Abstract
BACKGROUND:
Nearly half of adult acute myeloid leukemia (AML) patients were classified into cytogenetic normal acute myeloid leukemia (CN-AML). The expression level of Trophinin associated protein (TROAP) was proven to be associated with the prognosis of several cancers, but it is still unclear in the prognosis of patients with CN-AML.
METHODS:
We integrated CN-AML patient samples from 4 datasets to analyze the relationship between TROAP expression and the survival of CN-AML. In addition, we investigated 92 AML patients of The Cancer Genome Atlas (TCGA) database to analyze the relationship between TROAP expression and the survival of AML patients received chemotherapy. We investigated the relationship between the expression of TROAP and drug sensitivity in AML cell lines.
RESULTS:
CN-AML patients with high TROAP expression were related to good event-free survival (EFS) and overall survival (OS). In AML patients received chemotherapy, high TROAP expression was associated with good survival prognosis. Additionally, the expression of TROAP gene in leukemia stem cells (LSC)
CONCLUSION:
TROAP could serve as an independent predictor of CN-AML patients and could act as a potential biomarker for the prognosis of CN-AML. TROAP expression levels were closely correlated with the drug sensitivity of multiple drugs.
Background
Acute myeloid leukemia (AML) is a heterogeneous malignant disease that is classified by the abnormal proliferation of progenitor cells without the ability to differentiate [1]. Myeloid leukemogenesis driven by many genomic alterations is related to chromosome aberrations, such as reciprocal translocation, insertions, deletions, inversions, trisomies and monosomies [2]. Exploration of complexity of heterogeneity has been demonstrated that some certain chromosomal abnormalities could suggest the prognosis of patients who have received standard treatment (anthracycline and ara-C). However, a subgroup of AML patients (20% of pediatric and 45% of adult AML cases) [3] without any cytogenetic abnormalities is defined as cytogenetically normal acute myeloid leukemia (CN-AML). Characterized by intermediate-risk category [4], identifying novel prognostic determinants in patients with CN-AML has performed challenges.
In 14 AML patients with normal karyotype, 19 AML-associated mutated genes were detected by NGS, and only 9 mutations were found, including: DNMT3A, IDH2, IDH1, NRAS, NPM1, TET2, ASXL1, PTPN11, and RUNX1 [5]. According to different gene mutation combinations of CN-AML patients, the prognosis could be further predicted. For instance, mutations in CEBPA and NPM1 are associated with a good prognosis in CN-AML [6, 7], while DNMT3A, TET2, and RUNX1 mutations are associated with poor prognosis in CN-AML [8, 9, 10]. Besides, high expression of some genes including ITPR2, MAPKBP1, CPNE3, RUNX1 and ATP1B1 are associated with poor prognosis of CN-AML, while the high expression of LEF1 is regarded as a favorable prognostic factor for CN-AML [11, 12, 13, 14, 15, 16]. Therefore, the identification of new potential biomarkers can help further predict the prognosis risk of CN-AML.
Trophinin-associated protein (TROAP), formally known as tastin, was a protein first identified to mediate human embryo implantation and found to be related to microtubule regulation later. TROAP is a cytoplasmic protein that forms a complex with trophinin and bystin to mediate the initial attachment in the interface of trophoblasts and endometrium [17, 18, 19, 20, 21, 22]. This complex function in the cellular process of embryo implantation, which is accompanied by malignant tumor cell invasion and metastasis. Previous studies revealed that TROAP expression is located in the centrosomes, microtubules and the proper spindle during the mitosis. High level of TROAP mediates the formation of unipolar spindles. The loss of TROAP expression will lead to induction of multipolar spindles and mitotic blockade, indicating that TROAP has significance in proper spindle assembly [23, 24, 25].
Recently, some researchers have found that TROAP play a critical role in several cancers including gastric cancer [26], epithelial ovarian carcinoma [27, 28], colorectal cancer [29], prostate cancer [30], and hepatocellular carcinoma [31, 32]. However, the role of TROAP in the progression of heterogeneous malignant disease, especially the prognostic significance of TROAP expression in AML, remains unclear. In our study, through an analysis of data from The Cancer Genome Atlas (TCGA) and Gene Expression Omnibus (GEO), we investigated potential correlations between the expression of TROAP in patients with CN-AML and prognosis of cytogenetically normal acute myeloid leukemia and assessed the prognostic value of TROAP for event-free survival (EFS) and overall survival (OS) of CN-AML patients.
