Abstract
BACKGROUND:
Long non-coding RNAs have drawn increasing research interest in cancer biology. This study aims to investigate the function roles and the underlying mechanism of HnRNPU-AS1 in Hepatocellular carcinoma (HCC).
METHODS:
qRT-PCR was performed to detect the expression levels of HnRNPU-AS1, miR-556-3p, miR-580-3p in HCC tissues and cell lines. Western blot was used to determine protein levels of LC3-II, LC3-I, Beclin-1, P62, and SOCS6. Functional assays including CCK8 assay, colony formation assay, wound healing assay, Transwell assay were performed to evaluate the role of HnRNPU-AS1 in regulating the malignant phenotype of HCC cells. Dual luciferase reporter assay and RNA pull-down experiment were used to examined the RNA-RNA interaction.
RESULTS:
HnRNPU-AS1 expression was decreased in HCC tissues and cell lines, which was associated with poor prognosis in HCC patients. Overexpression of HnRNPU-AS1 could inhibit the proliferation, migration, invasion but promote autophagy in HCC cells. Two miRNAs (miR-556-3p and miR-580-3p) were identified as potential targets of HnRNPU-AS1 in lncBASE database, which were significantly upregulated in HCC tissues and cell lines. Cell experiments demonstrated the effects of HnRNPU-AS1 overexpression could be attenuated by miR-556-3p or miR-580-3p overexpression. We further revealed that SOX6 was the downstream target of HnRNPU-AS1/miR-556-3p or miR-580-3p axis. Xenograft mouse model validated the tumor-suppressor role of HnRNPU-AS1 overexpression in vivo.
CONCLUSIONS:
This study demonstrated the tumor suppressor function of HnRNPU-AS1 in HCC and identified the downstream molecules underlying its tumor suppressor function. Our results suggest that HnRNPU-AS1 suppresses HCC by targeting miR-556-3p and miR-580-3p/SOXS6 axis.
Introduction
Primary liver cancer is the sixth most common malignancy in the world [1]. Hepatocellular carcinoma (HCC) is the most prevalent pathological type, which accounts for 75%
Long non-coding RNAs (lncRNAs) are a class of non-coding RNAs with a length of more than 200 nucleotides [15]. Although lncRNAs do not encode proteins, they are implicated in a myriad of cellular processes including cell proliferation, cell metabolism, apoptosis, and cell differentiation [16]. Recent studies have revealed that a variety of lncRNAs are dysregulated and implicated in the progression of HCC. For instance, Zhao et al. demonstrated that LINC00174 could promote the cell proliferation of HCC cell lines by regulating miR-320/S100A10 axis [17]. Additionally, lncRNA MCM3AP-AS1 acts as an oncogenic factor to facilitate HCC growth by sponging miR-194-5p and maintaining the high expression of FOXA1 [18]. HnRNPU-AS1 is reported to be down-regulated in cervical cancer and it regulates cell proliferation and apoptosis [19]. Its potential functional role of HnRNPU-AS1 in HCC remains to be elucidated.
Suppressor of cytokine signaling 6 (SOCS6) has been reported as a tumor-suppressor in different cancers. For example, SOCS6 promotes the radio-sensitivity and decreases cancer cell stemness in esophageal squamous cell [20], and it can induce apoptosis and inhibit angiogenesis in prostate cancer [21]. Interestingly, SOCS6-CUL5 protein complex can regulate mTORC2 activity, and the knockout of SOCS6 gene in human pancreatic carcinoma cell lines PANC1 and BXPC3 impairs chaperon-mediated autophagy [22]. Since the functional role of SOCS6 in the progression of HCC is unclear, we attempted to investigate whether SOCS6 could modulate autophagy in HCC.
In our study, we first demonstrated the downregulation of HnRNPU-AS1 in HCC tissues and cell lines, which was associated with poor prognosis in HCC patients. HnRNPU-AS1 overexpression could suppress the malignant phenotype of HCC cells. Two miRNAs (miR-556-3p and miR-580-3p) were identified as downstream interactors of HnRNPU-AS1, which were upregulated in HCC tissues and cell lines. miR-556-3p and miR-580-3p overexpression attenuated the effects of HnRNPU-AS1 overexpression by targeting SOX6. We also found that the downregulation of HnRNPU-AS1 and SOCS6 in HCC impairs autophagy induction. Collectively, our results suggest that HnRNPU-AS1 suppresses HCC by targeting miR-556-3p/miR-580-3p/SOXS6 axis.
Materials and methods
Human samples
Tumor and para-tumor tissues were collected from 86 HCC patients at the Second Affiliated Hospital of Xi ’An Jiaotong University. All tissues were snap-frozen immediately in liquid nitrogen and stored at
Cell culture
Four Human HCC cell lines (Huh7, HepG2, HCCLM3 and Hep3B) and one normal liver cell line (L02) were obtained from the Chinese Academy of Sciences (Beijing, China). Cells were cultured in Dulbecco’s modified Eagle’s medium (DMEM, Gibco, USA) with 10% fetal bovine serum (FBS; Gibco, USA), 100 U/ml penicillin (Gibco, USA) and 100
qRT-PCR
TRIzol
GAPDH: F: GTCTCCTCTGACTTCAACAGCG; R: ACCACCCTGTTGCTGTAGCCAA. U6: F: CTCGCTTCGGCAGCACA; R: AACGCTTCACGAATTTGCGT. HNRNPU-AS1: F: GAGATTGCTGCCCGAAAGAAGC; R: TTCGCTGGAAGCCTGCAAACAG. hsa-miR-556-3p: F: GCGCAGATATTACCATTAGCTC; R: GGTCCAGTTTTTTTTTTTTTTTAAGATG. hsa-miR-580-3p: F: GCAGAAAAGTAATTGCGGTCT; R: GGTCCAGTTTTTTTTTTTTTTTACCA. Beclin-1: F: GGTGTCTCTCGCAGATTCATC; R: TCAGTCTTCGGCTGAGGTTCT. P62: F: GACTACGACTTGTGTAGCGTC; R: AGTGTCCGTGTTTCACCTTCC. SOCS6: F: ATCACGGAGCTATTGTCTGGA; R: CTGACTCTCATCCTCGGGGA.
Cell transfection
HnRNPU-AS1 overexpression vector, the empty pcDNA3.1, miR-566-3p mimic, miR-566-3p inhibitor, miR-580-3p mimic, miR-580-3p inhibitor, microRNA control (miR-NC), control for inhibitor (NC-inhibitor), control for siRNA (si-NC) and siRNA for SOCS6 (si-SOCS6) were purchased from GenePharma Co. Ltd. (Shanghai, China). Cell transfection was performed using Lipofectamine
CCK-8 cell proliferation assay
Cells were transfected with indicated molecules for 48 hours. Transfected ells were seeded in to a 96-well plate at a density of 1500 cell/well and cultured in a humidified cell culture incubator for 0, 24, 48, 72 and 96 hours, respectively. Subsequently, 10
Colony formation assay
Cells with indicated treatment were trypsinized and resuspended in culture medium. Cells were seeded into a 6-well plate (2000 cells/well) and the culture medium was changed every 3 days. After 14 days, cells were fixed with 4% paraformaldehyde at room temperature for 10 mins and stained with 0.5% crystal violet (Beyotime, Shanghai, China) for 20 mins. Subsequently, the number of colonies was counted and the morphology of the colonies was photographed under Leica AM6000 microscope (Leica, Wetzlar, Germany).
