Abstract
Background
The crucial event driving nasopharyngeal tumorigenesis is the hypermethylation of chromosome 3p-located tumor suppressor genes. This case-control study aims to investigate the methylation characteristics of RASSF1A, Blu, ADAMTS9, and DLEC1 to potentially develop effective diagnostic biomarkers for nasopharyngeal carcinoma, either individually or in combination.
Methods
The methylation of RASSF1A, Blu, ADAMTS9, and DLEC1 in the collection of 93 biopsy samples and 100 healthy swab specimens were evaluated by Nested methylation-specific polymerase chain reaction. The strength of the correlation between candidate genes and nasopharyngeal carcinoma was estimated by the evaluation of odds ratios (ORs).
Results
Promoter hypermethylation of RASSF1A, Blu, ADAMTS9, and DLEC1 were found in 60.22%, 80.65%, 62.37%, and 74.19%, respectively, in nasopharyngeal carcinoma tumors. A significant association between the methylation status of candidate genes with nasopharyngeal carcinoma was reported. The methylation of candidate genes significantly increased the risk of nasopharyngeal carcinoma in cancerous samples compared with control samples (OR > 1). Based on the value of the methylation index, methylation of at least one gene was found in 95.70% of nasopharyngeal tumors. Additionally, the methylation index among 93 tumors significantly correlated with advanced stage nasopharyngeal tumors.
Conclusion
The study explored a higher frequency of hypermethylation at least one candidate gene. Methylation of a panel of potential genes can be utilized to discriminate between nasopharyngeal carcinoma and non-cancer cells, particularly in the advanced stages of nasopharyngeal carcinoma. Thus, it could serve as a valuable marker for the diagnosis and monitoring of nasopharyngeal carcinoma.
Introduction
Nasopharyngeal carcinoma (NPC) is an epithelial malignancy of the head and neck region with a particular racial and geographic distribution toward the Asian population. 1 NPC is especially endemic in Vietnam, based on the Global Cancer Observatory, where it reached the high 5-year prevalence of 16.63 cases per 100,000 people. 2 Due to extremely variable and unclear symptoms, the initial misdiagnosis of NPC is still common. 3 Even though NPC therapy has advanced significantly in recent decades, advance-stage detection of tumors typically leads to treatment failure and reduced patient survival. 1 Therefore, early diagnosis provides the possibility to enhance treatment outcomes, resulting in a high cure rate. 4 Finding reliable biomarkers for early detection of NPC would help to increase survival and to guide treatment, especially in countries with a high incidence of NPC.
Previous studies suggested that NPC presents multiple etiological factors, including (a) Epstein–Barr virus (EBV) infection; (b) environmental factors; and (c) genetic disorders.1,4 Events of epigenetics have been reported, which play significant roles in human tumorigenesis, including NPC. The process of epigenetics change has been postulated as one of the beginning events of various human malignancies, including NPC, via modulating many pathways of cell signals.5–13 For the past few years, growing evidence demonstrates that aberrant methylated promoter regions of tumor suppressor genes (TSGs) lead to its inactivation and participates in various processes of nasopharyngeal tumorigenesis.6,9,11,14 The fact is that the epigenetic alteration, including DNA hypermethylation, could be reversible and restored to its origins, thus making such initiatives a biomarker for diagnosis as well as therapy.9,15 The current genomic analysis postulated the silencing of TSGs, which are located on chromosome 3p, by the phenomena of DNA methylation; these are the frequent events that play key roles in the nasopharyngeal tumorigenesis.7,12,16–20 The inactivation of various chromosome 3p-located TSGs have been reported, including RASSF1A, Blu, RAR-β, DLEC1, ADAMTS9, etc.5,6,8,10,12,14 These alterations had been observed in NPC cell lines biopsy samples in various countries.5,6,8,10,12,14,21,22 In our previous studies, conducting on the Vietnamese NPC patients, the status of RASSF1A and Blu methylation were individually investigated in NPC biopsy samples.21,22 Analysis of the panel genes’ methylation (located at chromosome 3p) to be inactivated by DNA hypermethylation in NPC will expand the prospects for the development of potential biomarkers for novel screening tests for NCP. Based on this notion, we refined the selected chromosome 3p-located genes—including RASSF1A (involved in the cell-cycle progression), Blu (involved in cell progression and stress response), ADAMTS9 (involved in the angiogenesis), and DLEC1 (involved in cell communication, signaling transduction, and cell proliferation)—to evaluate their methylation properties towards applying as an individual or in combination to develop promising diagnostic biomarkers for NPC.
