Abstract
BACKGROUND:
Long-non-coding RNAs, a class of transcripts with lengths
OBJECTIVE:
To examine the function of LINC00052 in MCF-7 breast cancer cells.
METHODS:
Loss-of-function studies were performed to evaluate LINC00052 role on MCF-7 breast cancer cells. Microarray expression assays were performed to determine genes and cellular functions modified after LINC00052 knockdown. Next, the impact of LINC00052 depletion on MCF-7 cell respiration and migration was evaluated.
RESULTS:
1,081 genes were differentially expressed upon LINC00052 inhibition. Gene set enrichment analysis, Gene Ontology and Key Pathway Advisor analysis showed that signalling networks related to cell migration and oxidative phosphorylation were enriched. However, whereas LINC00052 knockdown in MCF-7 cells revealed marginal difference in oxygen consumption rates when compared with control cells, LINC00052 inhibition enhanced cell migration in vitro and in vivo, as observed using a Zebrafish embryo xenotransplant model.
CONCLUSION:
Our data show that LINC00052 modulates MCF-7 cell migration. Genome-wide microarray experiments suggest that cancer cell migration is affected by LINC00052 through cytoskeleton modulation and Notch/
List of abbreviations
Introduction
Breast cancer is a heterogeneous group of diseases that represent the first cause of cancer-related death among women worldwide. Currently, breast cancer is classified on different subtypes based on the presence of oestrogen and, progesterone receptors and human epidermal growth factor receptor 2 [1]. Although great advances have been made in breast cancer research, patient mortality has remained high and still merits attention. Therefore, it is crucial to find novel relevant breast cancer regulators, among which the knowledge about epigenetic regulators has elicited high interest [2].
Genome-wide sequencing technology has shown that only 2% of the human genome encodes for proteins while the remaining genome includes a broad diversity of non-coding genes [3]. The latter have been linked to key cellular processes that are often altered in cancer. In particular, current research has focused in long (
lncRNA have been directly implicated in breast carcinogenesis, where they could play an important role in subgrouping breast cancer patients and are associated with known clinical phenotypes. Indeed, a recent study identified a group of 4 lncRNA that are specifically modulated in TNBC (triple-negative breast cancer) [9]. In addition, lncRNA differential expression in both tissue samples and cell lines has been associated with hierarchical groups related to subclassification system PAM50 [10]. Moreover, a differential co-expression between mRNA and lncRNA was observed when normal and tumoral breast tissues were compared [11]. In this sense, particular lncRNA have been shown to play a key role in MCF-7 breast cancer cells proliferation and in cell cycle-related gene expression upon oestrogen treatment [12]. In addition, lncRNA have been considered as predictors of breast cancer metastasis and recurrence risk. Accordingly, lncRNA-ROR has been related to Epithelial-Mesenchymal Transition (EMT) in breast cancer tissues [13]. NKILA, on the other hand, was identified as a lncRNA that suppresses breast cancer cell metastasis through regulation of the NF-
Previously, we evaluated the transcriptome of MCF-7 breast cancer cells in MCTS (Multicellular Tumoral Spheroids), a 3D model that mimics the tumour behaviour of cancer cells in vivo [15, 16]. Signalling pathways analysis identified gene signatures related to metastasis, oxidative mitochondrial metabolism and cell cycle. In these experiments, we found that LINC00052, an intergenic lncRNA, was strongly downregulated during MCTS growth. Interestingly, this lncRNA has also been associated to cancer cell proliferation, differentiation, and migration [17, 18, 19, 20, 21, 22]. LINC00052 promotes gastric and head and neck squamous cell carcinoma; however, it acts as a tumour suppressor in hepatocarcinoma, glioma, pancreatic as well as in colorectal cancer. Nevertheless, the role of LINC00052 in breast cancer still remains poorly understood. Salameh and colleagues have shown that HER3 expression correlates with LINC00052 levels and that when BC cells overexpress HER3, LINC00052 promotes growth and survival [17]. Though, LINC00052 is included within a lncRNA-non-TNBC cancer signature proposed by Lv et al. [9]. In the present study, microarray expression analyses showed that cell migration and mitochondrial metabolism-related pathways were enriched after LINC00052 knockdown. Consistently, we evaluated the function of LINC00052 in MCF-7 cells and found that its inhibition enhances cell migration in vitro and in vivo, and marginally affects mitochondrial metabolism. Our results suggest that LINC00052 regulates cellular processes that may allow tumour progression and provide further insight into the function of lncRNA in breast cancer.
Materials and methods
Analysis in silico of LINC00052 expression in breast cancer tissues
The Km-Plotter platform was used to analyse patient outcomes indicating LINC00052 best cut-off expression levels, using mRNA gene chip databases for breast cancer samples [23].
