Abstract
BACKGROUND:
Nasopharyngeal carcinoma (NPC), the common malignant head and neck cancer, is highly prevalent in southern China. The molecular mechanism underlying NPC tumorigenesis is unclear. We used 5-Aza-CdR, a DNA methyltransferase inhibitor, to treat NPC cell lines and discovered that the expression of TMEM130 changed significantly compared with the untreatment cells. This study aimed to identify the relationship between the DNA methylation status of TMEM130 and NPC, and to explore the function of TMEM130 in NPC cell migration.
METHODS:
qRT-PCR was performed to investigate the transcriptional expression of TMEM130 in NPC. Bisulfite sequencing PCR and 5-Aza-CdR treatment were used to detect the methylation level of the TMEM130 promoter. Gene Expression Omnibus (GEO) datasets were obtained to identifiy the methylation status and mRNA expression of TMEM130 in NPC and normal control tissues. Transwell and western blot analyses were used to detect cell migration ability after transfection of TMEM130/NC plasmids in NPC cells.
RESULTS:
The transcriptional expression of TMEM130 was decreased in NPC cell lines compared with in the NP69 cell line. TMEM130 promoter was significantly hyper methylated in three NPC cell lines (C666, CNE, and HONE) but hypo methylated in NP69 cells. The methylation level was higher in NPC than normal control tissues. Additionally, treatment of NPC cells with 5-Aza-CdR increased the TMEM130 mRNA expression level. Overexpression of TMEM130 in NPC cell lines suppressed cell migration ability and affected some epithelial-mesenchymal transition-associated gene expression.
CONCLUSIONS:
This study is the first to investigate the expression and function of TMEM130 in NPC. It was found that TMEM130 hyper methylation might contribute to NPC migration and this gene might act as a tumor suppressor gene. TMEM130 is a promising biomarker for NPC diagnosis.
Introduction
The primers for qRT-PCR and BGS
The primers for qRT-PCR and BGS
qRT-PCR, quantitative real-time PCR; BSP, bisulfite sequencing PCR; F, forward primer; R, reverse primer; bp, base pair.
Nasopharyngeal carcinoma (NPC) is a kind of malignant tumor occurring in nasopharyngeal epithelial cells. In the report of GLOBOCAN 2018 (International Agency for Research on Cancer), there were nearly 130,000 new cases of NPC and more than 70,000 deaths worldwide [1]. The risk of NPC in southern China is high, in contrast, NPC cases in western countries are better differentiated and have relatively lower malignant potential [2]. NPC is the most metastatic head and neck tumor, and although the use of intensity-modulated radiotherapy and combined chemoradiotherapy has improved locoregional control, distant metastasis is currently the main reason for treatment failure [3]. The etiology of NPC mainly includes three aspects: environmental factors, genetic susceptibility, and Epstein-Barr virus infection [4]. However, the exact mechanism of the progression of NPC has not yet been fully described.
DNA methylation is the most common epigenetic mechanism for cancer initiation and progression. The methylation changes in the promoter or the first exon region can enhance transcriptional silencing of genes, especially tumor suppressor genes, which participates in the progress of NPC development [5]. Many studies have detected certain genes with aberrant methylation status in NPC. DNMT3B could promote radio resistance in NPC through methylation of p53 and p21 [6]. The hyper methylation of the promoter silences the expression of PCDH20 and enhances cell migration and invasion ability of NPC cells [7]. NPC remains a significant public health problem in China and southeastern Asia. Finding and identifying specific DNA methylation genes could contribute to the understanding of pathogenetic mechanisms and improve the availability of biomarkers as regards the diagnosis or prognosis judgment NPC.