Materials and methods
Ethics statement and sample collection
The permission ID was 516/BVCR-HDDD given by the Medical Ethics Committee of the Cho Ray Hospital, Ho Chi Minh City, Vietnam. For the inclusion criteria: (a) all participants agreed to the use of their samples for laboratory testing and analysis by signing consent forms; (b) all samples were confirmed to be nasopharyngeal cancer by pathology; and (c) their records were up to date, complete, clear, accurate, and legible. A total of 93 biopsy samples were collected from NPC patients and 100 brushing samples from heathy people, collected from 2017 to 2018, in Cho Ray Hospital, Ho Chi Minh City, Vietnam, were enrolled in the current study. 23
Genomic DNA isolation, bisulfite modification, nested methylation-specific polymerase chain reaction
The phenol/chloroform procedure was used to extract total genomic DNA from biopsy and swab samples. Cells were lysed in a lysis solution that included 10 mM Tris-HCl pH = 8, 10 mM EDTA, 150 mM NaCl, 2% SDS, and 0.1 mg/mL proteinase. 23 Following that, ethanol precipitation was applied to purify the extracted genomic DNA. Bisulfite modification was performed by using 2 μg genomic DNA according to the guidelines of EpiJet Bisulfite Conversion Kit (Thermo Scientific, #1461).
The methylation status of candidate genes, including RASSF1A, Blu, ADAMTS9, DLEC1, were investigated by Nested-MSP assays. Supplemental Table S1 presents the sequences of primers used for stage 1 and stage 2 amplification. For polymerase chain reaction (PCR) assay, each reaction was performed in the volume of 15 μL, containing 200 ng bisulfite-modified template DNA (stage 1) or 200 ng stage 1 PCR product (stage 2), 7.5 l MyTaqTM Mix (Bioline, #25041), and 10 mM primers. The thermal cycling was set as: 95°C for 5 min, then following 40 cycles of denaturation at 95°C for 30 s, annealing at X°C for 30 s, extension at 72°C for 30 s, and a final extension at 72°C for 10 min. (X°C: each methylation or unmethylated primer annealing temperature is listed in Table S1). Electrophoresis was applied to visualize the products of Nested-MSP. Finally, MSP products were sequenced to confirm primer specificity and bisulfite modification effectiveness, and to determine the target gene's hypermethylation status.
Statistical analysis
The software of Medcalc® (Version 12.7.0.0) was used for statistical analysis. The Chi-square test was applied to evaluate the correlation between the status of methylation of each gene or combination thereof and clinicopathological variables. P ≤ 0.05 was considered a statistically significant difference of aberrant methylation frequencies among groups. The intensity of correlation between candidate gene promoters methylation and NPC were estimated by computing the value of odds ratios (ORs) with 95% confidence intervals (CIs).
Results
Characteristics of the participants
A total of 93 NPC biopsy samples were collected from patients who were diagnosed at Cho Ray hospital, Ho Chi Minh City, Vietnam. They included 68 male subjects (73.12%), and 25 female subjects (26.88%). The age of patients ranged from 20 to 81 (mean = 53.51 ± 1.43). The age group with the highest frequency was 40–60 years old (46.23%). They included 4 keratinizing squamous cell carcinoma subject (4.50%), 26 non-keratinizing carcinoma subjects (27.96%), and 63 undifferentiated carcinoma subjects (67.74%). NPC patients included 33 at stage II ( 35.48%), 15 at stage III (16.13%), and 45 at stage IV (48.39%). Also, 100 non-malignant swab samples were collected from heathy people. Among them, males accounted for 60.00% of the total and females for just 40.00%.
Methylation status in NPC biopsy samples and normal nasopharyngeal swab samples
The status of individual gene promoters methylation in the NPC samples and in normal nasopharyngeal swab samples are summarized in Table 1. The incidence of promoter hypermethylation in NPC samples was 60.22% (56 of 93) for RASSF1A, 80.65% (75 of 93) for Blu, 62.37% (58 of 93) for ADAMTS9, and 74.19% for (69 of 93) for DLEC1. Statistically, as shown in Table 1, a value of P < 0.0001 indicated a significantly higher association between individual genes with NPC.
