Abstract
BACKGROUND:
MicroRNAs can regulate tumor metastasis either as oncomiRs or suppressor miRNAs. Here, we investigated the role of microRNA 224 (miR-224) in lymphatic metastasis of non-small-cell lung cancer (NSCLC).
METHODS:
The expression of miR-224 was demonstrated by a validation cohort of 156 lung cancer patients (77 cases with lymphatic metastasis) by quantitative polymerase chain reaction (qPCR). In vitro and in vivo experiments were performed to study the malignant phenotype after upregulation and inhibition of miR-224 expression. Furthermore, the direct target genes of miR-224 were determined by a luciferase reporter assay.
RESULTS:
First, miR-224 was identified as a highly expressed miRNA in tumor tissues with lymphatic metastasis, with an area under the curve (AUC) of 0.57 as determined by qPCR analysis of a validation cohort of 156 lung cancer patients. Then, in vitro and in vivo experiments indicated that forced expression of miR-224 in H1299 cells promoted not only cell viability, plate colony formation, migration and invasion in vitro but also tumor growth and lung metastasis in vivo. Consistently, inhibition of miR-224 suppressed malignant characteristics both in vitro and in vivo. Moreover, molecular mechanistic research suggested that miR-224 enhanced NSCLC by directly targeting the tumor suppressor angiopoietin-like protein (ANGPTL).
CONCLUSIONS:
Overall, the collective findings demonstrate that miR-224 is a potential biomarker for the prediction of lymphatic metastasis of NSCLC.
Abbreviations
Background
Globally, lung cancer is the leading cause of cancer death [1], and its 5-year survival rate is approximately 4–17% [2]. Among the two pathological subtypes, non-small-cell lung cancer (NSCLC) is the most common type, accounting for approximately 80% of lung cancers [3]. Lymph node metastasis is an independent factor affecting survival and recurrence after surgical resection of the primary malignancy [4]. Patients with mediastinal lymph node dissection are associated with lower local recurrence rates and better survival [5]. Therefore, searching for lymph node metastasis-related microRNAs (miRNAs or miRs) and understanding the molecular mechanisms is deeply meaningful.
MiRNAs are defined as endogenous and small RNAs of
Previously, we identified differentially expressed miRNAs in patients with lymph node metastasis compared with patients without lymph node metastasis by miRNA microarray analysis [13]. In total, we obtained 39 upregulated miRNAs in tissues with lymph node-positive metastasis versus lymph node-negative tissues. miR-224 was upregulated 14.6-fold, ranking ninth among upregulated miRNAs. Moreover, the
Methods
Cell lines and culture conditions
Human lung cancer cell lines (H1299 and PC9) were purchased from the Chinese Academy of Medical Sciences & Peking Union Medical College (Beijing, China). Cells were cultivated in RPMI 1640 with 10% FBS, 100 U/mL penicillin, and 100 mg/mL streptomycin in a 5% CO2 humidified incubator at 37
Patients and samples
A cohort of 155 NSCLC patients (77 cases with lymphatic metastasis and 78 patients without lymphatic metastasis) were included from 2003 to 2019 to validate miR-224 expression in cancer tissues. Inclusion criteria: 1) definitive pathologic diagnosis; 2) no chemoradiotherapies before surgery; 3) completed a 5-year follow-up; and 4) provided written informed consent. Exclusion criteria: 1) other types of clinical disorders were observed; and 2) patients lost during follow-up or who died of other clinical disorders. This study was approved by the Ethics Committee of Peking University.
Reverse transcription quantitative real-time polymerase chain reaction (RT-qPCR)
MiRNAs were extracted from tissue samples from NSCLC patients or cells using an miRNeasy Mini Kit (Qiagen, Valencia, CA, USA) according to the manufacturer’s instructions. The purity and concentration of RNA were estimated using an ND-1000 microspectrophotometer (Thermo Fisher Scientific, Waltham, MA, USA). For mature miRNA expression detection, the polyA tailing method was used [14], and cDNA was synthesized from 100 ng RNA using Moloney murine leukemia virus reverse transcriptase (M-MLV RT) (Invitrogen, Carlsbad, CA). For mRNA expression detection, 2
Cell viability assay
5
Plate colony formation assay
Five hundred cells/well were plated in 6-well plates and cultured for 2 weeks. The number of cell colonies was counted after fixation with 4% formalin and staining with 1% crystal violet.
Transwell assay
For the cell migration and invasion (coated with extracellular matrix substitute) assay, mitomycin C-treated cells were loaded into the upper chamber of a Transwell insert with 8.0-
Wound healing assay
Cells were cultured in 24-well plates and scratched with a 20-
In vivo tumor growth and metastasis assay
Animal experiments were approved by the Animal Ethics Committee of Peking University Cancer Hospital & Institute. The in vivo tumor growth and metastatic characteristics of the cells were measured by a modified chick embryo chorioallantoic membrane assay (CAM). Briefly, 5
Analysis of predicted microRNA targets by online software
The predicted targets of the microRNAs were analyzed by an online database, miRecords (
Plasmid construction
The ANGPTL1 3’-UTR and YPEL1 3’-UTR or UTR mutant fragments containing the miR-224 complementary binding sites and corresponding mutants were inserted into the pGL3.0-control (Promega, Madison, WI) downstream of the luciferase coding sequence and were confirmed by sequencing.
