Abstract
Background
RASSF1A is a tumor suppressor gene. The methylation of RASSF1A has been reported to be associated with nasopharyngeal tumorigenesis. However, the heterogeneity was high among different studies. A meta-analysis was performed to evaluate the value of RASSF1A methylation for the diagnosis and early screening of nasopharyngeal carcinoma.
Methods
Relevant articles were identified by searching the MEDLINE database. Frequency and odds ratio (OR) were applied to estimate the effect of CDH-1 methylation based on random-/fixed-effect models. The meta-analysis was performed by using MedCalc® software. Subgroup analyses were performed by test method, ethnicity, and source of nasopharyngeal carcinoma samples to determine likely sources of heterogeneity.
Results
A total of 17 studies, including 1688 samples (1165 nasopharyngeal carcinoma samples, and 523 from non-cancerous samples) were used for the meta-analysis. The overall frequencies of RASSF1A methylation were 59.68% and 2.65% in case-group and control-group, respectively. By removing the poor relative studies, the heterogeneity was not observed among the studies included. The association between RASSF1A gene methylation and the risk of nasopharyngeal carcinoma was also confirmed by calculating the OR value of 30.32 (95%CI = 18.22–50.47) in the fixed-effect model (Q = 16.41, p = 0.36,I 2 = 8.62, 95% CI = 0.00–45.27). Additionally, the significant association was also found between the methylation of the RASSF1A gene and the subgroups.
Conclusions
This is the first meta-analysis that has provided scientific evidence that the methylation of RASSF1A is the potential diagnosis, prognosis, and early screening biomarker for nasopharyngeal carcinoma.
Introduction
Nasopharyngeal carcinoma (NPC) is a prevalent nasopharyngeal malignant tumor with remarkable differences in distribution, gravitating toward Southern Asia.1–7 According to the data from Global Cancer Observatory, updated to 2018, 129,079 new NPC cases were recorded worldwide, of which 72,987 NPC deaths occurred. Even though NPC treatment has been improved, the success rate of the treatment as well as the survival of patients reduces when it is diagnosed at an advanced stage. Early screening and diagnosis of the disease are very critical to improve the overall survival rate of patients. NPC is not detected and diagnosed early because there are non-specific symptoms in its early stage and because of the deep-seated location of the nasopharynx. 8 Therefore, effective biomarkers are truly needed. 9 Today, much effort has been made to identify early biomarkers by focusing on the etiological factors that lead to nasopharyngeal tumorigenesis. The progression of NPC is a multiple step associated with multiple factors, including the Epstein–Barr virus (EBV), environmental factors, plus alterations in genetics and epigenetics.10,11 Among them, DNA hypermethylation has been postulated as the best-studied epigenetic alteration in tumorigenesis. 12 The phenomenon of tumor suppressor gene (TSG) promoter causes transcriptional silencing, which leads to gene inactivation. This inhibits the functions of these genes, resulting in cancer development.13,14
The detection of EBV DNA in plasma samples was considered as the most useful method in screening for early asymptomatic NPC. 15 Until now, many TSGs have been recorded as being involved in the critical cancer-related cellular pathways. Thus, the patterns of TSG promoters’ methylation have been studied, and are reported to act as the potential epigenetic biomarkers for NPC.9,12 RASSF1A (RAS-association domain family 1 isoform A), located at 3p21.31, belongs to the family of RAS effectors, It is a well-described tumor suppressor gene, which encodes the scaffold protein member of the C-terminal RASSF family. 16 It functions as the signaling protein that plays important roles in cell signal transduction by regulating the complex signal networks, including ERK, Hippo, apoptotic, and p53 death receptor signal pathways. 17 The RASSF1A network modulates different biological functions, including apoptosis, cell cycle, DNA repair process, migration, and autography.15–20 The hypermethylation of two CpG islands located at the RASSF1A gene promoter mediated by DNA methyltransferase (DNMT), including DNMT1 and DNMT3A-3B, results in their loss of expression.17,18 Increasing evidence has demonstrated that this aberrant hypermethylation is one of the most common events in human diseases, including NPC, through the deregulation of its complex signal networks. It is considered to be necessary for the development of diseases. Increasing evidence has also indicated that the methylation of RASSF1A is strongly associated with NPC, but the value of RASSF1A methylation in NPC is uncertain. Challouf et al. reported that the methylation frequency of RASSF1A in NPC was 75.00%, but Tian et al. showed the methylation frequency was 17.5%.21,22 The high heterogeneity was observed in different studies due to the different sensitivities and intra-/inter-assay coefficients of variation of methods, the source of samples, and populations. Therefore, in the current study, we performed a meta-analysis to summarize previous studies and to identify the diagnosis, prognosis, and early screening value of RASSF1A for NPC.
