Abstract
Introduction
Nucleophosmin 1 (NPM1), FMS-like tyrosine kinase 3-internal tandem duplication (FLT3-ITD), and de novo methyl transferase 3 A (DNMT3A) triple-mutated acute myeloid leukemia (AML) represents a distinct entity with poor outcomes.
Methods
We explored the gene mutation spectrum and clinical characteristics of 165 AML patients retrospectively, particularly comparing patients with NPM1/FLT3-ITD/DNMT3A triple-mutations and those without.
Results
Our results demonstrated significantly elevated white blood cell counts (P < 0.001), bone marrow blast percentages (P = 0.037), and platelet counts (P = 0.007) in the triple-mutated cohort (6.7%) compared to the non-triple-mutated patients. Furthermore, all triple-mutated cases were classified as the M4/M5 subtype of the French-American-British classification (P = 0.017). Although no significant difference in complete remission rates was observed between the groups after initial treatment, the median overall survival for triple-mutated AML patients was only 4 months. Using the Gene Expression Omnibus (GEO) database and bioinformatics, we compared AMLNPM1mutFLT3-ITDmutDNMT3Amut and AMLNPM1mutFLT3-ITDmutDNMT3Awt. A total of 246 AML patients from the GEO dataset were included to evaluate the expression profiles of differentially expressed genes. The guanine nucleotide-binding protein subunit γ 4 (GNG4) was differentially expressed between AMLNPM1mutFLT3-ITDmutDNMT3Amut and AMLNPM1mutFLT3-ITDmutDNMT3Awt, which had the most adjacent nodes among hub genes. The prognostic value of GNG4 was further validated in AML patient samples through qRT-PCR.
Conclusion
Clinical validation indicated a substantial downregulation of GNG4 in AMLNPM1mutFLT3-ITDmutDNMT3Amut compared to AMLNPM1mutFLT3-ITDmutDNMT3Awt patients. Thus, GNG4 may play a role in the low survival rate of AMLNPM1mutFLT3-ITDmutDNMT3Amut patients, offering novel insights into the prognosis, therapeutic targets, and prognostic evaluation of AML.
Plain Language Summary
Acute myeloid leukemia (AML) is a type of blood cancer that can be influenced by various genetic mutations. In this study, we focused on a specific group of AML patients who have mutations in three genes: Nucleophosmin 1 (NPM1), FMS-like tyrosine kinase 3-internal tandem duplication (FLT3-ITD), and de novo methyl transferase 3 A (DNMT3A). These patients tend to have a poor prognosis. We compared 165 AML patients, analyzing those with these three mutations (referred to as the triple-mutated group) and those without. Our findings showed that patients with the triple mutations had higher white blood cell counts, more blast cells in the bone marrow, and higher platelet counts compared to patients without the triple mutations. All of the triple-mutated patients were also classified under a particular subtype of AML, the M4/M5 subtype. However, even though the two groups had similar remission rates after initial treatment, patients with the triple mutations had a much shorter overall survival rate, with a median survival of just 4 months. We also used a large dataset to examine gene expression patterns in these patients. One gene, called GNG4, was found to be significantly downregulated in patients with the triple mutations. This gene may help explain the poor survival outcomes for these patients. Our results suggest that GNG4 could serve as a potential target for future therapies and may help doctors predict the prognosis of AML patients with this triple mutation.

Introduction
Acute myeloid leukemia (AML) is a heterogeneous hematologic malignancy with a median overall survival (OS) of less than 1 year. 1 Recent advancements in high-throughput sequencing have emphasized the critical role of gene mutations in the diagnosis, prognostic risk stratification, and therapeutic decision-making of AML. 2 The most frequently observed mutations in AML include those in nucleophosmin 1 (NPM1) and FMS-like tyrosine kinase 3-internal tandem duplication (FLT3-ITD). 3 NPM1 mutations are present in approximately 25-35% of adult AML cases and about 8% of pediatric cases4,5 and are associated with higher complete remission (CR) rates, longer event-free survival, and extended OS.6-8 FLT3-ITD mutations, which occur in around 25% of AML patients, 9 are linked to increased leukemic cell counts, shorter OS, and a higher cumulative incidence of relapse. NPM1 and FLT3-ITD mutations frequently co-occur in adult AML patients. 10 Patients with NPM1 and FLT3-ITD mutations are classified as low-risk or intermediate-risk according to the ELN-2017 guidelines. 11 However, these patients with coexisting mutations of NPM1 and FLT3-ITD have variable clinical courses. 12 A sizable proportion of patients have a poor prognosis and short survival, 12 particularly when co-occurring with de novo methyl transferase 3 A (DNMT3A) mutations. 13
The DNMT3A gene encodes a 130-kDa protein that is involved in DNA methylation modification. 14 Aberrant DNA methylation patterns are a hallmark of AML.15,16 DNMT3A mutations are found in approximately 20% of newly diagnosed AML patients. 17 Notably, the co-occurrence of DNMT3A, NPM1, and FLT3-ITD mutations is relatively common, with approximately 7% of AML patients harboring these triple gene mutations.3,18 Co-occurrence of these three mutations is more frequent than can be explained by chance 8 and is associated with typical clinical features, such as significantly higher white blood cell counts. 13 Recent studies revealed genetic interaction of these three mutations in mice 19 and humans, 3 further suggesting that triple-mutated AML represents a distinct entity with very poor outcome. 13 However, there are relatively few studies investigating NPM1/FLT3-ITD/DNMT3A triple-mutated AML, 20 and little is known about its underlying molecular mechanisms. Therefore, further research is required to investigate the interactions between DNMT3A mutations and co-mutations in NPM1 and FLT3-ITD in triple-mutated AML patients.
