF1 compared to matched normal tissue samples. However, CRC tissues were significantly hypermethylated for BCAT1 (
Discussion
This study explored whether detection of methylated BCAT1 and IKZF1 DNA in blood was specific for CRC by comparing levels of these biomarkers in plasma from patients with breast, prostate or CRC, as well as control cases without evidence of any cancer. The blood test positivity rates in breast and prostate cancer patients were not significantly different from those of controls, whereas blood test positivity was significantly higher in CRC patients. These findings were consistent with levels of methylated BCAT1 and IKZF1 in cancer tissue from the three organs; levels were considerably higher in CRC tissues
The literature suggests a role for IKZF1 and BCAT1 in breast and prostate cancers, and therefore it was important to determine whether the biomarkers sequences applied within the assay for CRC detection are also hypermethylated in these other common cancers. This could have implications for the exclusive use of the circulating methylated BCAT1/IKZF1 biomarkers for CRC detection. The transcription factor IKZF1 has been implicated as a tumor suppressor in breast and prostate cancer [32, 33, 34], but also as a prostate cancer oncogene [17, 35], depending on the type of cancer cell-line being tested. Over-expression of BCAT1, a protein that catalyzes transamination of branched chain amino acids, leading to cell growth, is associated with poorer survival in breast cancer patients and enhanced malignancy of breast cancer cell lines in vitro [13, 14, 15, 36]. In prostate cancer, higher expression of BCAT1 promotes cancer cell survival [16] and decreases sensitivity to chemotherapy treatments of prostate cancer cells in vitro [18]. Although there is a suggestive role of BCAT1 and IKZF1 in breast and prostate cancer progression, less is understood whether this is driven by aberrant methylation in these genes. Our findings suggest this is not the case as there was no significant difference in the levels of methylation in tumor versus normal tissues at the CpG sites interrogated by the PCR assay studied here for either BCAT1 or IKZF1. Thus, this assay is not likely to have any translational potential as an effective screening tool for breast or prostate cancer.
Detection of methylated BCAT1/IKZF1 in CRC blood samples was strongly associated with higher TNM and AJCC stage, agreeing with our previous findings showing that the test is more sensitive for detection of advanced disease [9, 37]. A lower sensitivity for early-stage cancer, particularly T1 cancers, is a well-described limitation of ctDNA tests, including for breast and prostate cancer [38]. This study was limited by the number of advanced cancers cases for both breast and prostate cancer patients. However, the trends observed in blood testing were the same for these cancers and non-cancer controls in that we found circulating BCAT1 methylation was more frequently detected than IKZF1 methylation in blood. This aligns with our previous findings showing that methylation of BCAT1 is the main source of false positives in patients without colonoscopically-evident CRC [39]. The low frequency of methylated BCAT1/IKZF1 in blood collected from patients without CRC may stem from lysed white blood cells [30].
The main limitation of the current study was the small number of breast and prostate cancer cases. Additionally, statistical comparisons for breast and prostate cancer cases were constrained as there were zero cases for some subgroups (e.g. nodal stage and tumor stage). Larger-cohort studies are warranted to confirm the results reported herein. A further limitation of this study was that it did not assess sensitivity of the biomarkers in other common adenocarcinomas, such as lung cancer. This warrants further investigation as lung cancer is also commonly diagnosed in both men and women [1]. A potential weakness of the study was the use of adjacent normal tissues as opposed to using tissue samples from patients without cancer. Therefore, we cannot exclude the possibility that some cancer cells were in the “normal” samples or that there was a field effect in in adjacent non-neoplastic tissues. Despite these limitations, there was a clear differentiation between biomarker hypermethylation levels in cancer vs non-neoplastic tissue from CRC patients, in contrast to the low levels of methylation in these genes observed in tissues from prostate and breast cancer patients.
This study concludes that detection of circulating DNA methylated in BCAT1 and IKZF1 is both sensitive and specific for detection of CRC, supporting its use as a valuable non-invasive tool for detection of CRC. The likelihood of a positive test arising as a result of unsuspected prostate or breast cancer in a CRC screening population is very low.
Footnotes
Acknowledgments
Archival biological samples and data used in this research were provided by the Flinders Tissue Bank.
Author contributions
Jean M. Winter, Lorraine Sheehan-Hennessy, Beibei Yao, Susanne K. Pedersen, Molla M. Wassie, Michael Eaton, Michael Chong, Graeme P. Young, Erin L. Symonds.
Conception: Susanne K. Pedersen, Graeme P. Young, Erin L. Symonds Interpretation or analysis of data: Jean M. Winter, Beibei Yao, Susanne K. Pedersen, Molla M. Wassie, Graeme P. Young, Erin L. Symonds.
Preparation of the manuscript: Jean M. Winter, Lorraine Sheehan-Hennessy, Susanne K. Pedersen, Molla M. Wassie, Graeme P. Young, Erin L. Symonds.
Revision for important intellectual content: Jean M. Winter, Lorraine Sheehan-Hennessy.
Susanne K. Pedersen, Molla M. Wassie, Michael Eaton, Michael Chong, Graeme P. Young, Erin L. Symonds.
Supervision: Graeme P. Young, Erin L. Symonds.
Supplementary data
Breast cancer patients: circulating methylated BCAT1/IKZF1 positivity for different clinicopathological findings ER
All cases
BCAT1/IKZF1 test positivity
Sub-type
Non-invasive ductal carcinoma
9
0
Invasive ductal carcinoma
23
3 (13)
Histological Grade
Low (1–5)
7
1 (14.3)
High (6–9)
22
1 (4.5)
Hormone receptor status
ER
31
2 (6.5)
ER
1
1 (100)
PR
25
1 (4)
PR
7
2 (28.6)
Prostate cancer patients: circulating methylated BCAT1/IKZF1 positivity for different clinicopathological findings
All cases
BCAT1/IKZF1 test positivity
P
RR
95% CI
PSA
Low (4–9.99 ng/mL)
56
5 (8.9)
High (
43
2 (4.7)
0.662
0.70
0.15 – 3.4
Gleason score
Low/medium (6–7)
78
6 (7.7)
High (8–10)
21
1 (4.8)
0.776
0.73
0.08 – 6.4
