Abstract
BACKGROUND:
Acute myeloid leukemia (AML) is one of the most malignant hematopoietic system diseases. Interferon stimulated exonuclease gene 20 (ISG20) is a protein induced by interferons or double-stranded RNA, which is associated with poor prognosis in several malignant tumors. However its expression in AML is unknown.
OBJECTIVE:
To explore the expression of ISG20 in AML and its prognostic significance.
METHODS:
The expression of ISG20 in AML patients was analyzed by GEPIA database, detected by qRT-PCR and their prognosis was followed-up. Chi-square test was used to identify the association between ISG20 expression and clinical characteristics of the patients. Kaplan-Meier analysis was performed to draw survival curves and Cox regression analysis to confirm the independent prognostic factors of AML patients.
RESULTS:
Kaplan-Meier analysis revealed that whether to receive treatment, karyotype, and ISG20 expression were related to overall survival time of AML patients (
CONCLUSION:
The high expression of ISG20 acts as a poor prognosis indicator in AML patients.
Introduction
Acute myeloid leukemia (AML), one of the most malignant hematopoietic system diseases, accounts for 80% of acute leukemia in adults. AML is characterized by unlimited clonal proliferation and accumulation of myeloid progenitors [1, 2]. With the development of medical technology, the remission rate of initial chemotherapy in patients with AML has increased. However, 20–70% of patients relapse after achieving complete remission (CR), resulting in a poor prognosis of AML patients with the five-year survival rate less than 30% [3]. Emerging evidence has suggested that biomarkers related to disease progression and prognosis would contribute to individualize therapy and improve prevention of aggressive disease or treatment outcomes in AML patients [4].
Genetic abnormality is one of the most powerful prognostic factors and the discovery of specific molecular aberrations is critical to predict the clinical outcome of the AML patients [5]. Interferon stimulated exonuclease gene 20 (ISG20) is a protein induced by interferons or double-stranded RNA [6, 7]. Level of ISG20 expression differ in some diseases (such as rheumatoid arthritis) in comparison to those in healthy individuals [8]. Furthermore, previous studies indicated that ISG20 played a vital role in tumorigenesis and progression of neoplasms [9]. It has been reported that ISG20 might serve as a potential biomarker in clear cell renal cell carcinoma [9] and contributes to poor survival in human glioma [10]. To the best of our knowledge, the function of ISG20 has not been reported in AML. In this study, we focused on the expression and prognostic significance of ISG20 via bioinformatics analysis and detection of expression in clinical samples.
Methods
Data source
GEPIA, a web-based tool to deliver fast and customizable functionalities based on The Cancer Genome Atlas (TCGA) and Genotype-Tissue Expression (GTEx) project provide key interactive functions including differential expression analysis, profiling plotting, correlation analysis, and patient survival analysis [11]. UALCAN (
Patients
We collected 27 Bone marrow blood from newly diagnosed AML patients in the Affiliated Hospital of Southwest Medical University between December 2019 and September 2020. The research was approved by the Ethics Committee of our institution and was in agreement with the Declaration of Helsinki. The diagnosis was according to the 2008 WHO criteria [13]. The exclusion criteria for selected AML patients are as follows: (1) with other malignant tumors, (2) with severe hepatic or renal insufficiency, (3) Patients who have received hematopoietic stem cell transplantation. DA (daunorubicin 45 mg/m
Quantitative real-time PCR analysis (qRT-PCR)
Fresh bone marrow samples were collected. Total RNA was extracted from fresh bone marrow samples using the TRIzol Reagent (TIANGEN) according to the manufacturers’ protocol. RNA was Reverse-transcribed using a Rever Tra Ace qPCR RT Kit (Toyobo). The concentration and quality of RNA samples were examined by Nanodrop (Thermo Fisher). RT-qPCR was conducted using a Light Cycler 96 (Roche) with the following amplification protocol: denaturation at 95
Statistical analysis
SPSS 25.0 software was used for statistical analyses. AML patients were divided into high expression group and low expression group according to the median of ISG20 expression level. The relationship between ISG20 expression and clinicopathologic features was calculated by Chi-square. Kaplan-Meier method was used to draw survival curves while group comparison was performed by long-rank test. The Cox-regression analyses were used to confirm independent prognostic factors of AML patients. Meanwhile, hazard ratio (HR) and their corresponding 95% confidence interval (CI) were calculated. The probability value of less than 0.05 indicated statistical significance.