Methylation of multiple genes in NPC and normal nasopharyngeal swab samples.
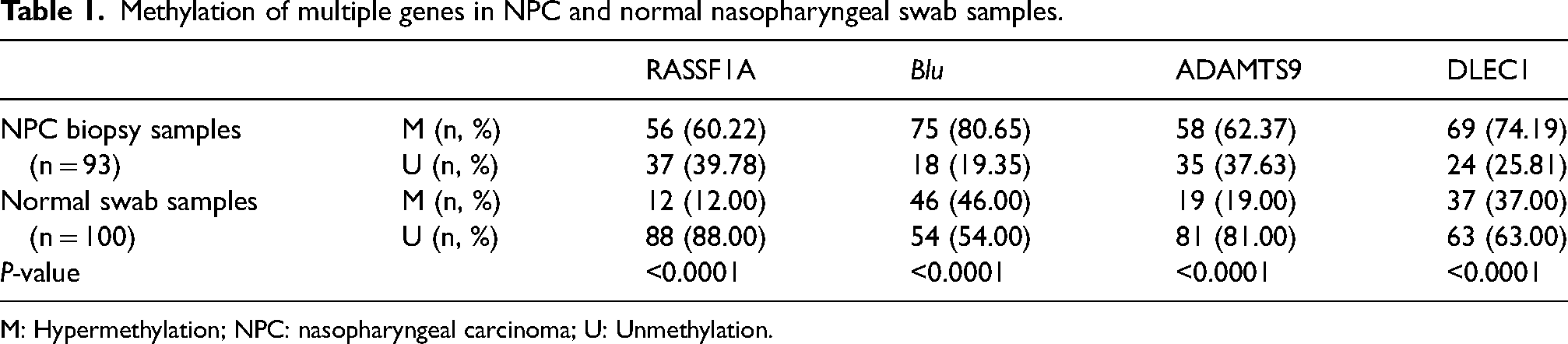
M: Hypermethylation; NPC: nasopharyngeal carcinoma; U: Unmethylation.
The ORs were 11.10 (95% CI 5.34, 23.08), 4.89 (95% CI 2.56, 9.35), 7.07 (95% CI 3.68, 13.57), and 4.90 (95% CI 2.64, 9.07) for reflecting the strength of correlation between the RASSF1A, Blu, ADAMTS9, DLEC1 gene methylation, and NPC, respectively.
The association between the methylation of RASSF1A, Blu, ADAMTS9, and DLEC1 with different clinical parameters are summarized in Table 2. The results showed that the highest frequencies of candidate gene promoters were observed in the advanced stages (III, IV). In detail, the percentages of RASSF1A, Blu, ADAMTS9, and DLEC1 methylation were 53.33% (32 of 60), 80.00% (48 of 60), 68.33% (41 of 60), and 68.33% (41 of 60) of advanced staged tumors, respectively. However, there was no significant correlation between RASSF1A, Blu, ADAMTS9, and DLEC1 methylation profiles with clinicopathological features, which was in accord with previous studies.
Correlation between methylation index with clinical parameters.