Dual-luciferase reporter assays
HEK-293FT cells cultured in 24-well plates were transfected with pGL-3 construct (300 ng), miR-224 mimic or negative control (NC: 50 nM) and Renilla luciferase construct (20 ng) using Lipofectamine 2000 (Invitrogen). Firefly and Renilla luciferase activities were detected by a dual-luciferase assay (Promega) 24 h after transfection, and firefly luciferase activity normalized to Renilla luciferase activity was calculated.
Western blotting analysis
Cells were lysed with RIPA lysis buffer with protease inhibitors, and the protein was quantified by the BCA method. Equal amounts (30
MiR-224 was highly expressed in NSCLC tissues with lymphatic metastasis. (A) Heatmap for miR-224 expression in lymphatic metastasis-positive and lymphatic metastasis-negative tissues by microarray analysis. (B) MiR-224 expression in lymphatic metastasis-positive (LN+) and lymphatic metastasis-negative (LN-) tissues by q-PCR analysis. (C) ROC curve analysis for the prediction of lymphatic metastasis by miR-224 expression. (D) Survival analysis according to miR-224 expression using the Kaplan-Meier method.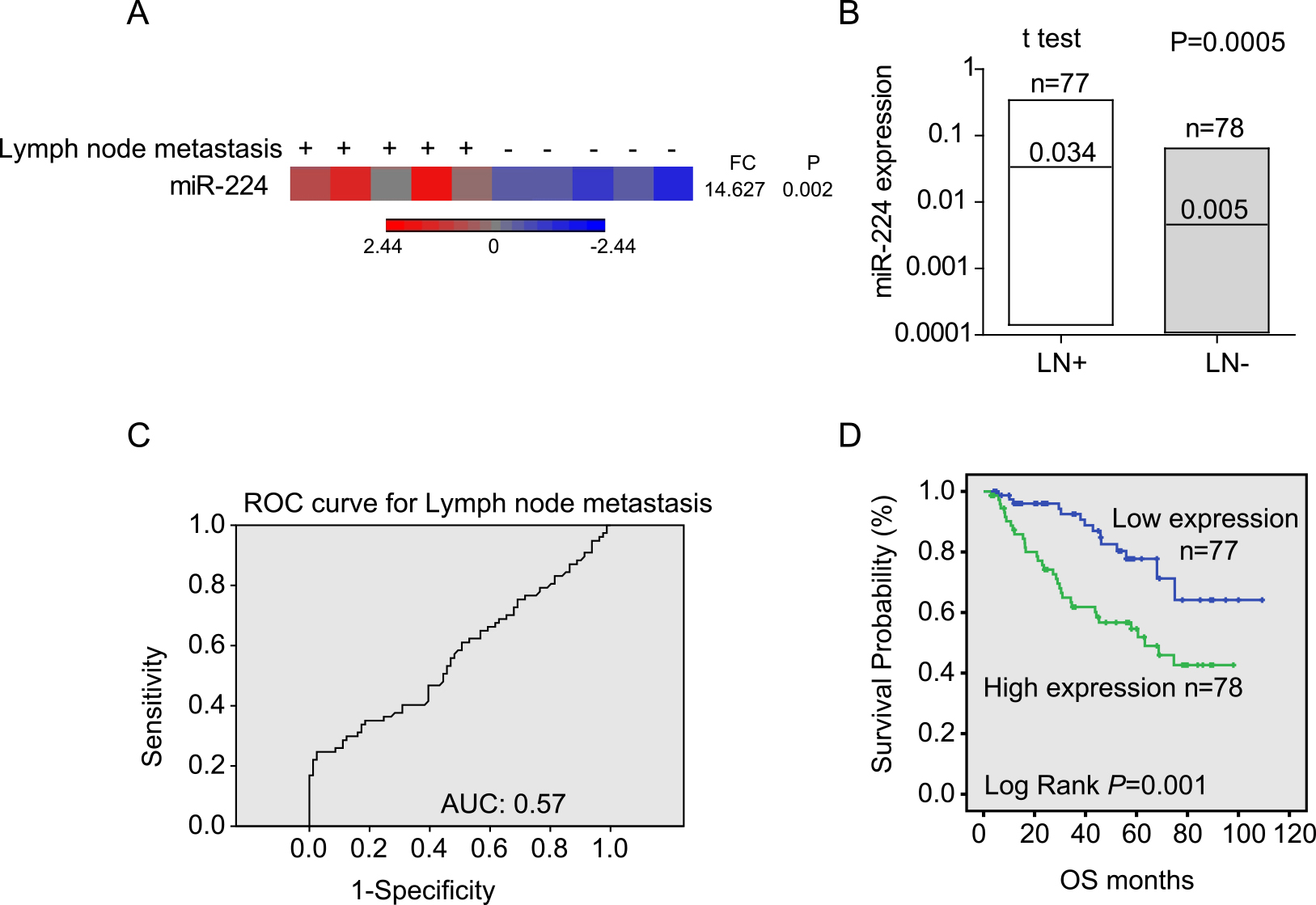
Statistical analyses were calculated with SPSS 16.0 (IBM, Armonk, NY, USA). Receiver operating characteristic (ROC) curves were established to discriminate metastatic lymph nodes and noncancerous lymph nodes. A two-tailed chi-squared test (
Results
Validation of miR-224 expression in NSCLC with lymphatic metastasis
Previously, 50 miRNAs were identified to be differentially expressed by miRNA microarray analysis in lymph node tissues with metastasis from five NSCLC patients and were compared with the expression levels in the corresponding noncancerous lymph node tissue [13]. Among these miRNAs, miR-224 was significantly upregulated in metastatic lymph nodes with a 14.6-fold change (Fig. 1A). Then, we validated miR-224 in a cohort of 155 patients. Consistently, miR-224 expression in samples with lymphatic metastasis was significantly higher than that in samples without lymphatic metastasis (approximately 6.8-fold change, Fig. 1B,
