Abstract
Objective:
Although microRNA-103a (miR-103a) dysfunction has been implicated in various cancers, its relevance to non-small cell lung cancer (NSCLC) has not been clarified. This study was conducted to examine the molecular mechanism underlying the regulatory role of miR-103a in NSCLC.
Methods:
Kaplan–Meier analysis was carried out to assess the relationship between overall survival of NSCLC patients and miR-103a expression. Reverse-transcription quantitative polymerase chain reaction and western blot analyses were applied to evaluate the expression of relevant genes in tissues and cells. Sphere formation, MTS, flow cytometry, and Transwell assays were performed to characterize stemness. Dual luciferase reporter gene assays were used to clarify the binding relationship between miR-103a and ovarian tumor domain-containing ubiquitin aldehyde binding protein 1 (OTUB1). Finally, western blot analysis was used to assess the involvement of the Hippo pathway in NSCLC.
Results:
In NSCLC tissues and cells, miR-103a was expressed at low levels, whereas OTUB1 was expressed at high levels. Higher miR-103 expression levels were associated with a better prognosis for patients with NSCLC. When miR-103a was overexpressed, cell viability and stemness decreased, whereas apoptosis and cell cycle arrest were facilitated. The expression of phosphorylated YAP also decreased significantly. Opposite trends were observed after miR-103a silencing. OTUB1 expression and YAP phosphorylation decreased in the presence of miR-103a, and OTUB1 overexpression blocked the inhibitory effects of miR-103a on NSCLC cells.
Conclusion:
The miR-103a/OTUB1/Hippo axis may play a role in modulating the malignant behavior and stemness of cancer stem cells and thus could be a potential therapeutic target for the management of NSCLC.
Introduction
Lung cancer, which comprises heterogeneous tumors, can be divided into non-small cell lung carcinoma (NSCLC; which is further divided into adenocarcinoma, large cell carcinoma, and squamous cell carcinoma) and small-cell lung carcinoma. 1,2 Of all cancers, lung cancer was estimated to be the most commonly diagnosed (7,333,000 new diagnoses) and led to the most fatalities (6,102,000 deaths) in China in 2015. 3 In general, early-stage lung cancer patients are treated with surgery, whereas patients at an advanced stage or with metastases undergo chemotherapy. 4 The 5-year survival rate for patients with lung cancer in the United States is 15.6%, and although some enhancement in survival has been achieved over the past few decades, the advancements have not been as substantial as in other malignancies. 5 Cancer stem cells (CSCs) are considered a reservoir of tumor cells because they exhibit self-renewal properties and competence for reestablishing heterogeneous cancer cell populations. 6 Moreover, it is well established that CSCs are resistant to treatment and can cause tumor relapses, and can be newly generated under therapeutic pressure and in an altered microenvironment. 7 Therefore, further clarification of the molecular mechanisms associated with CSCs might offer novel insights for the treatment of NSCLC.
MicroRNAs (miRNAs) are noncoding RNAs approximately 22 nucleotides in length; miRNA dysfunction has been linked to various tumorigenesis events through mediation of changes in the cell cycle, apoptosis, and migration. 8 Interestingly, several miRNAs, including miR-124a and miR-181b, have been reported to inhibit the stemness of CSCs and help overcome drug resistance in NSCLC. 9,10 In addition, miR-103a-3p expression was observed to be remarkably reduced in NSCLC tissues and cell lines, and low expression levels were tightly correlated with dismal survival in NSCLC patients. 11 Moreover, miR-103a-3p plays a tumor suppressive role together with linc00152 depletion in glioma stem cells. 12 However, its effects on CSCs in lung cancer have yet to be clarified. Here, we postulate that miR-103a exerts regulatory functions over CSCs in NSCLC. Located within locus 11q13.1, ovarian tumor domain-containing ubiquitin aldehyde-binding protein 1 (OTUB1) is expressed in a wide range of human tissues, and OTUB1 knockdown in A549 cells was shown to suppress soft agar colony formation and xenograft tumor growth. 13 Moreover, OTUB1 is a biomarker for glioma pathogenesis and has the potential to be a clinical biomarker in glioma. 14 In this report, the expression pattern of miR-103a in NSCLC patients was investigated to determine if miR-103a has a prognostic value for NSCLC. Furthermore, we elucidated the probable molecular mechanism underlying the regulation of CSCs in NSCLC by miR-103a, which likely involves regulation of OTUB1.