The expression levels of ISG20 based on common data base. A): The expression levels of ISG20 in AML and Normal based on GEPIA dataset. B): The expression levels of ISG20 in AML based on FAB classification (UALCAN).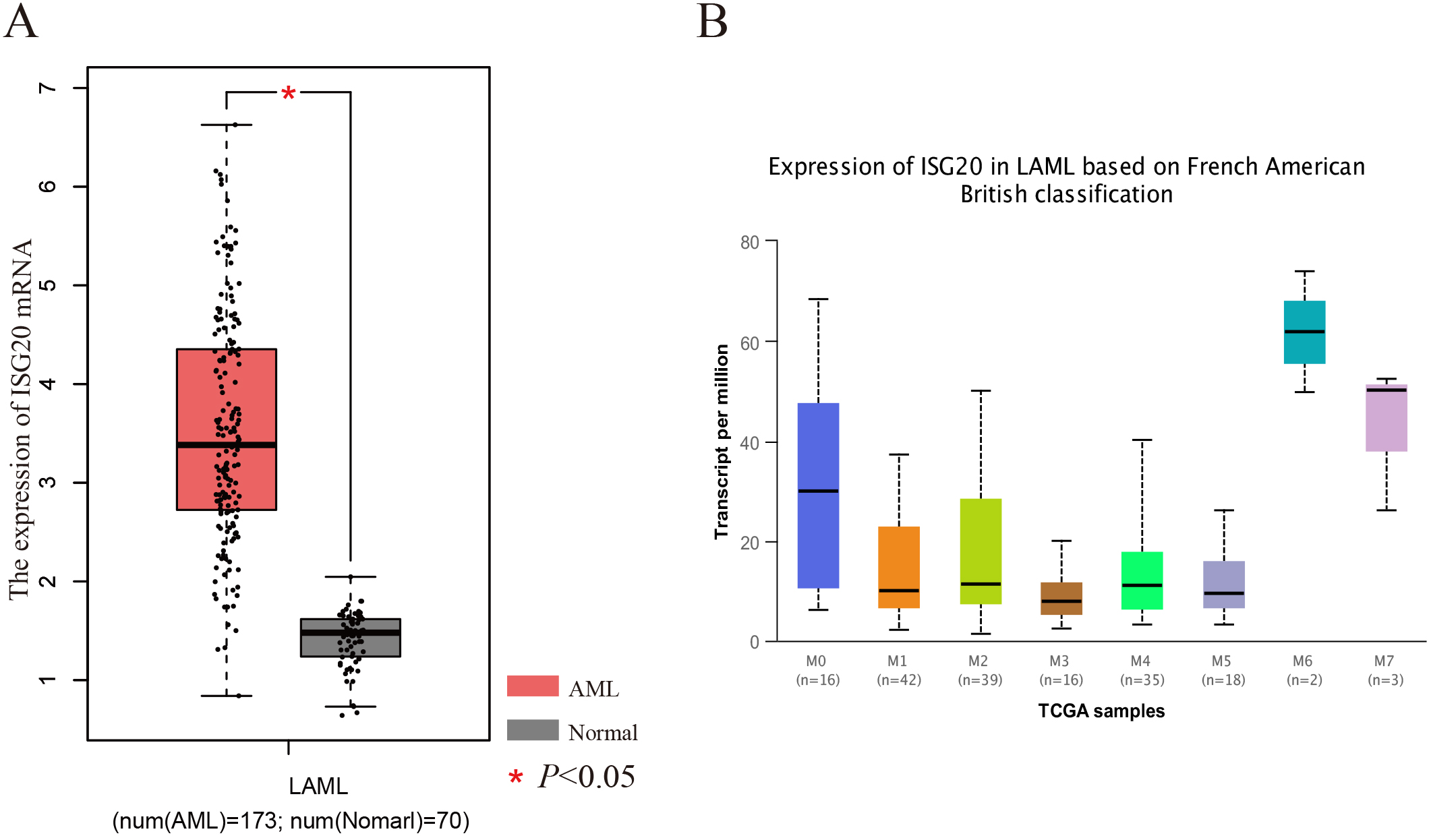
The effect of ISG20 expression level on survival outcomes of patients was analyzed based on datasets. A) The relationship between ISG20 expression and prognosis in patients with AML based on UALCAN datasets. B) The relationship between ISG20 expression and prognosis in patients with AML based on GEPIA datasets, The dotted line represents a 95% confidence interval.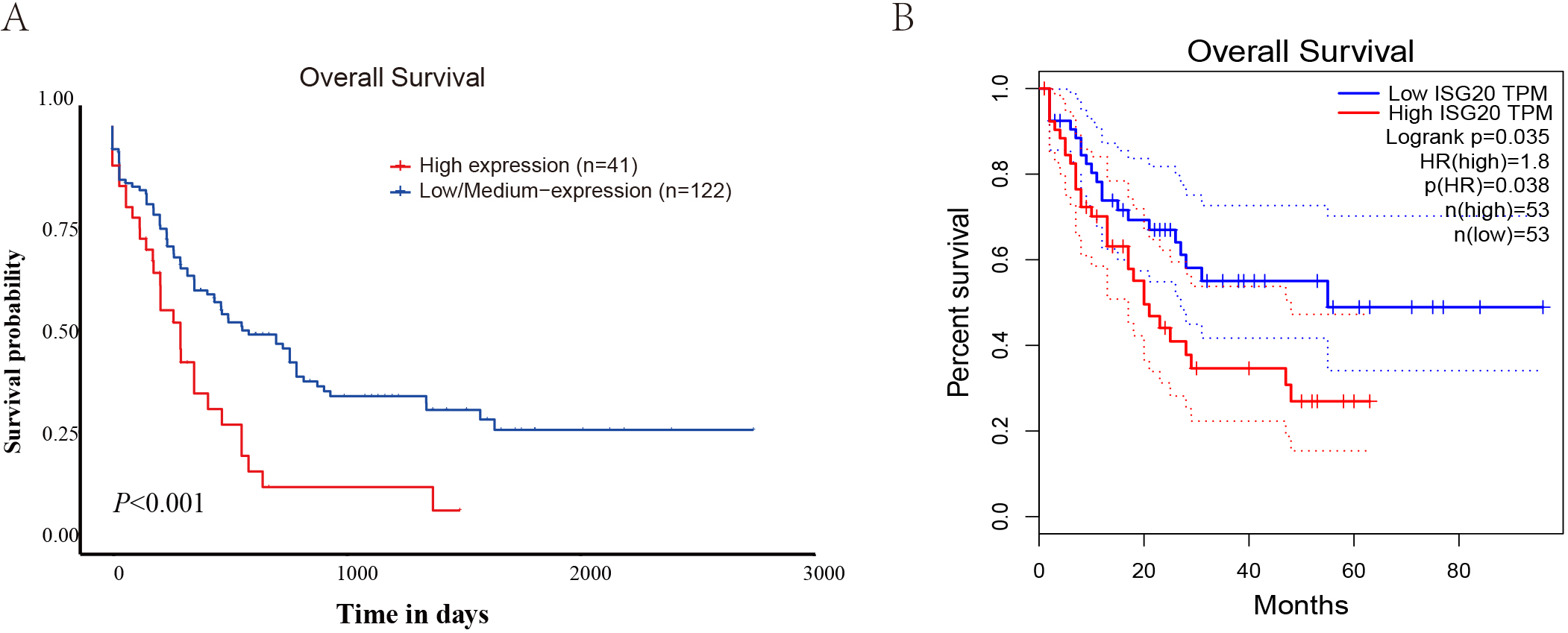
The expression of ISG20 was upregulated in AML patients
To clarify the expression of ISG20 genes in AML and normal tissues, we performed the differential expression analysis by using GEPIA tools. From the box-profile in Fig. 1A, the results indicated that ISG20 was higher expressed in AML than normal tissues (