Forced expression of miR-224 enhanced cell viability, colony formation, migration and invasion in H1299 cells in vitro
First, the expression of miR-224 was detected by qRT-PCR in six lung cancer cell lines. As shown in Fig. 2A, miR-224 expression levels were relatively high in PC9, A549 and H520 cells but low in GLC82, H1299 and H1975 cells. Therefore, we used H1299 cells for overexpression experiments and PC9 cells for knockdown experiments to reveal the role of miR-224 in the malignant phenotype of lung cancer cells. As shown in Fig. 2B, miR-224 expression was increased by approximately 40-fold after transfection with mimic at 24 hrs. Forced expression of miR-224 markedly promoted cell viability (Fig. 2C,
Overexpression of miR-224 enhanced cell viability, colony formation, migration and invasion in H1299 cells in vitro. (A, B) miR-224 expression in lung cancer cells was detected by q-PCR. (C) Cell viability assay for miR-224 overexpression and NC control cells was determined by CCK-8 assay. (D) A plate colony formation assay was performed for miR-224-overexpressing and NC control cells. (E) Cell spreading ability calculated as wound closure of monolayers was determined by wound healing assay for different cells. (F) Cell migration and invasion ability quantified by cell numbers were assessed by Transwell assay without or with Matrigel. Scale bars, 200 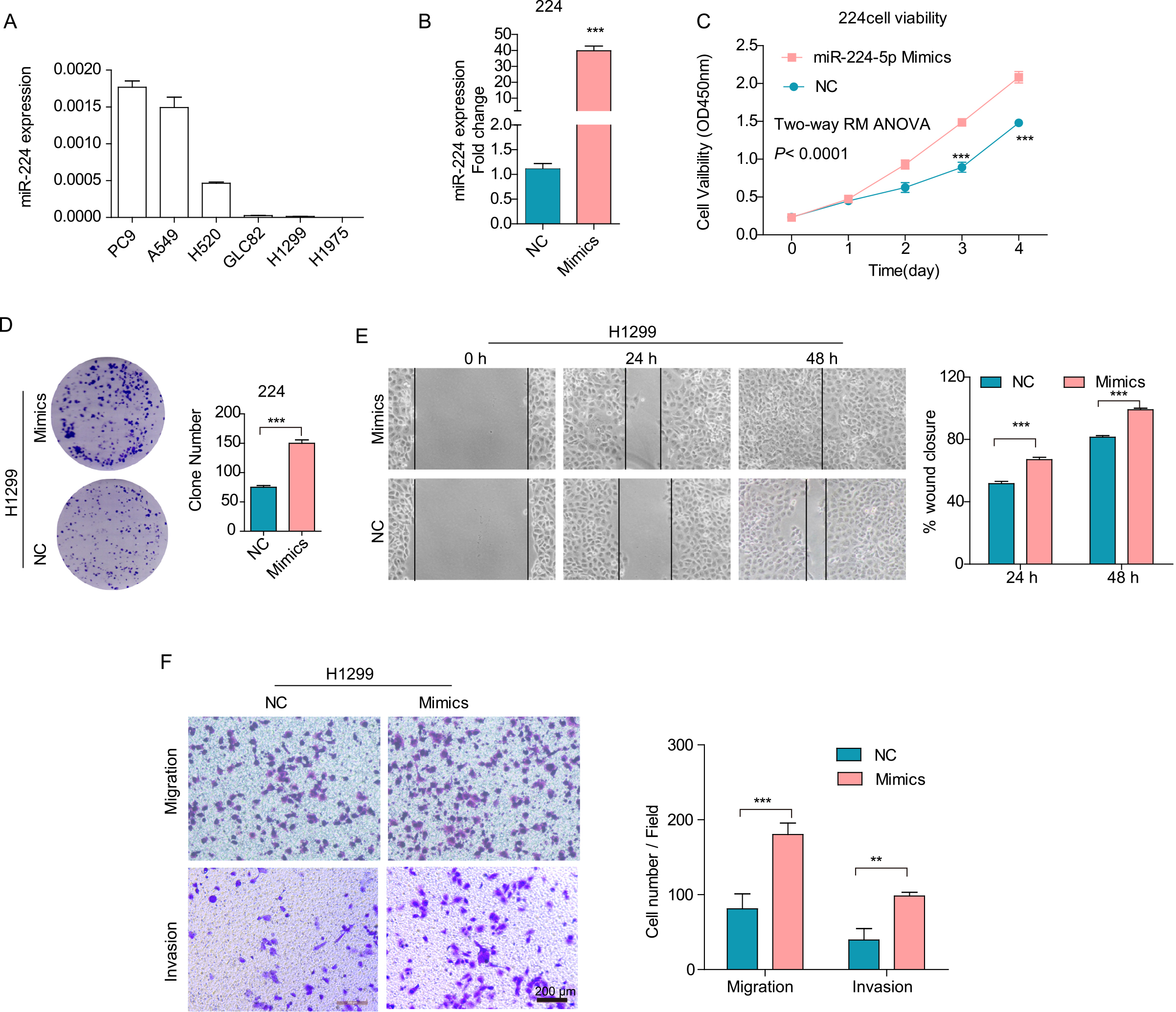
Inhibition of miR-224 suppresses PC9 cell migration and invasion in vitro. (A) miR-224 expression in different cells was detected by q-PCR. (B) CCK-8 assays were performed to assess cell viability. (C) Plate colony formation assays were performed to monitor cell growth. (D) A wound healing