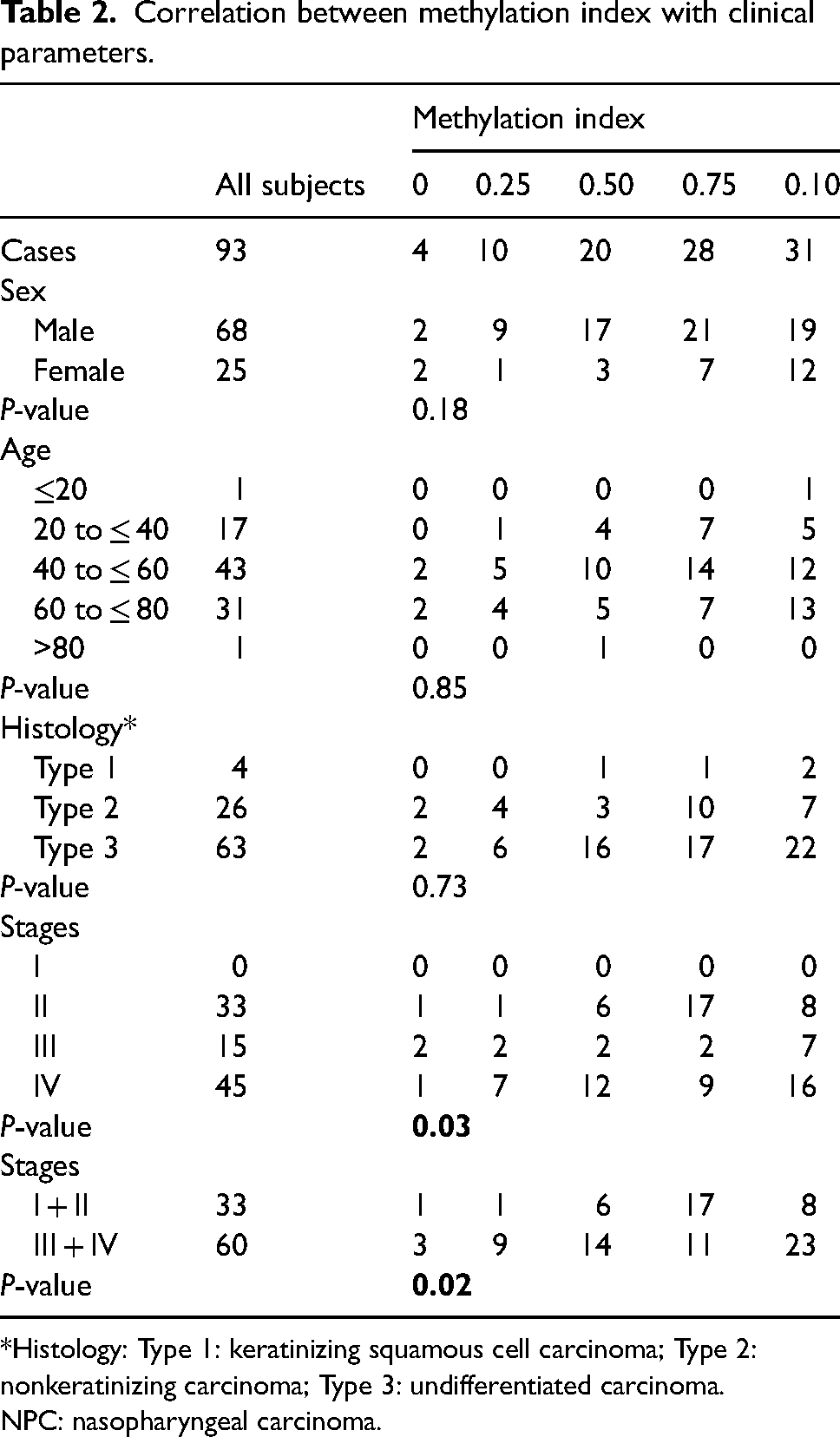
*Histology: Type 1: keratinizing squamous cell carcinoma; Type 2: nonkeratinizing carcinoma; Type 3: undifferentiated carcinoma.
NPC: nasopharyngeal carcinoma.
A panel of markers could be defined as an abnormality in NPC through detecting their aberrant methylation
The methylation index (MI) was calculated as the total of methylated genes per sample. The value of MI was computed in all 93 NPC samples, indicating that at least one of these genes had abnormal methylation (MI ≥ 0.25). Methylation of at least one gene (MI ≥ 0.25) was found in 95.70% of tumors (89 of 93). The percentage of tumors having methylation in one (MI = 0.25), two (MI = 0.50), three (MI = 0.75), and four genes (MI = 1.00) was 10.75% (10 of 93), 21.51% (20 of 93), 30.11% (28 of 93), and 33.33 (31 of 93), respectively. Only 4.30% (4 of 93) of tumors were unmethylated in all genes examined (Figure 1). Moreover, methylation of at least one gene (MI ≥ 0.25) was found in 73.00% (73 of 100) of non-cancerous samples; 27.00% (27 of 100) of non-cancerous samples were unmethylated in all genes tested. The strong correlation was investigated in the relationship between the profile methylation of at least one gene with NPC through the determination of P < 0.0001.
