Abstract
Long non-coding RNAs have been implicated in human cancer but their mechanisms of action are mainly undocumented. In this study, we found that HOXA11-AS expression was upregulated in non–small cell lung cancer tissues and cell lines. High levels of HOXA11-AS expression were correlated with larger tumor size and lymph node metastasis. Functional analysis revealed that HOXA11-AS promotes non–small cell lung cancer cell proliferation and invasion. In particular, HOXA11-AS functions as a competing endogenous RNA to regulate transcriptional factor Sp1 expression via sponging miR-124. Collectively, our findings reveal an oncogenic role for HOXA11-AS in non–small cell lung cancer tumorigenesis.
Introduction
Lung cancer is the most common cancer worldwide in terms of both incidence and mortality. 1 Non–small cell lung cancer (NSCLC) accounts for approximately 85% of all lung cancer cases. Despite recent advancements in clinics, the prognosis of NSCLC remains poor, with a 5-year overall survival rate of around 11%. 2 Because NSCLC is a multi-gene disorder, the biological processes involved in NSCLC tumorigenesis are complicated. Thus, a detailed understanding of the mechanisms and molecular pathways underlying NSCLC development and progression is essential to improve diagnosis and treatment of NSCLC patients.
Long non-coding RNAs (lncRNAs) are non-coding transcripts with over 200 nucleotides in length. In recent years, lncRNAs have drawn attention as potential biological regulators involved in a range of cellular activities such as cell proliferation, apoptosis, and invasion.3–5 The underlying mechanisms of lncRNAs are varied and include acting as competitive endogenous RNAs similar to microRNA (miRNA) sponges, chromatin remodeling, and histone–protein modification.6–9 For example, the lncRNA MALAT1 and miR-205 could interact with and repress each other, finally acting as crucial regulators for function of renal cancer. 10 PVT1 suppressed cell growth and induced apoptosis by binding to the EZH2 protein, a histone methyltransferase of the PRC2 complex, in lung cancer. 11 However, limited knowledge is available concerning the roles of lncRNA in NSCLC, which needs to be well documented.
In this study, we investigated the relationship between HOXA11-AS and NSCLC. We found that HOXA11-AS was significantly upregulated in NSCLC tissues and cells; HOXA11-AS could regulate cell growth and invasion and act as a molecular sponge for miR-124. These studies will help clarify the role of HOXA11-AS in NSCLC progression and its potential as a therapeutic target.
Materials and methods
Tissue collection and cell culture
We obtained 78 paired NSCLC and adjacent non-tumor tissues from patients who underwent surgery at Yinzhou Hospital Affiliated to Medical School of Ningbo University (Ningbo, China) between January 2012 and March 2015. Informed consent was obtained from all subjects, and this study was approved by the Review Board of Hospital Ethics Committee. All cases were confirmed as NSCLC based on histopathological evaluation.
The lung adenocarcinoma cell lines, including A549, H1299, H460, and Calu1, and one cultured human lung epithelial cell (BEAS-2B), were grown in Dulbecco’s modified Eagle’s medium (DMEM; Gibco, Grand Island, NY) containing 10% fetal bovine serum (FBS) with 100 μg/mL of penicillin/streptomycin at 37°C with 5% CO2.
Quantitative real-time polymerase chain reaction
Total RNA was extracted using TRIzol reagent (Invitrogen Carlsbad, CA). A volume of 1 μg of total RNA was reverse transcribed to complementary DNA (cDNA) using random primers under standard conditions with the PrimeScript RT Reagent Kit (TaKaRa, Dalian, China). We performed real-time polymerase chain reaction (PCR) analyses using SYBR Premix Ex Taq (TaKaRa) according to the manufacturer’s instructions. The primers used are listed in Supplementary Table S1.
Plasmid construction and cell transfection
The full-length complementary cDNA of human HOXA11-AS was synthesized by Invitrogen (Shanghai, China) and cloned into the expression vector pCDNA3.1 (Clontech Laboratories, Inc., San Francisco, CA). The miR-124 mimics and negative control (NC) were synthesized by GenePharma (Shanghai, China). Small-interfering RNAs (siRNAs) targeting human HOXA11-AS were designed according to previously validated oligonucleotides 12 and synthesized by GenePharma. Transfection was performed with Lipofectamine 2000 (Invitrogen, CA, USA) according to the manufacturer’s protocol.