Wound healing assay
Cells were seeded in 6 well plates to reach about 80% confluence. A scratch wound was created using a sterile 200
Transwell invasion assay
The transwell invasion experiment was carried out using transwell upper chamber (Corning, NY, USA, # 3421, pore size 5.0
Western blot
Western blotting was conducted as in a previous study [23]. Total protein was extracted from tissues and cells using RIPA lysis buffer containing protease inhibitor cocktail (Thermo Fisher Scientific 78429, Waltham, MA, USA). Cells suspended in RIPA buffer were lysed on ice for 10 mins and lysed cells were centrifuged at 14000 rpm for 10 mins. The supernatant containing total protein lysate was quantified by a BCA Protein assay kit (Beyotime Biotechnology P0009; Shanghai, China). 15 ug of protein was used for in SDS_PAGE gel and was then transferred onto the PVDF membrane. After blocking with 5% skimmed milk for 1 hour, the membrane was then incubated with primary antibodies: LC3-II/LC3-I (1: 1000; ab48394, Abcam), Beclin-1 (1: 1500, ab62557), p62 (1: 1000, ab155686), SOCS6 (1: 1000, ab197335) and GAPDH (1: 2000; Cell Signaling Technologies #2118) for overnight at 4
Dual luciferase reporter assay
To demonstrate the functional interaction between microRNAs and the interaction partners, the sequence containing the wild type binding site (WT) at HnRNPU-AS1 3’-UTR and SOCS6 3’-UTR or the sequence with mutated binding site (MUT) were cloned into the PmirGLO vector expressing firefly luciferase respectively (Promega, E1330). The reporter plasmid and Renilla luciferase (hRlucneo) control plasmid were co-transfected into cells in the presence of miR-556-3p/miR-580-3p mimic or miR-NC in a 12-well plate (1
RNA pull-down
Cells lysates were collected by IP lysis buffer (Beyotime, P0013) and were incubated with biotinylated HNRNPU-AS1-probe oligo and Control oligo. 10% of the lysates was saved as the input. The mixture was further incubated with M-280 streptavidin magnetic beads (Sigma-Aldrich, 11205D) at 4
Xenograft tumorigenesis in nude mice
Animal experiments were approved by Animal Care and Experimental Committee of The Second Affiliated Hospital of Xi ’An Jiaotong University. 0.2 mL of HCC cells transfected with HnRNPU-AS1 overexpression vector or empty vector (1
Association between clinical characteristics and HNRNPU-AS1 expression in 86 patients with hepatocellular carcinoma
Association between clinical characteristics and HNRNPU-AS1 expression in 86 patients with hepatocellular carcinoma