Material and Methods
Collection of Tissue Samples
NSCLC tissues and adjacent normal lung tissues (at least 5 cm away from tumor tissues) were harvested from 73 patients diagnosed with primary NSCLC who underwent tumor resection at Jiangxi Tumor Hospital (Jiangxi, China) from December 2013 to June 2015. None of the patients received radiotherapy or chemotherapy before surgery. Patients who had other chronic diseases were excluded. The obtained tissues were snap-frozen and kept in liquid nitrogen at −80°C until further analysis. The patients were visited every month to collect information on their prognosis and survival. Detailed information pertaining to all patients is listed in Table 1.
Detailed Characteristics of the 73 Patients.

Cell Treatment and Transfection
Two cultured normal lung epithelial cell lines (Beas-2B and HBE), 5 NSCLC cell lines (95D, A549, NCI-H520, NCI-H460, and H1299), and human embryonic kidney 293 T (HEK293 T) cells were obtained from ATCC (Manassas, VA, USA). We confirmed that there was no mycoplasma contamination using the online Cellosaurus query tool (https://web.expasy.org/cellosaurus/). The normal lung epithelial cell lines (Beas-2B and HBE) were cultured in LHC-9 medium. The 95D, A549, NCI-H520, NCI-H460, and H1299 lines and HEK293 T cells were grown in Dulbecco’s modified Eagle’s medium (DMEM; Invitrogen, Carlsbad, CA, USA) with 1% penicillin/streptomycin (Invitrogen) and 10% fetal bovine serum (HyClone, Logan, UT, USA) in a 5% CO2 incubator at 37°C.
The A549 cells were trypsinized and resuspended at a concentration of 1 × 10 3 cells/mL in serum-free DMEM/F12 culture medium containing epidermal growth factor, basic fibroblast growth factor, insulin, bovine serum albumin, B27, and glucose to make up a single-cell suspension. The culture flask was placed vertically in an incubator and shaken 5 times a day to allow microspheres to form. Half of the medium was replaced every 2 days, and the cells were passaged every 6 days. Microspheres in the logarithmic growth phase were collected, detached with Accutase, triturated into single cells, and incubated with Hoechst 33342. Flow cytometry was used to detect side population (SP) cells. When the proportion of SP cells reached 25%, all spheres were collected and placed in phosphate-buffered saline (PBS) to produce a single-cell suspension. The cell suspension was incubated with a fluorescence-labeled monoclonal antibody to sort A549 stem cells (CD133+ and CD44+) with a fluorescence-activated cell sorter.
Vectors that contained the negative control (NC) mimic, miR-103a mimic, NC inhibitor, and miR-103a inhibitor; and NC- and OTUB1-overexpression (oe) vectors were generated by Life Technologies (Grand Island, NY, USA) and delivered into cells at a final concentration of 20 nM. In accordance with the instructions for using FuGENE6 transfection reagent (Promega, Madison, WI, USA), A549 cells were cultured overnight and transfected once they reached 60% confluence. Cells were collected at 24 h or 48 h after transfection.
Reverse-Transcription Quantitative Polymerase Chain Reaction (RT-qPCR)
Total RNA was extracted from cells and tissues using TRIzol reagent (Invitrogen). A NanoDrop Spectrophotometer 2000 (1011U; NanoDrop Technologies, Wilmington, DE, USA) was used to determine the concentration and purity of total RNA. Reverse transcription was performed with TaqMan MicroRNA Assays Reverse Transcription primer (4427975; Applied Biosystems, Inc., Foster City, CA, USA) to produce cDNA in accordance with the manufacturer’s instructions. The qPCR primers (Table 2) were all synthesized by Sangon Biological Engineering Technology & Services Co., Ltd. (Shanghai, China). Real-time fluorescence qPCR detection was performed using an ABI 7500 qPCR instrument (ABI, Oyster Bay, NY, USA). U6 and glyceraldehyde-3-phosphate dehydrogenase were used as endogenous controls for miRNA and gene expression levels, respectively. The relative quantification (2-ΔΔCt) method was used to calculate fold changes in gene expression.
Primer Sequence.

Note: miR-103a, microRNA-103a; OTUB1, ovarian tumor domain-containing ubiquitin aldehyde binding protein 1; GAPDH, glyceraldehyde-3-phosphate dehydrogenase.