Cell proliferation and invasion assays
Cell viability was performed using a Cell Counting Kit-8 (CCK-8; Dojindo Molecular Laboratories, Kumamoto, Japan) according to the manufacturer’s protocol. For colony-forming assays, the cells were seeded on six-well plates at 1000 cells per well for 2 weeks. Then, the cells were washed twice with phosphate-buffered saline (PBS), fixed, and stained with 0.5% crystal violet, and the numbers of colonies were counted. Cell invasion assays were performed using transwell chambers as described previously. 13
Luciferase reporter assay
Cells were cultured on 24-well plates, receiving transfection with either pGL3-HOXA11-AS vector or pGL3-HOXA11-AS-Mut vector, together with miR-124 or NC. After 48 h, the cells were harvested, and the luciferase activity was measured using the Dual-Luciferase Reporter Assay System (Promega, Madison, WI). The primers used are listed in Supplementary Table S1.
Western blot analysis
Proteins from cells were separated by sodium dodecyl sulfate–polyacrylamide gel electrophoresis (SDS-PAGE) and then immunodetection was performed with standard techniques. Antibodies to Sp1, E-cadherin, vimentin, β-catenin, Snail, Slug, and β-actin were purchased from Santa Cruz Biotechnology, Inc. (Santa Cruz, CA). Signals were visualized with SuperSignal® West Pico Chemoluminescent Substrate (Pierce, Rockford, IL) by exposure to films.
RNA immunoprecipitation assays
The EZMagna RIP Kit (Millipore, Billerica, MA) was used following the manufacturer’s protocol. H1299 and A549 cells were lysed in complete RNA immunoprecipitation (RIP) lysis buffer, and the cell extract was incubated with magnetic beads conjugated with specific antibodies or control immunoglobulin (IgG; Millipore) for 6 h at 4°C. Beads were washed and incubated with Proteinase K to remove proteins. Finally, purified RNA was subjected to quantitative real-time polymerase chain reaction (qRT-PCR) analysis.
Statistical analysis
Statistical analyses were performed using SPSS 16.0 software (SPSS, Inc., Chicago, IL, USA). All data from three independent experiments were expressed as mean ± standard deviation (SD). Differences were assessed by two-tailed Student’s t test. The relationship between HOXA11-AS, miR-124, and Sp1 was tested with two-tailed Pearson’s correlation. p < 0.05 was considered statistically significant.
Results
HOXA11-AS expression is upregulated and correlated with tumor size and lymph node metastasis of NSCLC
The level of HOXA11-AS was detected in lung cancer cell lines by quantitative PCR (qPCR). The results showed that HOXA11-AS expression was significantly higher in all four lung cancer cell lines compared with that in human lung epithelial cells (Figure 1(a)). We also detected the expression of HOXA11-AS in 78 paired NSCLC tissues and corresponding adjacent normal tissues. As shown in Figure 1(b), HOXA11-AS expression was significantly upregulated in cancerous tissues compared with normal counterparts (Figure 1(b)). To understand the significance of HOXA11-AS overexpression in NSCLC, we used the median expression of HOXA11-AS as a cutoff point to divide patients into high HOXA11-AS expression (n = 41, fold change ⩾ 6.6) and low HOXA11-AS expression (n = 37, fold change ⩽ 6.6) groups. Statistical analysis revealed that high HOXA11-AS expression was significantly correlated with tumor size and lymph node metastasis. However, HOXA11-AS expression was not associated with other factors including smoking status and histological subtype in NSCLC (Table 1). These findings suggest that HOXA11-AS overexpression may play an important role in NSCLC progression.

HOXA11-AS expression was upregulated in NSCLC cells and tissues. (a) qRT-PCR analysis of HOXA11-AS expression in four NSCLC cells and one cultured human lung epithelial cells. (b) HOXA11-AS relative expression levels were determined in 78 paired NSCLCs and their corresponding normal samples. The level of HOXA11-AS expression was normalized to GAPDH (**p < 0.01).
Association between HOXA11-AS expression and clinicopathological characteristics.

p < 0.05.