Statistical analyses were performed with SPSS 20.0 software (IBM SPSS, Armonk, NY, USA). All the experiments were repeated three times. The association between HNRNPU-AS1 expression level and clinic pathological parameters was evaluated with Chi-square analysis. The statistical difference between two groups was compared using unpaired Student’s
Results
HnRNPU-AS1 is down-regulated in HCC tissues and HCC cells, and is linked with impaired autophagy
We first investigated the mRNA expressions of HnRNPU-AS1 in HCC tumor tissues and cell lines by qRT-PCR. Figure 1A and B showed that compared with adjacent normal tissues and normal liver cell line L02, HnRNPU-AS1 mRNA level was significantly decreased in tumor tissues and HCC cell lines (Huh7, HepG2, HCCLM3 and Hep3B). Next, 86 HCC patients were divided into HnRNPU-AS1 high expression (
HnRNPU-AS1 is down-regulated in HCC tissues and HCC cells. (A–B) The expression level of HnRNPU-AS1 in HCC tissues and cell lines (Huh7, HepG2, HCCLM3 and Hep3B) was compared with normal tissues or normal liver cell lines (L02). (C–D) Low HnRNPU-AS1 expression in HCC patients was associated with reduced overall survival and progression-free survival, as analyzed by Kaplan-Meier method. (E-F) The protein levels of autophagy-related proteins were compared between HCC tissues and adjacent normal tissues, and between HCC and normal cell lines by Western blot. 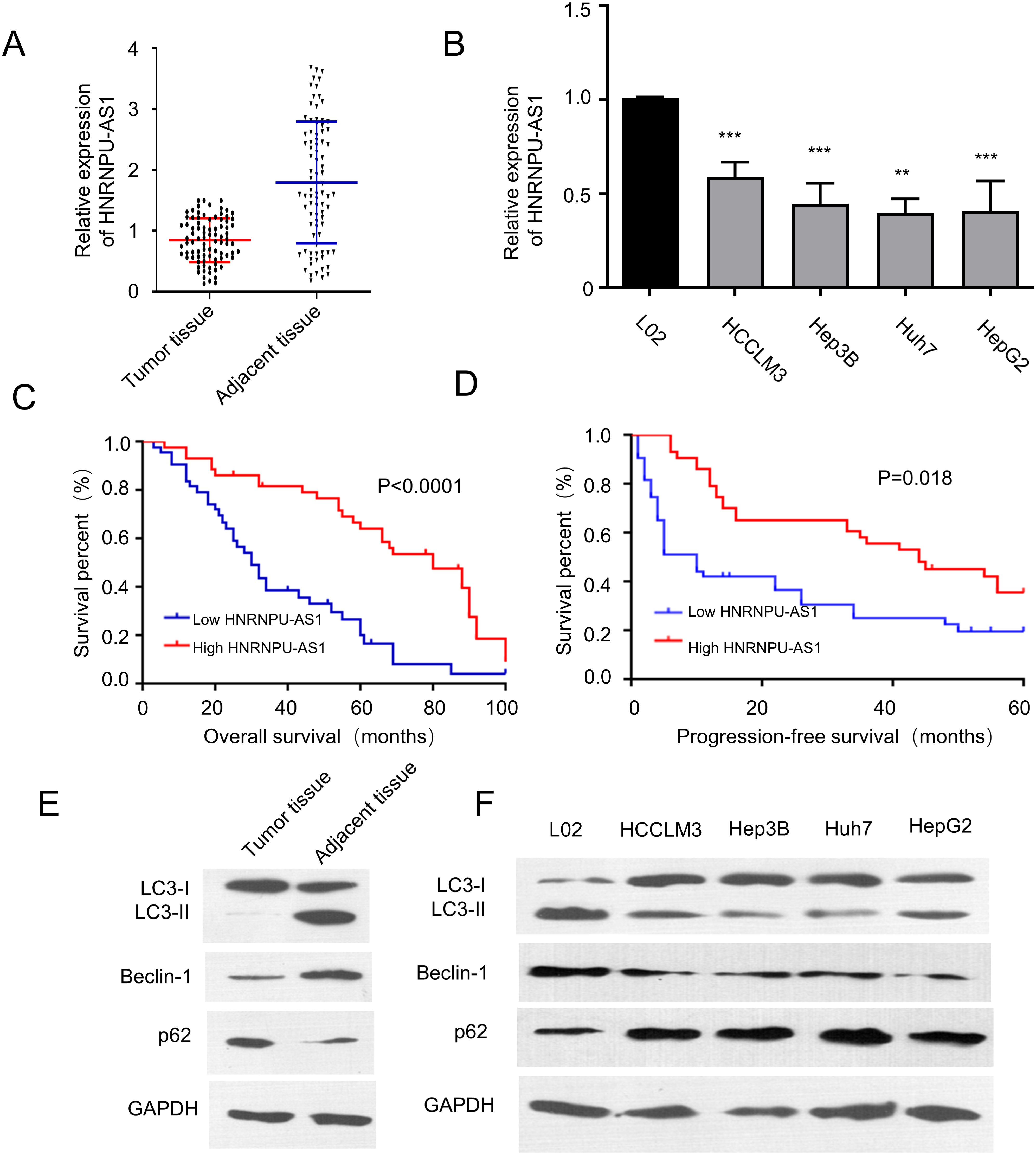
Overexpression of HnRNPU-AS1 inhibits cell proliferation, migration and invasion, and induces autophagy. (A) HnRNPU-AS1 is upregulated in Huh7 and HepG2 cells after the transfection with HnRNPU-AS1-pcDNA3.1 overexpression plasmid. Empty vector was used as control. (B) CCK-8 cell proliferation assay, (C) colony formation assay, (D) would-healing migration assay, and (E) Transwell invasion assay were performed in cells transfected with HnRNPU-AS1 overexpression vector in the presence or absence of autophagy inhibitor 3-Methyladenine (3-MA, 10 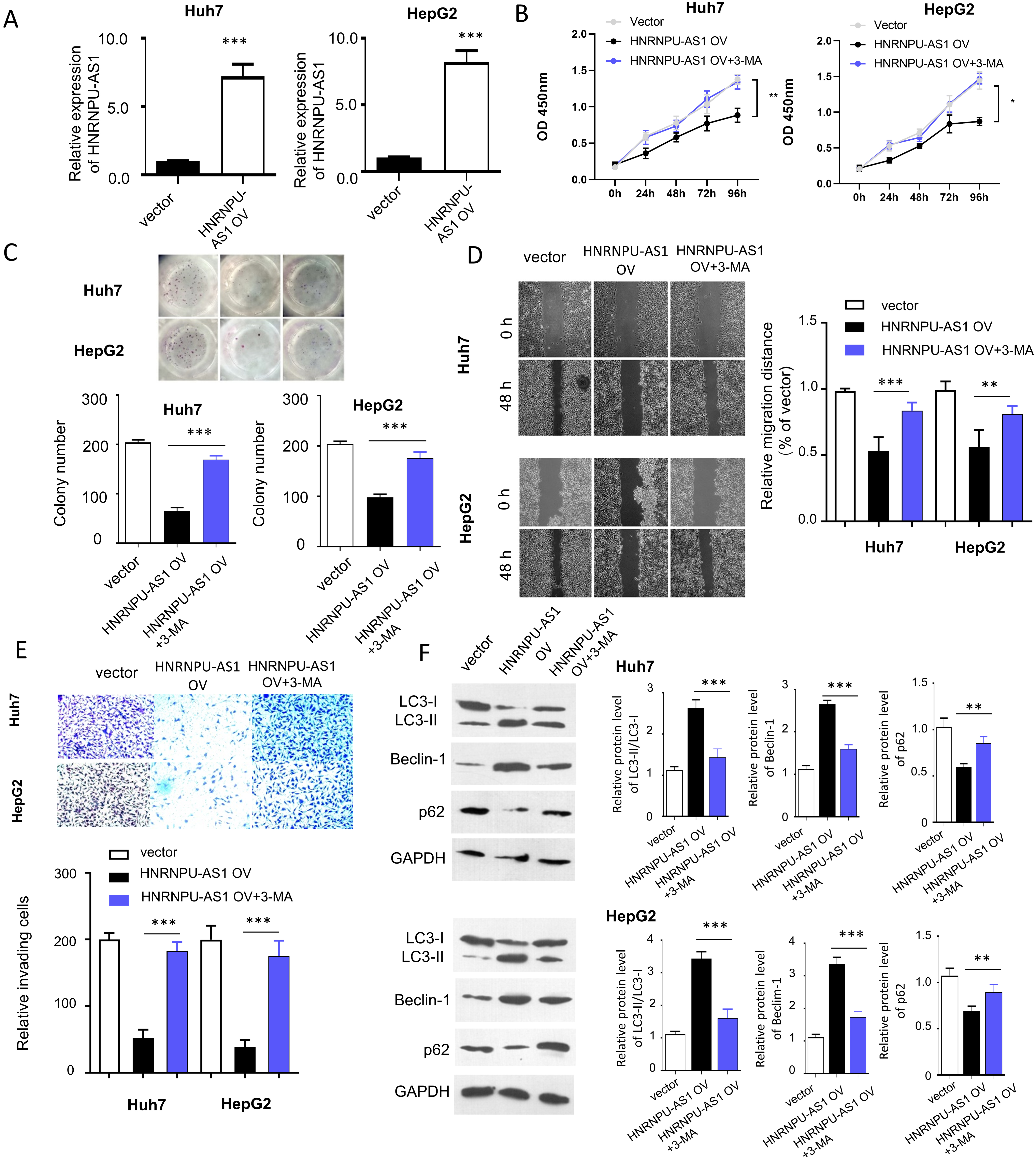
HnRNPU-AS1 targets both miR-556-3p and miR-580-3p. (A) The localization of HnRNPU-AS1 in nucleus and cytoplasm detected by qRT-PCR after nucleo-cytoplasmic fraction in Huh7 and HepG2 cells. (B) The binding sites between HnRNPU-AS1 and miR-556-3p/miR-580-3p predicted by lncBASE online database. (C–D) The expression level of miR-556-3p and miR-580-3p detected by qRT-PCR after transfecting with miR-556-3p inhibitor and miR-580-3p inhibitor. (E) Dual luciferase reporter assay using wild type (WT) reporter containing