GNG4 encodes a G-protein γ subunit involved in transmembrane signaling, and recent studies suggest it acts as a context-dependent regulator of leukemogenic pathways. Notably, GNG4 is frequently epigenetically silenced (eg, via promoter hypermethylation) in various cancers, including hematologic malignancies, and this loss is associated with more aggressive disease and poorer outcomes. 21 Mechanistically, GNG4 normally modulates GPCR signaling – for example, restoring GNG4 expression can dampen SDF1α/CXCR4-driven migration signaling in tumor cells and help maintain MHC class I expression via NF-κB signaling, an effect that bolsters anti-tumor immune surveillance. 22 Thus, downregulation of GNG4 may promote leukemogenesis and immune evasion, while high GNG4 levels have been linked to improved therapy responses in certain contexts.21,22 GNG4’s prognostic impact is under active investigation as a potential biomarker in leukemia, given its influence on signaling pathways and correlation with patient outcomes. 21 One recent example is Kawase et al, 2023, who identified GNG4 as a crucial NF-κB–dependent mediator of immune checkpoint efficacy. 22
In acute myeloid leukemia (AML), recurrent driver mutations commonly affect genes regulating epigenetics, signaling, or transcription, notably NPM1, FLT3-ITD, DNMT3A, CEBPA double, ASXL1, RUNX1, TET2, IDH1/2, KIT, and TP53. 23 These mutations often co-occur in non-random patterns that define molecular subgroups. For instance, NPM1-mutated AML frequently harbors concurrent FLT3-ITD and DNMT3A mutations (often alongside IDH1/2 or TET2), reflecting a common genotype in normal-karyotype AML. 24 By contrast, mutations associated with secondary or adverse-risk disease (such as ASXL1 and RUNX1) – as well as TP53 mutations – tend to cluster together and rarely overlap with NPM1 or CEBPA mutations. 25 Likewise, c-KIT mutations are typically restricted to core-binding factor AML subsets (eg, cases with t(8;21) or inv(16) translocations) and are known to co-occur in those leukemias. 23 Collectively, the presence or combination of these mutations has strong prognostic and biologic significance.
In this study, we explored the gene mutation spectrum of 165 AML patients, focusing on distinct co-mutation patterns, particularly comparing AMLNPM1mutFLT3-ITDmutDNMT3Amut and AMLNPM1mutFLT3-ITDmutDNMT3Awt. We compared the clinical characteristics of these two subsets and used bioinformatics to validate their prognoses. And we have focused on DNMT3A in order to determine whether the presence of DNMT3A mutation, alongside NPM1 and FLT3-ITD, defines a particularly high-risk subset. Furthermore, we investigated the underlying mechanisms and found that guanine nucleotide-binding protein subunit γ 4 (GNG4) may be the key to AMLNPM1mutFLT3-ITDmutDNMT3Amut.
Materials and Methods
Patient Samples
This retrospective study included 165 patients newly diagnosed with AML between January 2018 and August 2021 at the First Hospital of Lanzhou University. Patients diagnosed with acute promyelocytic leukemia were excluded from the study. All patients were diagnosed based on a comprehensive evaluation including peripheral blood cell count, bone marrow blast percentage, bone marrow cell morphology, cytochemical staining, flow cytometry (FCM) immunophenotyping, chromosomal karyotype analysis, and molecular biological assessment. All diagnoses were made in accordance with the World Health Organization diagnostic criteria. 26 Clinical and survival data were collected through electronic medical records and telephone follow-ups. This study adhered to the principles of the Declaration of Helsinki and was approved by the Ethics Committee of the First Hospital of Lanzhou University (Approval No. LDYYLL-2024-803; Date of approval: December 30, 2024). All patients or their family members provided written informed consent for bone marrow aspiration, biopsy, and chemotherapy-related procedures. For this retrospective analysis of fully de-identified data, the requirement for obtaining informed consent was formally waived by the above Ethics Committee (waiver date: 30 Dec 2024). We did not perform a formal prospective power calculation due to the exploratory, retrospective nature. Because AML patients are relatively uncommon, our sample size was determined by the availability of patients meeting these criteria rather than a prospective power calculation.
Cytogenetic Analysis
The karyotypes were described according to the International System for Human Cytogenetic Nomenclature 2009. 12
Mutation Detection
Mutations were detected using targeted next-generation sequencing and Sanger sequencing. Targeted sequencing of the following 11 genes was performed for all samples: NPM1, FLT3-ITD, DNMT3A, CEBPA double, ASXL1, RUNX1, TET2, IDH1, IDH2, c-KIT, and TP53.Genomic DNA was extracted from bone marrow aspirate samples and used for targeted NGS on an Illumina MiSeq platform to screen for mutations in 11 genes. A custom panel covering 11 recurrently mutated genes in acute myeloid leukemia (AML) – including DNMT3A, FLT3 (internal tandem duplication, FLT3-ITD), NPM1, CEBPA (double mutations), ASXL1, RUNX1, TET2, IDH1, IDH2, c-KIT, and TP53 – was sequenced for all samples. Library preparation and target enrichment were performed using standard protocols (eg, multiplex PCR-based amplification of target regions), and paired-end sequencing was carried out on the Illumina MiSeq platform. Sequence reads were aligned to the human reference genome (GRCh37/hg19), and variants in the target genes were identified using standard bioinformatics pipelines. Key mutations detected by NGS were validated by Sanger sequencing. Specifically, 11 gene mutations identified through NGS were confirmed by polymerase chain reaction (PCR) amplification and direct Sanger sequencing of the relevant gene regions. Gene-specific primers flanking each mutation site were used to amplify the target regions from genomic DNA, and the PCR products were purified for sequencing. Sanger sequencing was performed in both forward and reverse directions using BigDye® Terminator chemistry on an ABI capillary DNA sequencer. The resulting chromatograms were analyzed and compared to reference gene sequences, confirming the presence of the mutations initially observed in the NGS panel.