Methods
Search strategy and inclusion/exclusion criteria
We conducted a comprehensive search strategy towards the guidelines of Preferred Reporting Items for Systematics Reviews and Meta-Analyses. 23 The separation or combination of the following keywords: “nasopharyngeal carcinoma”, “methylation”, “RASSF1A”, “diagnosis”, “prognosis”, “epigenetic”, “hypermethylation” were used to search for published articles in the MEDLINE database. Additional studies were also identified via the references listed in the articles.
Studies were considered eligible only when they met all of the following inclusion criteria if the articles: (a) were in English; (b) had a case-control study design; and (c) included data about the frequency of RASSF1A methylation plus the sample size in both the case and control groups. Exclusion criteria were as follows: (a) articles written in other languages; (b) abstracts, case reports, letters to the editor or unpublished articles; (c) studies related to other tumors and not specific for NPC; and (d) studies lacking vital information for analysis.
Data extraction
The eligibility of each study, and the relevant data from the eligible studies were independently retrieved by two authors. Disagreements were resolved through discussion with the third author or our research team. The relevant data were extracted from each study according to the data form, including first author's last name, year of publication, country where the study was performed, sample type, the experimental methods used to assess the methylation of RASSF1A, and the number of case and control subjects.
Statistical analysis, publication bias and sensitivity analysis
MedCalc® software, by MedCalc Software Ltd, was used to statistically analyze the data extracted (https://www.medcalc.org/). The frequency of RASSF1A methylation was calculated in both the case and control groups. The strength of association between RASSF1A methylation and NPC was evaluated by odds ratio (OR) with 95% confidence intervals (CI). In the current study, the heterogeneity among the included studies was estimated by the Cochran Q test and I2 statistics. 24 The cut-off point of p = 0.05 for the Q test and I2 were used to test the heterogeneity between the studies. The scale of I2 value was classified as follows: I2 < 25%: no heterogeneity; 25% ≤ I2 ≤ 50%: moderate heterogeneity; and I2 > 50%: strong heterogeneity. 24 The random-effects model was applied if there was heterogeneity among the studies (p < 0.05 for Q test, I2 > 50%). In the case of no between-study heterogeneity, a fixed-effects model was applied to compute the pooled ORs. In order to determine the presence of publication bias, the symmetry of the funnel plots in which ORs were plotted against their corresponding standard errors was assessed by the Begg's funnel plot and the Egger's test (p < 0.05); it was statistically significant.27,28 Sensitivity analysis was also performed by sequential omission of individual studies to evaluate the stability of the results.
Results
Study characteristics
After excluding the studies that did not meet the inclusion criteria, 17 studies published from 2001 to 2020 including 1688 samples (1165 NPC samples, and 523 non-cancerous samples) were used for the meta-analysis. The characteristics of the studies included in RASSF1A methylation and the risk of NPC are summarized in Table 1.
The characteristics of studies included in the meta-analysis of RASSF1A methylation and risk of NPC.