binding site of miR-556-3p/miR-580-3p or mutated (MUT) reporter in the presence of microRNA inhibitor or inhibitor control. (F) RNA pull-down assay using biotin-conjugated HnRNPU-AS1 probe or control probe. The precipitated micro RNA was normalized to the input. (G) The expression level of miR-556-3p and miR-580-3p in HCC tissues was compared with that of normal tissues. (H) The relationship between HnRNPU-AS1 level and miR-556-3p/miR-580-3p level in HCC tissues were analyzed by Spearman correlation coefficient analysis. (I) The expression level of miR-556-3p and miR-580-3p in HCC cell lines was compared with normal cell line. 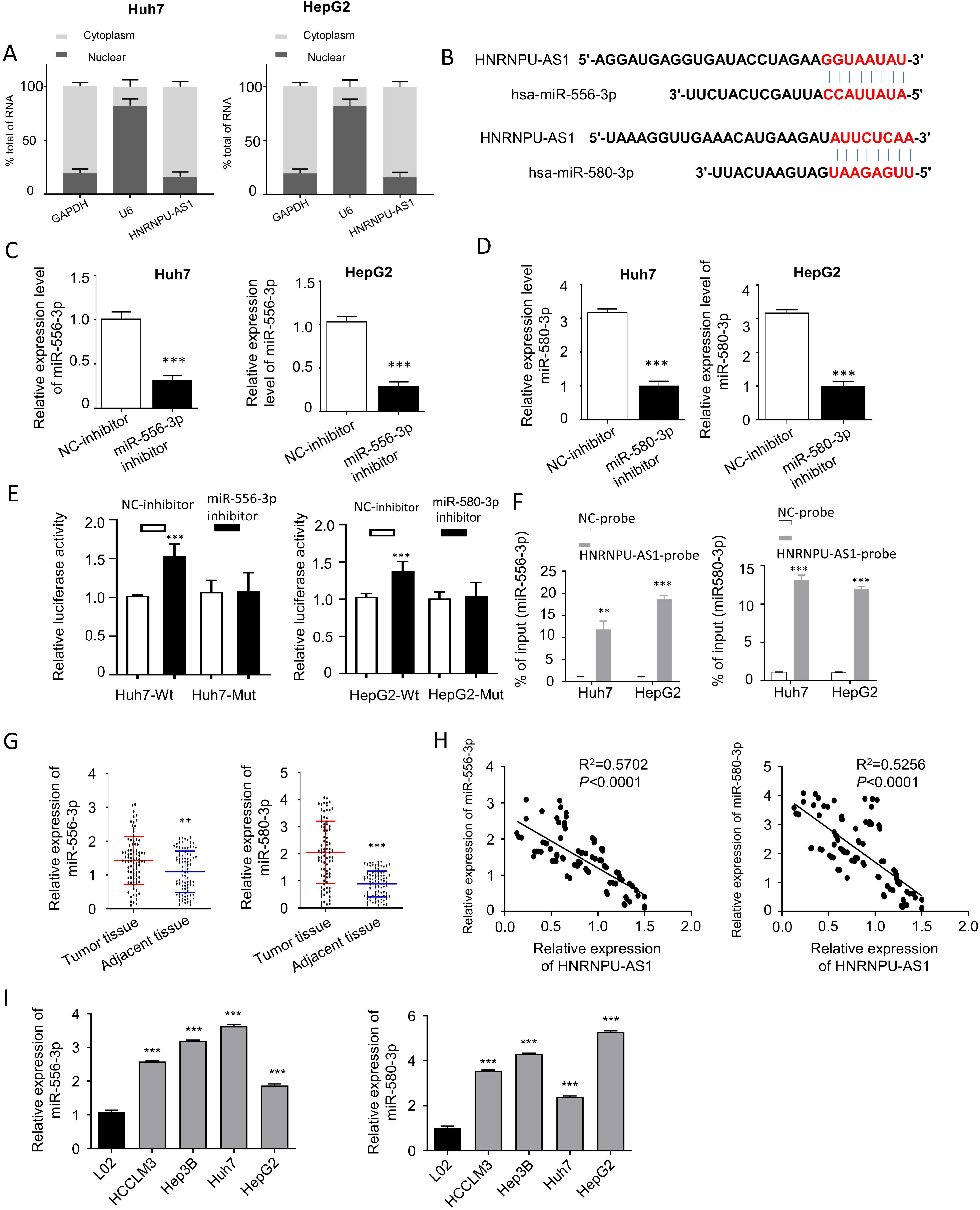
HnRNPU-AS1 regulates the malignant phenotype of HCC cells by targeting miR-556-3p and miR-580-3p. Cells were transfected with empty vector, HNRNPU-AS1 expression vector, HNRNPU-AS1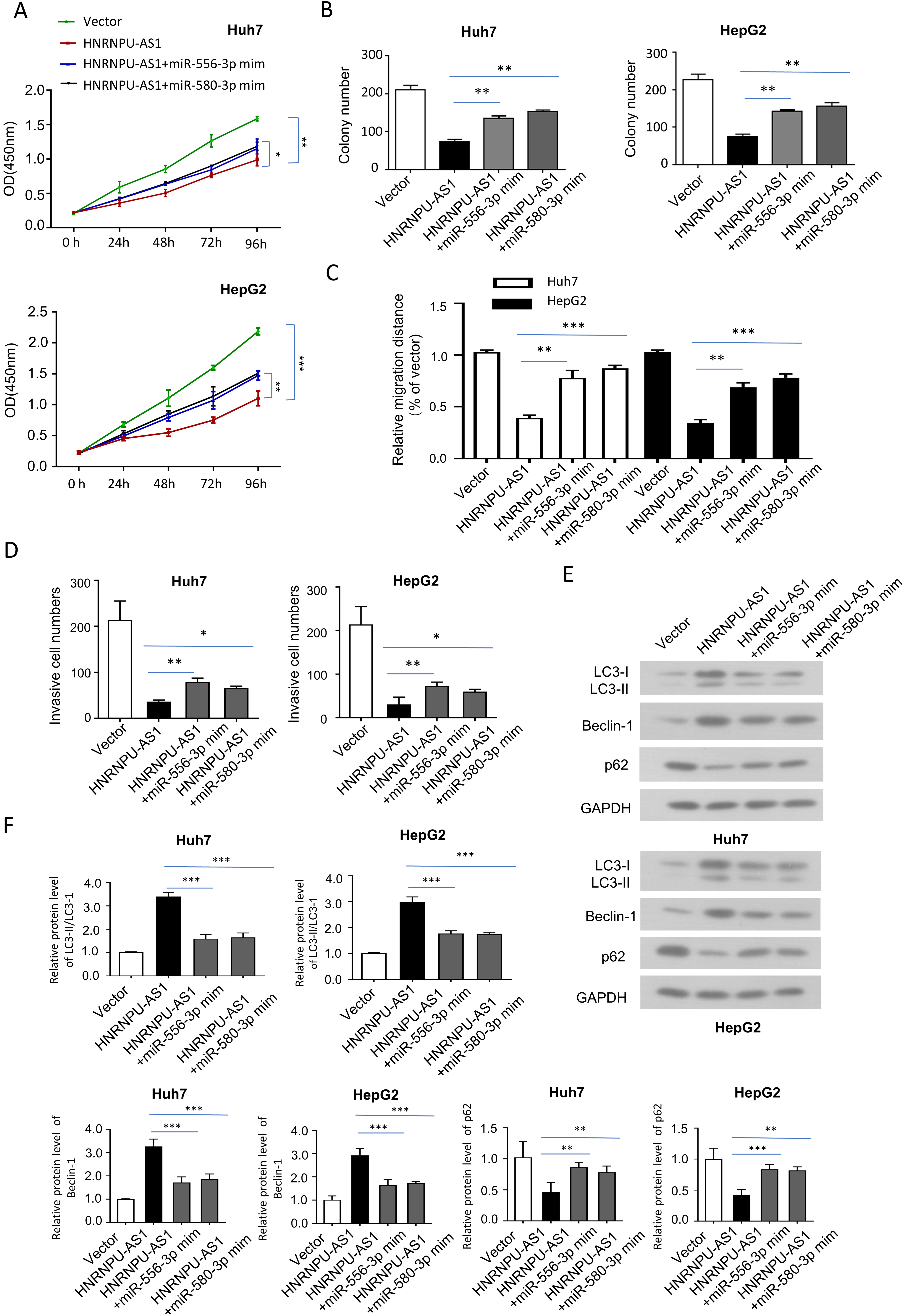
SOCS6 is a target gene of both miR-556-3p and miR-580-3p. (A) The bind sites between SOCS6 mRNA 3’UTR and miR-556-3p/miR-580-3p were predicted by StarBSAE online database, and their functional interaction was validated by dual-luciferase reporter assay in Huh7 cells. (B) The protein level of SOCS6 was analyzed by Western blot after transfecting with microRNA mimic or inhibitor. (C) The protein level of SOCS6 were determined by Western bot in different groups: Vector, HNRNPU-AS1, HNRNPU-AS1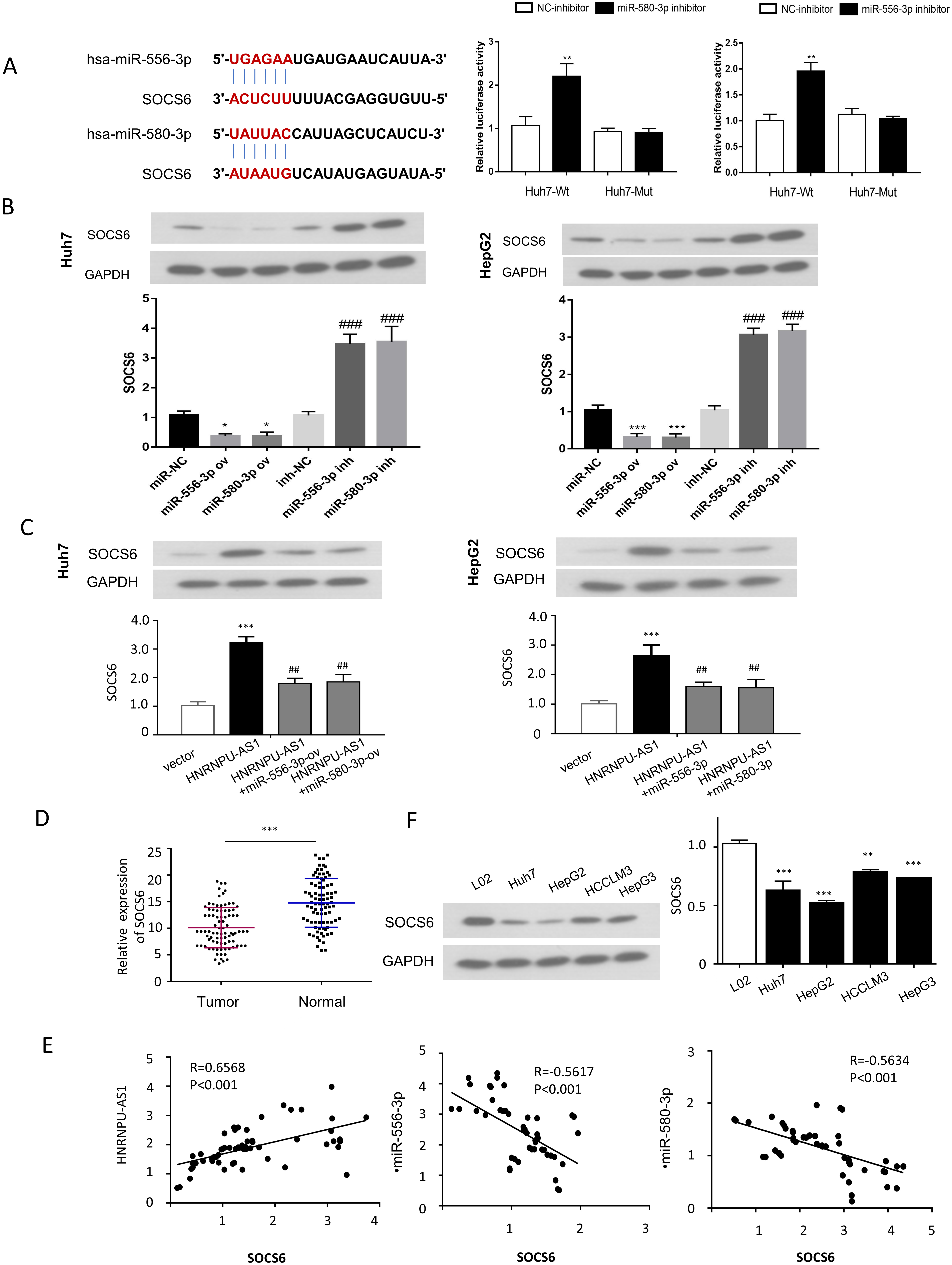
Knocking down SOCS6 partially reverses the effect of hnRNPU-AS1 overexpression in HCC cells. (A) SOCS6 protein level was measured by Western blot in cells transfected with control siRNA (si-NC) and siRNA targeting SOCS6 (si-SOXS6). (B–E) CCK-proliferation assay, colony formation assay, would-healing migration, and Trasnwell invasion assay were performed different groups: Vector, HNRNPU-AS1, HNRNPU-AS1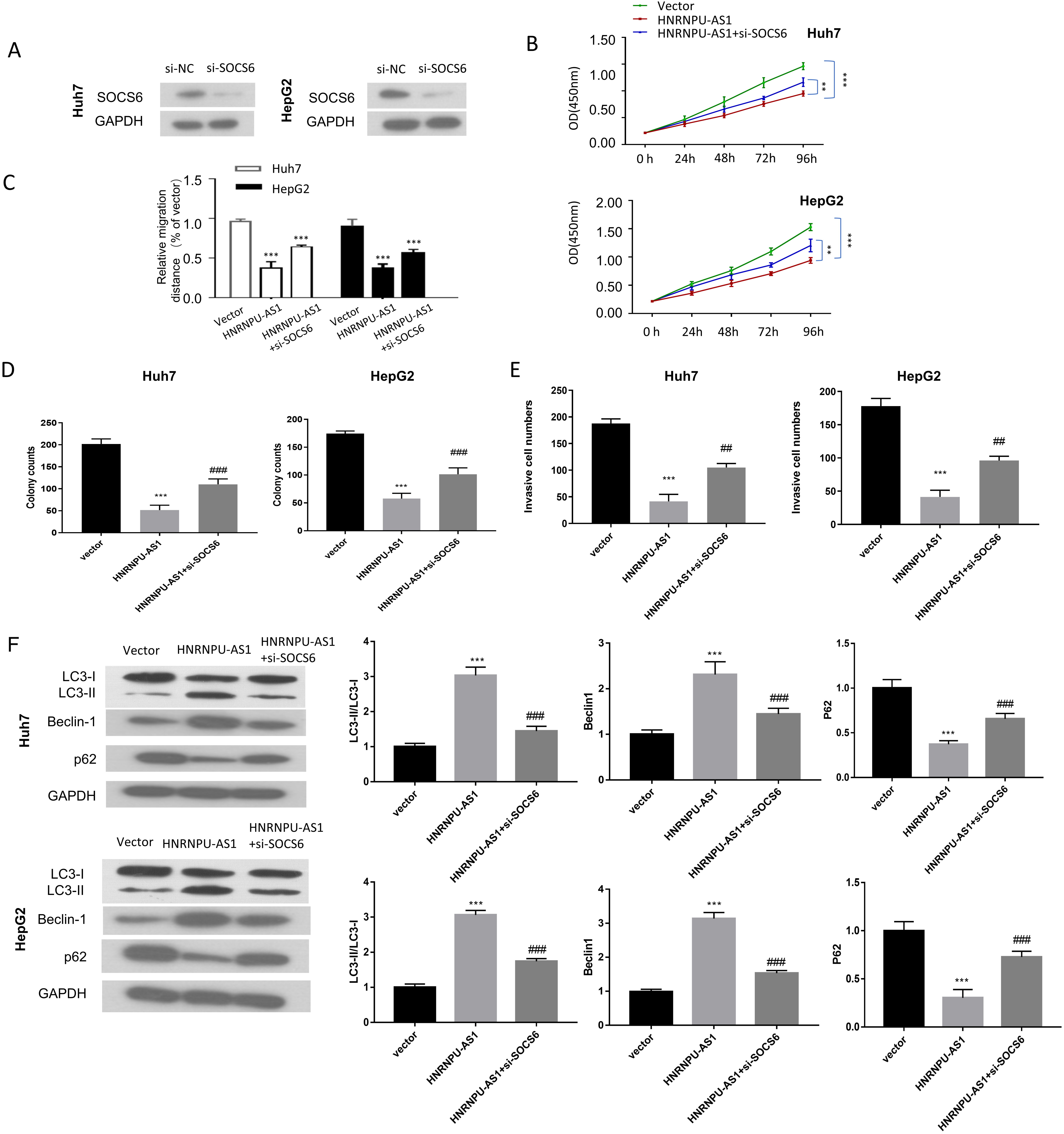
To confirm the expression level of HnRNPU-AS1 regulates phenotype and autophagy in HCC cells, we transfected HnRNPU-AS1 expression vector into two HCC cell lines (HUH7 and HepG2) with the lowest HnRNPU-AS1 expression. Transfection of HnRNPU-AS1 expression vector significantly increased the level of HnRNPU-AS1 in the two cell lines (Fig. 2A). We then performed CCK8 proliferation and colony formation assay and found HnRNPU-AS1 overexpression suppressed cell proliferation, which could be rescued by an autophagy inhibitor 3-Methyladenine (3-MA) (Fig. 2B and C). Similarly, wound healing assay and Transwell assay demonstrated that, cell migration and