Knockdown of HOXA11-AS inhibits NSCLC cell proliferation and invasion
To investigate the functional role of HOXA11-AS in NSCLC, a pcDNA/HOXA11-AS vector was transfected into A549 cells and si-HOXA11-AS was transfected into H1299 cells. Satisfactory transfection efficiency was achieved at 48 h after transfection (Figure 2(a) and (b)). Then, CCK-8 and colony formation assays showed that knockdown of HOXA11-AS impaired H1299 cell proliferation (Figure 2(c)), while HOXA11-AS overexpression promoted A549 cell proliferation (Figure 2(d)). Furthermore, invasion assays were performed to investigate the roles of HOXA11-AS on NSCLC cell invasion ability. The results showed that knockdown of HOXA11-AS dramatically decreased H1299 cell invasion, while HOXA11-AS overexpression promoted A549 cell invasion (Figure 2(e)). Epithelial–mesenchymal transition (EMT) plays important roles in cancer cell invasion, and lncRNAs were also involved in regulation of EMT process in our previous studies. 8 In this study, western blot assays showed that mesenchymal marker proteins including vimentin, β-catenin, and the EMT-mediating transcription factor Snail and Slug were decreased in HOXA11-AS-knockdown H1299 cells, while epithelial marker E-cadherin levels were increased. On the contrary, overexpression of HOXA11-AS promotes EMT in A549 cells (Figure 2(f)).
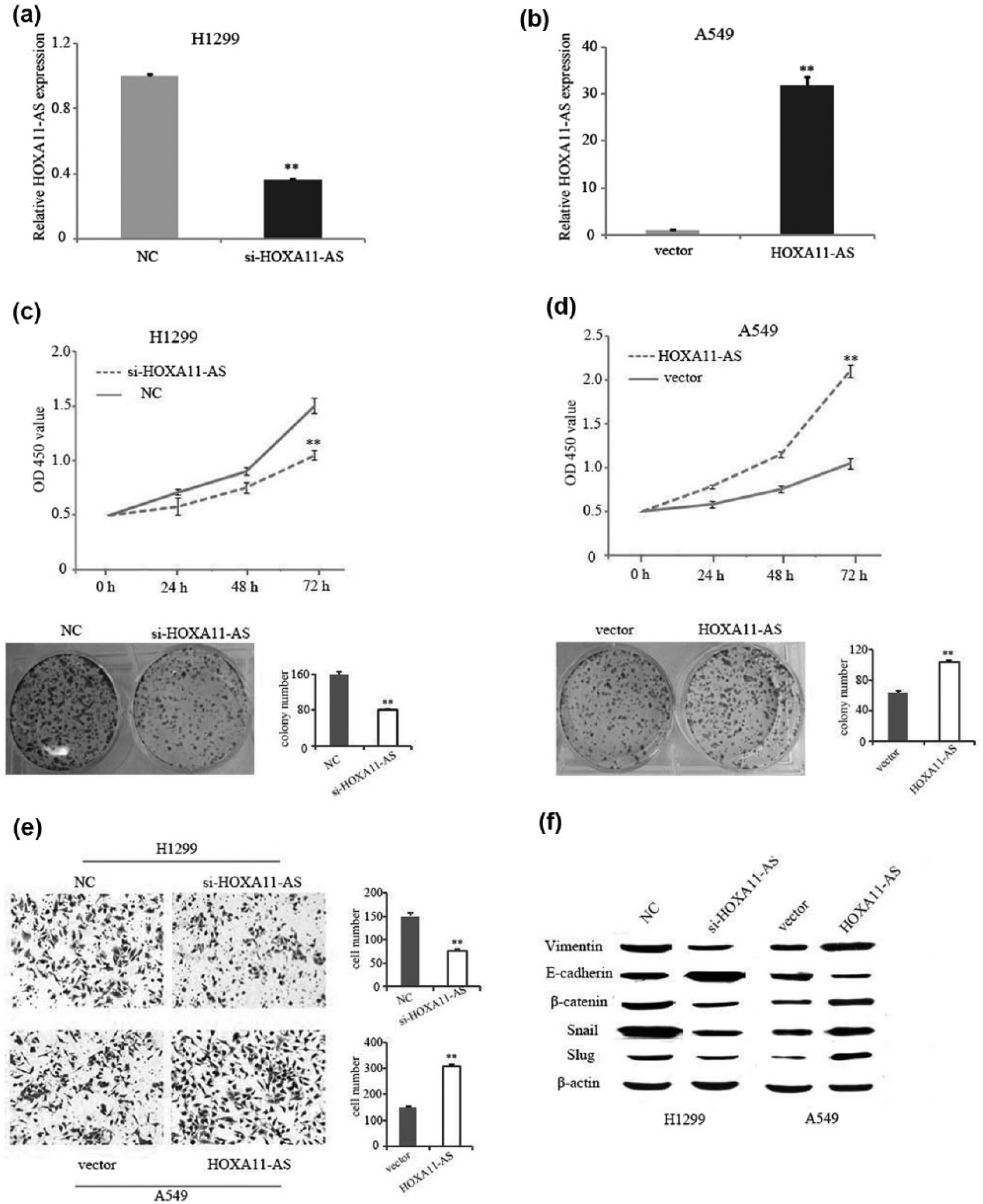
The effects of HOXA11-AS on NSCLC cell proliferation, invasion, and EMT. qRT-PCR analyses of HOXA11-AS expression level following (a) treatment of H1299 cells with siRNAs targeting HOXA11-AS and (b) treatment of A549 cells with pcDNA/HOXA11-AS vector. CCK-8 and colony formation assays were used to detect the proliferation of (c) si-HOXA11-AS-transfected H1299 cells or (d) HOXA11-AS vector–transfected A549 cells. (e) Invasion assay was used to measure invasion ability. (f) Western blotting was used to detect expression of EMT markers; β-actin was used as an internal control (**p < 0.01).