In our previous research, we used 5-Aza-CdR, a DNA methyltransferase inhibitor, to treat NPC cell lines and sequenced them. It was discovered that the expression of some genes significantly changed compared with the untreatment groups. From that, TMEM130 gene was observed to have a significant up-regulation. TMEM130 gene was discovered in 2003, located on human chromosome 7, and belongs to the transmembrane protein superfamily [8]. There are a few studies on the TMEM130 gene; the functions and expressions are unknown in cancer. In this research, we focused on the methylation status of the promoter region of TMEM130 gene, analyzed the association of TMEM130 expression and DNA methylation, and its role in cancer development.
NPC cell lines and culture
Three human NPC cell lines were used in this research. The C666 cell line was established from an undifferentiated NPC and a kind gift from Lunquan Sun (Hsiang-Ya Medical College). CNE was derived from a female patient with NPC and a gift from Wei Zhu (Guangzhou Medical Collage). HONE and the immortalized nasopharyngeal epithelial cell line NP69 were purchased from the Cell Bank of the Chinese Academy of Sciences. The cells were maintained in DMEM supplement with 10% fetal bovine serum (Biological Industries, BI), and 2% penicillin/streptomycin at 37
RNA extraction and quantitative real-time PCR (qRT-PCR)
Total RNA was extracted from the cell lines using TRIzol (Invitrogen, USA) according to the manufacturer’s instruction. 1-
Protein extraction, western blot, and reagents
Western blot was conducted to assess the semi quantitative expression of proteins. The collected cells were washed with cold phosphate-buffered saline and extracted using radio immunoprecipitation (RIPA) Buffer (GBCBIO Technologies Inc.) and 1% phenylmethanesulfonylfluoride (PMSF) on ice. The cell lysates were centrifuged at 12,000 rpm for 15 minutes, and the supernatant was collected. A bicinchoninic acid assay kit (CWBIO, China) was used to measure the total protein concentration. Then, the protein sample was mixed with 5
Gene expression omnibs (GEO) microarray dataset ananlysis
To validate the methylation levels of TMEM130 promoter between NPC and normal control tissues, we obtained the GSE52068 and GSE62336 microarray from the GEO database and analysed the methylation levels in NPC and normal control cases. Then we got the GSE61218 and GSE64634 microarray from the GEO database and analysed the mRNA expression levels of TMEM130 in NPC and normal control cases.
Plasmids and transfection
The TMEM130 cDNA was cloned into the pcDNA3.1-3Flag-T2A-EGFP vector, while the pcDNA3.1-3Flag-T2A-EGFP vector was used as a negative control (NC). Then, the plasmids were transfected into CNE and HONE using lipofectamine 2000 (Invitrogen, Germany) according to the manufacturer’s protocol. After 48 h transfection, the cells were collected for the following test.
Treatment of cell lines with a DNA demethylation agent (5-Aza-CdR)
According to our pre-experiments, appropriate treatment time and concentration of DNA demethylation agent were chosen. When the cell confluency was 50%–60%, NPC cells were treated daily with 10-
DNA extraction and bisulfite treatment
The genomic DNA extraction method with SDS-proteinase K digestion and phenol-chloroform purification was used as described previously for cell lines, then modified with bisulfite as described previously [9]. 5-
TMEM130 promoter methylation analysis (BSP)
Bisulfite treatment could convert the unmethylated cytosine to uracil, but does not affect the methylated cytosine. The methylation status of the TMEM130 CpG locus in cell lines was determined using bisulfite sequencing PCR (BSP). The primer sequences and conditions for BSP are presented in Table 1. The purified PCR products were cloned into the pU18-T vector, and six clones from each sample were selected and sequenced in Mapbioo (Shanghai, China).