invasion ability was reduced after overexpressing HnRNPU-AS1 in both HUH7 and HepG2 cell lines, which was also rescued by 3-MA (Fig. 2D and E). The impact of HnRNPU-AS1 overexpression on autophagy was detect by Western blot, which revealed that HnRNPU-AS1 overexpression could promote autophagy by driving the conversion of LC3-I to LC3-II, increasing Beclin-1 protein level and suppressing P62 protein level (Fig. 2F). These effects were suppressed by autophagy inhibitor 3-MA. Collectively, these data suggest that high expression level of HnRNPU-AS1 regulates cell proliferation, migration and invasion, and induces autophagy.
HnRNPU-AS1 targets both miR-556-3p and miR-580-3p
We next attempted to fin potential downstream targets of HnRNPU-AS1. The subcellular localization of HnRNPU-AS1 in HUH7 and HepG2 cells was detected in cytoplasm and nucleus by qRT-PCR after nucleo-cytoplasmic fraction. Similar to GAPDH mRNA, HnRNPU-AS1 was mainly located in the cytoplasm of HUH7 and HepG2 cells (Fig. 3A). Using lncBASE online database, we identified potential binding sites between HnRNPU-AS1 and miR-556-3p or miR-580-3p (Fig. 3B). To validate these functional interactions, we performed luciferase reporter assay using synthetic inhibitors targeting these two miRNAs. The expression level of miR-556-3p and miR-580-3p were considerably suppressed by transfecting miR-556-3p inhibitor and miR-580-3p inhibitor (Fig. 3C and D). The luciferase activity assay showed that the knockdown of miR-556-3p or miR-580-3p could increase the luciferase activity of the WT reporter, while this effect was abolished when the biding sites at 3’UTR of HnRNPU-AS1 were mutated (Fig. 3E).
To investigate whether miR-556-3p and miR-580-3p interacts directly with HnRNPU-AS1, RNA pull-down assay was performed using biotinylated HnRNPU-AS1 probe or control probe. The results showed that compared with control, HnRNPU-AS1 probe significantly enriched both miR-556-3p or miR-580-3p (Fig. 3F). Moreover, the expression levels of miR-556-3p and miR-580-3p were significantly increased in HCC tumor tissues as compared with adjacent normal tissues (Fig. 3G). Spearman correlation coefficient analyses further revealed the negative correlations between hnRNPU-AS1 and miR-556-3p or miR-580-3p (Fig. 3H). Finally, we also showed that the expression levels of miR-556-3p and miR-580-3p were significantly higher in HCC cell lines (Huh7, HepG2, HCCLM3 and Hep3B) as compared to normal cell line L02 (Fig. 3I).
The above data indicate that HnRNPU-AS1 targets both miR-556-3p and miR-580-3p.