Response to Treatment
All patients were classified according to the ELN 2017 criteria and received cytotoxic chemotherapy, except those who received supportive therapy only. Chemotherapy responses were assessed using the International Working Group criteria. Overall responses included CR, partial remission (PR), and non-remission (NR). CR was defined as less than 5% bone marrow blasts with no Auer rods or evidence of persistent extramedullary leukemia. PR was defined as a reduction in bone marrow blasts to below 20% after treatment, with a decrease of more than 50% from pretreatment levels. NR was defined as the number of bone marrow blasts remaining >20% after the treatment. OS was defined as the time from initial diagnosis to the date of death or last follow-up.
Bioinformatics Analysis
The data used for bioinformatics analysis in this study were extracted from the Gene Expression Omnibus (GEO) database (GSE146173
27
), focusing on AML patients harboring NPM1/FLT3-ITD co-mutations. We downloaded the RNA sequencing data from 246 AML samples and found that 27 samples had NPM1/FLT3-ITD co-mutations. Data processing and presentation were performed using R packages (Version 4.0.3). Pan-cancer RNA-seq data in TPM format from the TCGA and GTEx databases, uniformly processed through the Toil pipeline, were downloaded via UCSC Xena (https://xenabrowser.net/datapages/). Patients with missing OS data were excluded. We included patients with complete clinical data and normalized gene expression data (Figure 1). AML patients were divided into subgroups based on the mutation status of NPM1, FLT3-ITD, and DNMT3A. Survival differences among these subgroups were analyzed and compared using Kaplan–Meier survival curves. AMLNPM1mutFLT3-ITDmut patients were further stratified into two groups based on their DNMT3A mutation status: the control group AMLNPM1mutFLT3-ITDmutDNMT3Awt and the experimental group AMLNPM1mutFLT3-ITDmutDNMT3Amut. Differential expression analysis of RNA-seq data between these two groups was performed using the limma package in R. Genes were considered differentially expressed if they met the threshold criteria of |log2(FC)| > 1 and P-value <0.05. Differentially expressed genes (DEGs) were subjected to Gene Ontology (GO) annotation, Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway enrichment analysis, and protein-protein interaction (PPI) network analysis. Hub genes were defined based on node degree (the number of adjacent nodes in the network), as this provided a straightforward indicator of each gene’s connectivity in the PPI analysis. Workflow Diagram
RNA Isolation, cDNA Preparation, and Quantitative PCR (qPCR)
RNA was isolated using the miRNeasy Kit (QIAGEN, Hilden, Germany), and reverse transcription for cDNA was performed utilizing the SuperScript® III First-Strand Synthesis System (#18080-051, Invitrogen, Carlsbad, CA). SYBR Green qPCR (#4309155; Applied Biosystems, Foster City, CA) was used for all genes. The sequences of the primer sets were as follows: GNG4 forward primer, 5′-GAGGGCATGTCTAATAACAGCAC-3′; GNG4 reverse primer, 5′-AGACCTTGACCCTGTCCATAC-3′; GAPDH forward primer, 5′-GTCTCCTCTGACTTCAACAGCG-3′; GAPDH reverse primer, 5′-ACCACCCTGTTGCTGTAGCCAA-3′. GNG4 expression was determined through the comparative Ct method using GAPDH levels for normalization.
Statistical Analyses
This study was conducted and reported in accordance with the STROBE (Strengthening the Reporting of Observational Studies in Epidemiology) guidelines. 28 All patient information was anonymized and de-identified prior to analysis to ensure confidentiality. We included all consecutive patients diagnosed with AML at our institution between January 2018 and August 2021 at the First Hospital of Lanzhou University who met the inclusion criteria. General clinical data, such as peripheral blood count, bone marrow blasts, and age, are expressed as median values and compared using the non-parametric Mann–Whitney U-test. The frequency of gene mutations, proportions of French-American-British (FAB) categories, and remission rates are expressed as percentages and compared using the Pearson chi-square or Fisher’s exact test. Statistical analyses were performed using the statistical package SPSS 25. GO annotation and KEGG pathway analysis were performed employing the clusterProfiler package in R. PPI network analysis was conducted using the STRING online database. The network was further analyzed, and hub genes were selected using the Cytoscape software. R package (version 4.0.3) and Microsoft Office Excel were used to draw the figures.
Results
Patients Characteristics
We retrospectively analyzed the clinical and survival data of 165 patients (Figure 1) with newly diagnosed AML and found that 141 (85.5%) had de novo AML, while 24 patients (14.5%) had secondary AML, primarily owing to myelodysplastic syndrome and chronic myelomonocytic leukemia.
Patient Characteristics

Abbreviations: WBC white blood cell, s-AML secondary AML, FAB French-American British, HMAs hypomethylating agents, CR complete remission, PR partial remission, NR non-remission, MRD minimal residual disease.