Migration assay
The migration ability of NPC cells was measured by Transwell plates with 8-
The transcriptional expression and DNA methylation in NPC cell lines. (A) TMEM130 mRNA expression detected by qRT-PCR in three NPC cell lines and immortalized nasopharyngeal epithelial cell line NP69. The 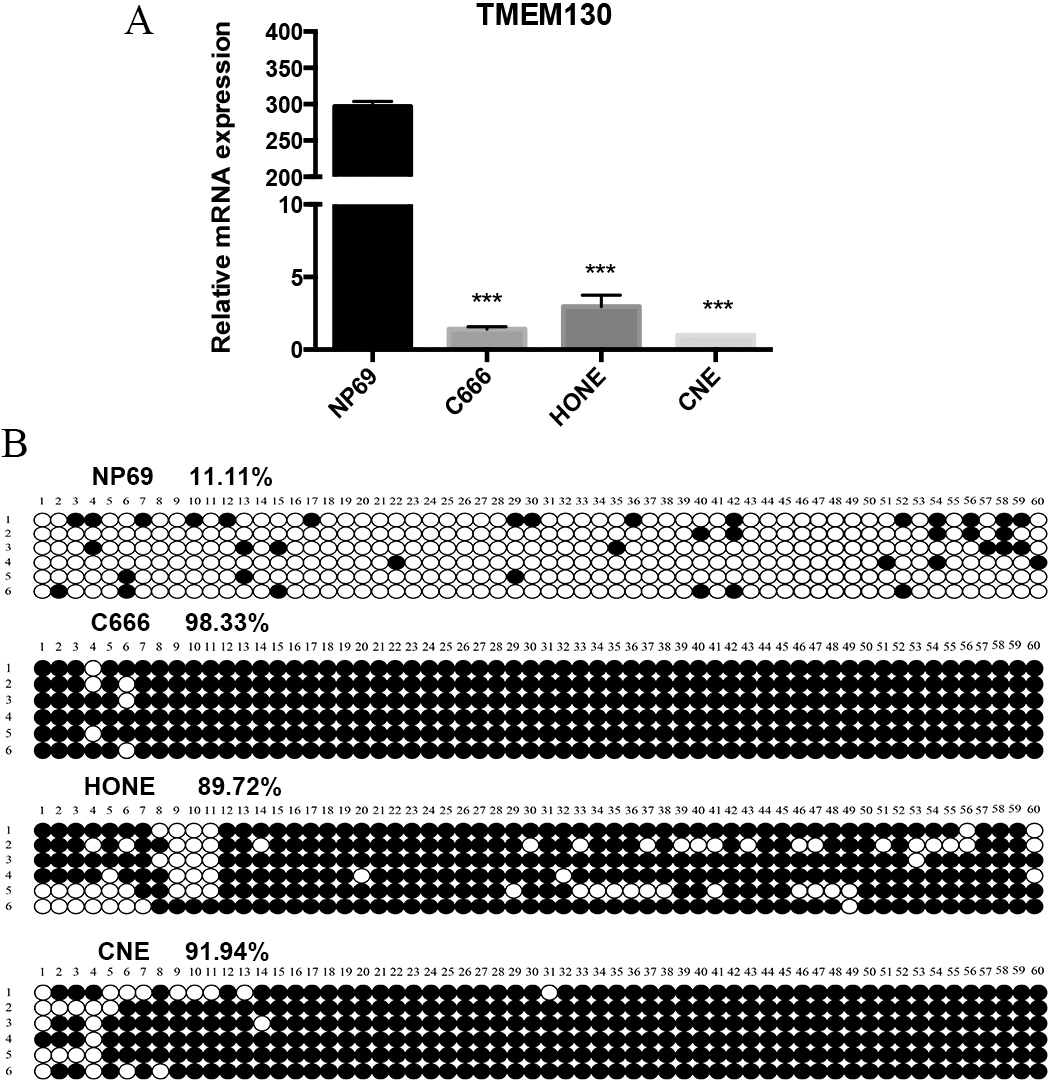
The DNA methylation level and transcriptional expression of TMEM130 in NPC and normal tissues. (A) The matrix map of TMEM130 methylation level in GSE52068, the blue group represents normal tissues, and orange group represents NPC groups. The methylation levels decrease from red to green. (B) The methylation levels of TMEM130 gene in NPC and normal tissues in GSE52068. 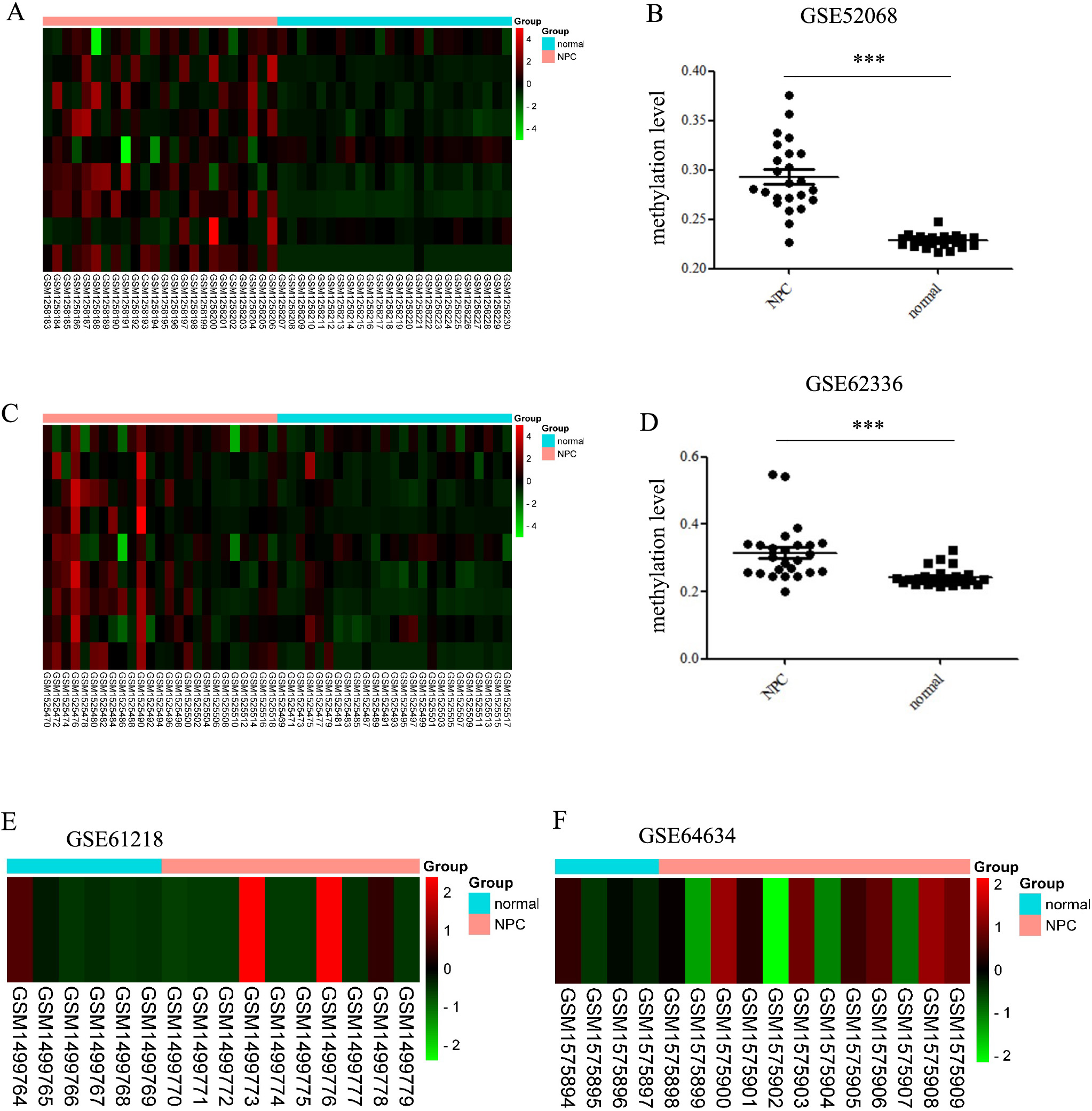
TMEM130 mRNA expression was restored after treatment with the demethylation reagent. (A) The expression of DNMT1 was reduced after the treatment of 5-Aza-CdR by western blot. NC, negative control; aza, 5-Aza-CdR treatment; 3D, three day’s treatment; 