Cell culture
The MCF-7 cell line was obtained by ATCC (Cat. No. HTB-22) and was genotyped at INMEGEN high-tech sequencing unit. Cells were grown at 37
Multicellular tumour spheroid generation
MCTS were formed as previously described [16, 24]. Briefly, cells were detached once they reached 80% confluence, washed with PBS 1X and separated with trypsin (Cat. No. 25200-056 Gibco) in PBS 1X EDTA 0.05 mM. Pelleted cells were resuspended in 3 mL of L-15 medium supplemented with 5% FBS and were counted in Neubauer chambers. One million cells were seeded in 12.5 cm
shRNA design and stable cell line establishment
Two shRNAs for LINC00052 (lnc-AGBL1-4:1) were designed using the E-RNAi software [25]. The sequences used were: A) 5
Total RNA extraction and reverse transcription (RT) reaction
Total RNA was extracted from shLuc, shA and shB MCF-7 cells grown as monolayers using TriReagent (Cat. No. AM9738 Ambion), as suggested by the provider. Isolated RNA was resuspended in 40
qPCR
Specific primers were designed to analyse the expression of TBP and LINC00052 genes (Supp. Table 1). qPCR was performed with SYBR
Transcriptome analysis by microarrays
HuGene-2_0-st-v1 Arrays from Affimetrix were used to analyse gene expression of shLuc and shB MCF-7 cells. RNA was extracted as previously described from 3 biological samples of each cell line [24]. RNA quality was evaluated with a Bioanalyzer from Illumina (RIN
Gene expression bioinformatic analyses
Microarray data (.cel files) were analysed with Transcriptome Analysis Console (TAC 4.0) software from Affimetrix (Fold change
Data intensity from differentially expressed genes (FC
Additionally, differentially expressed genes data were explored with Key Pathway Advisor (KPA, Version 17.4) platform (
Finally, to analyse miRNA and mRNA associations, fold change and
Microarray data validation by qPCR
Gene expression changes identified by microarray experiments were additionally evaluated by qPCR as previously described, employing specific primers (Supp. Table 1). For each gene, mRNA relative expression was calculated normalizing to TBP expression and fold change (FC) was estimated on reference to the shLuc MCF-7 cells found expression. Gene expression changes were assessed in three independent biological samples.
Oxygen consumption and determination of OxPhos-dependent flux
To measure the rate of oxygen consumption, shLuc, shA and shB-MCF-7 (80% monolayer confluence) cells were detached with 0.025 mM of trypsin PBS 1x and 0.05 mM EDTA for 5 minutes at 37
In vitro migration assay
shLuc, shA and shB MCF-7 cells were grown as monolayers, detached with Trypsin and counted. 30,000 cells of each cell clone were seeded in serum-free RPMI medium inside upper chambers of Transwell cell culture plates (Cat. No. 3422 Corning, USA). The lower chambers of plates were filled with RPMI medium with 15% FBS as a chemoattractant. After 48 h cell migration was assessed by staining the cells at filter bottom surfaces with 0.05% Crystal Violet. Five fields from each filter were imaged with a Leica microscope (10x) and a mean cell number was calculated for each condition. Three measurements were performed for all cell lines in all experiments. Assays were performed three independent times.
Gelatine degradation assay
shLuc, shA and shB MCF-7 cells were seeded on FITC-conjugated gelatine (Cat. No. G-13187 Molecular Probes) coated coverslips for 7 h. Cells were fixed and stained for F-actin to quantify gelatine degradation as previously described [24, 31]. Three independent experiments were performed.
Zebrafish husbandry
Zebrafish (Danio rerio) of the Tab Wik genetic background (donated by Dr. Ernesto Maldonado from the Institute of marine sciences and limnology-UNAM and by Dr. Francisco Carmona from the Instituto de Fisiología Celular-UNAM), were maintained at a temperature of 28.5
Care and feeding of fish and larvae followed the standard procedures described in “The zebrafish book. A guide for the laboratory uses of zebrafish (Danio rerio)”. All experiments and procedures in animals are carried out following the guidelines of “THE INSTITUTIONAL ANIMAL CARE AND USE COMMITTE (IACUC) UNIVERSITY OF OREGON”, available for consultation in “The zebrafish book. A guide for the laboratory use of zebrafish (Danio rerio)” [32]. In turn, at the Instituto Nacional de Medicina Genómica, all procedures in zebrafish followed the guidelines established by the local committee on ethics and management of laboratory animals (CICUAL, Comite Interno para el Cuidado y Uso de Animales de Laboratorio; Approved Verdict No. 427). Zebrafish embryo and larvae were euthanized with 0.4% tricaine (MS-222; Sigma) for 1 h (until heart beating stops).