The 165 patients received various therapies. Of these, 23 (13.9%) were discharged with supportive therapy only because of factors such as poor performance status, advanced age, and/or limited family economic resources. Among the remaining 142 patients, 86 (52.1%) received conventional chemotherapy alone; 11 (6.7%), hypomethylating agents (HMAs) alone; 26 (15.8%), conventional chemo- and HMAs; 16 (9.7%), venetoclax and HMAs; and three (1.8%), other therapies. After the first cycle of treatment (n = 115), 73 (63.5%) patients achieved CR, with 45.2% of them being minimal residual disease-negative. Additionally, 16 (13.9%) patients achieved PR, and 26 (12.6%) patients were evaluated as having NR.
Mutational Landscape: NPM1, FLT3-ITD, and DNMT3A Were the Most Frequent Mutated Genes
The mutational landscape of 165 patients with newly diagnosed AML is shown in Figure 2. The number of coexisting gene mutations in each patient ranged from 0 to 4. Mutations in NPM1, FLT3-ITD, and DNMT3A were the most frequent. NPM1 mutations were observed in 37 (22%) patients; FLT3-ITD mutations, in 31 (19%); and DNMT3A mutations, 31 (19%) patients. Other gene mutations were identified in TET2 (19%), IDH2 (16%), ASXL1 (16%), CEBPA double (15%), RUNX1 (13%), IDH1 (9%), c-KIT (8%), TP53 (8%), CEBPA other (7%), and FLT3 other (7%). Diagram of Gene Mutations Landscape of 165 Newly Diagnosed Non-APL AML Patients. Age, Sex, FAB Category, Risk Stratification, Gene Mutation Frequency/Number, Gene Mutation Status, Coexisting Gene Mutations, and Co-mutation Numbers in AML Patients at the Initial Diagnosis. Abbreviations: CEBPA Double: Biallelic CEBPA Mutation; CEBPA Other: No Biallelic CEBPA Mutations; FLT3 Other: No ITD FLT3 Mutations