assay was used to study the cell spreading ability, and wound closure of the monolayer was calculated. (E) Transwell assays were performed to determine cell migration and invasion. Cell numbers were counted in four different fields. Scale bars, 100 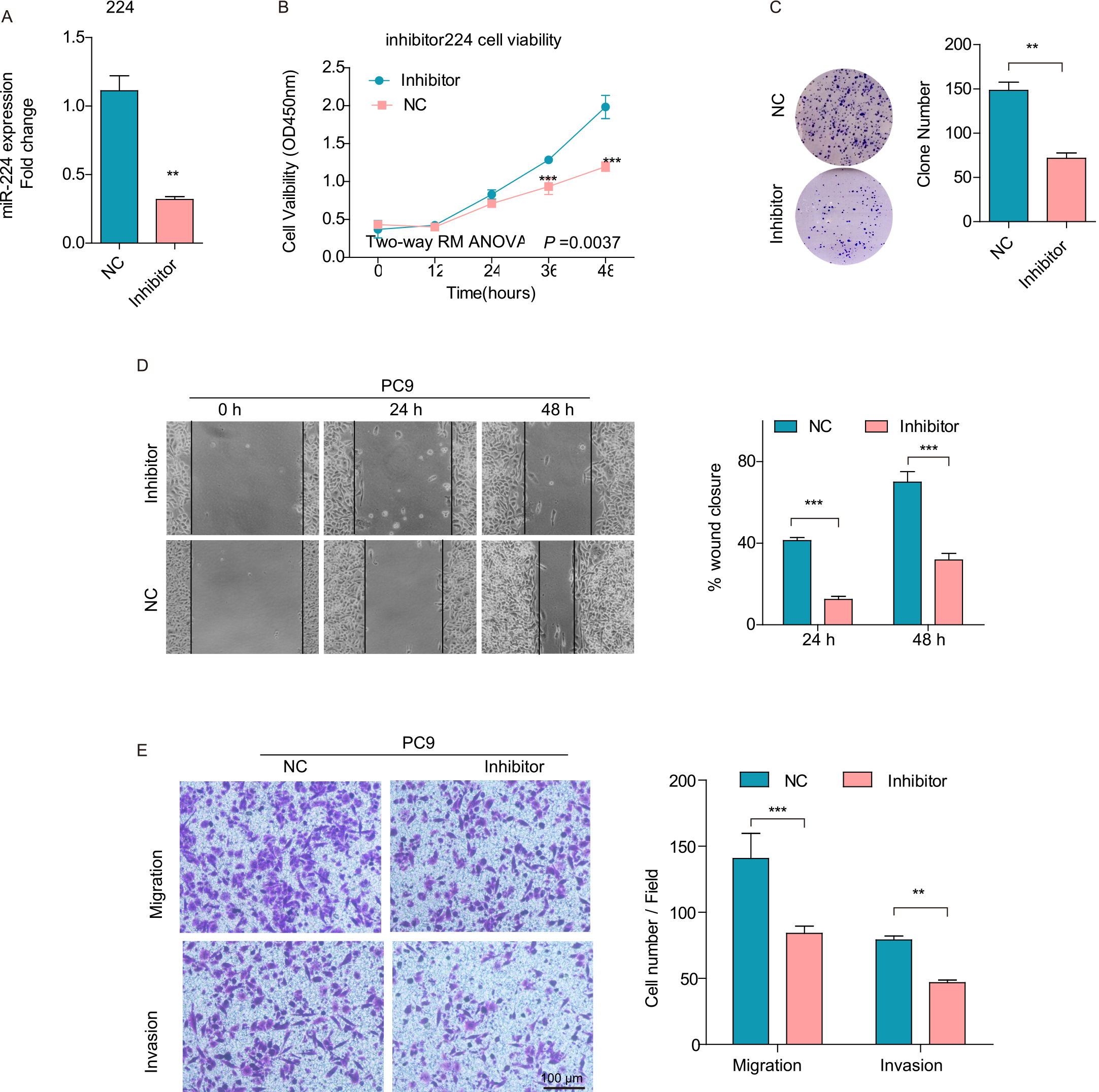
MiR-224 inhibitor was transfected into PC9 cells with relatively high endogenous expression of miR-224. MiR-224 expression was successfully decreased by 71% after cells were transfected with inhibitor (Fig. 3A). Consistent with the results above, inhibition of miR-224 decreased cell viability (Fig. 3B,
MiR-224 functions as an oncomiR in vivo. (A, C) Control cells or cells with miR-224 mimics or inhibitors were inoculated onto the CAM. The effect of miR-224 on tumor growth was evaluated based on the tumor weight of each group. (B, D) Representative pictures of lung metastasis lesions by laser scanning confocal microscopy. The effect of miR-224 on lung metastasis was evaluated by the number of metastatic lesions (tumor cells were labeled with the red fluorescent dye CM-Dil). The data in the bar graphs were calculated as the mean 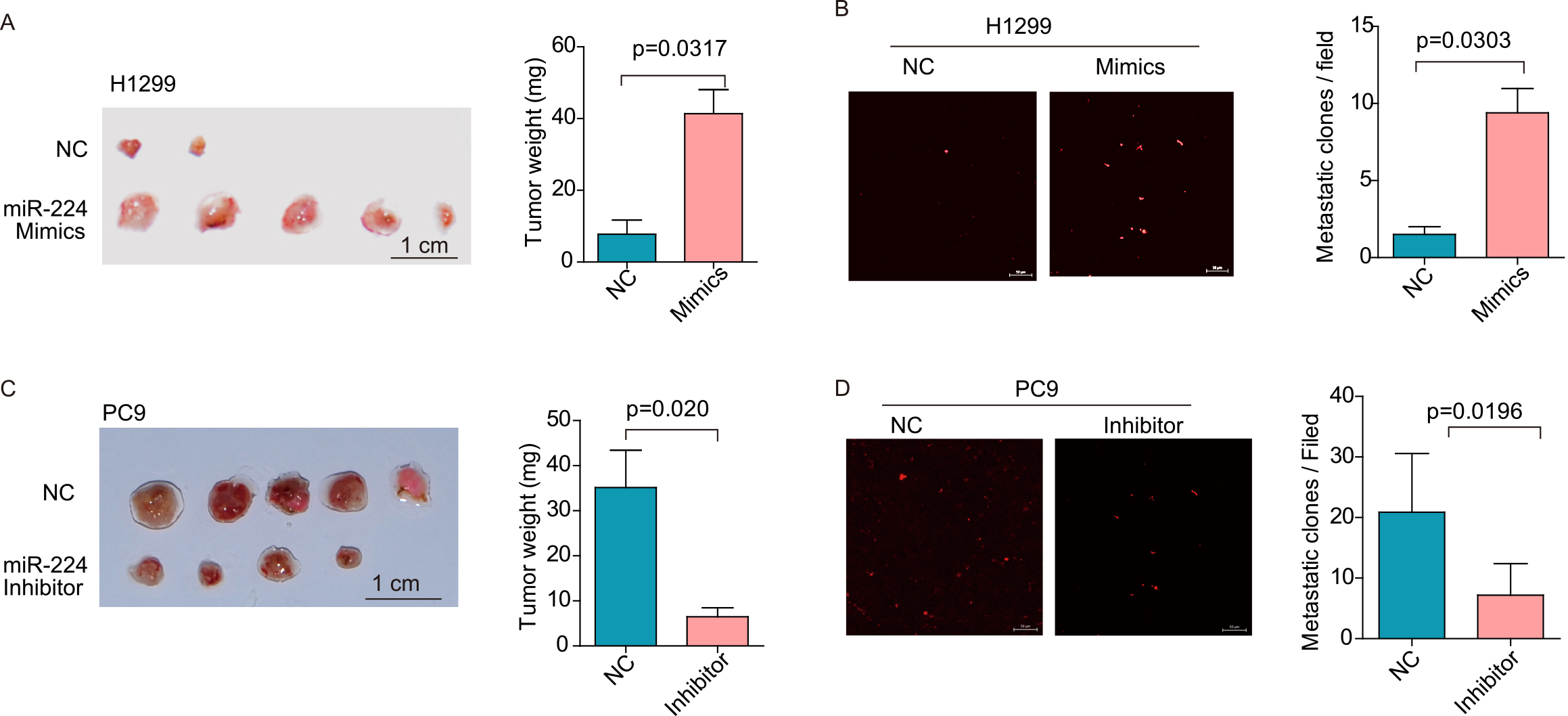
To validate the results above, we employed a modified chick embryo chorioallantoic membrane (CAM) assay to assess the role of miR-224 in tumor growth and metastasis in vivo. As shown in Fig. 4A, overexpression of miR-224 in H1299 cells resulted in a 5.2-fold increase in tumor weight compared to control cells (tumor weight,
ANGPTL1 is a direct target of miR-224 and is involved in its oncogene function
The biological significance of miRNAs relies on the regulation of their targets. First, we analyzed the miRNA targets using the online database miRecords. Given the hypothesis that miR-224 correlated with lymphatic metastasis, we analyzed the differentially expressed genes (DEGs) of lung cancer tissues at the N1-3 stage with lymph