Abstract
PURPOSE:
To explore the exact molecular mechanisms underline osteosarcoma (OS) patients with lung metastases.
METHODS:
The differentially expressed gene (DEG) as well as differentially expressed miRNAs (DEMs) for OS lung metastases were deeply investigated with two independent sources of databases (GEO dataset and clinical participants); The enriched biological processes and signaling pathways were explored; the miRNAs-mRNAs network was constructed; the functions of potential DEGs and DEMs were also verified with external analysis.
RESULTS:
The OS patients with lung metastases displayed 323 DEGs as C-C motif chemokine ligand 3 (CCL3), sorting nexin 10 (SNX10), alpha-2-macroglobulin (A2M), carboxypeptidase E (CPE), Rap guanine nucleotide exchange factor 4 (RAPGEF4), PDZ domain containing 2 (PDZD2), calpain 10 (CAPN10), four and a half LIM domains 2 (FHL2), alkaline phosphatase, biomineralization associated (ALPL), interleukin 6 (IL6), solute carrier family 26 member 1 (SLC26A1) as well as smoothened, frizzled class receptor (SMO) were significant differentially expressed. At the same time, 21 DEMs were potential for the progress of OS lung metastasis with hsa-miR-638, hsa-miR-451, hsa-miR-486-5p, hsa-miR-134 and hsa-miR-648 were significant distinct. It could been shown that hsa-miR-638 manipulated the largest number of target genes. The functions of hsa-miR-638 and target mRNAs for the development of lung metastasis in OS could be confirmed by quantitative Real-time PCR analysis.
CONCLUSION:
This integrated study hypothesized several miRNA dependent signaling pathway for OS patients with lung metastases and initiated a potential strategy for better understanding the lung metastases in clinic.
Introduction
Osteosarcoma (OS) is characterized as the most common primary bone cancer for children as well as adolescents [1]. At the same time, it is also the third most primary cancer in adults, just behind chondrosarcoma and chordoma [2]. The current reported overall incidence for OS is 3.4 per million per year worldwide [3]. The worldwide incidence of OS is not homogeneous based on the variable prevalence of the risk factors [4]. OS represents a primary malignant neoplasm of the skeleton, which is related with long bones of the human body [3]. Accumulating evidences have suggested that OS is indicated with a bimodal age distribution as the first peak generally at 15–19 years of age (8 cases/million/year) while the second peak normally at 75–79 years (6 cases/million/year) [4]. Clinically, the diagnosis of OS is relied on the a classical X-ray administration following a subsequent X-ray CT scan confirmation [5].
Various studies have focused on the genetic molecular mechanisms underline OS [6]. The tumor protein p53 (TP53) is the well-accepted one, which is also the most frequently altered in OS [7]. Other risk factors including MYC proto-oncogene, bHLH transcription factor (c-Myc) and retinoblastoma (RB) gene clusters as well as micro RNAs (miRNAs) such as miR-21, -34a, -143, -148a, -195a, -199a-3p and -382 have been also suggested for OS initiation [8]. Moreover, these factors have been demonstrated to interact with the pathogenic activity of MAPK and PI3K/Akt to some extent, supporting the connection between OS and these signaling pathway [9]. To date, a bench of chemical therapies for OS have been generated targeting the risk genes as well as potential molecular pathways [10]. Some of them have been evaluated in clinical trials such as bevacizumab, sorafenib, regorafenib etc [11].
Due to the early metastatic potential of OS, 20% of OS patients have already developed metastases at the time of diagnosis [12]. The 5-year survival rate of patients with distant metastases decreases to only 15–30% [13]. It is worthy noting that 90% of OS metastases are lung metastases. Even with decades of study, the knowledge of the genetic basis for OS metastases especially lung metastases is still inferior. It must be admitted that the mechanisms underline OS metastasis progression are complicate and also require a complex network for multiple genes and signaling pathways [14]. Currently, there are only few factors including TP53 and chemokine (C-X-C motif) receptor 4 (CXCR4)/chemokine (C-X-C motif) ligand 12 (CXCL12) pathway have been hypothesized as the key regulators for OS invasion and metastasis [15, 16]. At the same time, Yu and his colleagues claimed that Let-7a suppressed osteosarcoma cell growth and lung metastasis by targeting AURKB/NF-
Based on the truth of unclear mechanisms for OS lung metastases, the survival rate of OS patients is far from satisfaction. To address these issues, a systematically differential expression analysis was generated on OS lung metastasis specimens in this study. With online data source as well as self-generated clinical dataset, we comprehensively investigated the multiple miRNA dependent signaling pathways for OS lung metastasis patients, which provided a beneficial guidance for the future OS clinical study.