The expression level of ISG-20 and its relationship with clinicopathological characteristics. A) AML patients were divided into high expression group and low expression group according to the medium risk expression value of ISG20. B–H) The correlation between ISG20 expression level and the clinicopathological characteristics of chromosome, FLT3-ITD, NPM1, CEBPA, DNMT3A, IDH1/2 and ASXL1.ns: 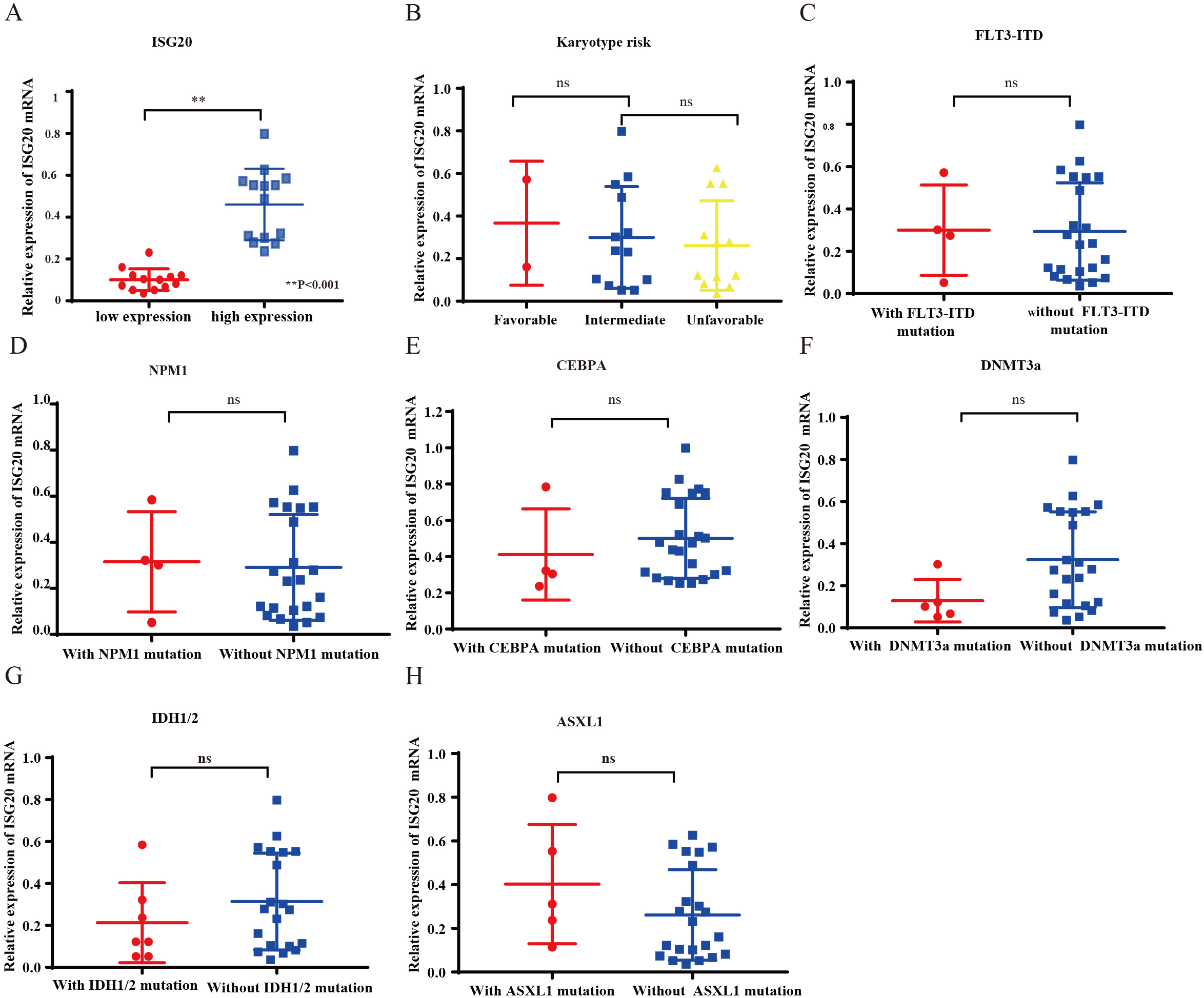
In order to verify the data results in GEPIA, We collected bone marrow specimens from 27 newly diagnosed AML patients and determined the mRNA level of ISG20 by using qRT-PCR. To further evaluate the clinical relevance of ISG20, 27 AML patients were stratified into ISG20-low (