Methods
Data source
In our study, we detected the TROAP expression level of CN-AML patients from TCGA and GEO databases (including GSE12417, GSE22778, GSE71014 datasets) [33, 34, 35, 36]. We analyzed the relationship between survival and TROAP expression through 75 CN-AML patients from TCGA database. Moreover, the analysis of relationship between TROAP expression and OS from GSE12417 (78 patients from GSE12417 U133 plus dataset, 162 patients from GSE12417 U133A dataset), GSE22778 (55 patients from GSE22778 GPL8653 dataset), and GSE71014 datasets (104 patients from GSE71014 dataset) were done. In the TCGA database, 92 patients with AML received chemotherapy were received analysis of the correlation between TROAP gene and survival (OS and EFS). We also analyzed relationship between TROAP expression and OS in 417 newly diagnosed AML patients from GSE37642 GPL96 [37]. By using the GSE76004 dataset [38], we examined 227 CD34/CD38 cell fractions from samples of 78 AML patients and detected TROAP expression of each group. Besides, we detected TROAP gene expression of those patients in different LSC activity groups (Supplementary Fig. 1). Meanwhile, we did univariate and multivariate analysis for EFS and OS of 75 CN-AML patients from TCGA dataset. The risk classification of TCGA 75 CN-AML patients are based on 2010 European Leukemia Net (ELN) guideline. We investigated the expression of TROAP of 29 tumor tissues and normal tissues, normal tissues from GTEx, and normal hematopoiesis cells from GSE42519 [39, 40, 41]. We also analyzed the dependence of cell lines of 23 tumor on TROAP using RNAi (Achilles
The informed consents of this research were consistent with the Ethics Committee of Peking University Third Hospital and did not violate the Declaration of Helsinki. All persons gave their informed consent prior to their inclusion in the study.
The 75 CN-AML were selected from TCGA database with the criteria as follows: 1) AML patient with normal karyotype. 2) All patients received gene expression detection with RNA-seq. 3) AML patients received chemotherapy regimen or transplant regimen.
The 92 AML received chemotherapy were selected from TCGA database with the criteria as follows: 1) AML patients received chemotherapy regimen but not transplant regimen. 2) All patients received gene expression detection with RNA-seq. 3) AML patients with normal or abnormal karyotype.
Gene expression analysis
We calculated the RNA expression from RNA-seq data of TCGA dataset using RPKM (Reads Per Kilobase per Million mapped reads). We calculated the RNA expression from microarray of each AML sample from GEO databases by using the method of RMA (robust multiarray averaging). Then, the value of TROAP expression level of each gene was further transformed with log2. The
Statistics
Through using R software v3.1.3 (ggplot2 and survminer v0.4.6 package), we finished statistical analysis in this research. Meanwhile, we used Kaplan-Meier estimation to plot survival curves and present survival analysis by the log-rank test. One-way analysis of variance (Anova test) was used in comparing TROAP expression levels between four CD34/CD38 subgroups. Additionally, we adopted unpaired
Multivariate analysis for TCGA CN-AML patients’ EFS and OS
Multivariate analysis for TCGA CN-AML patients’ EFS and OS
HR, hazard ratio; CI, confidence interval. TROAP (TROAP gene expression, high vs. low expression). DNMT3A (mutation vs. wild type), FLT3 (mutation vs. wild type).