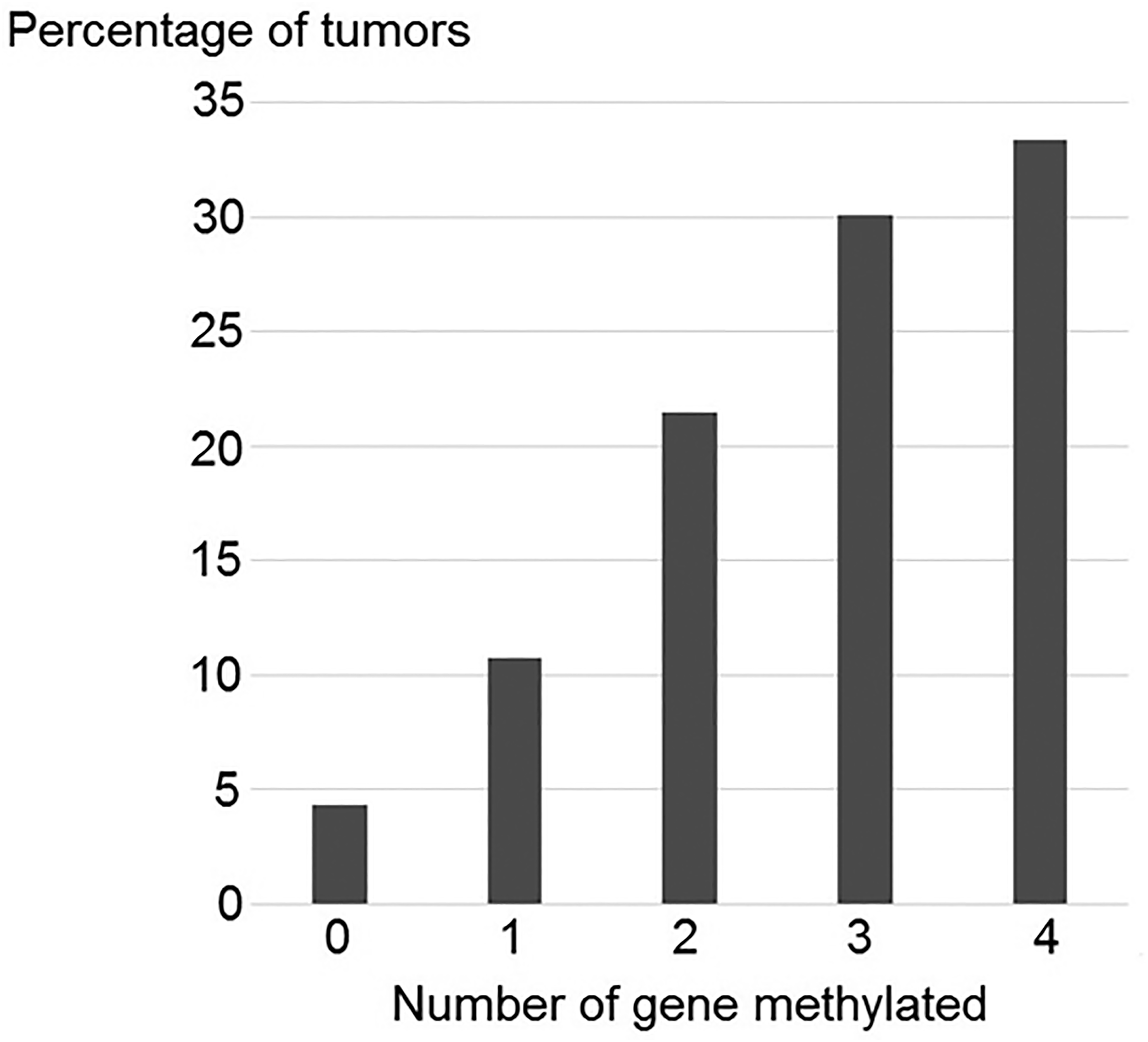
Percentage of the number of methylated candidate genes in 93 NPC samples.