node metastasis compared with those at the N0 stage without lymph node metastasis, and we performed correlation analysis between the mRNA and miRNA expression profiles from The Cancer Genome Atlas (TCGA) database. A three-set Venn diagram showed 41 intersections of candidate targets (Fig. 5A). Of the 41 predicted target genes, we selected two tumor suppressor genes, ANGPTL1 and YPEL1, for further validation. We detected the expression levels of ANGPTL1 and YPEL1 in miR-224-overexpressing and miR-224-inhibited cells. As shown in Fig. 5B, the expression levels of the two candidate genes were markedly decreased in miR-224-overexpressing H1299 cells but increased in PC9 cells transfected with miR-224 inhibitor at the protein level. Both ANGPTL1 and YPEL1 had one consensus binding site in their 3’-UTR. Luciferase reporters containing either the wild-type or mutant type 3’-UTR fragment were constructed (Fig. 5C). Dual-luciferase reporter assays showed that the miR-224 mimic significantly reduced the reporter activity of the wild-type ANGPTL1 luciferase construct (Fig. 5D,
ANGPTL1 is a direct target of miR-224. (A) Venn diagram predicting the candidate targets of miR-224. (B) Western blot analysis of ANGPTL1 and YPEL1 expression in miR-224 mimic or inhibitor cells. (C) Schematic model indicating complementary binding sites and corresponding mutations of miR-224 in the ANGPTL1 and YPEL1 3’UTRs. (D) Wild-type or mutant PGL3 control plasmids were transfected into HEK293T cells together with miR-224 mimic or NC. Firefly fluorescence intensity (FL) normalized by Renilla fluorescence (RL) intensity was calculated 24 h after transfection. (E, F) Spearman correlation analysis of miR-224 and ANGPTL1 expression in the TCGA-LUAD database (E) and in our cohort of 155 cases of lung cancer (F). The data in D are presented as the means 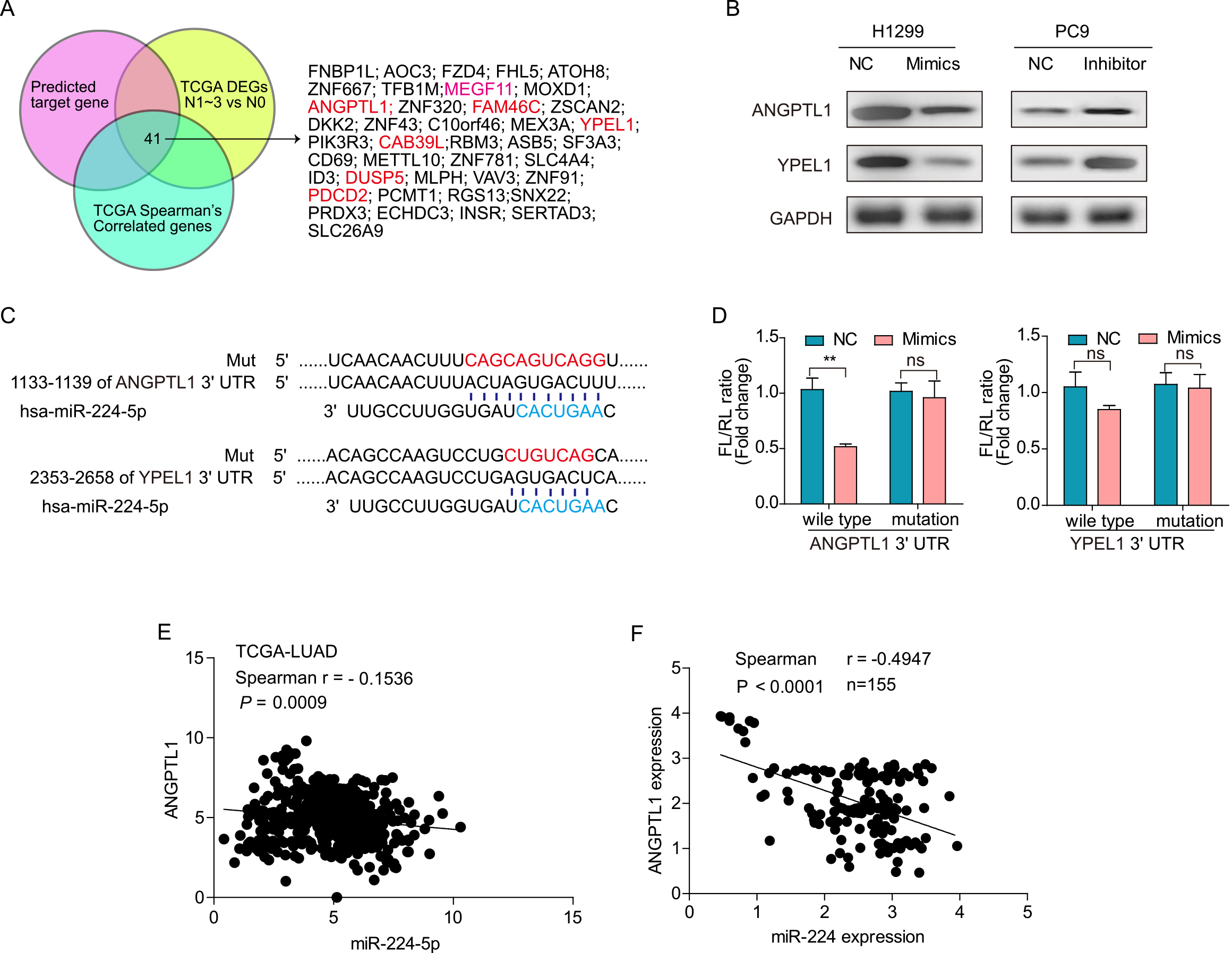