HOXA11-AS functions as competing endogenous RNA and sponges miR-124 in NSCLC cells
Bioinformatics analysis of miRNA recognition sequences on HOXA11-AS revealed the presence of miR-124 binding site (Figure 3(a)). Transfection with HOXA11-AS siRNA markedly increased the expression level of miR-124 in H1299 cells, while ectopic expression of HOXA11-AS decreased miR-124 expression in A549 cells (Figure 3(b)). Moreover, we observed that miR-124 expression was negatively correlated to HOXA11-AS expression in NSCLC cancer tissues (Figure 3(c)). Dual-luciferase reporter assays showed that miR-124 reduced luciferase activity in the HOXA11-AS wild-type reporter gene but not in the mutant type (Figure 3(d)). Furthermore, RIP experiments showed that HOXA11-AS and miR-124 were preferentially enriched in Ago2-containing micro-ribonucleoproteins (miRNPs) relative to control IgG (Figure 3(e)).

HOXA11-AS acts as a ceRNA for miR-124. (a) The HOXA11-AS cDNA containing the putative miR-124 recognition sites was cloned downstream of the luciferase gene. (b) The ectopic expression of HOXA11-AS decreased miR-124 expression and of si-HOXA11-AS increased miR-124 expression. (c) Expression levels of HOXA11-AS were negatively correlated with miR-124 among NSCLC samples (n = 30) as indicated by two-tailed Pearson’s correlation analysis. (d) Luciferase reporter assays were used to determine the interacting activity between miR-124 and HOXA11-AS. (e) RNA levels in immunoprecipitates are presented as fold enrichment in Ago2 relative to IgG immunoprecipitates. Human FOS was used as a positive control (**p < 0.01).
MiR-124 reverses the promoting effect of HOXA11-AS on proliferation and invasion
To dissect the importance of miR-124 binding in HOXA11-AS-promoting NSCLC progression, we ectopically expressed miR-124 in stable HOXA11-AS overexpressing A549 and H460 cells, and by using CCK-8 assays, we found that the overexpression of miR-124 attenuated the promoting proliferation effect of HOXA11-AS (Figure 4(a)). Moreover, the effect of HOXA11-AS on invasion was also reversed by miR-124 (Figure 4(b)). These results showed that HOXA11-AS promotes tumor cell proliferation and invasion in part via competitively binding with miR-124.

MiR-124 reverses the promoting effect of HOXA11-AS on NSCLC cell proliferation and invasion. (a) The introduction of miR-124 mimics attenuated the promoting proliferation effect of HOXA11-AS in A549 and H460 cells. Cell number was determined by the CCK-8 assay. (b) The introduction of miR-124 mimics attenuated the promoting invasion effect of HOXA11-AS in A549 and H460 cells (*p < 0.05).