Co-Existing Pattern of Gene Mutations of NPM1, FLT3-ITD and DNMT3A

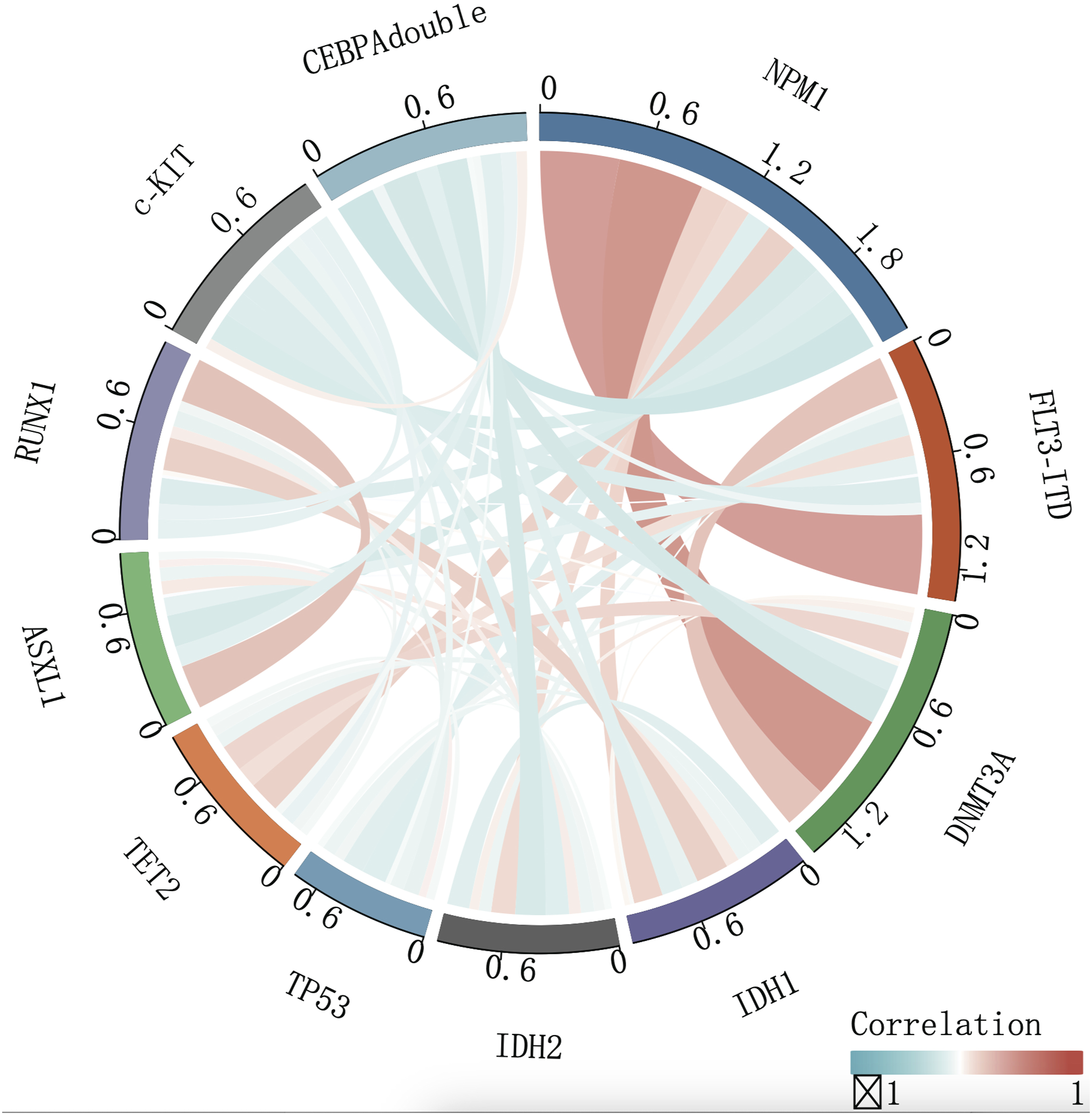
Co-Mutation Patterns of 165 Newly Diagnosed Non-APL AML Patients. Red and Blue Indicate Positive and Negative Correlations, Respectively, and the Depth Signifies the Correlation Strength
AML NPM1mutFLT3-ITDmutDNMT3Amut Patients had Reduced OS and Higher WBC Count and Were all in the M4/M5 FAB Category
The NPM1/FLT3-ITD/DNMT3A Triple-Mutated AML Patient Characteristics
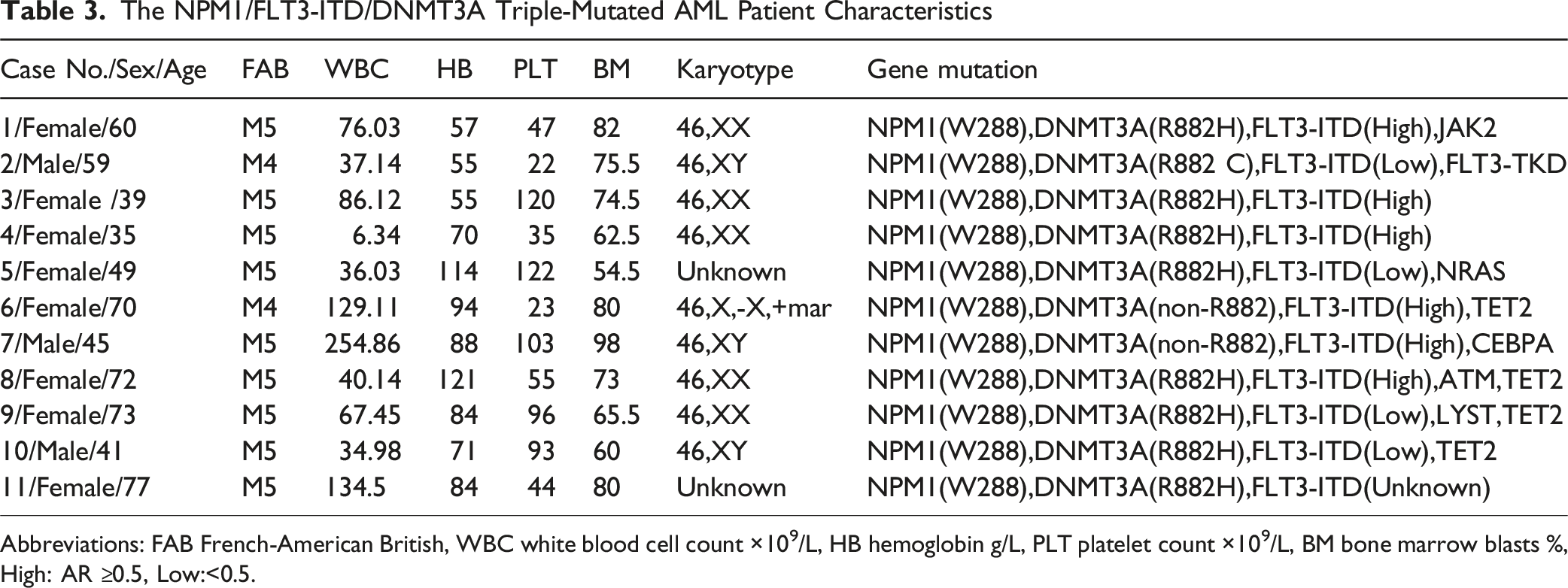
Abbreviations: FAB French-American British, WBC white blood cell count ×109/L, HB hemoglobin g/L, PLT platelet count ×109/L, BM bone marrow blasts %, High: AR ≥0.5, Low:<0.5.

Swimmer Plots of AMLNPM1mutFLT3-ITDmutDNMT3Amut Patients. Abbreviations: HMAs: Hypomethylating agents, CR: complete remission, PR: partial remission, NR: non-Remission, Ambiguous: Bone Marrow Morphology is Hypoplastic
Patient Characteristics Comparison According to the NPM1/FLT3-ITD/DNMT3A Mutations

*P < 0.05. p-values have been presented in bold to indicate statistical significance (p < 0.05).
The Hub Gene of AML NPM1mutFLT3-ITDmutDNMT3Amut –– GNG4
As illustrated in the workflow (Figure 1), in addition to analyzing the AML clinical data obtained from our hospital, our study extended the examination of the prognostic outcomes of AMLNPM1mutFLT3-ITDmutDNMT3Amut patients identified in the GEO database.