3-(4,5-dimethylthiazol-2-yl)-5-(3-carboxymethoxyphenyl)-2-(4-sulfophenyl)-2H-tetrazolium (MTS) Assay
A549 cells were plated into a 96-well plate (Corning, Corning, NY, USA) at a density of 1 × 10 3 cells per well and cultured at 37°C and 5% CO2. Cell viability in each group was analyzed using a CellTiter96 Aqueous One Solution Cell Proliferation Assay Kit (Promega) after 24 h of cell transfection. The cells were grown for 2 h with MTS reagent at 37°C and 5% CO2. Optical density at 490 nm was measured using a SpectraMax 340PC384 microplate reader to assess cell proliferation.
Flow Cytometry
After 48 h of transfection, cells at 80–90% confluence were detached with trypsin and fixed with 70% ethanol in phosphate-buffered saline (PBS) at −20°C overnight. The cell suspension (1 × 10 6 cells/mL) was then incubated with 50 μg/mL of propidium iodide in PBS in the absence of light for 10 min. Apoptosis and cell cycle distribution were analyzed using an Attune NxT flow cytometer (Thermo Fisher Scientific, Waltham, MA, USA).
Assessing Cell Migration and Invasion
Matrigel (Corning) was diluted with Roswell Park Memorial Institute (RPMI)-1640 medium (Solarbio, Beijing, China) and added into a chamber dropwise. Cells suspended in RPMI-1640 medium (1 × 10 4 cells/mL) were seeded into 6-well plates, whereas 500 µL of RPMI-1640 medium supplemented with 10% fetal bovine serum was placed in a basolateral chamber. After incubation for 24 h at 37°C and 5% CO2, cells in the basolateral chamber were washed with PBS, fixed in 4% paraformaldehyde for 30 min at room temperature, and stained with 0.5% crystal violet staining solution for 30 min. After fixation with neutral gum (Sigma-Aldrich, St. Louis, MO, USA), 5 visual fields were randomly selected under an inverted microscope (Eclipse Ti, Nikon, Tokyo, Japan) to photograph and count invaded cells. Except for the omission of Matrigel, the cell migration assay was performed using the same steps as in the invasion assay.
Sphere Formation Assay
Pretreated cells were resuspended in DMEM/F12 medium (500–1000 cells/mL) and cultured for 12 days in 6-well ultra-low-attachment petri dishes (Corning). Tumor spheres were counted and measured using a contrast microscope (EVOS M7000; Nikon) on day 12 to assess the growth of the tumor spheres.
Western Blot Assay
After 48 h of cell transfection, the cells were lysed on ice with radioimmunoprecipitation assay solution and centrifuged at 20,000 × g for 10 min at 4°C before the supernatant was collected. Subsequently, 15 µg of total protein was subjected to 12% sodium dodecyl sulfate-polyacrylamide gel electrophoresis and electroblotted onto a nitrocellulose membrane. Protein concentrations were determined with a bicinchoninic acid assay kit (Abcam Inc., Cambridge, UK). The proteins were then incubated with primary rabbit antibodies against OTUB1 (ab175200), YAP (ab52771), and serine-127-phosphorylated YAP (p-YAP; ab76252) as well as goat anti-rabbit IgG (ab97051) at 4°C. All antibodies were purchased from Abcam. Immunoreactive bands were measured using an enhanced chemiluminescence detection kit (Thermo Fisher Scientific).
Dual Luciferase Reporter Assay
We synthesized oligonucleotides in the OTUB1 mRNA 3′untranslated region (3′UTR) containing miR-103a-targeting sequences. The miR-103a-oe vector and vectors containing a miR-103a inhibitor and mutation were also designed. Targeting sites were mutated to construct a miR-103a mutant. The pGLO vector was used to construct the fluorescence reporter vector pGLO-OTUB1. The vectors were extracted using a plasmid purification kit (Invitrogen). Additionally, 293 T cells were cultivated in a 24-well plate. Then, 200 ng of pGLO-OTUB1 plasmid and 20 nM of the miR-103a mimic, inhibitor, or mutant were co-transfected for 1 day. Subsequently, the cell lysates were collected and luciferase activity was determined using the dual luciferase reporter system (E1910; Promega). Firefly luciferase activity was used to normalize activity levels.
Statistical Analysis
SPSS 22.0 software (IBM Corp., Armonk, NY, USA) was used for all statistical analyses. All data are displayed as the means ± standard deviations. Comparisons between 2 groups were analyzed using paired (between normal and tumor tissues) or unpaired (other groups) t-tests, whereas comparisons among multiple groups were assessed using 1-way or 2-way analysis of variation, followed by Tukey’s post hoc test. Patient survival was evaluated using Kaplan–Meier analysis. A p-value < 0.05 was taken to denote statistical significance.