HOXA11-AS positively regulates Sp1 expression
A recent study has reported that miR-124 inhibits invasion of hepatocellular carcinoma cells by targeting transcriptional factor Sp1. 14 Here, we observed that ectopic expression of HOXA11-AS upregulated Sp1 expression in A549 cells, while si-HOXA11-AS decreased Sp1 at the transcript and protein levels in H1299 cells (Figure 5(a)). Furthermore, the expression of Sp1 in 30 paired samples of NSCLC and corresponding non-carcinoma tissues was determined by qRT-PCR. The relative levels of Sp1 in NSCLC tissues were significantly higher than those in the corresponding normal tissues (Figure 5(b)). More importantly, the expression level of Sp1 was inversely correlated with miR-124 (Figure 5(c)). On the contrary, Sp1 and HOXA11-AS levels were positively correlated (Figure 5(d)).

HOXA11-AS positively regulates the miR-124 target gene Sp1. (a) The ectopic expression of HOXA11-AS upregulated Sp1 expression at both transcript and protein levels in A549 cells; si-HOXA11-AS decreased Sp1 at both transcript and protein levels in H1299. (b) Sp1 relative expression levels were determined in 30 paired NSCLC and their corresponding normal samples. Expression levels of Sp1 were (c) negatively correlated with miR-124 and (d) positively correlated with HOXA11-AS among NSCLC samples as indicated by two-tailed Pearson’s correlation analysis (*p < 0.05 and **p < 0.01).
Discussion
Recent research has revealed that lncRNAs play pivotal roles in tumorigenesis of cancers including colorectal cancer (CRC), gastric cancer, and lung cancer. 15 However, the knowledge about the function of lncRNAs in tumorigenesis of NSCLC is far from defined. In this study, we demonstrated that HOXA11-AS was upregulated in the NSCLC tissues and cells. Increased HOXA11-AS expression is associated with tumor size and lymph node metastasis. In addition, HOXA11-AS knockdown markedly suppressed NSCLC cell proliferation, colony formation, and invasion. These data indicated that HOXA11-AS may serve as an oncogene and play a crucial role in NSCLC development and progression.
HOXA11-AS is an lncRNA that is located in the 5′ region of the HOXA gene cluster. The HOXA genes not only determine the identity of body segments but are also reported to be abnormally expressed in cancers.16,17 The well-known lncRNA in HOXA clusters is HOTAIR, which plays important and various roles in cancer.18,19 A recent study has shown that HOXA11-AS was upregulated and closely correlated with tumor grade, poor prognosis, and molecular subtype in glioma. 20 Moreover, Sun et al. 21 reported that HOXA11-AS promotes proliferation and invasion of gastric cancer by scaffolding the chromatin modification factors PRC2, LSD1, and DNMT1. However, some studies indicated that HOXA11-AS also have tumor suppressor function in other cancers. For instance, Li et al. 22 reported that HOXA11-AS expression was decreased in the CRC tissues and cells and significantly associated with CRC patients poor prognosis. Besides, HOXA11-AS expression was significantly downregulated in human epithelial ovarian cancer (EOC), and an exonic variant within HOXA11-AS, rs17427875 (A > T), marginally inhibited the oncogenic phenotype of EOC. 23 These findings suggest that HOXA11-AS has tissue-specific expression pattern and could function as either oncogenes or tumor suppressors depending on the circumstance.
Increasing evidence has revealed that lncRNAs also contribute to tumorigenesis by silencing tumor suppressors or activating oncogenes through sponging miRNAs.6–8 Here, we demonstrated that HOXA11-AS directly binds with miR-124. MiR-124 was proved to be a tumor suppressor in a variety of cancers including NSCLC.14,24–26 Our results also showed that ectopic miR-124 expression could reverse the effects of HOXA11-AS on proliferation and invasion. In addition, HOXA11-AS modulated expression of endogenous miR-124 target Sp1. Sp1 is a well-known member of a family of transcription factors. Evidences suggest that the Sp family of proteins plays pivotal roles in cell proliferation and metastasis of various tumors.27–29 In patients with a variety of cancers, high levels of Sp1 protein are considered as a negative prognostic factor.30–32 In this study, we observed Sp1 to be positively correlated with HOXA11-AS expression.
Collectively, our results clearly identified a novel oncogenic role for HOXA11-AS in NSCLC. HOXA11-AS promotes NSCLC cell proliferation and invasion partially through regulating miR-124/Sp1 pathway. Our findings provide additional insight into molecular mechanisms of NSCLC carcinogenesis.
Footnotes
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by Zhejiang Provincial Natural Science Foundation of China (Grant No. LY14H160002) and Zhejiang Provincial Traditional Chinese Medicine Foundation of China (Grant No. 2017ZA128).
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