Regulation of LINC00052 expression. LINC00052 expression was analysed by KMplots bioinformatic tool. (A) Graph shows relapse-free survival and (B) graph depicts distant metastasis-free survival, regarding LINC00052 expression. (C) LINC00052 expression decreases early during MTCS formation. MCF-7 cells were grown as a monolayer (M) and as multicellular aggregates during 4 h, 2 and 4 days. LINC00052 expression was evaluated in these growth conditions by RT-qPCR. Graph shows LINC00052 average fold change in three independent experiments. (D) LINC00052 expression knockdown. MCF-7 cells were stably transfected with 2 independent shRNA, shA and shB as well as with a control shRNA, shLuc. LINC00052 expression was assessed in these cell lines by RT-qPCR in 3 independent samples. Statistical difference was calculated by a One-way ANOVA test (PRISMA Graph software). Asterisks indicate a 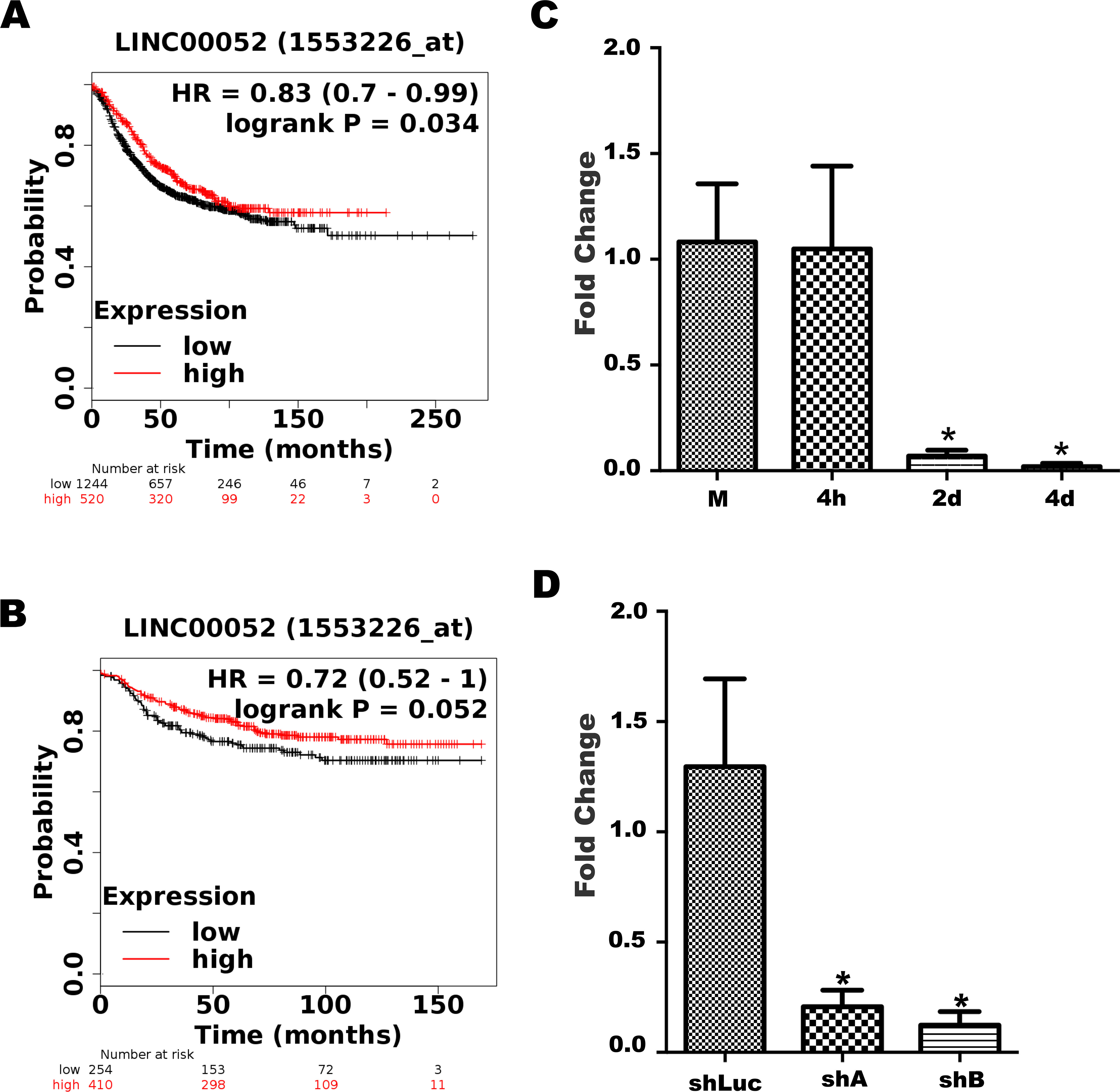
Two days post-fertilization (dpf) zebrafish embryos were dechorionated and anesthetized with 0.04% tricaine, and then microinjected with MCF7 cancer cells into the middle of the embryonic yolk sac region. For the subsequent detection in zebrafish body, either MCF7-shLuc, MCF7-shA or MCF7-shB cells were transiently transfected with pSiren-EGFP vector 48 hours prior to injection. On the day of the xenografts, cells were re-suspended at a concentration of 10
Statistical analysis
All experiments were performed at least by triplicate and included at least two technical replicates. One-way ANOVA test was employed in all assays for independent groups to compare mean values (
LINC00052 knockdown affects MCF-7 cells gene expression. (A) Gene expression analysis was done in shLuc-MCF-7 and shB-MCF-7 cells using Affymetrix chip HuGene-2_0-st-v1. Graph shows the number of differentially regulated transcripts in shB-MCF-7 cells compared to control shLuc-MCF-7 cells. (B) Number of LINC00052 modulated coding and non-coding transcripts. (C) A heatmap was generated using a Euclidean correlation method and shows differentially expressed genes in all biological samples.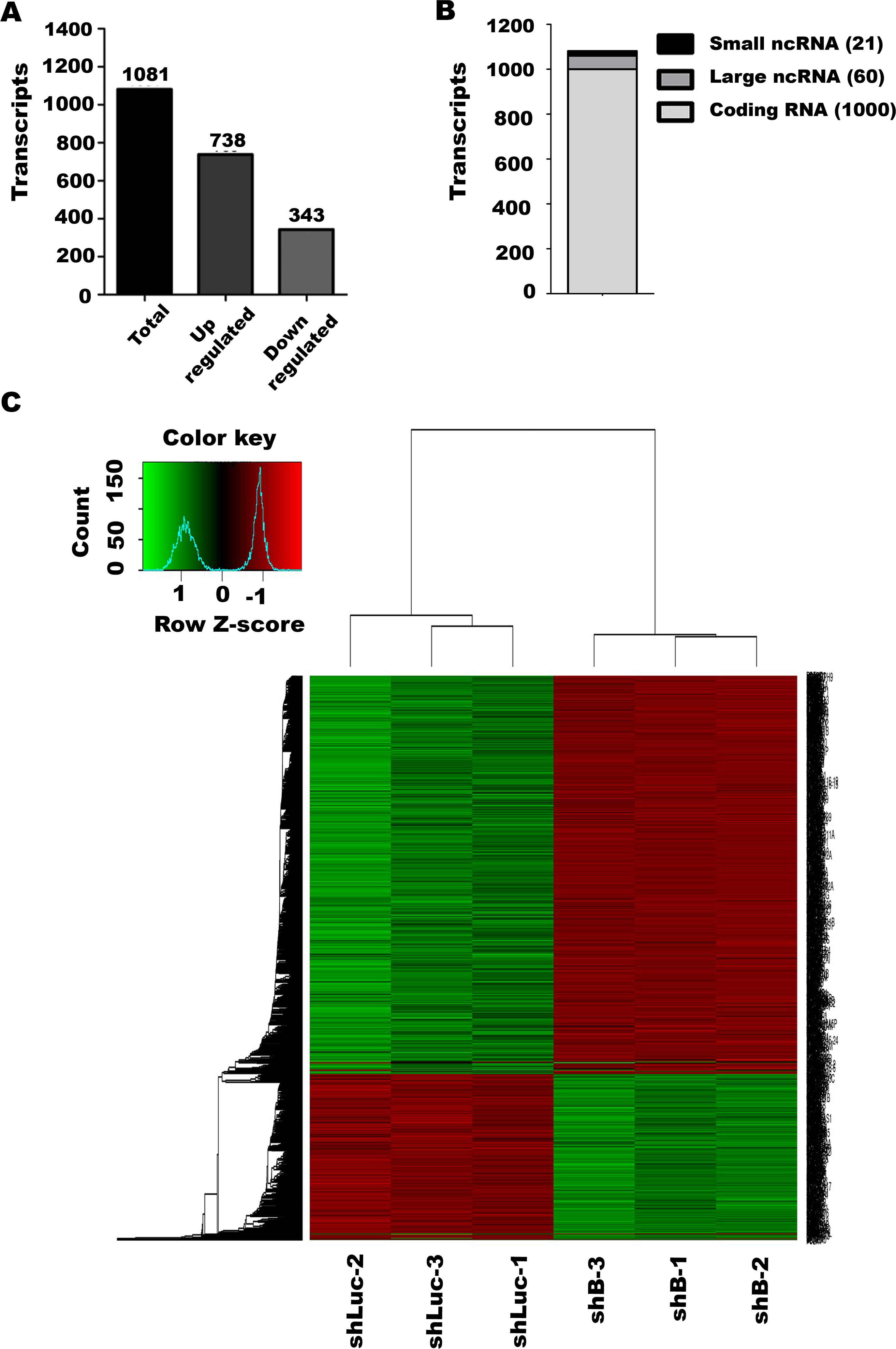