ANGPTL1 rescue expression partially diminishes the miR-224-mediated promotion effect on the malignant phenotype of H1299 cells. (A) Western blot determines ANGPTL1 rescue expression. (B) CCK-8 assays were performed to determine cell viability. (C) Plate colony formation assays were performed to detect cell growth. (D) Transwell assays were used to monitor cell migration and invasion. Cell numbers were counted in four different fields. Scale bars, 200 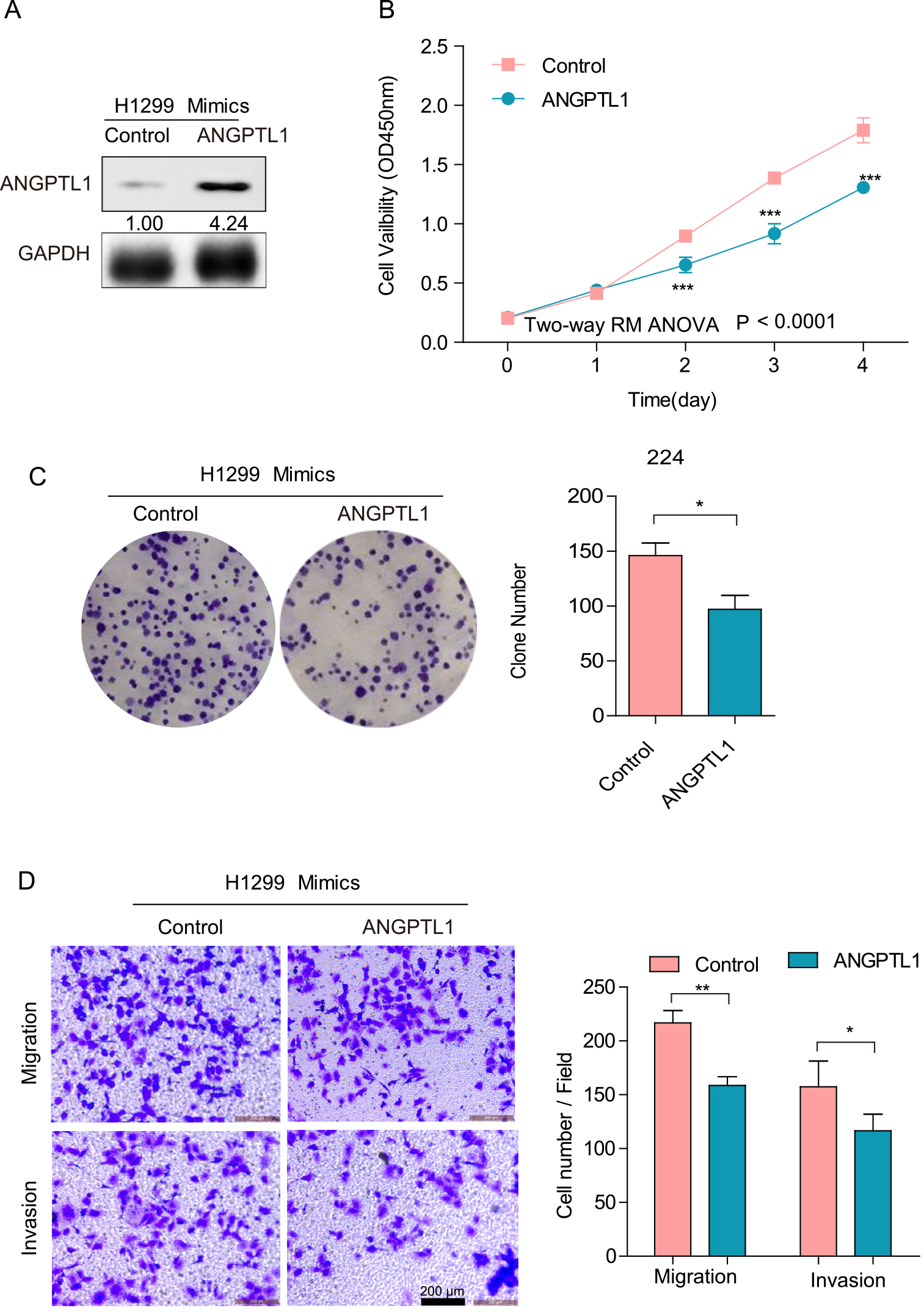
To further determine the role of ANGPTL1 in the promotion effect on the malignant phenotype mediated by miR-224, we reintroduced ANGPTL1 expression in miR-224-transfected H1299 cells (Fig. 6A,
Discussion
Lymph node status is of great importance in NSCLC, not only for prognosis but also to guide postoperative therapeutic strategies [16]. Previous reports showed that some miRNAs as molecular markers were correlated with either lymph node metastasis or distant metastasis. MiRNA-130a [17] and miR-196a [18] expression were positively associated with lymph node metastasis, while miRNA-451 [19] expression was negatively associated with lymph node metastasis. MiR-328 [20] was validated to have predictive value for brain metastasis with a sensitivity and specificity of 75% and 81.8%, respectively. These studies revealed that miRNAs might be correlated with and used to predict the lymphatic metastasis of NSCLC. In the present study, we aimed to evaluate the role of miR-224 in the lymph node metastasis of NSCLC. Previously, miRNA microarray analysis identified the differentially expressed miRNAs between lymphatic metastatic nodes and noncancerous lymph nodes from five NSCLC patients [13]. One miRNA, miR-224, exerted higher expression in tissues with lymph node metastasis. Here, a validation cohort consisting of 155 NSCLC patients was recruited to validate the positive correlation of miR-224 and lymph node metastasis.