We correlated the value of MI with clinical indexes from the NPC patients. We found no association between methylation index and clinicopathological characteristics, including sex, age, histological types (P > 0.05). This could explain that the NPC tumors exhibit the heterogeneity of clinical parameters of age and histological types. Moreover, the development and progression of NPC are influenced by a complex interplay of genetic, environmental, and lifestyle factors. Age and histological characteristics might be affected by these factors more strongly than by the methylation status of specific genes. In contrast, the significant clinical correlation of the methylation index among 93 tumors with NPC stages was recorded (Table 2). In our results, 46.67% (7 of 15 stage III tumors), 35.56% (16 of 45 stage IV tumors), and 38.33% (23 of 60 advanced stage tumors) were methylated in all genes. Moreover, 95.7% (57 of 60 advanced stage tumors) were methylated in at least one of four candidate genes (MI ≥ 0.25). This considered correlation indicated that the abnormal methylation of at least one of four genes was achieved in advanced stage NPC.
Discussion
Growing advances in the area of cytogenetic and molecular studies, the loss of expression of a variety of TSGs is now well understood to be caused by aberrant methylation of CpG islands in the promoter region, which causes the onset and progression of human malignancies.6,9,11,14 Many previous studies, as well as our systematic review, have identified that the silencing of chromosome 3p-located TSGs by the phenomenon of hypermethylation is the key crucial event in nasopharyngeal tumorigenesis.24–27 In the present study, we have sought to evaluate the methylation profile of several chromosome 3p-located genes in NPC patients, and to determine the correlation to clinicopathological characteristics. We investigated the epigenetic changes of four genes that were chosen, including RASSF1A, Blu, ADAMTS9, and DLEC1, as the target genes for methylation screening of Vietnamese NPC patients. Additionally, this is the first case-control study to ascertain whether there is a correlation between the methylation of candidates with NPC, as well as clinical characteristics, in Vietnamese NPC patients. For the four tested genes, methylation on RASSF1A and Blu have been previously published with a limited number of NPC samples; therefore, in the present study, the methylation frequencies of RASSF1A and Blu were again evaluated and combined with ADAMTS9 and DLEC1 on larger sample numbers to develop promising diagnostic biomarkers for NPC.
As TSGs, Dammann et al. 18 were the first to map RASSF1A on chromosome 3p21.3. The inactivation of its expression was recorded in lung tumors through the phenomenon of hypermethylation. 18 Recent growing evidence has reported that the RASSF1A is highly inactivated in NPC by DNA methylation.14,21,28–33 The biological function of RASSF1A has been reported. RASSF1A is a member of RAS effectors involved in the cell cycle, cell adhesion, apoptosis, and microtube stability. 31 It was suggested that RASSF1A is possibly a key tumor suppressor protein, which acts at the level of the G0/G1 phase cell cycle by inhibiting the cyclin D1 accumulation, which blocks the progression of cells. 34 Previous studies have strongly demonstrated that the methylation frequency of RASSF1A gene in NPC was observed in many type of samples with various ranges of frequencies, such as tumor biopsies (from 46.43% to 91.18%), nasopharyngeal brushing (from 52.22% to 59.18%), plasma/serum (from 4.88% to 17.50), etc. 24 In the present study, 56/93 (60.22%) of NPC tumor specimens presented with the hypermethylation, as did 12/100 (12.00%) of the non-cancerous swab samples. In comparison with their previous reports, we observed the similar frequency of RASSF1A methylation and higher frequency in the case group. Additionally, the strong correlation between the RASSF1A hypermethylation with NPC was observed based on the statistically significant difference in status of aberrant methylation (P < 0.0001).
Involvement of the disorder of 31p21.3, the hypermethylation of Blu has been previously reported.12,14,19,33,35 Blu, also known as ZMYND10 (zinc finger, MYND-type containing 10), encodes a 50-kd protein, which has a function in the cell cycle. 36 Additionally, Blu protein contains a conserved MYND zinc finger DNA binding domain located in its C-terminus, which is commonly found in the transcription repressor; thus, it is suggested that it plays a role of transcription regulation.19,37 The expression of Blu was commonly found to be down-regulated in NPC through the aberrant methylation. The present study utilized Nested-MSP to determine the methylation status of Blu in NPC tumors. Our results pointed the methylation of Blu in 75/93 (80.65%) of NPC tumors. This result was similar to previous findings stated that the hypermethylation was recorded in the samples of NPC. However, a higher frequency of aberrant methylation in the current study was observed in the comparison with 66.00%–74.00% in the Chinese NPC16,34 and 34.10% in the Tunisian NPC samples. 12 We also found that a significant difference in aberrant methylation was observed between the above two groups (P < 0.0001). In fact, the function of Blu protein is an inhibitor of colony formation of cancer cells, and could be activated by environmental stresses, such as heat shock and it is regulated by E2F. 33 Moreover, the functional suppression of a forming tumor was demonstrated by the in vivo study of Yoo and Jones. 15 They reported that Blu expressed an absence of doxycycline, which could lead to the suppression of tumor in nude mice. 19 Taken together, and concurring with previous studies, our study also suggested that Blu was a candidate TSG involved in NPC.