Material and methods
Data source
There were two data sources for this study. The first one was from GEO database, numbering GSE14359 (
The second data source was from clinical specimens. 10 conventional OS patients and 10 OS lung metastasis patients as well as 5 control participants without OS were recruited from the second hospital of Tianjin medical university. This study was given official approval by the institutional review boards of the hospital. The second data source was utilized for miRNA differentially expression analysis using miRNA-seq method (since there is no available online miRNA data source for OS lung metastasis tissue), quantitative Real-time PCR analysis.
miRNA expression analysis
Areas of interest tissues for each individual were dissected and harvested for total RNA isolation. As previously reported [19], total RNA was digested overnight with RecoverAll Total Nucleic Acid Isolation digestion buffer (Life Technologies, USA) and proteinase K. Next day, the RNAs were isolated using Qiagen FFPE miRNeasy kit (Qiagen, USA) and the concentrations were measured using a NanoDrop 2000 spectrophotometer.
The miRNA expression analysis was performed by Geneseed Co (Guangdong, China). The NanoString human v2 array was applied, containing 800 miRNA probes. A total of 100 ng RNA input was used per sample and conditions were set up based on the manufacturer’s recommended protocol. The miRNAs were quantified using the nCounter Digital Analyzer, normalizing using the geometric mean. The filtered geometric mean normalized data was then log2 transformed for further analysis.
Differential gene analysis
The limma package in R language (version:3.5.2) was developed to analyze the differentially expressed mRNAs as well miRNAs between different groups, taking the absolute value of the log-transformed differential expression multiple (Log2FC)
Functional enrichment analysis
For the obtained differentially expressed genes, we used the “clusterProfiler” function package in R language (version:3.5.2) for enrichment analysis of GO (including Biological Process, Molecular Function and Cellular Component) and KEGG Pathway. When
Prediction of miRNA target genes
The target genes of miRNAs were predicted through the miRDB (
Quantitative real-time PCR
The total RNA of different groups of patients was extracted using RNAiso Plus (Takara, Beijing, China). The real-time PCR was performed with BrightGreen 2