LINC00052 expression decreases in breast tumour from patients with worse outcome
LINC00052 is an intergenic lncRNA located at chromosome 15, between the AGBL1 and NTRK3 genes and intronic-antisense to lnc-NTRK3. LINC00052 was selected for further investigation based on its low expression profile during MCTS development, as previously reported [15, 16]. To determine the clinical relevance of LINC00052, we queried breast cancer patient samples data using the Kaplan-Meier plotter (KM-plotter) web-tool [23]. KM-plotter analysis revealed a worse prognosis for patients with low LINC00052 expression, showing a decreased relapse-free and distant metastasis-free survival (Fig. 1A and B).
LINC00052 expression is downregulated upon MCTS formation
To further explore LINC00052 regulation, we analysed its transcript levels by RT-qPCR in MCF-7 cells growing as a monolayer (M) and during the early phases of MCTS formation, at 4 h, 2 and 4 days. Interestingly, we found that its expression was maintained 4 h after cell detachment (Fig. 1C). However, after two days of MCTS growth, when cell-cell interactions began to be established, LINC00052 expression was significantly downregulated (FC-15 and
Our previous research demonstrated higher migration capacity and increased respiration rates when MCF-7 cells were grown as MCTS since early stages (6 days) [15, 16]. As previously mentioned, LINC00052 expression was also downregulated, suggesting that this lncRNA could facilitate a more aggressive breast cancer cells phenotype. To assess these possibilities and to mimic LINC00052 downregulation in breast cancer cells as in MCTS, we utilized two specific shRNA targeting LINC00052 to generate stably transfected MCF-7 cell clones, hereafter named shA and shB. These cell clones displayed significantly lower levels of LINC00052 expression in comparison with control MCF-7 cells (control shRNA, shLuc) when cells are grown as monolayers (Fig. 1D).