MiR-224 has been revealed to be involved in the regulation of malignant characteristics in human cancers. It was reported to promote tumor growth and metastasis in gastric [21], colorectal [22], and papillary thyroid cancer [23]. In addition, miR-224 was reported to be associated with aggressive progression and poor prognosis in human cervical [24] and colorectal cancer [25]. In colonic cancer, miR-224 was identified as one of the miRNAs related to lymph node metastasis [26]. As for lung cancer, in 2014, Zhu et al. reported that decreased miR-224 expression was found in cancer tissues and tissues with lymph node metastasis by q-PCR analysis of 115 patients, and overexpression of miR-224 in A549 cells transfected with mimics resulted in suppression of cell migration and invasion by Transwell and wound healing assays [27]. However, contrary to the findings of Zhu et al., in 2015, Cui et al. published their results in the PNAS that miR-224 was significantly upregulated in NSCLC tissues, particularly in resected NSCLC metastasis by both q-PCR and in situ hybridization analysis, and increased miR-224 expression promoted cell migration, invasion, and proliferation [28]. In 2020, miR-224 was predicted to be one of the miRNAs associated with lymph node metastasis in lung adenocarcinoma only by bioinformatics analysis of TCGA and GEO datasets [29]. Here, consistent with the report from Cui et al., our microarray also identified miR-224 as an upregulated miRNA in tissues with lymph node metastasis. More importantly, we also validated its high expression of approximately 7-fold change in tissues with metastasis compared to nonmetastasis in our own cohort consisting of 155 patients. Although miR-224 alone was only weakly predictive for lymphatic metastasis with AUC
Previously, some direct targets of miR-224 were found to be involved in its function, such as by directly targeting the tumor suppressors TNF