Resultant values of mRNAs were normalized against the internal control gene GAPDH for each replicate. For miRNA expression analysis, the relative expression levels of hsa-miR-638 were calculated by normalization with hsa-miR-638 expression in conventional OS patients tissues. All the experiments were performed in triplicate.
Results
Differentially expressed analysis of mRNA and miRNA for OS lung metastasis
Using the dataset of GSE14359, we did a differentially expressed gene (DEG) pairwise comparison for non-neoplastic primary osteoblast cells, conventional OS tissue as well as OS lung metastasis tissue. Compared with control (non-neoplastic primary osteoblast cells), conventional OS tissue displayed 2503 DEGs, while OS lung metastasis tissue demonstrated 2425 DEGs (Fig. 1A and B). The 2102 DEGs were in the intersection of conventional OS and OS lung metastasis tissue, which indicated 323 DEGs were primary for lung metastasis progression of OS patients (Fig. 1E). In the case of
The analysis of DEGs and DEMs for OS lung metastasis. (A) The volcano map of DEGs between two groups of OS patients. The horizontal axis represents the multiple of differential expression (Log2FC), the vertical axis represents -log10 (FDR). The red dots indicate up-regulated genes, and the green dots indicate down-regulated genes respectively. (B) The heat map of DEGs. The horizontal axis represents sample, vertical axis represents different gene. (C) The volcano map of DEMs between two groups of OS patients. (D) The heat map of DEMs. (E) Venn diagram of DEGs. (F) Venn diagram of DEMs.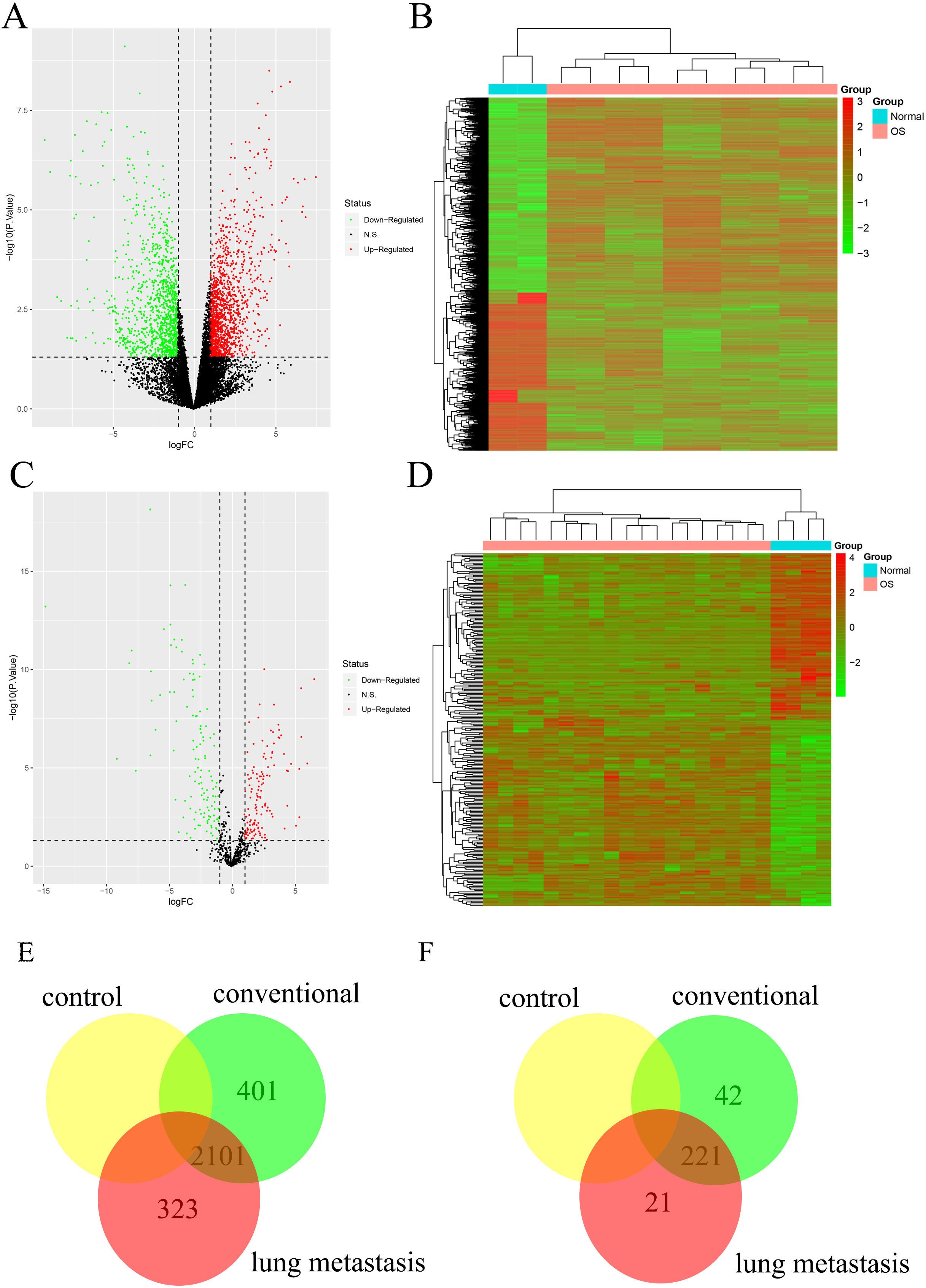
At the same time, based on the miRNA-seq results of clinical specimens, conventional OS tissue showed 263 differentially expressed miRNAs (DEMs), while OS lung metastasis tissue presented 242 DEMs (Fig. 1C and D). Taken a intersection between conventional OS and OS lung metastasis tissue (221 DEMs), 21 DEMs were potential for the progress of OS lung metastasis (Fig. 1F). With further investigation, hsa-miR-638, hsa-miR-451, hsa-miR-486-5p, hsa-miR-134 and hsa-miR-648 were significant distinct with top absolute value of Log2FC (as increased expression for hsa-miR-451 as well as hsa-miR-134 and decreased expression for hsa-miR-638, hsa-miR-486-5p as well as hsa-miR-648).