Gene expression changes in LINC00052-knockdown MCF-7 cells
We performed microarray expression analyses (Affimetrix chip HuGene-2_0-st-v1) to gain further insight regarding the genes and signalling pathways modulated by LINC00052 in MCF-7 cells. We found 1,081 differentially regulated genes (Fold change
Enrichment network analysis upon LINC00052 inhibition. Enrichment nodes were generated with GO:BP database and obtained with significant 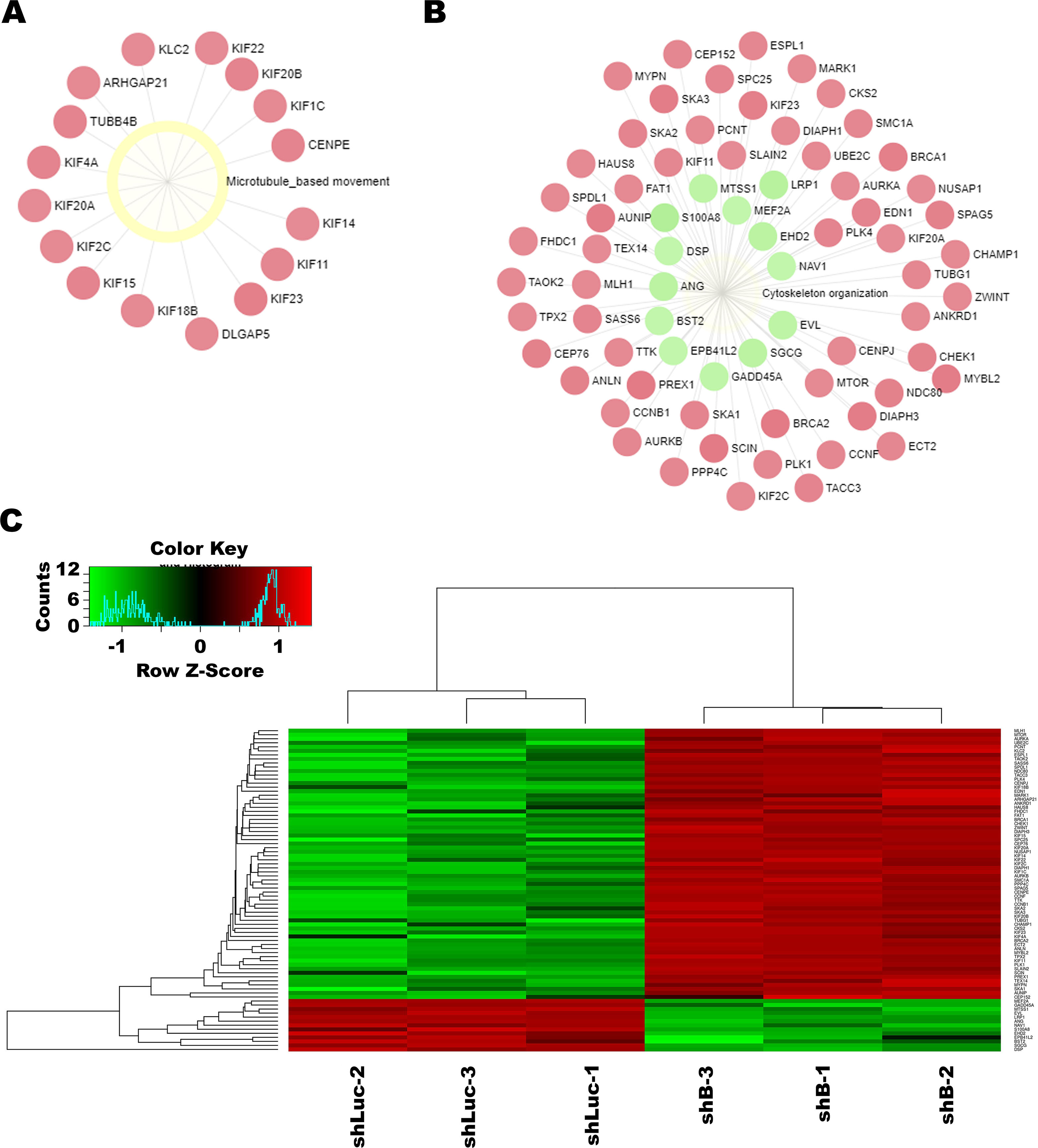
We observed that the top up- and downregulated coding genes were associated with known cellular processes modulated in cancer cells (Table 1). Among these, the highest upregulated gene was FOLH1 (FC 16.790) whereas the lowest was SERPINA3 (FC-13.010). Gene expression changes were confirmed by RT-qPCR in shLuc, shA and shB MCF-7 clones for 5 of the top modulated genes (Table 2). We observed that the sense of expression change was the same as in microarrays.
Differentially expressed coding genes
Validation of gene expression changes
Gene expression changes (mRNA accumulation) were analysed by RT-qPCR using specific primers for each gene in shLuc, shA and shB MCF-7 cell lines. Values are expressed as Fold Change (FC) and TBP gene expression was employed as an internal expression control. Gene expression changes were statistically significant (
“Cytoskeleton organization” node molecules modulated by LINC00052 knockdown
MCF-7 cells oxygen consumption rates
Basal respiration rates and OxPhos-dependent flux was evaluated in shLuc, shA and shB MCF-7 cells grown as monolayers. Values are shown in natgO/min
The microarray data was first explored using the Gene Set Enrichment Analysis (GSEA) tool [28], looking for enriched cellular processes. The KEGG platform evaluation demonstrated the positive enrichment of oxidative phosphorylation and mitochondrial protein complexes in shB MCF-7 cells (
Gene Ontology microarray data analysis
In parallel, a network enrichment analysis was performed with Gene Ontology (BP: Biological Processes) database [33]. On the modelling of microarray data this analysis showed that two of the main significant nodes were related to “Microtubules-based movement” (Fig. 3A) and “Cytoskeleton organization” (Fig. 3B). Among node components we observed key molecules for cell motion that were positively enriched in both processes. As well, and in consistence with our functional assays, these molecules participate in breast cancer cell motility and include: Receptors/ligands (ANG, MARK1); Scaffolding proteins (ANL, EBPL41L2, SKA2), RHO-GAP/GEF (ARHGAP1, ECT2, PREX1), Junction components (CLDN1, END1, MYBL2), Trafficking protein (EHD2) and Kinases (CKS2, PLK1 and 4) (Table 3). Moreover, this analysis showed that 11 Kinesins, key for breast cancer pathogenesis were modulated by LINC00052 inhibition (Table S2). Gene expression changes related to these nodes are more specifically depicted by a heatmap, generated by the Shiny heatmap tool (Fig. 3C).