High TROAP expression predicts better survival of cytogenetic normal acute myeloid leukemia (CN-AML) and AML patients received chemotherapy from the TCGA database. The X axis represents time (months) and the Y axis represents survival probability. A, Kaplan-Meier curves for event-free survival (EFS) and overall survival (OS) in different TROAP expression groups of CN-AML in the TCGA database (Cut off value 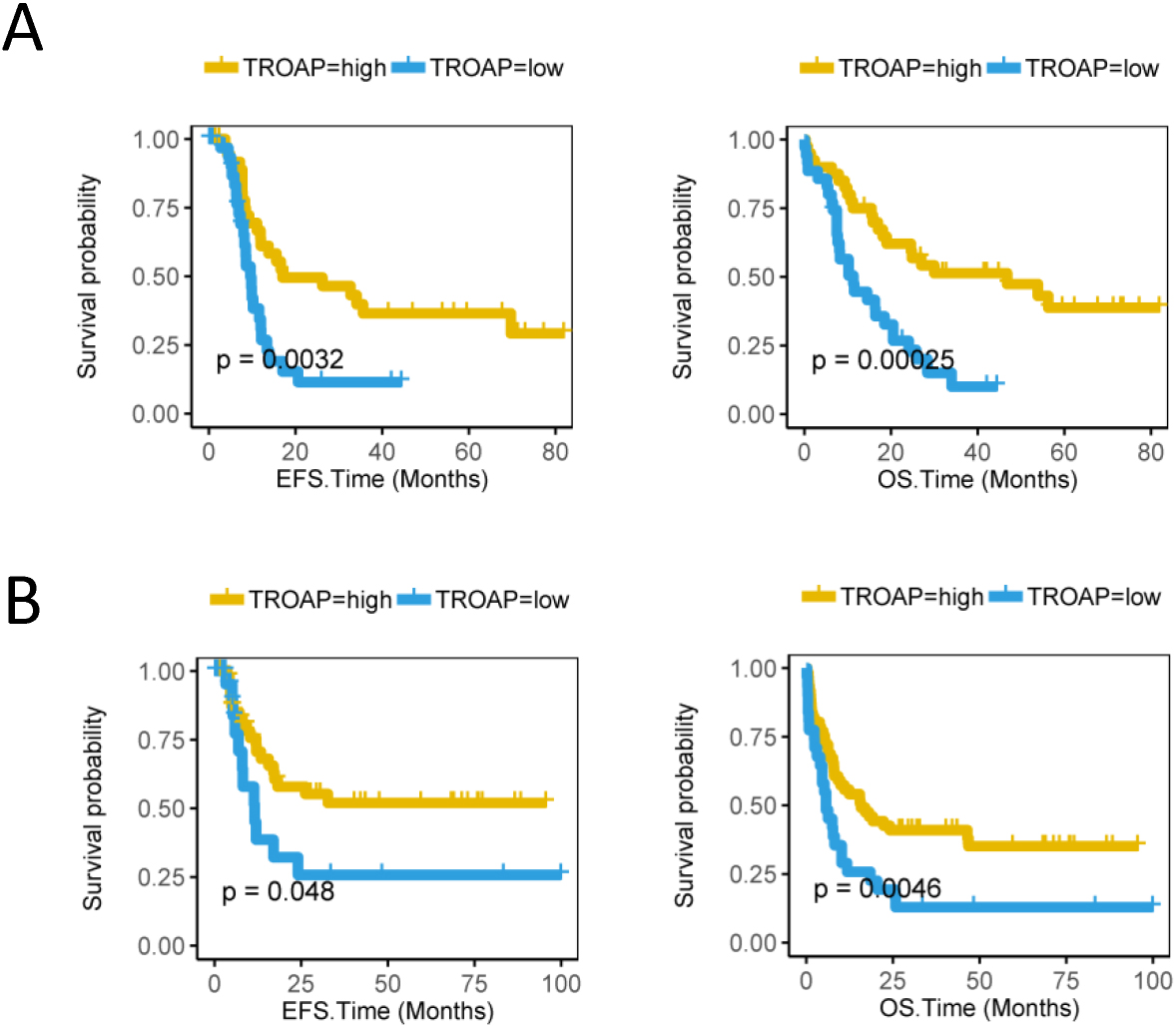
Clinical baseline characteristics of patients with CN-AML from TCGA database
We divided 75 CN-AML patients into TROAP-high group and TROAP-low group based on TROAP expression level (see method for the standard of each group). Then we compared the molecular and clinical characteristics of two groups (Supplementary Table 1). WBC of TROAP-low group was higher than the group of high TROAP expression (