Results
Levels of miR-103a Are Lower in NSCLC Tissues and Cells
We collected 73 matched pairs of NSCLC and normal lung tissue samples and used RT-qPCR to assess miR-103a expression in the samples. Levels of miR-103a were lower in tumor tissues than in normal tissues (Figure 1A). Thereafter, we divided patients into groups with high and low miR-103a expression based on the median miR-103 expression level of the 73 patients. Using these 2 groups, we analyzed the survival of NSCLC patients and found that miR-103a expression was closely correlated to overall survival rate, with a higher survival rate observed in patients who express high levels of miR-103a (Figure 1B). Multivariate analysis of the effects of age, sex, smoking history, tumor node metastasis (TNM) stage, and miR-103a expression on overall patient survival showed that only miR-103a was significantly associated with patient survival (Table 3). Additionally, the TNM stage of NSCLC patients was negatively correlated with miR-103a expression (Figure 1C). Next, we examined the expression of miR-103a in the Beas-2B, HBE, 95D, A549, NCI-H520, NCI-H460, and H1299 cell lines. Compared with the Beas-2B and HBE cell lines, miR-103a expression was significantly lower in NSCLC cell lines (Figure 1D). Therefore, miR-103a expression may be linked to NSCLC occurrence. To further explore the regulatory relationship between miR-103a and CSCs in NSCLC, we selected A549 cells for screening and identifying SP cells in subsequent experiments (Figure 1E-G).
A Cox Proportional Hazards Model Analysis on Different Factors on NSCLC Prognosis.

Note: SE, standard error; CI, confidence interval; TNM, tumor-node metastasis; NSCLC, non-small cell lung cancer.

miR-103a is reduced in NSCLC tissues and cells and predicts good prognosis. (A) The miR-103a expression in NSCLC and adjacent normal tissues determined by RT-qPCR (*p < 0.05 according to the 2-way ANOVA); (B) Kaplan-Meier analysis of the survival rate of NSCLC patients with high (blue) or low (red) miR-103a expression; (C) correlation analysis of miR-103a expression with TNM stage of NSCLC patients; (D) the miR-103a expression in Beas-2B, HBE, 95D, A549, NCI-H520, NCI-H460 and H1299 cells determined by RT-qPCR (*p < 0.05 according to the 2-way ANOVA); (E) observation of SP cells and non-SP cells morphology under a contrast microscope; (F) SP and non-SP cells sorted by flow cytometry; (G) the statistical analysis of panel E (*p < 0.05 according to the unpaired t test). Each reaction was run in triplicate.
miR-103a Inhibits CSC Proliferation and Facilitates Apoptosis in NSCLC
To confirm whether miR-103a expression affects the biological properties of CSCs in NSCLC, we overexpressed and silenced miR-103a in sorted CSCs and analyzed miR-103a expression in each group of cells using RT-qPCR. Compared to the respective controls, miR-103a expression increased and decreased markedly in the presence of its mimic and inhibitor, respectively (Figure 2A). We then assessed cell proliferation activity using the MTS assay. Cell proliferation was repressed by the miR-103a mimic and promoted by the miR-103a inhibitor (Figure 2B). In addition, relative to the NC-mimic treatment, the cell cycle in miR-103a mimic-transfected cells was blocked at the G0/G1 phase, and apoptosis rates increased sharply (Figure 2C and D). Compared to cells transfected with the NC inhibitor, the proportion of cells treated with the miR-103a inhibitor in G0/G1 decreased significantly, and less apoptosis was observed.

Ectopic expression of miR-103a attenuates the proliferation, while accelerates apoptosis of NSCLC cells. CSCs were delivered with miR-103a mimic or inhibitor with NC mimic or inhibitor as controls. (A) The miR-103a expression in cells after transfection determined by RT-qPCR (*p < 0.05 according to the 1-way ANOVA); (B) OD value of cells at the 0th, 24th, 48th, 72nd, and 96th h measured by MTS assay (*p < 0.05 according to the 2-way ANOVA); (C) cell cycle distribution measured by flow cytometry (*p < 0.05 according to the 1-way ANOVA); (D) cell apoptosis evaluated by flow cytometry (*p < 0.05 according to the 1-way ANOVA); the experiment was repeated 3 times independently.