To clarify the prognostic impact of DNMT3A on AMLNPM1mutFLT3-ITDmut patients, we divided the 27 AMLNPM1mutFLT3-ITDmut patients from GSE146173 into two groups based on DNMT3A status: wild-type or mutated. A comparison of survival analysis between the two groups is shown in Figure 5C. We found that AMLNPM1mutFLT3-ITDmutDNMT3Amut patients had a significantly lower OS than AMLNPM1mutFLT3-ITDmutDNMT3Awt patients (P < 0.05). DEGs between AMLNPM1mutFLT3-ITDmutDNMT3Amut and AMLNPM1mutFLT3-ITDmutDNMT3Awt patients were visualized using a heatmap (Figure 6A). In total, 480 DEGs were identified, including 196 upregulated and 284 downregulated genes. The results of the biofunctional analyses, which included GO and KEGG pathway enrichment analyses, are shown in Figure 5D–I. We found that the DEGs played important roles in (1) biological processes: positive regulation of cell adhesion, cell-cell junction organization, granulocyte migration, and granulocyte chemotaxis; (2) cell composition: extrinsic components of the membrane, collagen-containing extracellular matrix, and extrinsic components of the plasma membrane; and (3) molecular function: receptor ligand activity, signaling receptor activator activity, cytokine activity, and cytokine receptor binding. The main signaling pathways involved were cytokine-cytokine receptor interactions, osteoclast differentiation, and human T-cell leukemia virus 1 infection. The PPI network was obtained from the STRING database using the Cytoscape software (version 3.8.0) (Figure 6B). The number of adjacent nodes is shown in Figure 6C. Notably, we identified hub genes using a degree-centrality approach; by this measure, GNG4 showed the highest connectivity with 13 adjacent nodes, marking it as the top hub gene, although we acknowledge that alternative centrality metrics (eg, maximal clique centrality) could also be applied for hub selection. Bioinformatics Analysis of GEO data: GSE146173 Dataset. a-c. Kaplan–Meier Survival Curves for the Patients From Different Groups. d-f. Results of the GO Analyses—Bar Plot, Bubble, and Circus. g-i. Results of the KEGG Analyses—Bar Plot, Bubble, and Circus Bioinformatics Analysis of GEO data: GSE146173 Dataset a. Heat Map of the Differentially Expressed Genes (DEGs) Between the NPM1 + FLT3-ITD + DNMT3A + AML (Red) Patients and the NPM1 + FLT3-ITD + DNMT3A-(Blue) Patients. (B) Protein-Protein Interaction of DEGs Using STRING. (C) Number of Adjacent Nodes of the DEGs. (D) Kaplan–Meier Survival Curves for the LAML Dataset Between High GNG4 Expression Group and Low GNG4 Expression Group. (E) Kaplan–Meier Survival Curves for the AMLFLT3-ITDmut Patients Between High GNG4 Expression Group and Low GNG4 Expression Group. (F) Validation of the GNG4 Expression Through qPCR Using Human Bone Marrow Samples From AMLNPM1mutFLT3-ITDmutDNMT3Amut and AMLNPM1mutFLT3-ITDmutDNMT3Awt Patients. Data are Presented as Mean ± Standard Deviation (SD), n = 10. **P < 0.01, vs the AMLNPM1mutFLT3-ITDmutDNMT3Amut Group

We found that GNG4, a hub gene, had 13 adjacent nodes. Based on the expression level of GNG4 (with the top 50% expression serving as the threshold), AML patients were categorized into two groups: a high GNG4 expression group and a low GNG4 expression group. Using the TCGA database LAML dataset, we extracted OS data to conduct survival analysis for these two groups. Across the entire AML cohort, patients in the high GNG4 expression group exhibited longer OS (P = 0.027) (Figure 6D). Further stratification of AML patients revealed that among AMLFLT3-ITDmut patients, the high GNG4 expression group showed slightly improved OS compared to the low expression group, although the difference was not statistically significant (P = 0.3) (Figure 6E). Using the TCGA, TIMER and TIDE database, we analyzed the GNG4 expression in pan-cancer and immune infiltration. We explored the differential expression levels of GNG4 in AML and analyzed the correlation between GNG4 expression levels and clinical features (Supplemental Figure S1).
Low GNG4 Expression may Contribute to the Low Survival Rate of AML NPM1mutFLT3-ITDmutDNMT3Amut Patients
To investigate the role and clinical significance of GNG4, identified as a potential distinguishing factor between AMLNPM1mutFLT3-ITDmutDNMT3Amut and AMLNPM1mutFLT3-ITDmutDNMT3Awt patients in our previous bioinformatics analysis, we conducted qPCR experiments to further validate the expression of GNG4 using the bone marrow samples from AMLNPM1mutFLT3-ITDmutDNMT3Amut patients (n = 4) or AMLNPM1mutFLT3-ITDmutDNMT3Awt patients (n = 6). All samples were obtained from patients who were newly diagnosed with AML at the First Hospital of Lanzhou University. RNA was extracted from these samples, and PCR was performed to quantify the transcript levels of GNG4.
Our results indicated that the transcript level of GNG4 in AMLNPM1mutFLT3-ITDmutDNMT3Amut patients was significantly lower than that in AMLNPM1mutFLT3-ITDmutDNMT3Awt patients (P = 0.0095) (Figure 6F). This finding is consistent with our bioinformatic predictions, suggesting that GNG4 expression may serve as a critical molecular marker for differentiating between these two patient groups. The lower expression of GNG4 in AMLNPM1mutFLT3-ITDmutDNMT3Amut patients underscores its potential role in the pathogenesis and progression of AML, warranting further investigation into its clinical implications.
Discussion
This study is among the first to investigate the hub genes of AMLNPM1mutFLT3-ITDmutDNMT3Amut patients. We identified 11 AMLNPM1mutFLT3-ITDmutDNMT3Amut patients from a cohort of 165 newly diagnosed AML patients, yielding a mutation frequency of 6.7%, which was consistent with previous reports.18,20 These AMLNPM1mutFLT3-ITDmutDNMT3Amut patients exhibited distinct clinical features, including elevated WBC counts, higher platelet counts, increased BM blasts, and predominantly monocytic leukemia (an increased number of monocytes in the bone marrow and peripheral blood), 29 particularly within the M4/M5 subtypes of the FAB classification system.9,20,30 These findings align with earlier studies and provide further insight into the clinical presentation of this subtype of AML.