An additional microarray data analysis was performed with the Key Pathway Advisor (KPA) tool. KPA predicted 12 significantly modified signalling pathways related to cell migration based on the observed differentially expressed genes (Table S3). Among them, “Signal transduction_Angiotensin II/AGTR1 signalling via Notch,
Finally, we hypothesized that LINC00052 could influence breast cancer cells behaviour by miRNA regulation (S1A Fig). In this sense, the analysis regarding the observed set of miRNAs and mRNAs showed that hsa-miR-130b, which targets the DIAPH3 mRNA, was downregulated. In parallel, hsa-miR-21, a microRNA that binds to FOXM1 mRNA, was also inhibited (S1B Fig).
Effects of LINC00052 downregulation on cell respiration rates
Microarray data analysis suggested that LINC00052 downregulation triggered gene expression changes associated with mitochondrial functions. This result is in agreement with our previous findings where we observed that mitochondrial respiration increased rates were associated with more aggressive MCTS characteristics, together with LINC00052 downregulation [15, 16]. Afterwards, we analysed the oligomycin sensitive respiration rate in shA and shB MCF-7 cells, which represents mitochondrial ATP synthesis by complex V coupled with the respiratory chain activity [34]. We found a small increment in shA and shB clones’ basal respiration when compared to control shLuc MCF-7 cells (Table 4). The OxPhos-dependent flux, determined by exposing cells to oligomycin (a specific F1 FO-ATP synthase inhibitor), revealed that shA and shB cells had a higher ATP synthesis-coupled respiration rate when compared to control shLuc cells (Fisher poshoc analysis indicated statistically significant differences between shLuc and shA (
LINC00052 knockdown results in greater MCF-7 cell migration capacity in vitro and in vivo
Enhanced cell migration capacity was observed previously in cells of MCTS model [16]. In agreement, transcriptome changes triggered upon LINC00052 knockdown were associated with cell migration. Therefore, we tested whether LINC00052 inhibition could change this phenomenon. Using in vitro Boyden chamber assays, we observed a significant migration increment in shA and shB cells in comparison to shLuc control cells after 48 h of culture (Fig. 4A). In parallel, we analysed the ability of these cells to degrade the extracellular matrix (ECM) through a fluorescent-gelatine assay. We observed that both shA and shB cells had a significantly greater capacity to degrade ECM than shLuc control cells (Fig. 4B). Finally, we examined the participation of LINC00052 in cell migration in vivo using a zebrafish model (2-day old embryos), comparing the migration rate of fluorescent shA, shB, and shLuc MCF7 cells to the posterior trunk and tail region (Fig. 4C). We counted the number of cells inside the tail area of all embryos [35], finding a significantly higher number of migrating cells in the embryos injected with shA and shB MCF-7 clones than in those inoculated with shLuc MCF-7 control cells (Fig. 4D). Of note, we examined whether LINC00052 inhibition affected MCTS formation in shA, shB cell lines in comparison with control shLuc and we did not observe differences between the three cell lines (Fig. 4E).