HnRNPU-AS1 regulates the malignant phenotype of HCC cells by targeting miR-556-3p and miR-580-3p
We next examined whether miR-556-3p and miR-580-3p mediates the functional role of HnRNPU-AS1. In HUH7 and HepG2 cells, we overexpressed HnRNPU-AS1 in the presence of miR-556-3p mimic or miR-580-3p mimic. The suppressive effects of HnRNPU-AS1 overexpression on cell proliferation, colony formation, migration and invasion were partially rescued by miR-556-3p mimic or miR-580-3p mimic (Fig. 4A–D). Besides, the effects on the conversion of LC3-I to LC3-II, Beclin-1 and P62 protein levels by HnRNPU-AS1 overexpression were also attenuated by miR-556-3p mimic and miR-580-3p mimic in HUH7 and HepG2 cells (Fig. 4E and F).
SOCS6 is a target gene of both miR-556-3p and miR-580-3p
To further search for the protein-coding mRNA target of miR-556-3p and miR-580-3p, we used StarBSAE online database and identified the putative binding sites between miR-556-3p or miR-580-3p and SOCS6 (Fig. 5A). Luciferase reporter assay was performed to determine the functional interaction between miR-556-3p or miR-580-3p and SOCS6 in HUH7 cells. The results showed that both miR-556-3p inhibitor and miR-580-3p inhibitor increased the luciferase activity of WT reporter, which was not observed in the MUT reporter containing mutated 3’-UTR sequence of SOCS6 mRNA (Fig. 5A). In addition, SOCS6 protein level was reduced upon the overexpression of miR-556-3p and miR-580-3p using microRNA mimics, while microRNA inhibitors enhanced their expression (Fig. 5B). We also examined whether HnRNPU-AS1/microRNA axis regulates SOCS6 protein level using the following experimental groups: Vector, HnRNPU-AS1 overexpression, HnRNPU-AS1
Knocking down SOCS6 partially reverses the effect of hnRNPU-AS1 overexpression
We next investigated whether SOCS6 mediates the functional role of hnRNPU-AS1 overexpression, by knocking down SOCS6 using siRNA. Compared with si-NC group, si-SOCS6 efficiently reduced SOCS6 protein level in HUH7 and HepG2 cells (Fig. 6A). When co-transfected with si-SOCS6, the suppressive effects of HnRNPU-AS1 overexpression on cell proliferation, colony formation, migration and invasion were partially reversed (Fig. 6B–E). Besides, the effects on the conversion of LC3-I to LC3-II, Beclin-1 and P62 protein levels by HnRNPU-AS1 overexpression were also attenuated by knocking down SOCS6 in HUH7 and HepG2 cells (Fig. 6F). The above results suggest that SOCS6 is a downstream mediator of hnRNPU-AS1 in HCC cells.