Based on the GO and KEGG enrichment analysis of the 323 DEGs and 21 DEMs, it could be demonstrated that the differentially expressed genes were enriched in GO terms including immunoglobulin complex etc. (Fig. 2A). At the same time, the signaling pathways related to the spliceosome as well as RNA transport were significantly enriched in KEGG analysis (Fig. 2B).
GO and KEGG enrichment results for OS lung metastasis. (A) The top GO term enrichment results with the largest number of genes. In the figure, the horizontal axis represents the number of enriched genes, and the vertical axis represents the name of each GO term respectively. (B) The enrichment results of the KEGG pathways with the largest number of genes. The horizontal axis in the figure indicates the number of genes enriched, and the vertical axis indicates the name of each KEGG pathway respectively. 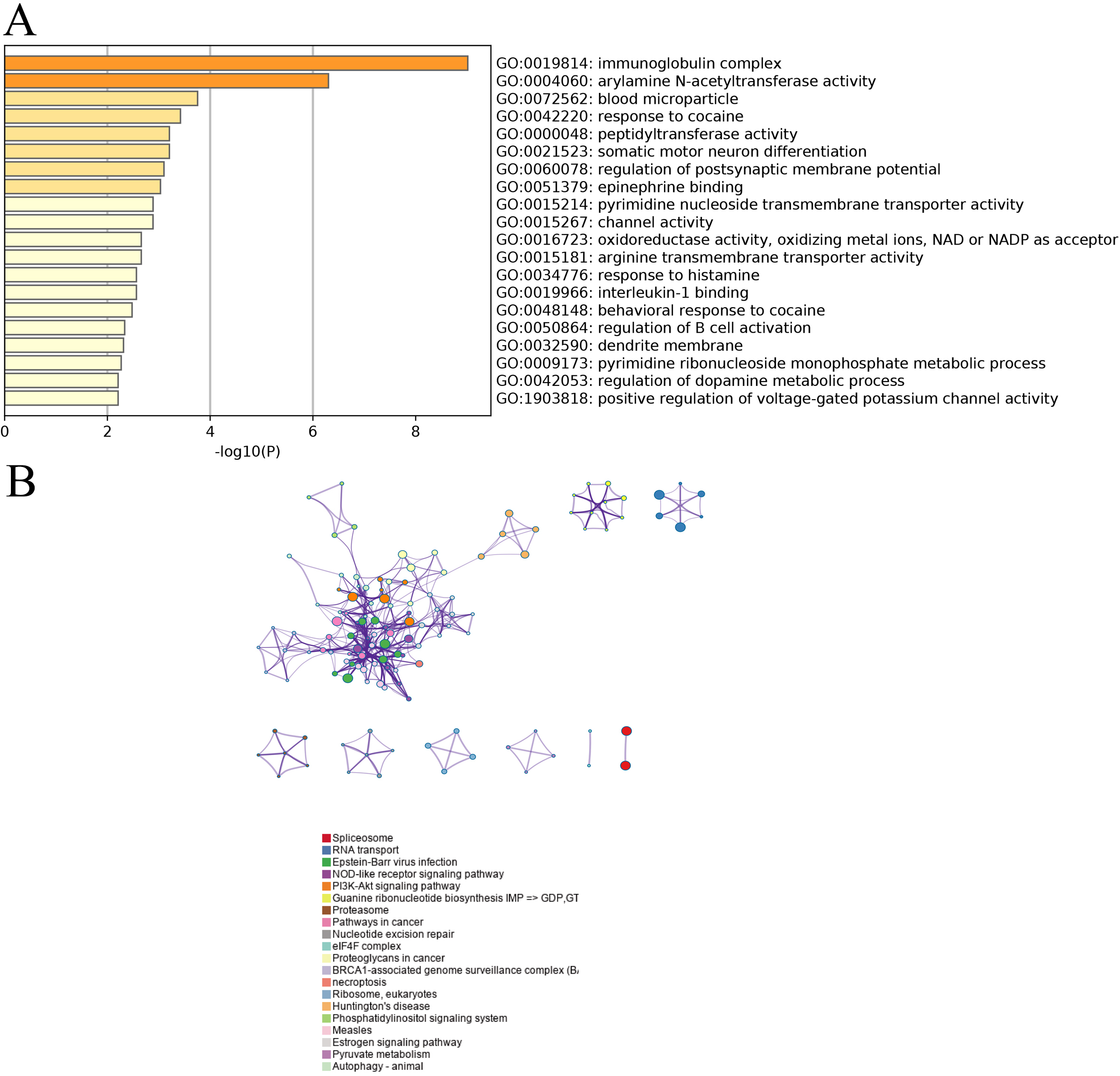
The 21 DEMs and 323 DEGs were further developed for a regulatory network which was visualized with Cytoscape software. The number of interacted target genes greater than 5 was further screened out, which are 5 of them (Fig. 3). It could been shown that hsa-miR-638 manipulated the largest number of target genes [12]. At the same time, the