4D, four day’s treatment. (B) The methylation status of CpG sites in the TMEM130 gene promoter in CNE and HONE cell lines after the treatment of 5-Aza-CdR. (C) TMEM130 mRNA expression level in CNE and HONE cell lines after the treatment of 5-Aza-CdR. Expression levels were calculated by qRT-PCR. The 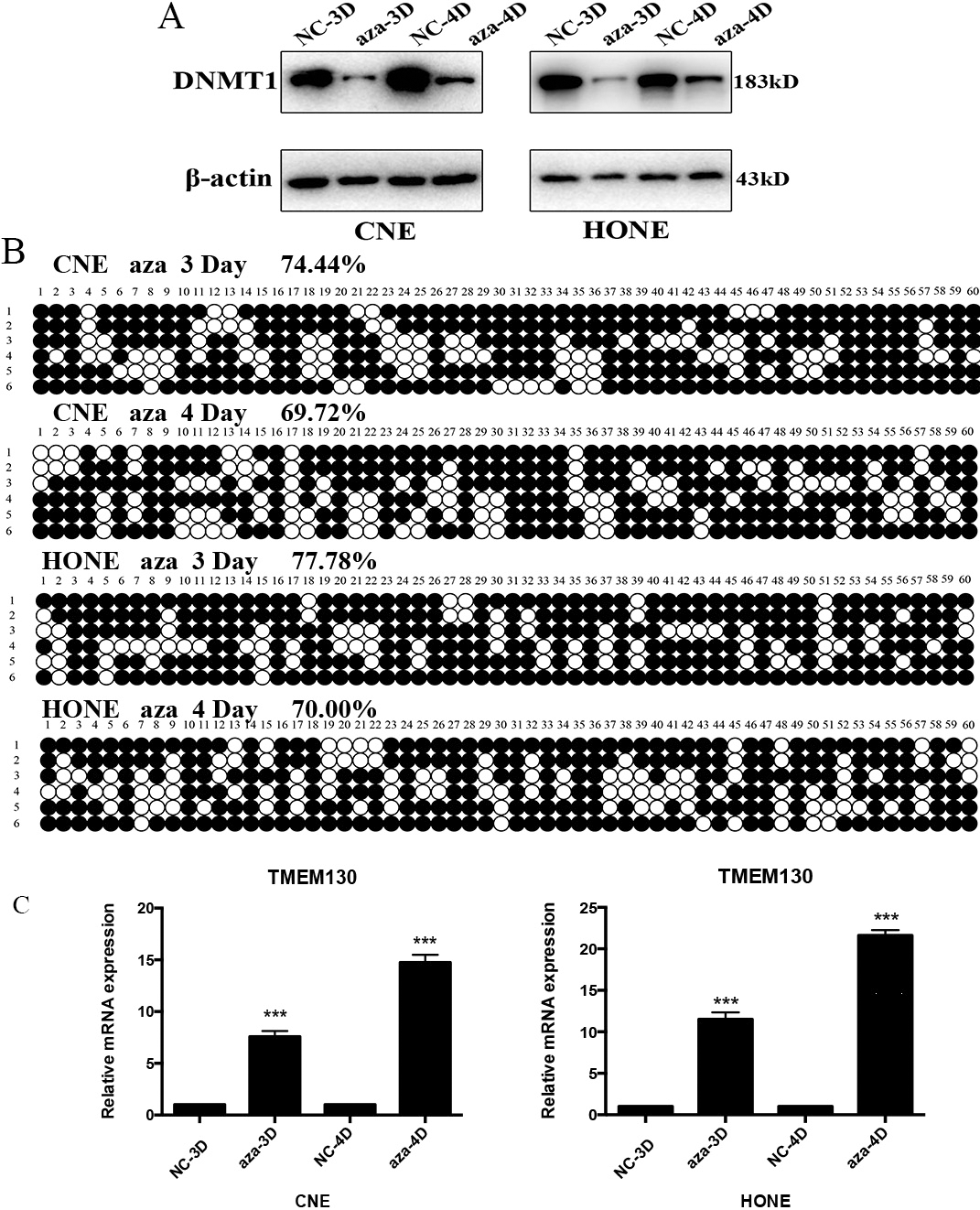
TMEM130 inhibition of NPC cell migration. (A) qRT-PCR was used to detect the expression of TMEM130 in CNE and HONE cells after 48 h transfection. 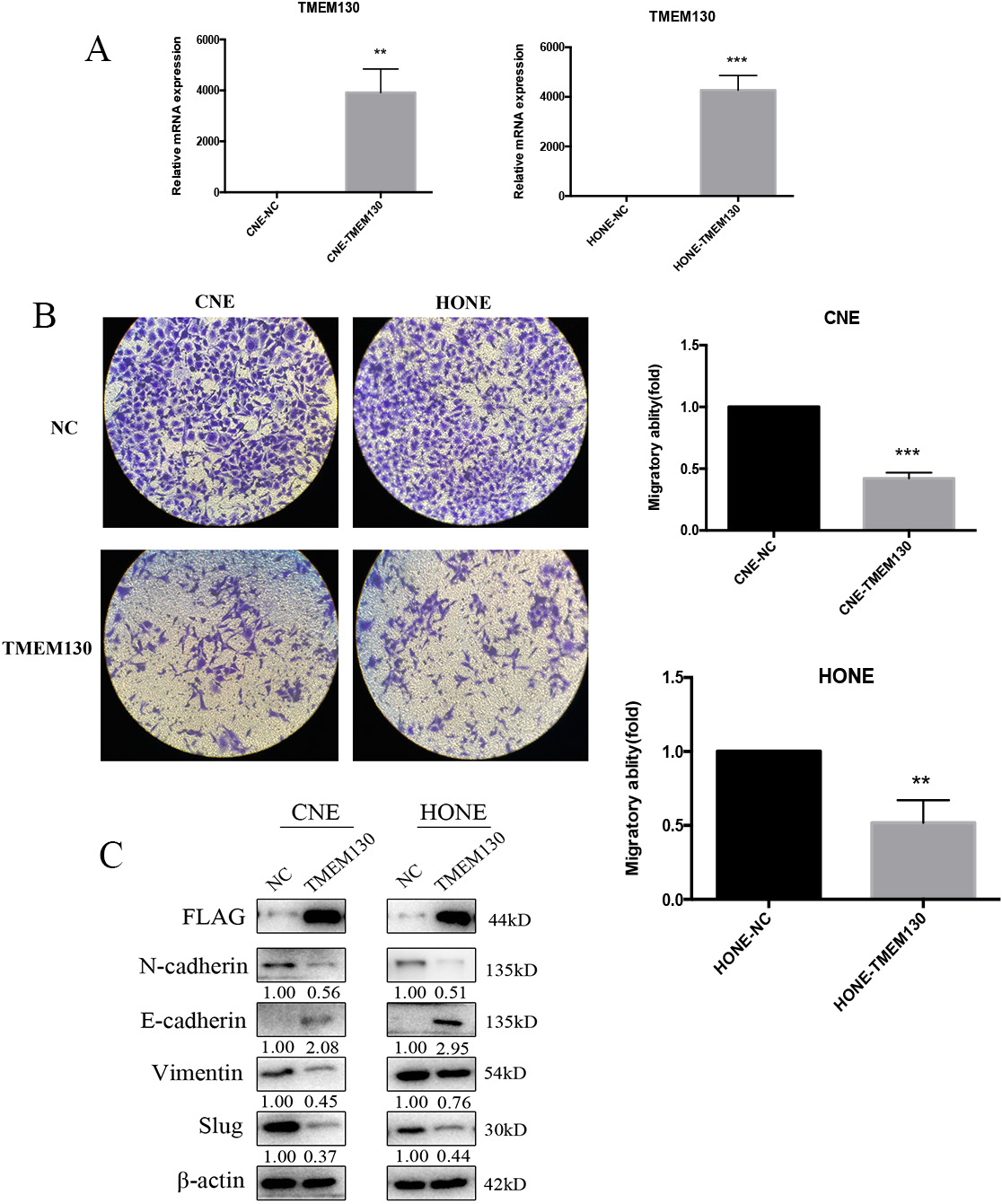
The statistical significance of the two groups was determined using Student’s
Results
The transcriptional expression of TMEM130 and its promoter methylation status
The transcriptional expression of TMEM130 was evaluated among the three NPC cell lines (C666, CNE, HONE) and an immortalized nasopharyngeal epithelial cell line NP69. Through qRT-PCR analysis, it was shown that NPC cell lines had a nearly silencing TMEM130 expression, while the expression of NP69 