In the study by Dai et al., 8 they identified a number of TSGs, including ADAMTS9, which was an important one, by utilizing a method of microcell-mediated chromosome transfer approach. It encodes a protein that functions in cell migration and the inhibition of angiogenesis. 17 The levels of ADAMTS9 expression was identified to be critically correlated with tumor suppression in NPC. The hypermethylation of ADAMTS9 contributes to the loss of its expression in several tumorigenic NPC cell lines: HK1, JNE1, HONE1, CNE1, and CNE2. 17 In accordance with the concept of their report,8,17 in present study, we also suggested that ADAMTS9 was methylated in 58/93 (63.37%) of NPC tumors. Such a rate of ADAMTS9 promoter hypermethylation as well as a strong association between its methylation with NPC indicated the important role of this TSG in NPC.
The gene of DLEC1 was mapped to the region of 3p22—the alterations of which have been identified as highly prevalent in NPCs by the phenomenon of hypermethylation. 38 The hypermethylation of DLEC1 was observed in Tunisian, Chinese, and Hong Kong NPC patients with frequencies of 86.3%, 71.43%, and 60.42%, respectively.12,39,40 Aberrant methylation of DLEC1, in the present study, was also often found in our samples within the frequency of 69/93 (74.19%). However, the mechanism of DLEC1 related to nasopharyngeal tumorigenesis was unclear. Since there was a strong relationship between methylation of the DLEC1 promoter with NPC involvement, screening DNA hypermethylation of DLEC1 may be a promising and valuable biomarker for the diagnosis of NPC.
In an agreement with the previous findings, we found that the high frequency methylation of RASSF1A, Blu, ADAMTS9, and DLEC1 in 60.22%, 80.65%, 62.37%, and 74.19% of NPC tumor samples, respectively, compared to non-cancerous samples.12,16,29,41 The methylation of each candidate was strongly associated with the nasopharyngeal tumorigenesis based on the calculation of OR value. The increased odds of the methylation of RASSF1A, Blu, ADAMTS9, and DLEC1 were 11.10, 4.89, 7.07, and 4.90 times, respectively, among the samples from NPC patients than among the non-cancerous samples (P < 0.0001). The ORs > 1 indicated that the promoter methylation of individuals is strongly associated with higher odds of NPC risk. In the current study, no significant association between the profiles of each methylated candidate with clinical characteristics of NPC patients was found.
Tumor markers, which have the characteristics of significantly clinical value in screening as well as monitoring for NPC, are necessary. Rapid examination of a panel of three or four genes may be enough to get a high hitting rate and serve as hypermethylated promoters for human cancer screening and monitoring.8,12,24,42,43 A panel of methylated genes for NPC has been studied. For example, Kwong et al. 14 examined the hypermethylation of eight genes—p16, p15, p14, RASSF1A, RAR-β, DAPK, MGMT, and GSTP1—in primary tumors and normal epithelium. In their study, at least one of the seven genes was detected in 28/28 (100%) of tumor samples except for the gene of GSTP1. DNA hypermethylation of a panel of genes were also studied.6,25,44–46 In their studies, given the evidence of at least one gene of a gene panel suggested that setting a panel of hypermethylation-based markers may be useful for NPC screening and monitoring. The bias might exist due to the small sample size of tumors in their studies. In comparison with their results, we also investigated the characteristics of at least one methylated gene of a gene panel in our samples. All genes in our study are located at chromosome 3, which is different from previous studies, theirs were located at different chromosomes, including chromosome 3, and 9. Considering of the value of MI ≥ 0.25, 95.70% tumors were methylated within at least one of four genes examined, which was strongly associated with NPC (P < 0.0001). In our sample collection, most patients—accounting for 64.52% (60 of 93)—were diagnosed at the advanced stages (III and IV). As shown in Table 2, 57 of 60 advanced stage tumors (95.70%) were identified as at least one gene methylation, which is strongly associated with the advanced stage of NPC tumors. In comparison to individual markers, a panel of methylation biomarkers consisting of RASSF1A, Blu, ADAMTS9, and DLEC1 revealed a high prevalence of at least one methylated gene of NPC, giving effective differentiation between NPC and non-cancer, especially in the later stages of malignancies. This combination increases the likelihood of detecting tumor-specific methylation patterns. The cumulative effect of multiple biomarkers enhances sensitivity and specificity, making it more robust for distinguishing between cancerous and non-cancerous tissues. This is especially valuable in later stages when genetic and epigenetic alterations are more pronounced. Promoter hypermethylation of a current panel of genes may be useful as a promising biomarker for NPC diagnosis and monitoring.