number of target genes regulated by hsa-miR-451, hsa-miR-486-5p, hsa-miR-134 and hsa-miR-648 were 8, 6, 6 and 5 respectively (Fig. 3). Base on these facts, the 5 DEMs (hsa-miR-638, hsa-miR-451, hsa-miR-486-5p, hsa-miR-134 and hsa-miR-648) were suggested as the key factors for the progression of lung metastasis in OS patients.
miRNA-mRNA regulatory network. The orange node is miRNA and the blue node is mRNA. The line type represents the interaction between miRNA and mRNA. The solid line represents positive regulation, and the dotted line represents negative regulation. 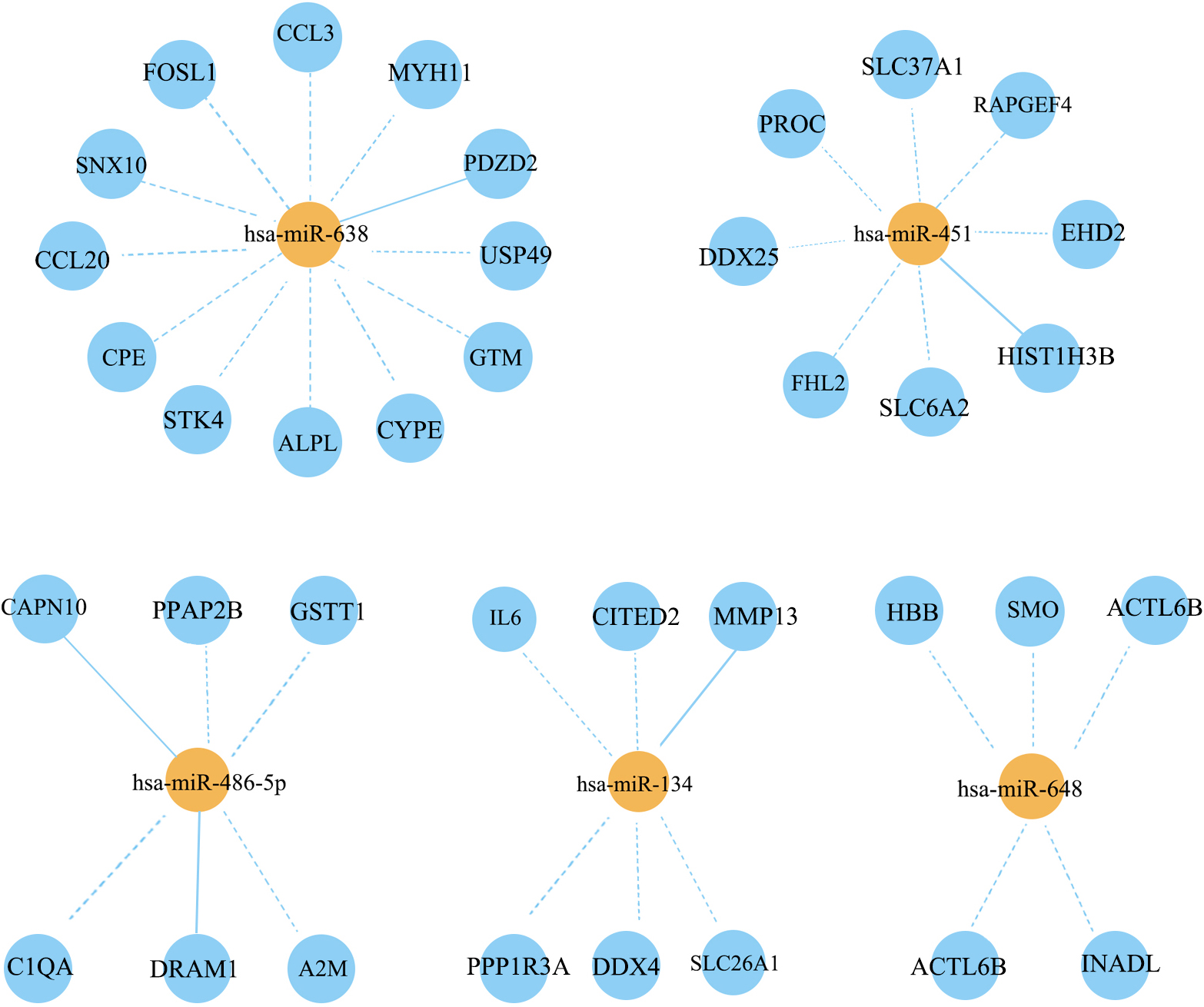
Next, we sought to verify the functions of potential DEGs and DEMs for OS lung metastasis. The hsa-miR-638 was considered as the primary DEM since it modulated the largest numbers of DEGs. At the same time, CCL3, SNX10, CPE, ALPL and PDZD2 were selected since they all significant differentially expressed DEGs (Fig. 1) and the targets of hsa-miR-638 (Fig. 3). Compared with conventional OS tissues, the expression levels of hsa-miR-638 and PDZD2 declined significantly while CCL3, SNX10, CPE and ALPL kept an upward tendency in the OS tissues with lung metastasis. These outcomes were consistent with the differentially expression analysis results.
Discussion
Metastasis is a term referring to the spread of tumor cells from primary sites to surrounding structures and distant sites [21]. This process requires a complicate procedure which includes original detachment from the primary tumor sites, subsequent invasion into vessels as well as extravasation into the appropriate secondary site plus the development of surrounding microenvironment establishment. Nowadays, tumor metastasis has been believed as the striking cause for morbidity and mortality. Metastasis has been considered as a tremendous factor for survival rate of tumor patients. For instance, in breast cancer, the overall 5-year survival drops significantly from 96% to 21% with distant metastasis. At the same time, patients of colorectal cancer with lung or liver metastasis have a 5-year survival of less than 10% compared with 91% with conventional colorectal cancer [22]. Based on the facts that patients with metastasis encounter distinct clinical prognosis and treatment options compared as others with primary tumor of origin, almost 1500 people die per day from metastatic tumors worldwide [23]. This highlights the major limitations of the modern-day treatment options once the disease is widespread.