miR-103a Inhibits CSC Sphere Formation, Migration, and Invasion in NSCLC by Impairing YAP Phosphorylation
Next, we assessed cell invasion (Figure 3A) and migration (Figure 3B) in each group. Ectopic expression of miR-103a diminished cell migration and invasion rates, whereas the opposite trend was observed with miR-103a depletion. Furthermore, the overexpression of miR-103a led to a decrease in sphere size and number, whereas the miR-103a inhibitor facilitated sphere formation in terms of number and size (Figure 3C). It is widely accepted that the Hippo signaling pathway influences the development of NSCLC. To explore whether miR-103a affects the Hippo signaling, we analyzed YAP expression and the extent of YAP phosphorylation in cells using western blot assays (Figure 3D). YAP expression did not differ significantly among groups. However, relative to the respective controls, YAP phosphorylation was significantly attenuated by the miR-103a mimic and enhanced by the miR-103a inhibitor. These results indicate that miR-103a suppresses the biological activities of CSCs through engagement of the Hippo signaling pathway in NSCLC.

Ectopic expression of miR-103a attenuates the migration, invasion and sphere formation of NSCLC cells by regulating the Hippo pathway. (A) cell invasion tested by Matrigel invasion assay (*p < 0.05 according to the 1-way ANOVA); (B) cell migration examined by migration assay (*p < 0.05 according to the 1-way ANOVA); (C) sphere-formation activities of cells detected by sphere-formation assay (*p < 0.05 according to the 1-way ANOVA); (D) the expression of YAP and the extent of YAP phosphorylation measured by western blot (*p < 0.05 according to the 2-way ANOVA); the experiment was repeated 3 times independently.
OTUB1 Is Directly Targeted and Inhibited by miR-103a in NSCLC Cells
We examined OTUB1 mRNA expression in NSCLC and adjacent normal lung tissue samples using RT-qPCR, and found higher OTUB1 expression levels in NSCLC tissue samples (Figure 4A). Furthermore, miR-103a expression and OTUB1 mRNA levels in the NSCLC tissue samples were significantly and negatively correlated (Figure 4B). As predicted by the bioinformatics website, miR-103a and OTUB1 share a complementary sequence (Figure 4C). To further confirm the effects of miR-103a on the translation of OTUB1 mRNA, a dual luciferase reporter assay was performed. Compared with the NC, the luciferase activity exhibited by the PGLO-OTUB1 reporter vector decreased in the presence of the miR-103a mimic, increased in the presence of the miR-103a inhibitor, and did not change significantly in the presence of the mutant vector (Figure 4D). OTUB1 mRNA and protein expression in transfected cells was also analyzed using RT-qPCR (Figure 4E) and western blot assays (Figure 4F and G). OTUB1 expression decreased in the presence of the miR-103a mimic compared to the NC mimic but increased significantly in the presence of the miR-103a inhibitor compared to the NC inhibitor.

miR-103a targets and negatively mediates OTUB1 in NSCLC cells. (A) the mRNA expression of OTUB1 in NSCLC and adjacent normal tissues determined by RT-qPCR (*p < 0.05 according to the 2-way ANOVA); (B) the correlation analysis of miR-103a and OTUB1 mRNA in tissue samples of NSCLC; (C) the sequences of miR-103a binding sites in OTUB1 3′UTR; (D) the analysis of relative luciferase activities of PGLO-OTUB1 in cells treated with miR-103a mimic, inhibitor or mutant (*p < 0.05 according to the 1-way ANOVA); (E), mRNA expression of OTUB1 determined by RT-qPCR (*p < 0.05 according to the 1-way ANOVA); (F) protein expression of OTUB1 determined by western blot assays; (G) the statistical analysis of panel F (*p < 0.05 according to the 1-way ANOVA); the experiment was repeated 3 times independently.