Additionally, there is a difference in age and sex distribution between triple-mutated and non-triple-mutated AML.9,20,31 We found a notable trend toward a higher incidence of triple mutations in patients aged >70 years and women, although these differences were not statistically significant. This study identified a novel characteristic—the platelet count in patients with triple mutations at the time of initial diagnosis was significantly higher than that in non-triple mutated AML patients. This suggests that triple mutated patients may not present with bleeding symptoms caused by thrombocytopenia in the early stages of diagnosis. Based on the results of the chromosomal karyotype analysis, the majority of the 11 triple-mutated patients included in the study exhibited a normal karyotype. Only one patient presented with a chromosomal deletion, while two patients showed no metaphase cells available for analysis. A previous study showed that NPM1 mutations are strongly associated with normal karyotype AML and are commonly accompanied by DNMT3A and FLT3-ITD mutations. 32
From a cytomorphological perspective, nearly half of the triple-mutated patients (Patient No. 5, 8, 9, 10, and 11) displayed bone marrow characteristics consistent with predominantly monocytic leukemia. The monocyte lineage showed abnormal proliferation, comprising mature monocytes as well as primitive and immature monocytes. The proportion of abnormal cell populations ranged from 65% to 85%. Although AMLNPM1mut has a wide range of morphologies that are not directly related to FAB classification, numerous studies have demonstrated a strong association between NPM1 mutations and both acute myelomonocytic and acute monocytic leukemia.33,34 FCM results have led to the diagnosis of acute monocytic leukemia (AML-M5). For example, in Patient No. 5, immune cell phenotyping indicated that abnormal cells comprised approximately 80% of the bone marrow nucleated cells. These cells predominantly expressed markers, such as HLA-DR, CD4, CD11b, CD13, CD14, CD15, CD33, CD38, CD56, CD58, CD64, and CD123. Among these, about 24.5% were CD64bri+CD14+ mature monocytes, while approximately 54.5% were CD64bri+CD14- naïve monocytes. Similar immune cell profiles were observed in Patient Nos. 8, 9, 10, and 11. These findings suggested that certain cases of AMLNPM1mutFLT3-ITDmutDNMT3Amut exhibit distinct monocyte characteristics.
All AMLNPM1mutFLT3-ITDmutDNMT3Amut patients showed rapid disease progression, a high tendency for recurrence, and a poor overall prognosis. Previous studies have indicated that AML patients harboring both NPM1 and FLT3-ITD mutations have a poor prognosis, especially in cases with a high FLT3-ITD AR. Additional research has confirmed the negative prognostic impact of DNMT3A mutations on AMLNPM1mutFLT3-ITDmut patients, suggesting that DNMT3A further exacerbates poor outcomes in this subset of AML. According to previous research findings, AML patients with NPM1/FLT3-ITD/DNMT3A triple mutations have shorter median OS and disease-free survival (DFS) than those with only FLT3-ITD mutation. 9 Further analysis showed no significant difference in the CR rate between patients with triple mutations and those without. However, the relapse rate was significantly higher in the triple mutated group, which may explain the poorer prognosis observed in these patients. 20 Allo-HSCT was found to improve OS and DFS in both groups (triple-mutated and non-triple-mutated AML patients). However, in triple-mutated AML, the benefit was less pronounced, with the 1-year OS rate remaining below 30%. 30
To understand the reasons behind the poor prognosis of AMLNPM1mutFLT3-ITDmutDNMT3Amut patients and to explore the underlying mechanisms, we used the GEO dataset GSE146173 to identify specific genes associated with poor outcomes. Our analysis identified the GNG4 gene as playing a significant role in the prognosis of triple-mutated AML patients. GPCR, comprising α, β, and γ subunits, responds to various extracellular stimuli, such as hormones, growth factors, and sensory stimulating signals. GNG4 is one of 14 γ subunit proteins of GPCR. 35 GNG4 is important in MHC-I expression and immunotherapy resistance. Previous studies have shown that GNG4 is involved in PI3K/protein kinase B (Akt) signaling pathway and NF-κB signaling pathways.36-40 In addition, NF-κB signaling pathways are reportedly related to MHC-I expression.41-45 Kawase et al. found that MHC-I expression was reduced by GNG4 knockdown via the NF-κB signaling pathway, and GNG4 knockdown induced resistance to PD-1 blockade. 46
In addition, Pal et al reported that the promoter region of GNG4 was significantly hypermethylated and that its transcript level was significantly downregulated in glioblastoma, renal cell carcinoma, and thymic carcinoma.36,47 It functions as a tumor suppressor gene. Relapse-free survival was significantly worse in tumors with higher GNG4 DNA methylation levels than in those with lower DNA methylation levels in all thymic epithelia tumors. 47 Indeed, it has been proposed that malignancies can hijack central neuroendocrine homeostatic mechanisms to reset systemic physiology in a manner that favors tumor growth at the host’s expense. 48
A systems biology- and machine learning-based study revealed the potential benefits of midostaurin in FLT3-mutated AML. GNG4 has been identified as a protein that is differentially modulated in response to midostaurin treatment in FLT3-mutated AML, specifically through midostaurin’s non-FLT3 target effects. The upregulated GNG4 protein was related to a beneficial effect from the treatment, suggesting that it may play a role in enhancing the drug’s therapeutic efficacy by impacting signaling pathways beyond FLT3 inhibition. 49 Thus, GNG4 expression is consistently downregulated across various solid tumors and in AML. Its promoter is often highly methylated in these tumors, contributing to reduced expression levels. Moreover, hypermethylation of GNG4 frequently compromises relapse-free survival in cancer patients. 47
Consistent with this, our findings confirmed that GNG4 was downregulated in AMLNPM1mutFLT3-ITDmutDNMT3Amut compared to AMLNPM1mutFLT3-ITDmutDNMT3Awt and survival analysis showed that the low-expression GNG4 group had lower OS in AMLFLT3-ITDmut, a finding that aligns with the survival analysis results of Reina et al. 47 Our PCR experiments confirmed that GNG4 is a significant prognostic factor distinguishing AMLNPM1mutFLT3-ITDmutDNMT3Amut from AMLNPM1mutFLT3-ITDmutDNMT3Awt.