Conclusions
Our findings highlight that miR-224 serves as an oncogene associated with lymph node metastasis in NSCLC patients. The promotion effect of miR-224 on the malignant phenotype might be due to its regulation of two tumor suppressors, ANGPTL1 and YPEL1. Thus, miR-224 may be a predictor for the lymph node metastasis of NSCLC.
Ethics approval and consent to participate
All methods and experiments were approved by the Ethics Committee of Peking University Cancer Hospital & Institute, and the informed consent was obtained from all the participants. All animal experiments were performed in accordance with guidelines and regulation of Animal Ethics Committee of Peking University Cancer Hospital & Institute. Written informed consent was obtained from the owners for the participation of their animals in this study.
Consent to publish
All authors give consent for publication.
Availability of data and materials
All relevant materials and data are available from corresponding author (Dr. Jinfeng Chen) for non-commercial purposes.
Funding
This study was supported by the National Natural Science Foundation of China (Grant No. 81773144, 81772632), Special funds of the Ministry of Finance for Reform and Development and Science Foundation of Peking University Cancer Hospital 2020-9.
Authors’ contributions
Study supervision and conception: JFC; Data acquisition: HBH; Analysis or interpretation of data: BP, FL, LNW, XJL and YY; Statistical analysis: HBH; Preparation of the manuscript: HBH. All authors approved the final manuscript.
Supplementary data
The supplementary files are available to download from http://dx.doi.org/10.3233/CBM-210376.
sj-docx-1-cbm-10.3233_CBM-210376.docx - Supplemental material
Supplemental material, sj-docx-1-cbm-10.3233_CBM-210376.docx
Footnotes
Acknowledgments
Not applicable.
Conflict of interest
All authors declare no competing interest.