OTUB1 Reverses the Anti-Proliferation and Pro-Apoptotic Effects of miR-103a on CSCs in NSCLC
To further verify whether the regulation of OTUB1 by miR-103a affects the development of CSCs in NSCLC, we transfected A549 cells with NC mimic + NC-oe, miR-103a mimic + NC-oe, NC mimic + oe-OTUB1, or miR-103a mimic + OTUB1-oe vectors. OTUB1 and miR-103a expression in the co-transfected cells was analyzed using RT-qPCR and western blot assays (Figure 5A and B). Elevated miR-103a expression and downregulated OTUB1 mRNA and protein expression were observed in cells transfected with miR-103a mimic + NC-oe compared with cells transfected with NC mimic + NC-oe, whereas elevated OTUB1 mRNA and protein expression was observed in cells transfected with NC mimic + OTUB1-oe. OTUB1 expression increased significantly following miR-103a mimic + OTUB1-oe treatment compared to that following miR-103a mimic + NC-oe treatment. As expected, miR-103a mimic + NC-oe transfection led to markedly reduced proliferation (Figure 5C), cell cycle arrest (Figure 5D), and accelerated apoptosis (Figure 5E) relative to NC mimic + NC-oe transfection. Opposite trends were observed with NC mimic + OTUB1-oe treatment.

Overexpression of OTUB1 expedites proliferation and prevents apoptosis in NSCLC cells in the presence of miR-103a. Cells were treated miR-103a mimic and/or oe-OTUB1. (A) miR-103a expression and mRNA expression of OTUB1 determined by RT-qPCR (*#& p < 0.05 according to the 2-way ANOVA); (B) the protein expression of OTUB1 determined by western blot assays (*#& p < 0.05 according to the 1-way ANOVA); (C) OD value of cells at the 0th, 24th, 48th, 72nd, and 96th h measured by MTS assay (*#& p < 0.05 according to the 2-way ANOVA); (D) cell cycle distribution measured by flow cytometry (*#& p < 0.05 according to the 2-way ANOVA); (E) cell apoptosis evaluated by of flow cytometry (*#& p < 0.05 according to the 1-way ANOVA). The experiment was repeated 3 times independently.
OTUB1 Reverses the Inhibitory Effects of miR-103a on CSC Migration and Invasion in NSCLC
The results of the cell invasion assay showed that the invasion ability of cells transfected with miR-103a mimic + NC-oe decreased significantly compared with the NC mimic + NC-oe group, whereas that of cells transfected with NC mimic + OTUB1-oe increased significantly. Compared with the miR-103a mimic + NC-oe treatment, cell invasion ability following miR-103a mimic + OTUB1-oe transfection increased significantly (Figure 6A). Similar results were obtained with the cell migration assay (Figure 6B). Sphere formation assays were performed to analyze the efficiency of sphere formation in each group of cells. Transfection with miR-103a mimic + NC-oe led to a significant decrease in the number and size of spheres formed compared with transfection with NC mimic + NC-oe, whereas transfection with NC mimic + OTUB1-oe led to a substantial increase in the number and size of spheres formed. Furthermore, transfection with miR-103a mimic + OTUB1-oe significantly increased sphere size and number compared with transfection with miR-103a mimic + NC-oe (Figure 6C). To further confirm the regulatory relationship between miR-103a and OTUB1 in the Hippo signaling pathway, we analyzed the expression of relevant members of the signaling pathway—YAP and p-YAP—in each group of cells using western blot assays (Figure 6D). YAP phosphorylation decreased upon transfection with miR-103a mimic + NC-oe but increased significantly upon transfection with NC mimic + OTUB1-oe. Moreover, levels of p-YAP were upregulated in miR-103a- and OTUB1-overexpressing cells. The results thus indicate that miR-103a acts as a tumor suppressor that inhibits CSC stemness in NSCLC by activating the Hippo signaling pathway via OTUB1.