The correlation between ISG20 expression and clinicopathologic characteristics in AML
The correlation between ISG20 expression and clinicopathologic characteristics in AML
The Kaplan-Meier was performed to evaluate prognostic factors including age, karyotype, gene mutation (FLT3-ITD, NPM1, ASXL1, CEBPA, and DNMT3a), whether to receive treatment and ISG20 expression in AML patients. Patients with high ISG20 transcript level of had a poor prognosis as compared to those with low ISG20 transcript level (
The Kaplan-Meier analyses of the overall survival of 27 AML patients
The Kaplan-Meier analyses of the overall survival of 27 AML patients
Note:
Cox regression analysis of multivariate analysis with AML patients
Note:
The Kaplan-Meier survival analysis was used to analyze the prognostic factors of 27 AML patients. A) The relationship of ISG20 expression and overall survival times in 27 AML patients. B) The relationship of treatment condition and overall survival times in 27 AML patients.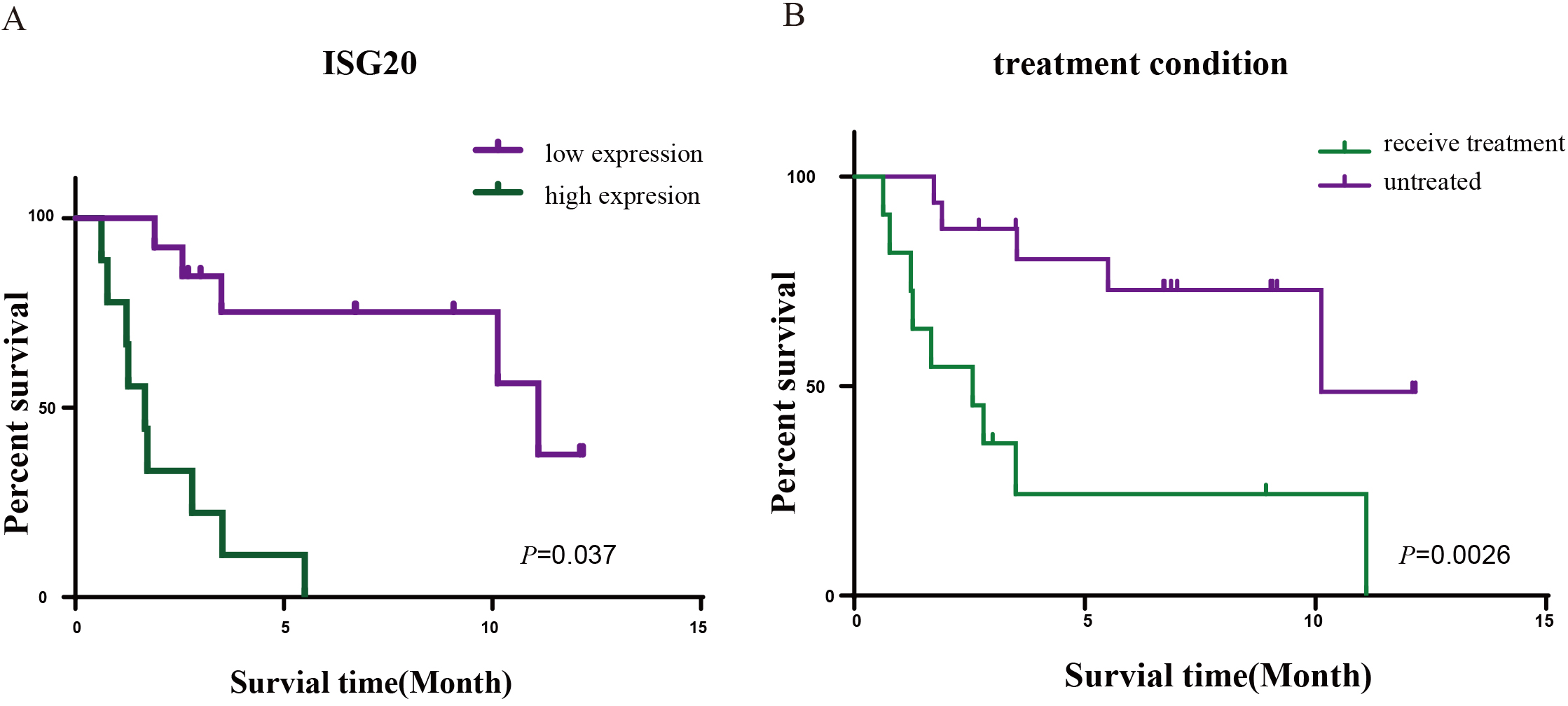
Acute myeloid leukemia (AML) is a clinically and biologically heterogeneous disease associated with genetic and epigenetic alterations which result in blocked dedifferentiation and uncontrolled proliferation of immature hematopoietic precursor cells in the bone marrow [2]. AML is a highly heterogeneous disease at the molecular level. There is a difference in terms of survival time among the patients with same grade AML. Despite substantial advances in recognizing the molecular heterogeneity of AML including hundreds chromosomal translocations and gene mutations, there remains a need for identifying newer biomarkers that would improve the early diagnosis and the prognostic evaluation of AML patients [16, 17, 18].