RT-PCR analysis of selected mRNAs and miRNAs functions for OS lung metastasis. (A-F) The quantitative Real-time PCR analysis of hsa-miR-638, CCL3, SNX10, CPE, ALPL as well as PDZD2 between conventional OS and OS with lung metastasis, respectively. * As a indication of 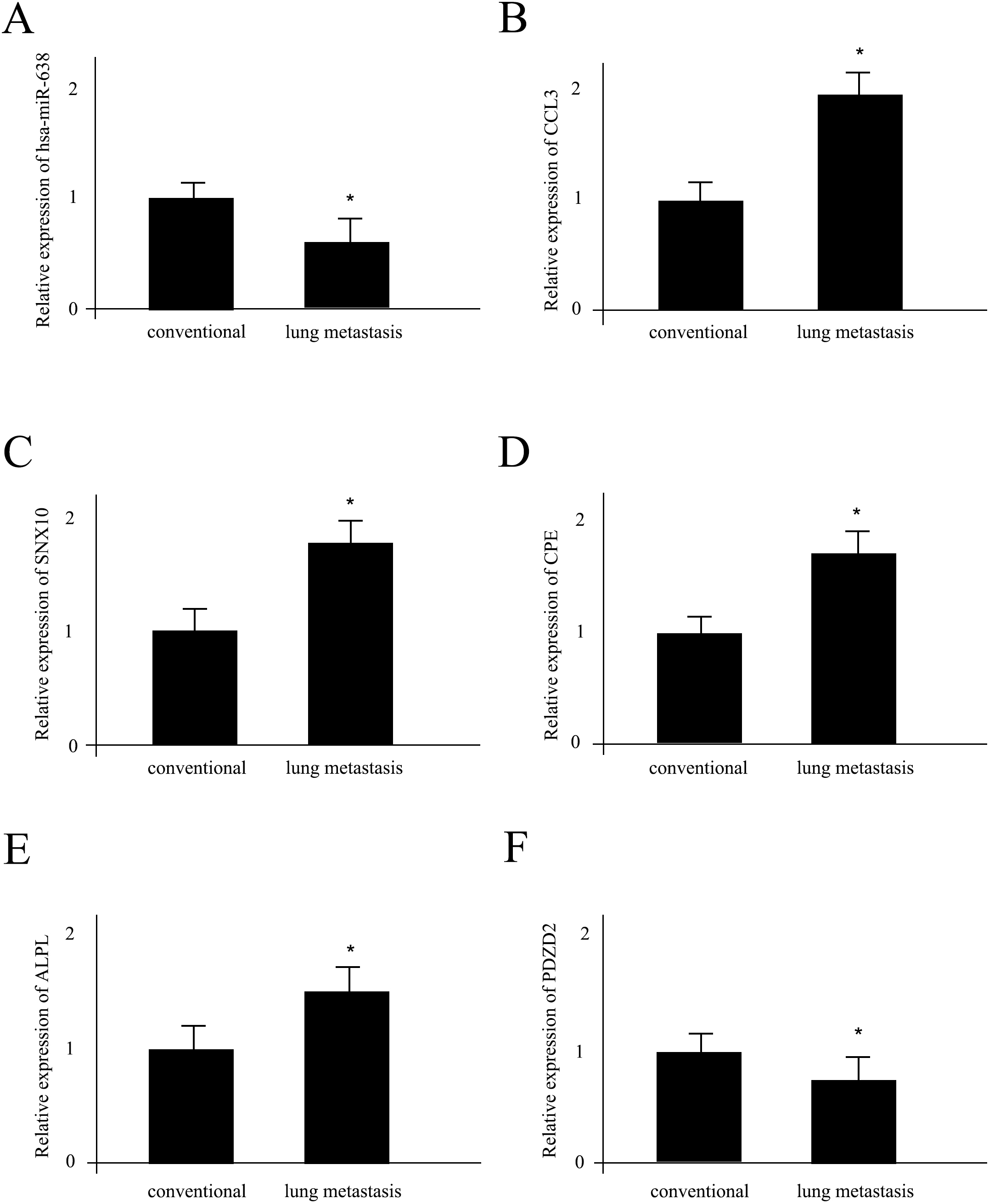
As far as tumor metastasis, the lung is the second most common site of metastatic focus [24], 20 to 54% of which is the primary region for malignant tumors developing elsewhere [25, 26]. As estimated, various cancer types beside OS such as colorectal cancer, head/neck squamous cell carcinomas, renal cancer and so on metastasize to lung parenchyma [27]. However, the molecular mechanism is still poorly understood. Actually, it has been suggested that small RNAs have a close relationship with OS initiation as well as development. For instance, overexpression of miR-421 has been shown to result in proliferation, migration and invasion of tumor cells [28]. The expression level of miR-421 is significantly higher in patients with OS than in healthy participants. Other miRNAs such as miR-21, -382, -143, -195a, -148a and -34a also play a critical role in OS formation through regulation of MAPK and PI3K/Akt signaling pathways [29]. As far as OS lung metastasis, a recent study by Wang and his colleagues reported that the dysregulated miR-491 played an important role in OS lung metastasis and chemoresistance [30]. This finding was based on a comparison of serum between OS patients and healthy control subjects. In our study, we also demonstrated 5 primary DEMs (hsa-miR-638, hsa-miR-451, hsa-miR-486-5p, hsa-miR-134 and hsa-miR-648) for the progression of lung metastasis in OS patients. The roles of hsa-miR-638 in tumor metastasis have been often mention. In one study, miR-638 was shown as a tumor suppressor gene in gastric cancer [31]. The miR-638 was down-regulated in human GC tissues and its expression level was negatively correlated to TNM stage and lymph metastasis. In another study, the author claimed that miR-638 was one of the most significantly overexpressed miRNAs in metastatic lesions of melanomas compared with primary melanomas via enhancing the tumorigenic properties of melanoma cells in vitro and lung colonization in vivo [32]. miR-451 has been suggested to indirectly affect tumor cell invasion and metastasis upon secretion into the tumor microenvironment via exosomes [33]. At the same time, using lung cancer cell line A549 as a model, the miR-451 may suppress the development and migration of lung cancerre through PSMB8/NOS2 inflammatory factors related signaling pathways [34]. The functions of miR-486-5p have been shown to be highly correlated with exosome activity. Moreover, the exosome-derived miR-486-5p could manipulate cell cycle, proliferation as well as metastasis in lung adenocarcinoma, which targeting NIMA related kinase 2 (NEK2) dependent signaling pathway [35]. The functions of miR-134 for tumor metastasis seems to be dynamic. On one hand, miR-134 inhibition or DAB2 restoration may be a novel strategy for inhibiting lung adenocarcinoma metastasis and overcoming lung adenocarcinoma cell resistance to chemotherapy [36]. On the other hand, miR-134 has been shown to suppress angiogenesis and the progression of OS [37].