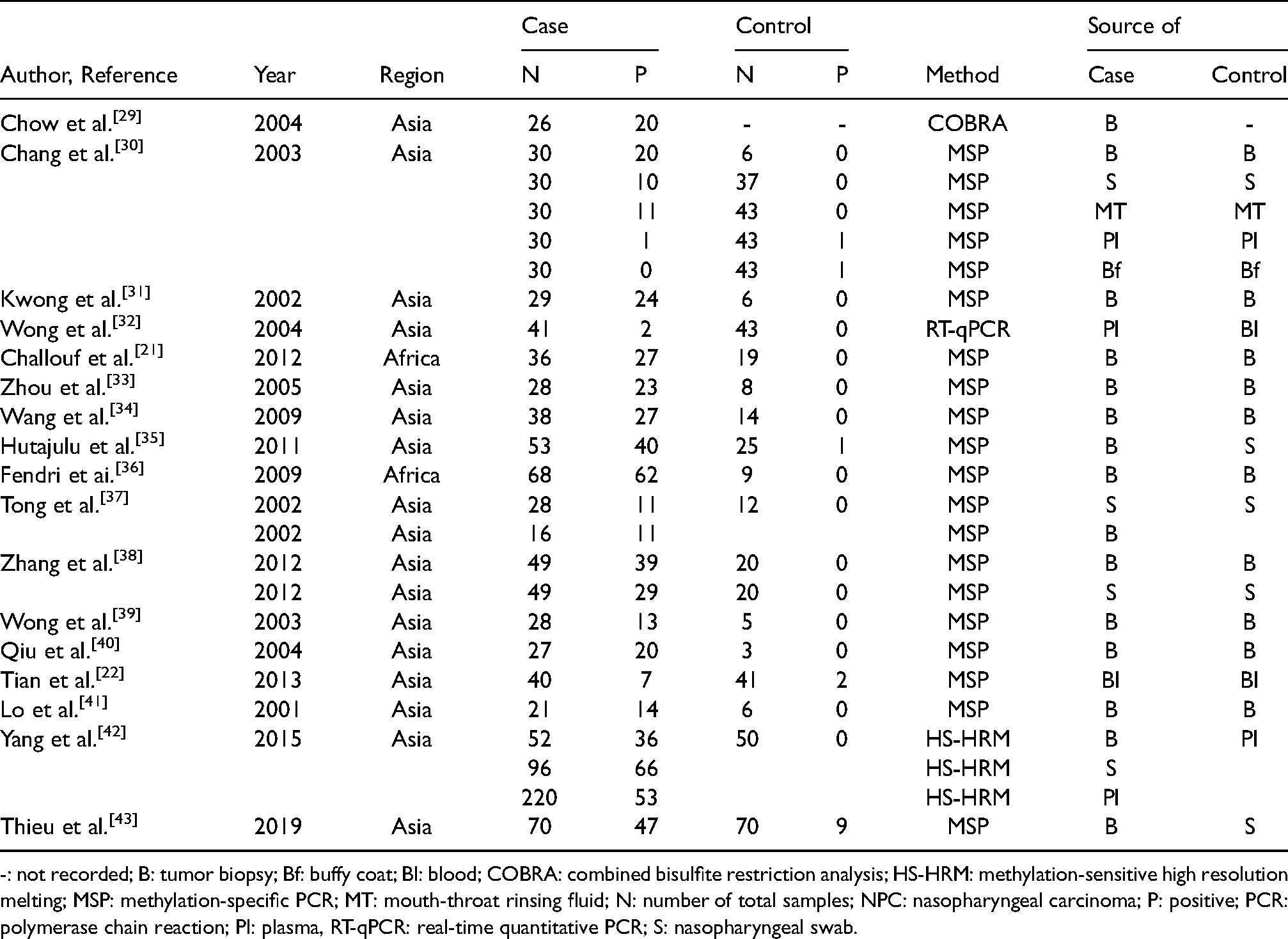
-: not recorded; B: tumor biopsy; Bf: buffy coat; Bl: blood; COBRA: combined bisulfite restriction analysis; HS-HRM: methylation-sensitive high resolution melting; MSP: methylation-specific PCR; MT: mouth-throat rinsing fluid; N: number of total samples; NPC: nasopharyngeal carcinoma; P: positive; PCR: polymerase chain reaction; Pl: plasma, RT-qPCR: real-time quantitative PCR; S: nasopharyngeal swab.
The number of NPC samples studies ranged from 21 to 368 (mean: 68.53 ± 83.75), and the number of control samples studies ranged from 3 to 172 (mean: 32.69 ± 42.11). Patient ethnicity in the 17 studies comprised 15 studies from Asian countries: China, Hong Kong, Singapore, Taiwan, Vietnam (accounting for 88.24%), and 2 studies from an African country: Tunisia (accounting for 11.76%). Regarding the test-method used for RASSF1A methylation, 14 studies used methylation-specific polymerase chain reaction (MSP) (counting for 82.36%), 1 study used combined bisulfite restriction analysis (COBRA) (accounting for 5.88%), 1 study used real-time quantitative PCR (RT-qPCR) (counting for 5.88%), and 1 study used methylation-sensitive high-resolution melting (MS-HRM) (accounting for 5.88%).