Overexpression of HnRNPU-AS1 inhibits tumor growth in vivo. (A) The tumor volume and (B) The tumor weight was measured in nude mice injected with Huh7 cells overexpressing HnRNPU-AS1 or control vector. (C) The expression levels of HnRNPU-AS1, miR-566-3p, and miR-580-3p were measured in tumor tissues of vector and HnRNPU-AS1 groups. (D) The protein level of SOCS6, LC3-II, LC3-I, Beclin-1, p62 were determined by Western blot in tumor tissues of vector and HNRNPU-AS1 groups. 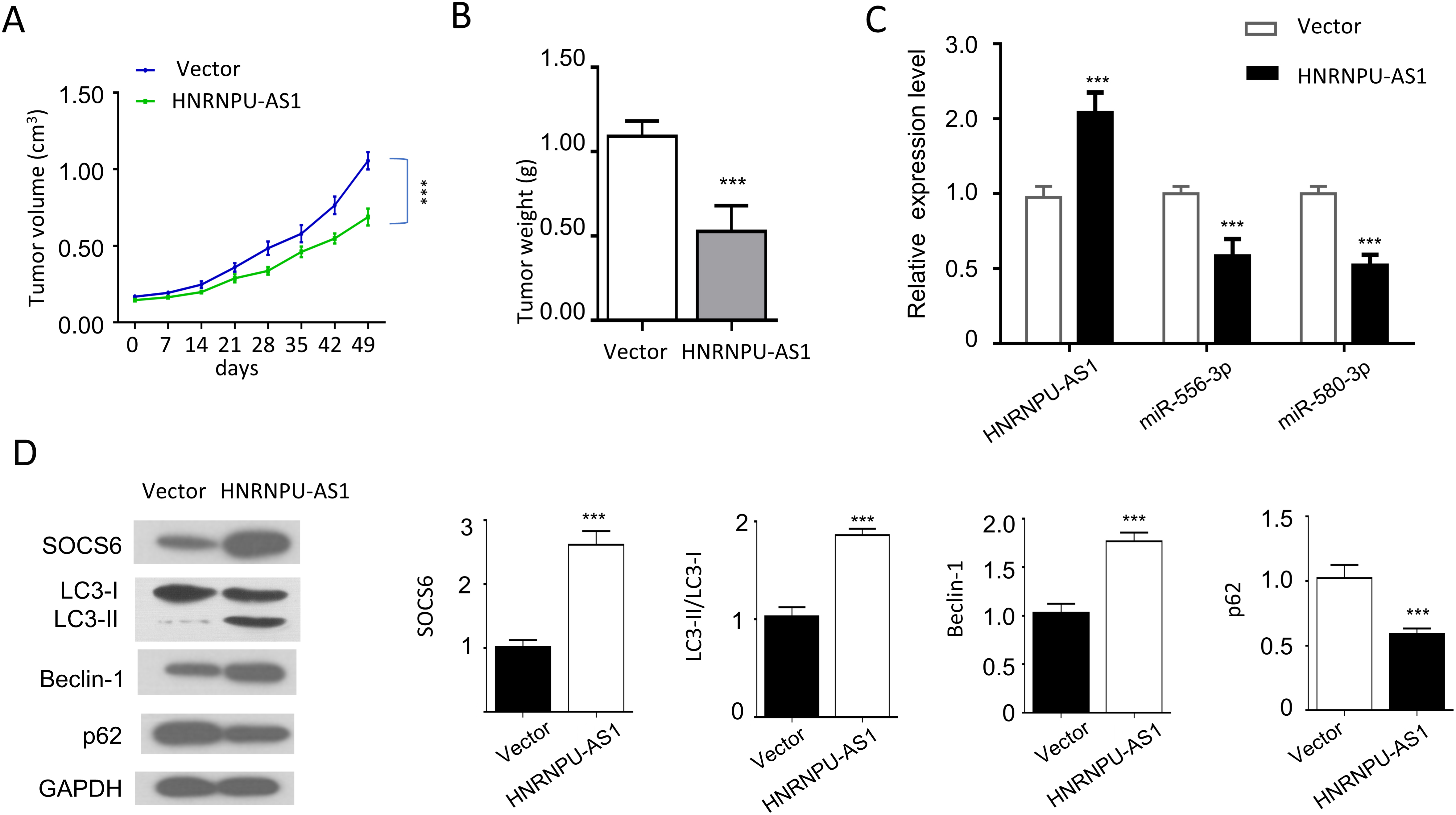
Schematics of molecular interplay among HnRNPUAS1, miR-566-3p, miR-580-3p and SOCS6 in HCC. In HCC, HnRNPU-AS1 is downregulated and its downregulation is correlated with the upregulation of miR-556-3p and miR-580-3p. miR-556-3p and miR-580-3p function as downstream mediators of HnRNPU-AS1 to suppress the expression of tumor suppressor SOCS6. The reduced SOCS6 expression contributes to the promotion of proliferation, migration and autophagy in HCC cells.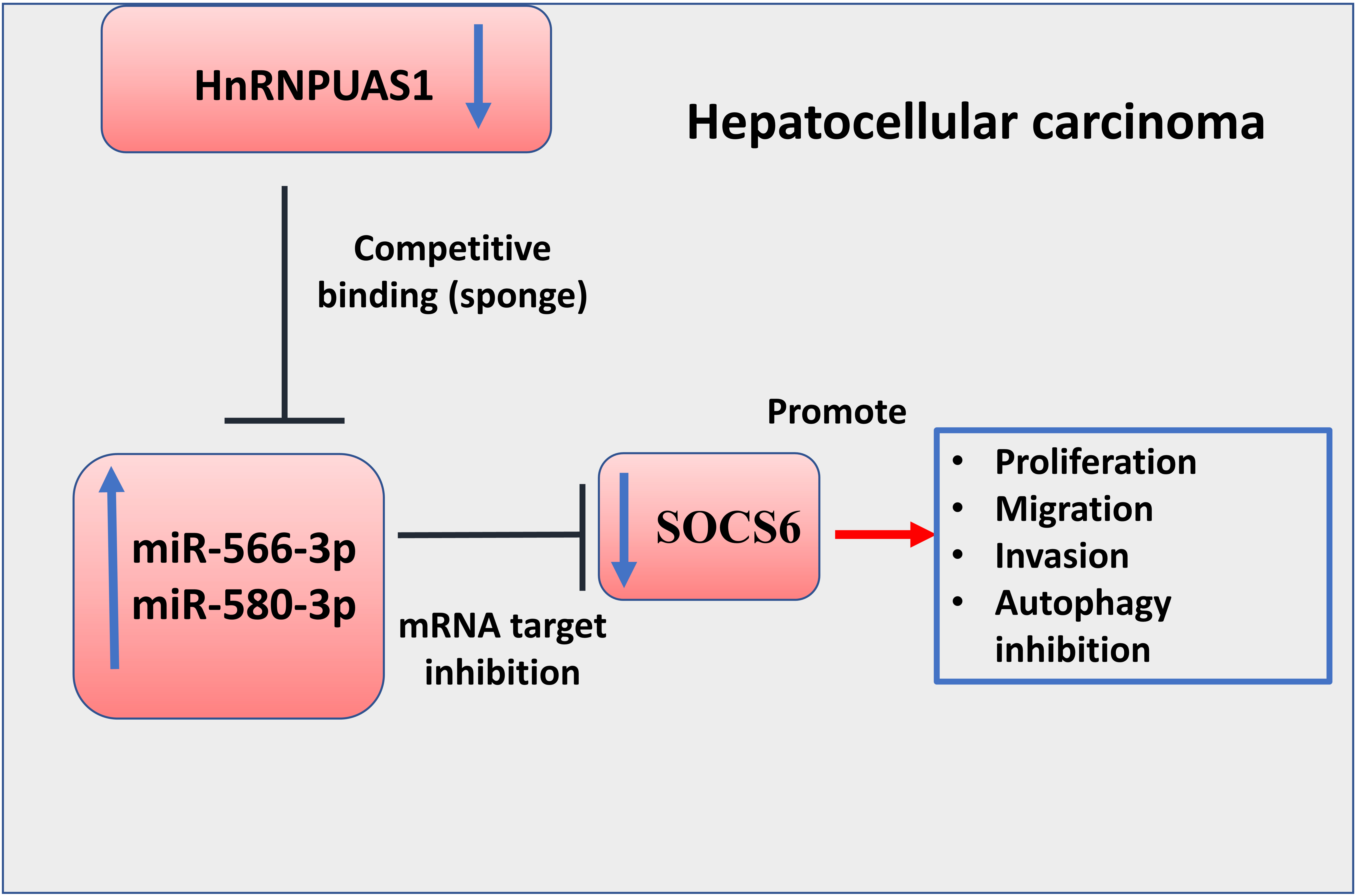
To explore the tumor-suppressor effects of HnRNPU-AS1 in vivo, HUH7 cells with empty vector or overexpressing HnRNPU-AS1 were subcutaneously injected into nude mice. Tumor volume and tumor weight in HnRNPU-AS1 overexpression groups were significantly reduced as compared to the control group (Fig. 7A and B). Besides, in tumor tissue of HnRNPU-AS1 overexpression group, the mRNA level of HnRNPU-AS1 was higher but the expression level of miR-556-3p and miR-580-3p was reduced (Fig. 7C). Furthermore, Western blot showed that HnRNPU-AS1 overexpression increased the protein level of SOCS6, LC3-II/LC3-I, and Beclin-1, while P62 level was downregulated in HnRNPU-AS1 overexpression group (Fig. 7D). Together, these data demonstrated that HnRNPU-AS1 is a tumor suppressor in HCC tumor model.