ISG20 is localized to human chromosome 15q26 [19]. It was first introduced in 1997 by Celine Gongora and colleagues as an IFN-induced promyelocytic leukemia nuclear body (PMLNB)-associated protein [7]. PML-NB is reported to be extensively involved in oncogenesis and gene transcription [20, 21] and the function differs according to its binding protein [22, 23]. The endogenous ISG20 protein is present both in the nucleolus and in the Cajal bodies, and participates in the maturation of small nucleolar RNAs and ribosomal RNAs. ISG20 is also an exonuclease preferentially degrades RNA at a rate 35-fold higher than it degrades sing lestranded DNA [24, 25]. ISG20 can be induced by type I and type II IFNs, [26, 27] and its expression has been shown to be elevated during infection [28] and in several types of cancers [29, 30]. However, the expression of ISG20 in AML remain elusive.
In the present study, we investigated the prognostic significance of ISG20 in AML, which has been rarely described and reported previously. GEPIA database analysis showed that the transcriptional level of ISG20 in patients with AML was higher than that in normal subjects, and the prognosis of patients with high transcriptional level of ISG20 was poor. We collected bone marrow samples from newly diagnosed AML patients for qRT-PCR detection and reached the same conclusion. Our analysis demonstrated that high ISG20 expression level is significantly associated with worse OS of AML, based on Kaplan-Meier curve and log rank test. Therefore, ISG20 may serve as a prognostic marker for AML patients.
The important role of ISG20 in other tumors has been reported [15]. Rajkumar [30] detected that ISG20 is up-regulated in cervical cancer. It was also reported that ISG20 expression level associated with various clinicopathological parameters in clear cell renal cell carcinoma. In addition, high expression of ISG20 predicted poor overall survival in glioma [10]. Similarly, patients with high expression of ISG20 had shorter overall survival time in our present study. However, the specific mechanism by which ISG20 affects the prognosis of cancer patients is still being explored. Xu [9] found ISG20 facilitated proliferation and metastasis of clear cell renal cell carcinoma via mediating the expression of CCND1/MMP9. Meanwhile, Lin et al. [31]. also demonstrated that ISG20 enhanced angiogenesis and supported progression of hepatocellular carcinoma (HCC) regulated by thyroid hormone. In human glioma, ISG20 was positively correlated to immune checkpoints (PD-1 and PD-L1) and suppressed the adaptive immune response. In addition, high expression of ISG20 predicted poor overall survival in glioma. Similarly, patients with high expression of ISG20 had shorter overall survival time in our present study [10]. Combined with the immune checkpoint effect of PD-1 and the role of ISG20 in the immune system, we speculated that in acute myeloid leukemia, the poor prognosis caused by the high expression of ISG20 may be related to the immune infiltration of tumor cells and the immune escape of tumor, and the specific mechanism needs to be further explored in basic experiments.
In conclusion, our research shows the transcriptional level of ISG20 is different between normal and AML patients, meanwhile, the high expression levels of ISG20 at diagnosis predict poor outcomes in AML patients. However, our study was restricted by the amount of samples. In our current work, we are expanding the sample size and researching related signaling pathways of ISG20-high AML.
Footnotes
Acknowledgments
This study was supported by Sichuan department of science and technology (grant no. 20YYJC0940) and Lu zhou people’s government- Southwest Medical University joint project (grant no. 2017LZXNYD-Z06).
Author contributions
Conception: Chunlan Huang, Hao Xiong, Xinwen Zhang.
Interpretation or analysis of data: Hao Xiong, Xinwen Zhang, Xiaomin Chen, Yang Liu, Jialin Duan.
Preparation of the manuscript: Hao Xiong, Xinwen, Zhang.
Revision for important intellectual content: Hao Xiong, Xinwen Zhang, Xiaomin Chen, Yang Liu, Jialin Duan.
Supervision: Chunlan Huang.