Meta-analysis: The frequency of RASSF1A promoter methylation, and the association between RASSF1A gene methylation and NPC
The differences of RASSF1A gene methylation between the case group and control group in the 17 studies (1165 samples from NPC samples, and 523 samples from non-cancerous samples) were accessed. There was heterogeneity across studies in the case group (Q = 266.10, p < 0.0001, I2 = 93.99%, 95% CI for I2 = 91.76–95.00), and moderate heterogeneity across studies in the control group (Q = 20.54, p = 0.15, I2 = 26.98%, 95% CI for I2 = 0.00–59.98). Thus, the random-effects model and fixed-effect model were applied to calculate the frequency of the RASSF1A gene in the case group and the control group, respectively (Figure 1). The frequency of RASSF1A gene methylation in the case group and the control group was 59.68% (ranging from 4.88% (95% CI = 0.59–16.53) to 91.18 (95% CI = 81.72–96.69)) and 2.65% (ranging from 0.00 (95% CI = 0.00–7.12) to 12.86 (95% CI = 6.05–23.01)), respectively. The meta-analysis result also indicated that the frequency of RASSF1A gene methylation in the case group was significantly higher than in the control group.
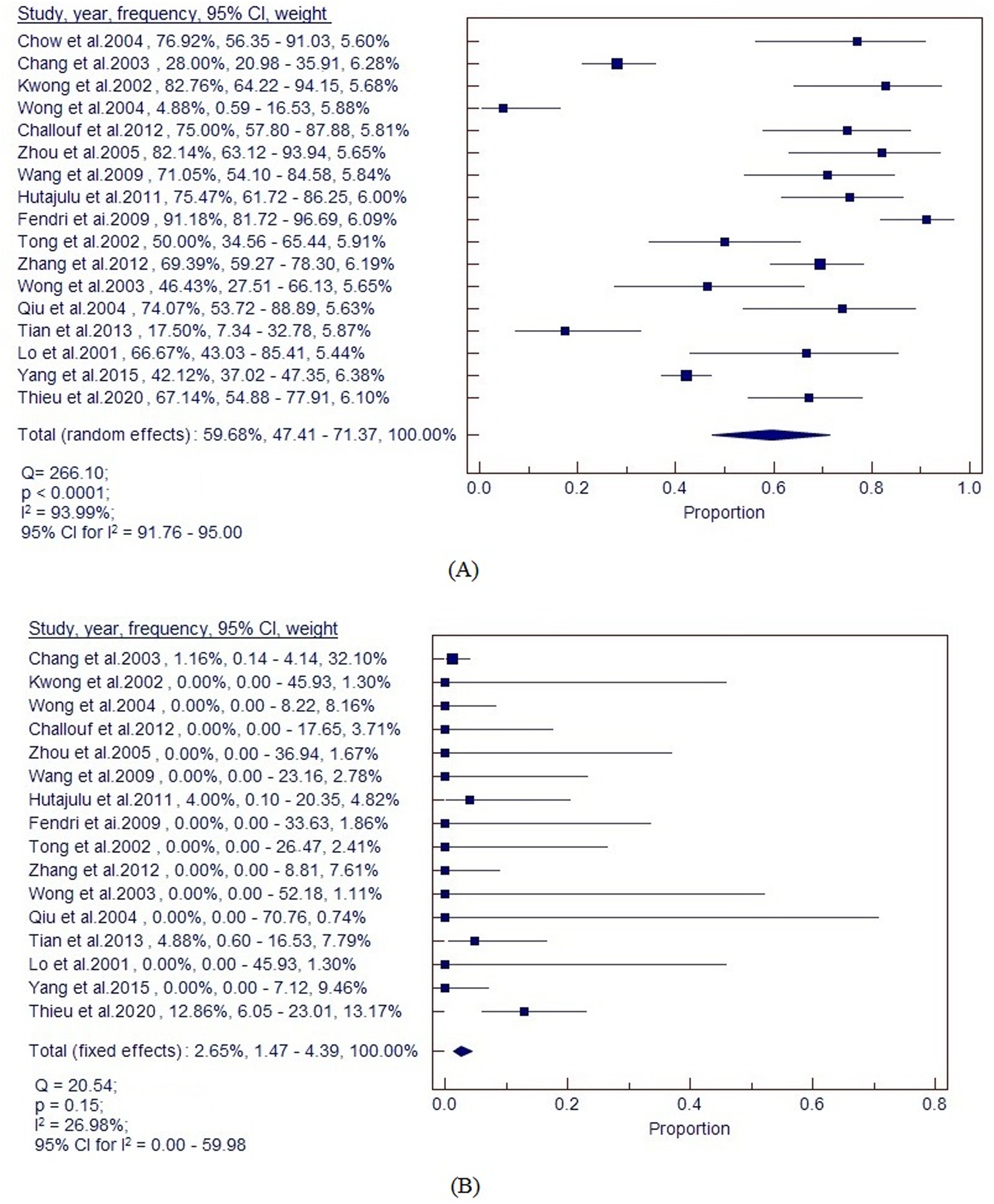
Forest plot of frequency of RASSF1A gene methylation detected in (a) NPC samples, and (b) control samples.