LINC00052 knockdown modulates MCF-7 cells migration. (A) shLuc, shA and shB MCF-7 cells in vitro migration capacity. Cells were allowed to migrate in commercial migration chambers for 24 h as indicated formerly. Graph displays cell average per field assessed in three independent experiments. (B) FITC-gelatine degradation capacity of shLuc, shA and shB MCF-7 cells. Cells were grown on top of FITC-gelatine during 7 h to assess their extracellular matrix degradation ability as previously described. Chart shows the average cell degradation area per field evaluated in three independent experiments. In (A) and (B) Statistical difference was calculated by a One-way ANOVA test (PRISMA Graph software). Asterisks indicate a 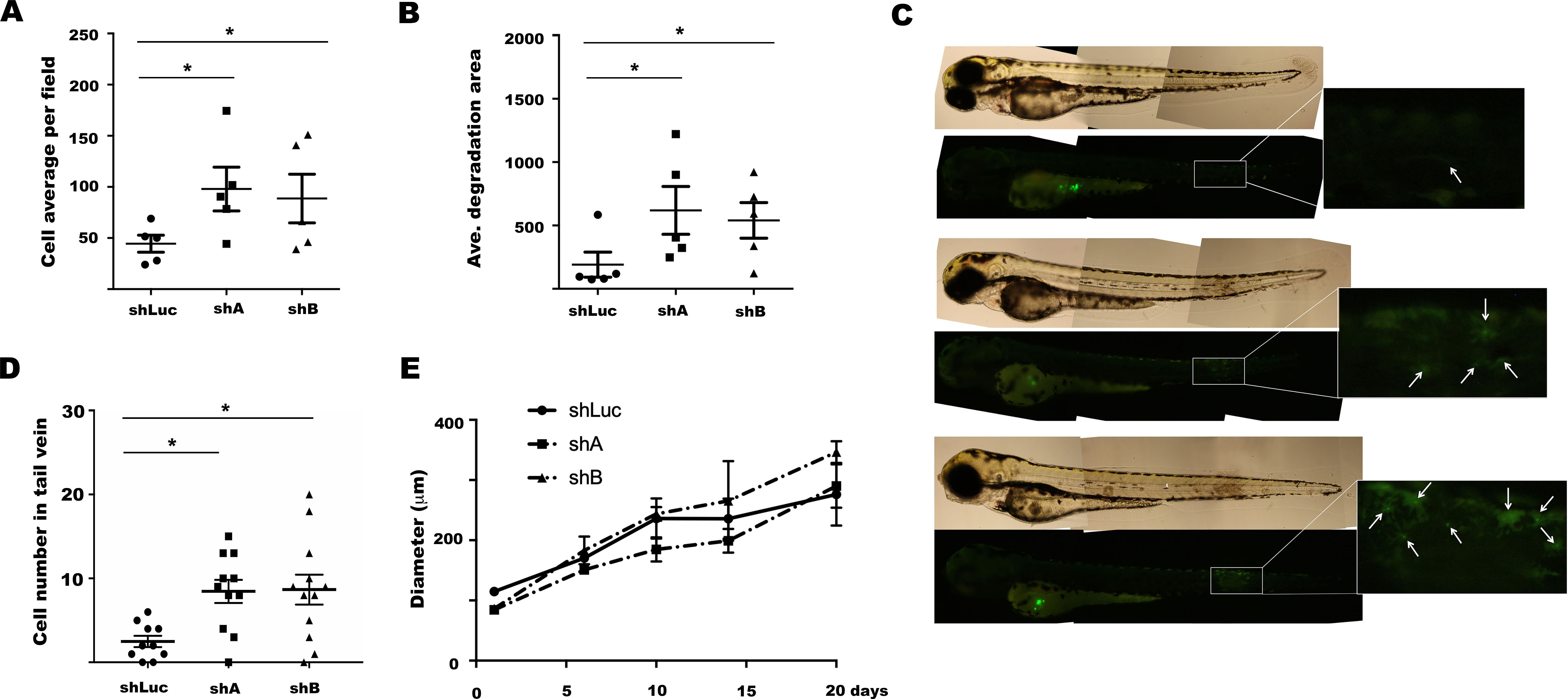
Altogether these findings suggest that downregulation of LINC00052 expression is associated with enhanced cell migration and invasion abilities and thus, to a more aggressive phenotype in MCF-7 cells.
The present study provides evidence that LINC00052 plays a regulatory role in altered cellular processes commonly observed in breast cancer. Previous work from our laboratory showed a transcriptome change when MCF-7 cells were grown as MCTS in comparison with a monolayer counterpart. Further, we observed an expression shift not only in coding genes but also in non-coding ones, which were subsequently reflected by the hyperactivation of cellular processes such as metabolism (oxidative phosphorylation and cholesterol signalling), cell cycle, and migration.
Interestingly, among the most inhibited molecules in MCTS we found a non-coding RNA, LINC00052 [15, 16]. These findings led us to evaluate whether LINC00052 inhibition triggers some of breast cancer cells characteristics. First, when we analysed LINC00052 expression in MCF-7 breast cancer cells at early time points of MCTS formation, we found that its expression becomes downregulated after 48 and 96 hours under non-hypoxic conditions and when the establishment of cell-cell junctions is a main process. This data suggests that the downregulation of LINC00052 could be associated with cell-cell interactions during the free substratum growth of 3D cultures.
In agreement, the analysis of a breast cancer patient database revealed that relapse-free survival and distant metastasis-free survival rates are poorer in breast cancer patients with lower LINC00052 expression. This observation is consistent with a previous report showing that LINC00052 expression was lower in basal breast cancer samples and cell lines compared with a non-basal counterpart [9] and support the idea that LINC00052 could act as a tumour suppressor during breast cancer development. Even though, further patient data analysis would be required to support this hypothesis. In this regard, the work of Salameh and colleagues also showed that LINC00052 together with HER3 mRNA expression is increased in luminal BC samples in comparison to normal tissue [17], supporting the idea that LINC00052 delicate regulation acts during breast cancer development and progression.
Nonetheless these findings [9], the function of LINC00052 in breast cancer cells remains poorly understood. In this sense, Xiong et al. [19] showed that the overexpression of LINC00052 results in the inhibition of cell migration, invasion and proliferation in hepatocellular carcinoma SMMC7721 cells [19]. Furthermore, there is evidence suggesting that LINC00052 acts as sponge for miR-128, miR-485-3p and miR-101-3p miRNAs, which respectively regulate the NTRK3, EPB41L3 and SOX9 expression, thus modulating cell proliferation and migration in several cancers including breast cancer [20, 21]. Consistent with these findings, we observed that some key miRNA-mRNA pairs were modulated in MCF-7 cells after LINC00052 knockdown. It is possible then to suggest that LINC00052 could regulate breast cancer cell migration and invasion, by working as a miRNA sponge, as has been the case for several other lncRNA.