The discoveries of methylated genes, as well as at least one methylated gene, have ignited hope for the application of various drugs, such as 5ʹ-aza-2ʹ deoxycytidine and 5-azacytidine, which precisely target the phenomena of DNA aberrant methylation. 8 The in vitro and in vivo studies ascertained that applying the 5ʹ-aza-2ʹdeoxycytidine significantly restored RASSF1A gene expression. Additionally, by treating with 5ʹ-aza-2ʹdeoxycytidine, demethylation of RASSF1A was also detected. 41 Therefore, our findings highlight the potential for using these 3p-lcoated TSGs in the development of in vitro assays for screening, monitoring, and treating routine nasopharyngeal carcinogenesis. However, the challenges and limitations of the current study include: (a) obtaining adjacent non-cancerous cells within the cancer samples as well as obtaining biopsies from non-cancerous tissues; and (b) only detecting the overall DNA methylation changes located on the promoter of target genes, ignoring global changes. In the current study, although it is very difficult to get tissue samples from healthy individuals, most case studies compare the epigenetic alterations found in tissue sample DNA with DNA from non-invasive cell sources.
Conclusion
This case-control study was carried out to identify the methylation profile of candidate tumor suppressor genes located at chromosome 3p, including RASSF1A, Blu, ADAMTS9, and DLEC1. The high frequencies of cases with each candidate gene promoter methylations were recorded in nasopharyngeal tumors. A significant association between the individual methylation with NPC was demonstrated. In addition, the high frequency of 64.52% was significantly diagnosed at the advanced stage tumors (III and IV). Therefore, these panels of current candidates may offer the promising methylated-biomarker for diagnosis and NPC monitoring. By providing a high prevalence of methylation in at least one of the selected genes, this panel enhances the ability to distinguish between NPC and non-cancerous tissues, particularly in later stages of the disease. For future studies, it is necessary to improve the invasion method in NPC diagnosis in NPC patients.
Supplemental Material
sj-docx-1-jbm-10.1177_03936155241268431 - Supplemental material for Hypermethylation of genes on chromosome 3p as a biomarker for nasopharyngeal carcinoma diagnosis: A Vietnamese case-control study
Supplemental material, sj-docx-1-jbm-10.1177_03936155241268431 for Hypermethylation of genes on chromosome 3p as a biomarker for nasopharyngeal carcinoma diagnosis: A Vietnamese case-control study by Thuan Duc Lao and Thuy Ai Huyen Le in The International Journal of Biological Markers
Footnotes
Acknowledgments
We would like to express our gratitude to Ministry of Education and Training, Ha Noi, Vietnam for their unwavering support during this research project, which was funded by grant number of B2023-MBS-01, and Ho Chi Minh City Open University, under the grant number of E2019.07.3.
Author contributions
Conceptualization: LD Thuan and LHA Thuy: Supervision, methodology, analysis. Writing—original draft: LD Thuan. Writing—original draft: LD Thuan. Writing—review and editing: LHA Thuy. Funding acquisition: LD Thuan LD and LHA Thuy. Both authors read and approved the final manuscript.
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This research was funded by Ministry of Education and Training, Ha Noi under the grant number of B2023-MBS-01, Ho Chi Minh City Open University, Ho Chi Minh City, Vietnam, under the grant number E2019.07.3; Đại học Mở Thành phố Hồ Chí Minh.
Statement of ethics
The ethics approval was obtained from the Medical Ethics Committee of the Cho Ray Hospital, Ho Chi Minh City, Vietnam. The decision number of the permission was 516/BVCR-HDDD.
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