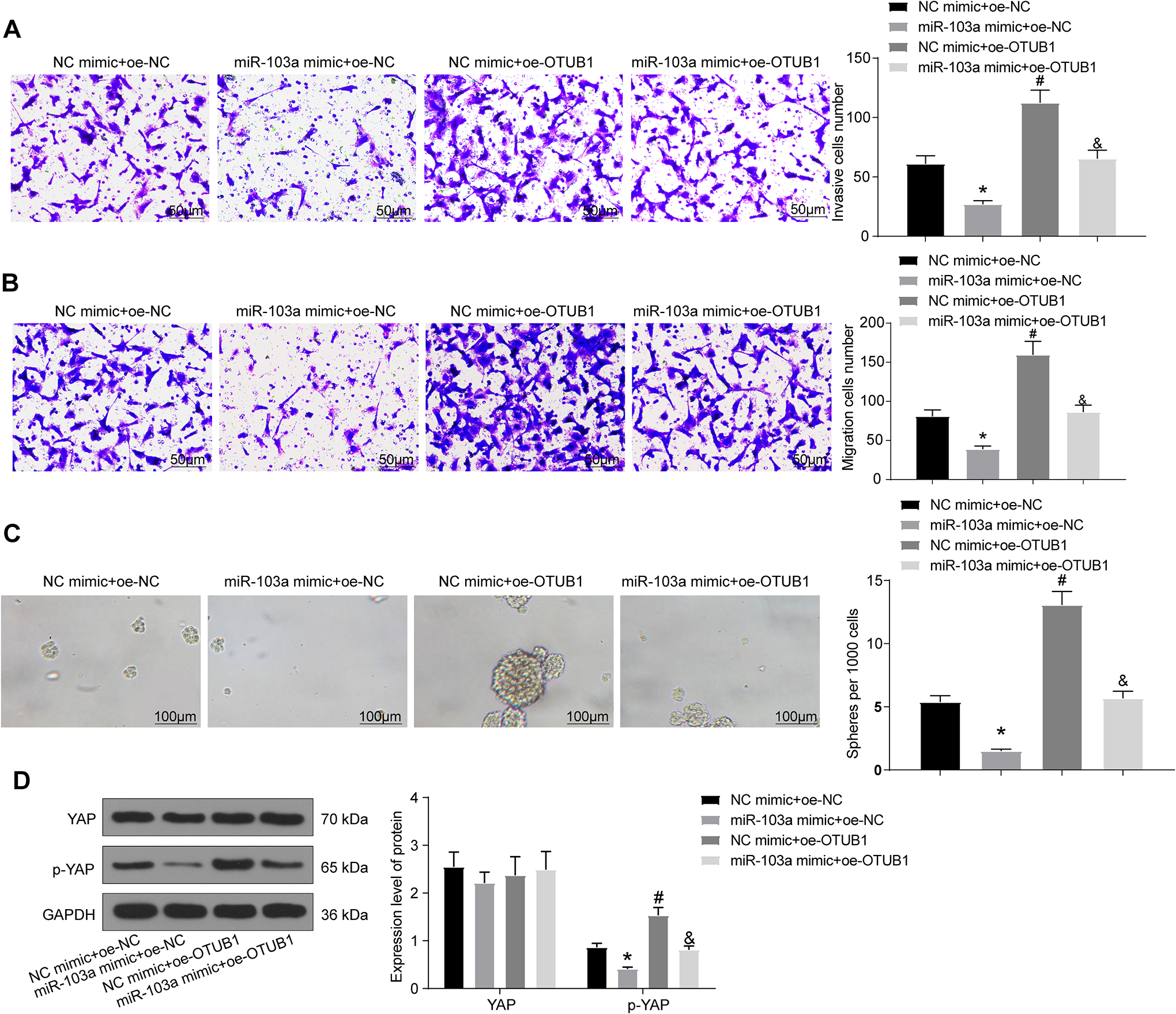
miR-103a exerts anti-migration and anti-invasive properties in NSCLC cells by regulating Hippo pathway via interacting with OTUB1. (A) Cell invasion tested by Matrigel invasion assay (*#& p < 0.05 according to the 1-way ANOVA); (B) cell migration examined by migration assay (*#& p < 0.05 according to the 1-way ANOVA); (C) sphere-formation activities of cells detected by sphere-formation assay (*#& p < 0.05 according to the 1-way ANOVA); (D) the expression of YAP and the extent of YAP phosphorylation measured by western blot (*#& p < 0.05 according to the 2-way ANOVA). The experiment was repeated 3 times independently.
Discussion
The projected overall survival rate for lung cancer in 2020 for all stages combined is one of the lowest (19%) of all cancers, followed by pancreatic (9%) and liver cancers (18%). 15 CSCs, which are characterized by their capacity for self-renewal and differentiation, play a significant role in the development of different cancers, such as melanomas and gliomas. 16 -18 Thus, a thorough understanding of the mechanisms that maintain CSC stemness is of great importance for the development of anti-cancer therapies. 19 Because there is still no effective treatment that targets CSCs to date, much research attention has been paid to CSC-targeting measures. 17 Here, we found that miR-103a expression was markedly reduced in NSCLC tissues and cells. In addition, the downregulation of miR-103a was correlated to low survival rates in NSCLC patients. Restoration of miR-103a suppressed CSC viability, migration, invasion, and sphere formation as well as promoted apoptosis in NSCLC. OTUB1 was revealed to be a target of miR-103a. The results further indicate that miR-103a represses CSC self-renewal and stemness in NSCLC through the Hippo signaling pathway by targeting OTUB1.