The microarray transcriptome analysis showed that LINC00052 regulated the expression of 1,081 genes. In agreement with our previous work, gene enrichment analysis of our microarray data suggested that LINC00052 knockdown affects oxidative phosphorylation pathways [15, 16]. In this study, we observed a minor increase in the aerobic metabolism after LINC00052 knockdown in MCF-7 cells as we observed acceleration of both the basal respiration and the oligomycin-resistant respiration. However, we currently do not know if these results are associated with higher efficiency of mitochondria in these cells as we did not observe any response to the uncoupling agent CCCP (data not shown), as has been reported for other cancer cell lines [36]. It remains to be answered if LINC00052 knockdown promotes an increase in the mitochondrial number and/or an increase in the concentration/activity of mitochondrial respiratory chain complexes.
Afterwards, we validated the expression changes of five of the top regulated genes that are known to have a role in breast cancer development and cell migration. Among them, we observed FOLH1 as the highest upregulated gene. FOLH1, also known as prostate-specific membrane antigen, modulates breast cancer progression and metastasis by PI3K signalling [37]. Another upregulated gene by LINC00052 inhibition was PEG10 (Paternally expressed gene 10). Overexpression of this gene correlates with decreased survival of breast cancer patients. Moreover, in breast cancer cell lines PEG10 overexpression promotes cell proliferation, migration and metastasis through ERK/MMP-7 signalling [38, 39]. Additionally, we validated a reduced S100A7 (Psoriasin) expression when LINC00052 is inhibited. Interestingly, when this calcium-binding protein is overexpressed by MCF-7 cells, Rac-1 pathway and MMP-9 secretion are inhibited, thereby cell migration and invasion are repressed [40]. We also confirmed the IGFBP5 downregulation upon LINC00052 knockdown. In agreement with our results, previous work has shown that IGFBP5 induces MCF-7 cells adhesion while it inhibits cell migration, by its interaction with
Moreover, in vitro and in vivo assays allowed us to determine that the major changes triggered by LINC00052 inhibition were associated with enhanced cell migration and invasion. Cell migration has been widely studied in zebrafish embryos, since cancer cells preserve their phenotype within the zebrafish [45, 46]. Remarkably, we observed that apparent cell groups moved together within zebrafish embryos. These observations are consistent with the fact that cancer cells commonly migrate as clusters from the primary tumour to reach other tissues [47]. Interestingly, LINC00052 inhibition induced the expression of genes linked to cell dissemination pathways, which was reflected by the increased cell migration capacity in the zebrafish embryo xenograft assay.
Consistent with our findings of a role of LINC00052 in cell migration, gene enrichment analysis showed a “Cytoskeleton organization” node that contains molecules directly related to metastasis. In agreement, within the Microtubule-based movement node, we observed that a group of these genes belong to the Kinesin superfamily (KIF). Kinesins role is mainly related to intracellular transport (vesicles and organelles as well as chromosomes); therefore, and as expected, alteration of Kinesins contributes to cell transformation, including invasive and metastatic behaviour [48, 49]. Particular or collective functions of these LINC00052-modulated genes associate with breast cancer progression and cell migration.
In agreement with these observations, integral bioinformatic analysis inferred signalling pathways that modulate cell migration. First, we observed the route related to PTHR1 and Angiotensin II, “Pro- oncogenic action of PTHrP in breast cancer”. Remarkably, PTHrP signalling pathway participates in breast cancer cell adhesion, migration and metastasis [50, 51]. In particular, we observed the upregulation of heterotrimeric G proteins, G-alpha after LINC00052 knockdown. Through this signalling pathway we could hypothesize that the observed increase in PLK1/4 expression could lead to its activation and the enrichment of cell protrusion, that phosphorylates and activates RacGAP1/ECT2 GEF [52], which is also upregulated by LINC00052 knockdown. GEF proteins lead to RhoA activation and thus trigger several signalling pathways that may promote cell migration [53, 54]. For instance, RhoA activates ROCK kinase and Diaphanous formins, from which DIAPH1 and DIAPH3 are upregulated after LINC00052 inhibition.
Interestingly, we noticed that downstream Angiotensin II, another signalling pathway, “Signal transduction_Angiotensin II/AGTR1 signaling via Notch,
In conclusion, the present study demonstrates that LINC00052 causes changes in the transcriptome of MCF-7 cells and inhibits cell migration. Although the precise LINC00052 action and molecular mechanisms await further investigation, these phenotypic changes are possibly associated with cytoskeleton modulation and signal transduction pathways NOTCH-mTOR/AKT-NF-
Supplementary data
The supplementary files are available to download from
Footnotes
Acknowledgments
José Manuel Sánchez López thanks the Postgraduate program of Biological Sciences (Posgrado en Ciencias Biológicas, Doctorado en Ciencias Biológicas) and Universidad Nacional Autonoma de Mexico (Mexico’s Autonomous National University), UNAM, Faculty of Medicine. JMSL additionally thanks CONACYT national fellowship program, for support during his graduate training (CVU No. 777479). We acknowledge Microarray High Technology Unit from Instituto Nacional de Medicina Genomica. We thank Dr. Julio Perez-Carreon and Dr. Marcela Lizano Soberon for kind manuscript review and for support during JMSL graduate training. We are in debt with Dr. Edgar Krötzch and Dr. J. Perez-Carreon for allowing the use of microscopes.