Discussion
In the present study, the results show that the lncRNA HnRNPU-AS1 and SOCS6 are downregulated, while by the expression of miR-566-3p and miR-580-3p is upregulated in HCC, which is implicated in promoting cancer progression via suppressing autophagy. HnRNPU-AS1 seems to function as a tumor suppressor, and its overexpression impairs the malignant phenotype pf HCC cells (cell proliferation, invasion and migration) and induces autophagy. The miR-556-3p and miR-580-3p/SOXS6 axis mediates the tumor-suppressor effect of HnRNPU-AS1.
The tumor-suppressor function of HnRNPU-AS1 is evidenced by the observation that low HnRNPU-AS1 expression is associated with a poor prognosis in HCC patients. In previous studies, HnRNPU-AS1 was reported to be downregulated in acute myocardial infarction (AMI) [24] and in rheumatoid arthritis [25]. However, the expression pattern and functional role of HnRNPU-AS1 in cancer progression remains enigmatic. We provided evidence that HnRNPU-AS1 is downregulated in both HCC tissues and cell line, and its overexpression inhibits HCC progression in vitro and in vivo. Future studies are required to further explore the mechanism underlying the downregulation of HnRNPU-AS1 in HCC.
We further demonstrated the role of HnRNPU-AS1 as a sponge for miR-566-3p and miR-580-3p, and miR-566-3p and miR-580-3p. Many reported lncRNAs are located in the cytoplasm and act as ceRNAs to competitively bind to the miRNA target [26, 27, 28, 29]. HnRNPU-AS1 is also primarily localized in the cytoplasm, and its functional interaction with miR-556-3p and miR-580-3p were validated by dual luciferase reporter and DNA-pull down assay. These two miRNAs are up-regulated and their expression level show negative correlation with HnRNPU-AS1 in HCC tissues, indicating that their upregulation is implicated in HCC progression. Indeed, a previous study has shown that miR-556-3p could promote the progression and development of HCC, and downregulating miR-556-3p in HCC cells could suppress cell proliferation and induce apoptosis [30]. In non-small cell lung cancer, Pang et al. found that the miR-580-3p was upregulated to support cell proliferation, which could be sponged by circular RNA hsa_circ_0072309 [31]. We further demonstrated that the effect of overexpressing HnRNPU-AS1 in HCC cells could be attenuated by miR-556-3p or miR-580-3p overexpression. These results support the oncogenic roles of miR-556-3p and miR-580-3p, which could be sponged by tumor-suppressor HnRNPU-AS1. In line with this notion, miR-556-3p and miR-580-3p could act as target and be sponged by lncRNAs or cirRNAs in a various of tumors, such as melanoma [32] and lung cancer [33].
We also identified SOCS6 as a target negatively regulated by miR-556-3p and miR-580-3p, and SOCS6 expression level is negatively correlated with that of miR-556-3p or miR-580-3p. Consistently, SOCS6 was reported as suppressor in HCC development, which is negatively regulated regulated by miR-21 and miR-183 [34]. Moreover, SOCS6 low expression is associated with a poor prognosis in HCC patients [35]. Together, these findings highlighted SOCS6 as a tumor suppressor in HCC. Based on the positive correlation between HnRNPU-AS1 and SOCS6 and their downregulation in HCC, our study further validated that SOCS6 acts as a downstream factor mediating the tumor-suppressor effect of HnRNPU-AS1 by silencing SOCS6 in cells overexpressing HnRNPU-AS1.
Interestingly, HnRNPU-AS1/SOCS6 axis seems to regulate HCC progression by mediating autophagy. A previous study showed that the knockout of SOCS6 gene in PANC1 and BXPC3 cells impairs chaperon-mediated autophagy [22]. In our data, we also observed the correlation of SOCS6 downregulation and impaired autophagy in HCC cells. In cancer cells, autophagy induction seems to be a double-edged sword, which can suppress tumorigenesis by inducing cell death, but it also facilitates tumor growth by promoting cancer cell survival under nutrient-restricted conditions [36, 37, 38]. It remains to be investigated how autophagy impairment caused by HnRNPU-AS1/SOCS6 downregulation specifically contributes to HCC progression. In addition, the mechanisms of autophagy regulation by HnRNPU-AS1/SOCS6 axis remains to be elucidated.
In conclusion, our study demonstrated the tumor suppressor function of HnRNPU-AS1 in HCC. HnRNPU-AS1 is downregulated in HCC tissues and cell lines, which is negatively correlated with the upregulation of miR-556-3p and miR-580-3p. miR-556-3p and miR-580-3p function as downstream mediators of HnRNPU-AS1 to suppress the expression of tumor suppressor SOCS6 (Fig. 8). These data suggest that HnRNPU-AS1 inhibits the progression of HCC by targeting miR-556-3p and miR-580-3p/SOCS6 axis.
Disclosure of competing interests
The authors declare that there are no competing interests associated with the manuscript.
Fund
This study was approved by the Shanxi Provincial Key Research and Development Program (2019SF-169).
Author contributions
Conception: Di Zhang.
Interpretation or analysis of data: Li Zhang, Yao Zhao, Hao Guan.
Preparation of the manuscript: Li Zhang.
Revision for important intellectual content: Li Zhang and Di Zhang.
Supervision: Di Zhang.