In the present study, we first verified that miR-103a is significantly downregulated in NSCLC clinical samples and cell lines. A previous study reported that miR-103a levels were low in both gastric cancer cells and clinical cancer specimens, and that levels were linked to the TNM stage of gastric cancer. 20 Additionally, miR-103a-3p was found to be weakly expressed in bladder cancer and to promote cell proliferation and migration via interactions with CDK6. 21 In our experiments, transfection with the miR-103a mimic led to reduced CSC viability, migration, invasion, and sphere formation, whereas opposite trends were observed following transfection with the miR-103a inhibitor. Similarly, miR-103a-3p was reported to repress growth and metastasis via regulation of KRAS signaling and the epithelial-to-mesenchymal transition in NSCLC cells. 22 The miR-103a mimic reduced the extent of YAP phosphorylation in NSCLC cells, indicating that miR-103a is linked to the Hippo signaling pathway. YAP, a mammalian homolog of Drosophila yorkie that acts as an effector in the Hippo signaling pathway, is often induced in malignancies and may exert tumor-promoting effects by activating CSCs. 23 Furthermore, YAP regulates key cellular functions, including proliferation control, suppression of apoptosis, and enhancement of metastasis, in many tissues, and the Hippo signaling pathway functions as an important tumor suppressor by negatively regulating oncogenic YAP. 24 Taken together, weakly expressed miR-103a is likely associated with the maintenance of CSC stemness through dysfunction of the Hippo signaling pathway.
An online tool was used to identify the molecular targets of miR-103a, and OTUB1, which was overexpressed in NSCLC cells and tissues, was found to harbor putative binding sites for miR-103a. OTUB1 expression has also been reported to be substantially higher in colon cancer tissues than in matched normal tissues and correlated with tumor size and lymph node metastasis in patients with colon cancer. 25 Wound-healing and Transwell assays conducted by Weng et al. revealed that OTUB1 exhibited pro-migration and pro-invasive properties in gastric cancer. 26 In addition, OTUB1 was identified as an oncogene that accelerates the progression of hepatocellular carcinoma pathogenesis, with OTUB1 expression correlated with pathogenic status. 27 Similarly, transfection with NC mimic + OTUB1-oe facilitated cell viability, migration, and invasion; cell cycle entry; and sphere formation but suppressed cell apoptosis. OTUB1 upregulation reversed the inhibition of cell proliferation, migration, and invasion and the enhancement of apoptosis induced by miR-524-3p, implying that miR-524-3p blocks colorectal cancer cell mobility by directly downregulating OTUB1. 28 The repression of esophageal cancer cell migration and invasion by miR-542-3p was also at least partly attenuated by the introduction of recombinant OTUB1. 29 Our rescue experiments provide strong evidence indicating that OTUB1 abrogates the carcinostatic function of miR-103a in NSCLC; i.e. the inhibition of CSC viability, migration, invasion, and sphere formation. Moreover, miR-103a can activate the Hippo signaling pathway by impairing YAP phosphorylation in an OTUB1-dependent manner. We postulate that OTUB1 directly or indirectly interacts with the WW domain of YAP1 to promote the translocation of endogenous YAP from the nucleus to the cytoplasm as well as YAP phosphorylation. This topic should be studied further. As only in vitro assays were performed in this study, additional in vivo experiments are needed to confirm our findings.
Conclusion
Briefly, we found that high levels of miR-103a expression led to a decrease in CSC proliferation, invasion, migration, and sphere formation as well as promoted apoptosis in NSCLC via the downregulation of OTUB1 and activation of the Hippo signaling pathway (Figure 7). Therefore, this axis is a potential therapeutic target for NSCLC.

The schematic cartoon of the mechanism of miR-103a as a tumor suppressor by modulating the OTUB1/Hippo signaling pathway in CSCs in NSCLC. After YAP phosphorylation, Hippo signaling pathway is blunted to promote proliferation, migration, invasion and sphere formation of CSCs and to inhibit their apoptosis in NSCLC. Overexpression of miR-103a inhibits the expression of OTUB1 and induces the Hippo signaling pathway, thereby inhibiting the stemness of CSCs in NSCLC.
Footnotes
Abbreviations
Authors’ Note
All patients approved to be enrolled by providing written informed consent. The study was accepted by the Ethical Committee of Jiangxi Tumor Hospital (approval number: 2013023), and complied with the Declaration of Helsinki.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