was high (Fig. 1A). To further assess the possible mechanism of down-regulation of TMEM130 expression in NPC cells, bioinformatics analysis indicated that TMEM130 contains a typical CpG island in its promoter region. Then, BGS was conducted. The results showed that methylation rates in the NPC cell lines were much higher than that of the NP69 cell line. The methylation rates of the NPC cell lines C666, CNE, HONE, were 98.33%, 89.72%, 91.94% respectively, while NP69 was only 11.11% (Fig. 1B).
GSE52068 (24 NPC tissues and 24 normal nasopharygeal epithelial tissues) and GSE62336 (25 NPC tissues and 25 non-tumor counterparts) datasets were downloaded from the GEO. GSE52068 and GSE62336 were genome-wide ananlysis of DNA methylation between NPC and normal nasopharygeal epithelial tissues/non-tumor counterparts. The results revealed that the methylation levels were much higher in NPC than in the normal controls (
TMEM130 expression is down-regulated by promoter hyper methylation in NPC
To further assess whether CpG island methylation directly mediated the TMEM130 expression, two NPC cell lines (CNE and HONE) were treated with the DNA methyltransferase inhibitor, 5-aza-2’-deoxycytidine (5-Aza-CdR). The expression of DNMT1 was detected to evaluate the efficacy of 5-Aza-CdR. BSP was used to verify the methylation status of TMEM130 promoter after the treatment of 5-Aza-CdR. After 3 or 4 days treatment (10-
Effects of TMEM130 on cell migration
To investigate the function of TMEM130 in NPC cells, we subsequently transfected CNE and HONE cells with TMEM130 plasmids and the vector control. The expression of TMEM130 was confirmed by qRT-PCR and western blot (anti-FLAG) after the plasmids were transfected for 48 h (Fig. 4A and C). Then, the cells were collected for Transwell, after another 48 h incubation, the number of migrated cells obviously decreased in the group of TMEM130 transfection compared to that of the control group (
Discussion
NPC is one of the most usual malignancies in southern China and southeastern Asia. Although considerable advancements have been made in NPC treatment, distant metastasis and local recurrences are frequently observed, this is the main challenge of NPC management. Therefore, there is a need for continuous studies to find specific tumor markers to predict tumor metastasis and improve patient prognosis.
To date, DNA methylation in the promoter region provides an important epigenetic mechanism for the initiation and progression of cancers. DNA hyper methylation in the promoter region could silence the transcriptional expression of genes, especially the tumor suppressor genes, causing carcinogenesis and the development of cancers [10]. DNA methyltransferases (DNMTs) plays an important role in DNA methylation and is involved in gene transcriptional silence. A series of experiments have been conducted to explore the role of DNMTs inhibitors in various cancers, such as 5-azacytidine and 5-aza-2’-deoxycytidine [11, 2, 13].