In the meta-analysis, there was no heterogeneity between the studies based on the Q-test (Q = 16.41, p = 0.36), and I2 (I2 = 8.62%) in the calculation of OR. Therefore, the methylation of the RASSF1A gene was significantly associated with an increased NPC risk with a pooled OR of 30.32 (95% CI = 18.22–50.47) based on the fixed-effect model (Q = 16.41, p = 0.36, I2 = 8.62%, 95% CI = 0.00–45.27) (Figure 2).
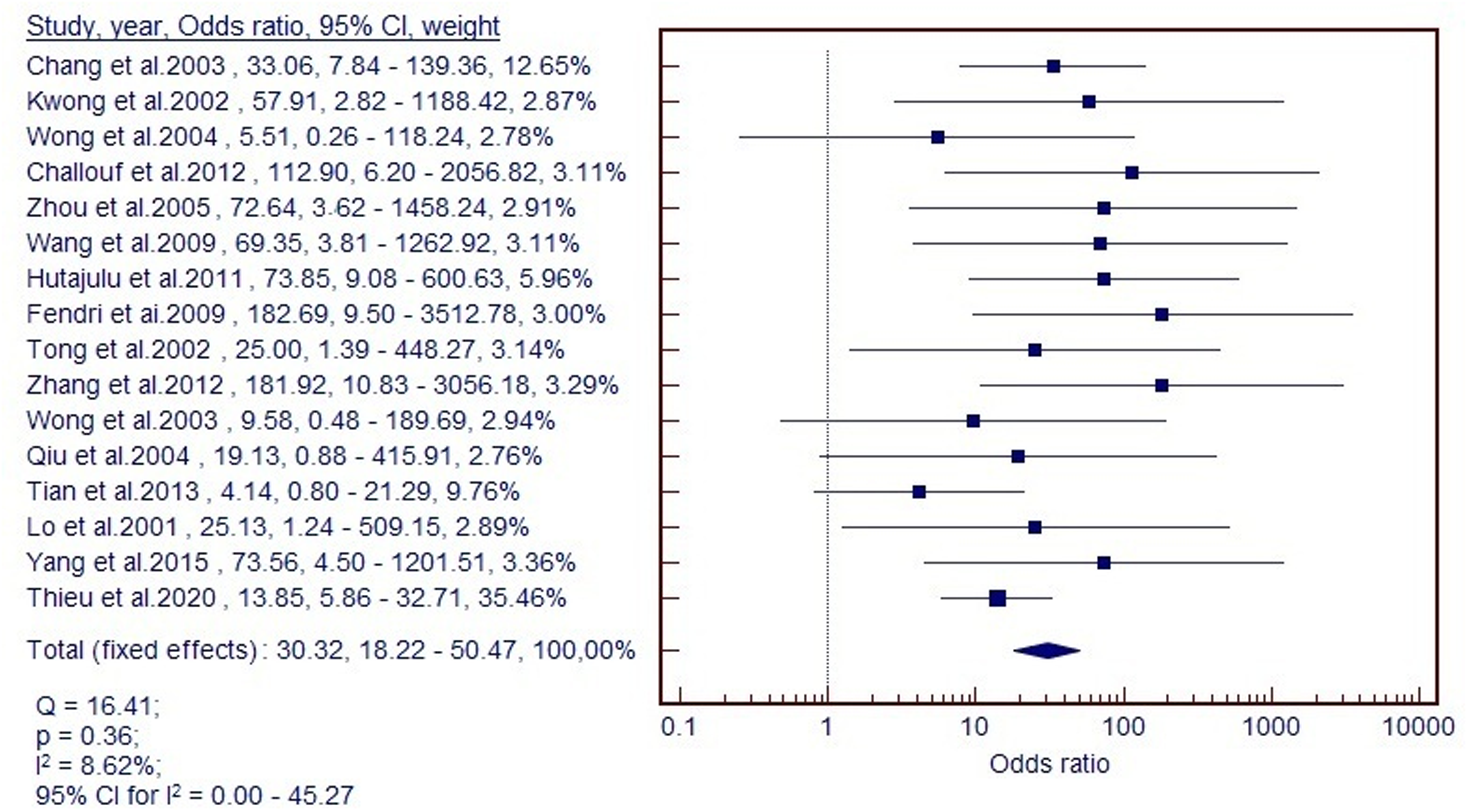
Forest plot of RASSF1A methylation and NPC risk using fix-effects model.