Except for the DEMs, we also implicated several DEGs for OS lung metastasis patients including CCL3 and SNX10 (shown as Figs 1 and 3). CCL3 represents the chemokine macrophage inflammatory protein-1 alpha (MIP-1alpha, CCL3), which is a special member of the chemokine family. During the establishment of tumor microenvironment for tumor metastasis, CCL3 plays a key role in immune cell trafficking regulation and immune landscape formulation. CCL3 has been approved to function in both immune surveillance and tolerance, emerged as a prognostic biomarker in both solid and hematological malignancies [38]. SNX10 refers to sorting nexin 10 in human, which has been reported as a inhibitor for colorectal cancer initiation and progression by controlling autophagic degradation of non-receptor tyrosine kinase SRC [39]. Except for these findings, the correlations between the progress of OS lung metastasis and other risk factors have not been fully understood yet, which raise a interesting direction for the future study.
Certain limitations should be acknowledged in this study. For instance, one significant aspect was that a large number of potential mRNAs and miRNAs have been suggested in this study, the central ones remained unclear which clearly required further comparison between them. Second, even we performed a in-depth analysis for the hub gene interactions, the internal molecular mechanisms behind these genes have not been fully explored yet, which required a in vivo study to verify. Third, the external verification of hub genes functions was collected from only one hospital center. The sample size was limited. In order to let a carry on the further promotion and the application in the clinical, more participates including different gender and personality traits should be recruited in the study. To address these issues, we would take further steps and more efforts to consider the aspects mentioned above.
Conclusion
All in all, based on the facts of unclear molecular mechanisms behind lung metastasis for OS patients, we systematically generated a differential expression profile for mRNAs and miRNAs from two sources of dataset. Multiple miRNAs dependent signaling pathways had been explored and verified, which offered a beneficial reference for future OS as well as other tumor metastasis studies.
Footnotes
Conflict of interest
The authors declare no competing interests.
Author contributions
Fengsong Liu: Interpretation of data.
Xiaojian Pang and Ziqi Yu: Preparation of the manuscript.
Kai Wang: Supervision.