Emerging evidence suggests that expression patterns of certain hub genes in leukemia can reflect key clinical aspects, including a propensity for central nervous system involvement, 50 differential responses to therapy, 51 and underlying developmental lineage biases. 52 In this context, GNG4 expression may hold promise as a diagnostic and prognostic biomarker for AML patients harboring NPM1/FLT3-ITD/DNMT3A triple mutations. However, we emphasize that this potential utility of GNG4 remains to be validated, and further studies are needed to determine its clinical significance.
Patients with triple mutations showed significantly lower transcript levels of GNG4, which is consistent with previous studies and our bioinformatics predictions. This suggests that lower expression of GNG4 may contribute to the pathogenesis and progression of AML. These findings highlight the clinical value of GNG4 expression for diagnosing and potentially guiding the treatment of AML, warranting further investigation into its therapeutic implications.
This study must be interpreted in light of several limitations. First, it was a retrospective observational analysis with a relatively small sample size – especially in the triple‐mutated subgroup (only 11 patients) – which reduces statistical power and limits the generalizability of our findings. Second, our identification of hub genes in the PPI network was based on node degree (adjacent node count) alone, a relatively simplistic criterion; using more robust centrality metrics (eg, betweenness or eigenvector centrality) might have revealed a different or more nuanced set of hub genes. Third, due to the limited size of our patient cohort and the low number of events, we assessed the prognostic value of GNG4 using ROC analysis in an external dataset rather than in our own patient data. Another limitation of our study is the use of qRT-PCR for gene expression analysis, which cannot capture the full complexity of transcriptional regulation. Fifth, due to the low number of NPM1/FLT3-ITD/DNMT3A triple-mutated cases in our cohort, we relied on external cohorts to assess GNG4 expression and its prognostic impact. Therefore, any conclusions regarding GNG4’s specificity as a marker for this subgroup should be interpreted with caution, and further validation in larger patient cohorts is warranted. Relying on external data provided an independent perspective but also introduces potential biases and differences in patient characteristics, which may limit the direct applicability of those results to our population. Taken together, these limitations mean that our conclusions are tentative and should not be overstated. Further validation in larger, prospective cohorts and additional functional studies are warranted to confirm and expand upon our findings.
Conclusion
In conclusion, our study confirmed a high frequency of NPM1/FLT3-ITD/DNMT3A triple mutations (approximately 6.7%). Patients with triple-mutated AML consistently had higher WBC counts, platelet counts, and bone marrow blast percentages, and all fall within the M4/M5 FAB category. Most importantly, they had a high relapse rate and shorter OS, although there was no significant difference in the CR rate. Using bioinformatics analysis, we found that poor prognosis may be associated with low expression of GNG4, and further research may explore the mechanisms linking GNG4 to the prognostic significance of AML. However, the main limitation of this study was the small sample size of triple-mutated AML patients. Further studies with a larger case series are warranted.
Supplemental Material
Supplemental Material - Clinical Characteristics and Outcomes of Acute Myeloid Leukemia Patients Harboring NPM1/FLT3-ITD/DNMT3A Triple Mutations and the Potential Prognostic Value of GNG4
Supplemental Material for Clinical Characteristics and Outcomes of Acute Myeloid Leukemia Patients Harboring NPM1/FLT3-ITD/DNMT3A Triple Mutations and the Potential Prognostic Value of GNG4 by Yujie Niu, MD, Xingchun Luo, MD, Xiaoxiao Yang, MD, Yuancheng Guo, Xiao Tang, Long Zhao, Jinli Jian, Bei Liu in Cancer Control.
Footnotes
Author Note
The authors declare that this study was conducted in the absence of any commercial or financial relationships that could be construed as potential conflicts of interest.
Acknowledgments
We thank the patients and investigators who submitted data to the GEO database for providing the high-quality data analyzed in this study. We also express our gratitude to the patients who provided clinical information and to the medical testing institutions for their support.
Ethical Considerations
The study protocol was reviewed and approved by the Ethics Committee of the First Affiliated Hospital of Lanzhou University (Approval No. LDYYLL-2024-803) and was conducted in accordance with the Declaration of Helsinki.
Author Contributions
Yujie Niu: Conceptualization, Investigation, Data curation, Visualization, Writing - original draft, Writing - review & editing. Xingchun Luo: Investigation, Methodology, Formal analysis, Validation, Writing - original draft. Xiaoxiao Yang: Resources, Investigation, Formal analysis, Visualization, Validation, Writing - original draft. Yuancheng Guo: Investigation, Methodology, Validation, Project administration, Writing - review & editing. Xiao Tang: Formal analysis, Methodology, Software, Writing - review & editing. Long Zhao: Resources, Visualization, Writing - review & editing. Jinli Jian: Data curation, Writing - review & editing. Bei Liu: Conceptualization, Investigation, Funding acquisition, Supervision, Writing - original draft, Writing - review & editing.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by The First Hospital of Lanzhou University under Grant number ldyyyn2023-12; The First Hospital of Lanzhou University under Grant number ldyyyn2018-64; The Natural Science Foundation of Gansu province under Grant number 18JR3RA356.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Data Availability Statement
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