The subgroup analysis, based on the fixed-effect model, was applied to evaluate the RASSF1A promoter methylation including ethnicity, test-method, and source of sample (p > 0.05 for Q test, I2 < 50%) (Table 2). With respect to the subgroups categorized by ethnicity, there was a significant association between RASSF1A methylation and risk of NPC among the Asian countries and non-Asian countries in the fixed-effect model (Asian countries: OR = 22.43, 95% CI = 13.97–36.03; non-Asian countries: OR = 135.96, 95% CI = 16.03–1153.41). For the subgroup analysis by source of NPC samples, there was a strong association between RASSF1A methylation and NPC among the NPC biopsy tissue group, swab samples, and other samples, including nasopharyngeal swab, mouth and rinsing fluid, plasma, and blood in the fixed-effect model (biopsy tissue: OR = 38.42, 95% CI = 21.72–67.96; swab sample: OR = 35.21, 95% CI = 6.72–184.36; others: OR = 5.56, 95% CI = 2.20–14.15). The overall sensitivity and specificity of RASSF1A methylation evaluated on the swab samples were 46.73% (95% CI = 37.02%–56.62%), and 100.00% (95% CI = 94.79%–100.00%), respectively. They were calculated in predicting NPC. Additionally, a significant association between RASSF1A methylation and NPC risk among the test-method subgroup was found in the fixed-effects model (MSP: OR = 19.81, 95%CI = 12.10–32.43; other methods: OR = 73.79, 95%CI = 17.93–303.73).
Summary of subgroup analysis in meta-analysis of RASSF1A methylation and NPC risk.
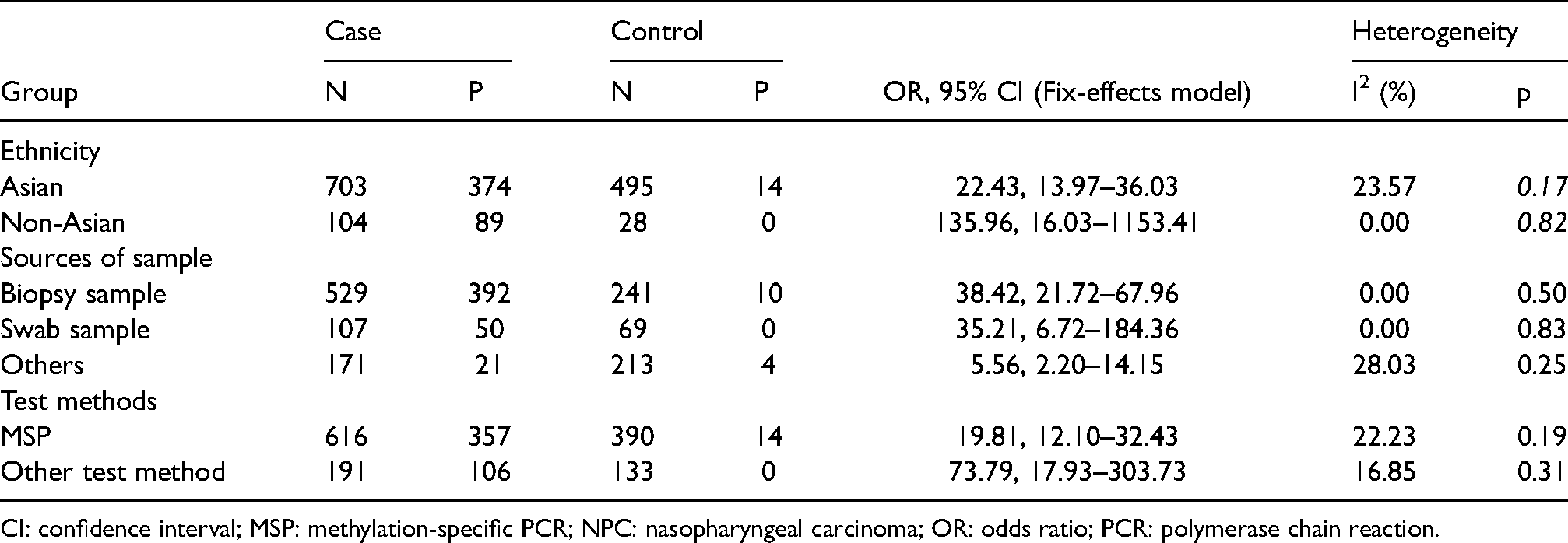
CI: confidence interval; MSP: methylation-specific PCR; NPC: nasopharyngeal carcinoma; OR: odds ratio; PCR: polymerase chain reaction.