In this research, we have investigated that TMEM130 transcriptional expression in NPC cell lines compared with NP69 cell line. By qRT-PCR, we found that the transcriptional expression of TMEM130 was silenced in NPC cell lines (C666, CNE, and HONE), while NP69 had a high expression. Then, the methylation status of the TMEM130 promoter was detected using the BSP technique to explore the cause of low expression of TMEM130 in NPC cells. Hyper methylation of the promoter was detected in NPC cell lines (98.33%, 91.94%, 89.72%), while the normal control cell line NP69 was hypo methylated in the promoter (only 11.11%). DNMTs inhibitor, 5-Aza-CdR, was treated in CNE and HONE cells and restored the expression of TMEM130. These results support the fact that the hyper methylation of the promoter accounted for the downregulation of TMEM130 in NPC cells. The GEO datasets showed that the methylation level was higher in NPC tissues than normal controls. However, the mRNA expression had no difference between NPC tissues and normal controls, it might be related to the low number of specimens in the dataset. There is no research on the DNA methylation of TMEM130 in cancer, however, some transmembrane protein expressions are closely related to DNA methylation. Transmembrane protein 88 (TMEM88) is transcriptionally regulated by promoter methylation and acts as an inhibitor of Wnt signaling, hence, contributing to the development of platinum resistance in ovarian cancer [14]. In 2014, Xu et al. discovered that transmembrane protein 106A was silenced by the promoter region hyper methylation and suppressed gastric cancer growth by inducing cell apoptosis [15]. TMEM220 is highly methylated in gastric cancer tissues and could be restored by treating with 5-Aza-2’-deoxycytidine [16].
TMEM130 is a newly discovered gene, and its expression and function in tumorigenesis are unknown. However, some genes of the transmembrane protein superfamily to which TMEM130 belongs were found to inhibit cell proliferation and migration. TMEM137 was identified as a growth inhibitor with major histocompatibility complex (MHC) class II, and inhibited the expression of TRAP, cathepsin K, MMP-9 [17]. TMEM100 expression could suppress metastasis and enhance sensitivity to chemotherapy in gastric cancer [18]. TMEM100 acts as a cancer suppressor gene in non-small-cell lung cancer and hepatocellular carcinoma [19, 20]. In this study, we investigated the biological function of TMEM130 silencing by DNA methylation in promoting tumorigenesis and development of NPC. We further transfected TMEM130 and control plasmids into NPC cells and found that the overexpression of TMEM130 inhibited cell migration using Transwell assay. To determine whether TMEM130 affected cell migration via the modulation of epithelial-mesenchymal transition-associated genes, several genes were detected by western blot and further studied. Compared with the NC group, N-cadherin, vimentin, and slug were significantly decreased, while E-cadherin was increased in both CNE and HONE cell lines. These anti-tumor effects indicated that TMEM130 acts as a tumor suppressor gene in NPC and is associated with cell migration, thus having a potential therapeutic effect. However, the exact mechanism by which TMEM130 modulating cell migration needs further research.
In conclusion, the key finding of our study is that TMEM130 is inactivated by hyper methylation of the promoter region in NPC. Moreover, TMEM130 may be a potential tumor suppressor gene in tumorigenesis via inhibition of cell migration. This is the first study to explore TMEM130 expression and function in NPC, and may lead to new options for cancer diagnosis and therapy in the future.
Competing interests
The authors have declared that no competing interest exists.
Author contributions
Conception: Wen Zhang.
Interpretation or analysis of data: Hong Liu, Yong Zhang.
Preparation of the manuscript: Wenqiang Chen, Yan Zhang.
Revision for important intellectual content: Hong Liu, Wen Zhang.
Supervision: Wen Zhang.
Footnotes
Acknowledgments
This work was supported by Natural Science Foundation of Shandong Province [ZR2020MC020]; China postdoctoral Science Foundation [2020M682126].