Sensitivity analysis and publication bias
Sensitivity analysis was done to evaluate the stability and reliability of the conclusions according to the leave-one-out method by excluding one study. The pooled OR ranged from 26.78 (95% CI = 15.88–45.14) to 37.74 (95% CI = 20.67–70.98) under the fixed-effects model within the I2 = 0.00–14.91 (p > 0.05) (Supplemental Table 1). This confirmed the stability and reliability of the results. The publication bias of the included studies was assessed through the Begg's funnel plot and the Egger's test The shape of the funnel plot did not reveal potential asymmetry (Supplemental Figure 1; for Egger's test p > 0.05). Therefore, the results and conclusion of the present meta-analysis, which was used to evaluate the association between methylation of RASSF1A and NPC risk, are stable and reliable.
Discussion
It is important to find out the effective biomarker for early screening of NPC. Based on the etiological factors of NPC, the methylation of TSGs promoter has been recognized as the common mechanism that inactivates TSG, leading to the tumorigenesis of NPC.13,44 Xiong et al. identified the susceptibility locus at the chromosome 3p21 linked to the NPC tumorigenesis based on the genome-wide linkage analyses of high-risk Chinese NPC patients. 45 RASSF1A, belonging to this region, is reported as the tumor suppressor gene. The inactivation of RASSF1A via methylation has been recognized as highly associated with the progression of NPC.33,46 Notably, the methylation of TSGs promoter is responsible for early molecular modification; it transforms the human epithelial cells to malignant cells, and it often occurs earlier before there are changes in the morphology of cancer. 47 Hence, the analysis of TSG hypermethylation, including RASSF1A, may serve as the potential epigenetic biomarker for early screening of NPC. As previously reported, the frequency of RASSF1A methylation in NPC varied greatly. To determine whether RASSF1A methylation could serve as potential biomarkers for NPC, the meta-analysis and subgroup analysis by ethnicity, source of samples, and test-methods were performed.
To the best of our knowledge, this is the first meta-analysis of previous published studies that revealed that the methylation of RASSF1A increased NPC; it evaluates the relationship between RASSF1A promoter and NPC tumorigenesis. In this meta-analysis, 1165 samples from NPC samples, and 523 from non-cancerous samples were used. The overall frequency of RASSF1A methylation in the NPC and control group was 59.68% (ranging from 4.88% to 91.18%) and 2.65% (ranging from 0.00% to 12.86%), respectively. It could be observed that the individuals with RASSF1A gene methylation were significantly associated with NPC based on the calculation of pooled OR (OR = 30.32; 95% CI = 18.22–50.47) using the fixed-effect model (Figure 2). The resulting OR indicated that a 30.32-fold increase escalated the odds of a positive outcome in RASSF1A methylation, and this increase was statistically significant at a 5% level. Therefore, the methylation of RASSF1A was significantly associated with NPC tumorigenesis. It suggests that the methylation of RASSF1A might play a crucial role in the pathogenesis of NPC.
In the subgroup analysis, there was a significant association between RASSF1A gene methylation and NPC risk in all the subgroups, including ethnicity, source of samples, and test method. Regarding ethnicity, a significant association between methylation of RASSF1A and NPC was found among both the Asian regions and the non-Asian regions. Additionally, most of the studies were performed in Asian regions (15 of 17 studies). Only 2 studies were done in Africa. This once again confirmed that NPC is native to the Asian region. The subgroup analysis by source of cancer samples revealed a significant correlation between subgroups: biopsy samples, swab samples, and other samples. It indicates that the type of biopsy was more suitable to evaluate the methylation of the RASSF1A gene. However, the sources of other samples, including nasopharyngeal swab, mouth, and rinse, and the plasma type of non-/less-invasive sources of samples could cause alterations in NPC and the facility of collecting NPC samples. These are a potential biomarker for diagnosis and early screening of NPC. Especially, for the swab-samples used in previous studies, the overall specificity of RASSF1A methylation reached 100%. It indicates that the methylated RASSF1A-based test could correctly identify people without NPC. However, the overall sensitivity was 46.73%, indicating that 46.73% of NPC patients experienced high RASSF1A methylation levels. Thus, to increase the sensitivity, it should be combined with other biomarkers, or other patterns of tumor suppressor genes. Therefore, the focus in the future should be on finding out the non-/less-invasive biomarker for NPC. The MSP method was used in 14 studies (accounting for 82.36%). This means MSP is the “gold-standard method” used for the evaluation of methylation. MSP is a useful tool for the qualitative DNA methylation analysis; it is easily designed and used, has high sensitivity, and can detect small quantities of methylated DNA. 48 Moreover, MSP products are run on a gel, and the results are reported as methylated or unmethylated at the target DNA sequence. 49 In recent years, the method of HS-HRM has been developed to detect the methylation status of RASSF1A in clinical samples. It is a cost-effective method that can be used for the reliable quantification of DNA methylation by unique primers, which are enabled to detect 0.1–1% methylated alleles in an unmethylated background. 50 Overall, these results are in accordance with the documented RASSF1A gene methylation, which is common, and an early epigenetic event in the progression of NPC tumorigenesis. Therefore, its epigenetic event could be used as the potential epigenetic biomarker for the diagnosis and early screening of NPC.
The current meta-analysis exhibited some limitations due to the number of studies used. The data of non-English language studies as well as the evaluation of the correlation between the methylation of the RASSF1A gene and clinicopathological features, age, and the TNM stage, which are the differences in RASSF1A methylation between the case group and the control group, were not included.
Conclusion
This meta-analysis highlights the significant association between the methylation of the RASSF1A gene and the risk of NPC based on the evaluation of frequency of both the case group (accounting for 59.68%) and the control group (accounting for 2.65%), as well as an OR value of 30.32. Also, our findings underscore the correlation between RASSF1A gene methylation and all subgroups, including region, source of samples, and test method. The findings in the present meta-analysis emphasized that the methylation of the RASSF1A gene was recorded as the early epigenetic event in the progression of NPC tumorigenesis on the literature-based meta-analysis. Here, we reported that RASSF1A methylation is the potential diagnosis, prognosis, and screening biomarker for NPC.
Supplemental Material
sj-docx-1-jbm-10.1177_17246008211065472 - Supplemental material for Hypermethylation of the RASSF1A gene promoter as the tumor DNA marker for nasopharyngeal carcinoma
Supplemental material, sj-docx-1-jbm-10.1177_17246008211065472 for Hypermethylation of the RASSF1A gene promoter as the tumor DNA marker for nasopharyngeal carcinoma by Thuan Duc Lao, Hue Hong Thieu, Dung Huu Nguyen and Thuy Ai Huyen Le in The International Journal of Biological Markers
Footnotes
Acknowledgments
We wish to express our thanks to the research project sponsored by Ho Chi Minh City Open University, Ho Chi Minh City, Vietnam, under the grant number E2019.07.3.
Author contributions
TDL conceived, designed, performed, analyzed data, wrote the draft manuscript and acted as primary author for the project. TAHL designed, performed, analyzed data, oversaw the research and edited the article. All authors have read and approved the manuscript.
Availability of data and materials
All data generated or analyzed during this study are included in this published article.
Ethics approval and consent to participate
This is an observation and review study; ethics and consent are not applicable.
Consent for publication
Not applicable.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the Ho Chi Minh City Open University, Ho Chi Minh City, Vietnam (grant number E2019.07.3).
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