High TROAP expression predicts better survival of CN-AML patients from the other independent datasets. The X axis represents time (days) and the Y axis represents survival probability. A, (left side) Kaplan-Meier curves of OS in different TROAP expression groups of CN-AML in GSE12417 U133 plus (Cut off value 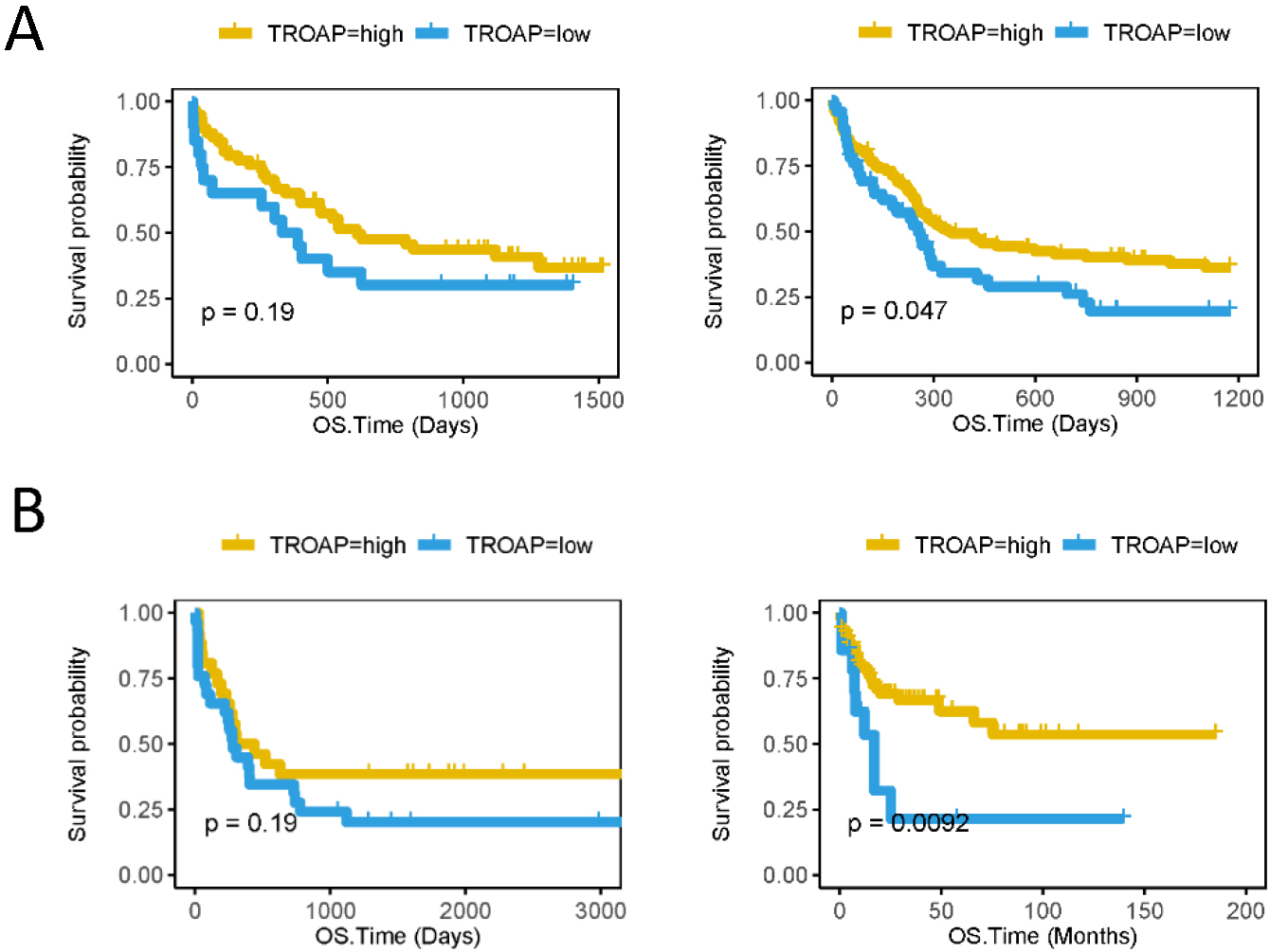
Expression level of TROAP gene in LSC. The X axis represents different group of AML cells and the Y axis represents gene expression. TROAP gene expression was transformed as log2. A, Comparison of TROAP expression levels in four subtypes of AML cells, 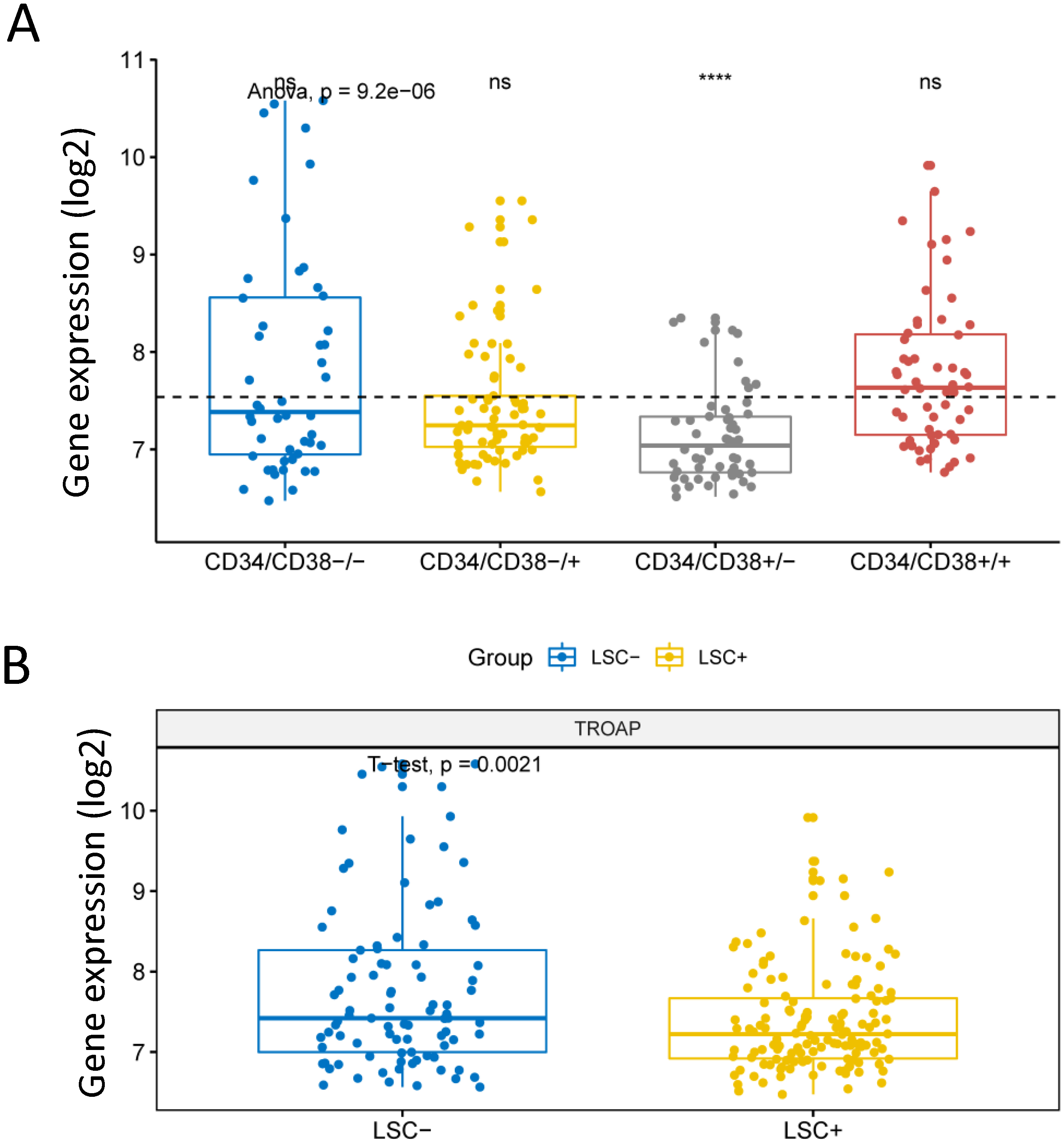
In our study, we examined the relationship between the different levels of TROAP gene expression and survival (including OS and EFS) from samples of 75 TCGA patients with normal karyotypes AML. The EFS and OS of group of high level of TROAP gene expression was better than the group of low level of TROAP (Fig. 1A, EFS,
TROAP expression predicts survival received chemotherapy in AML patients
To research the expression level of TROAP in AML patients received chemotherapy, we investigated the correlation of the gene expression of TROAP and the prognostic value of 92 received chemotherapy patients from TCGA database. By analyzing the EFS and OS in TROAP-high and TROAP-low groups of 92 CN-AML received chemotherapy patients, we found that the high expression of TROAP predicts better survival rate (Fig. 1B, EFS,
Expression level of TROAP in LSC
Leukemia Stem Cells (LSC) were divided into 4 cell populations according to fractions of the CD34 and CD38 phenotypes. The expression level of TROAP was significantly different in each subgroup (Fig. 3A,
Univariate and multivariate analysis of TCGA CN-AML patients’ EFS and OS
Age (
We selected TROAP expression levels, age
TROAP expression level in various tumors, normal tissue, and hematopoiesis cell
We studied the expression levels of TROAP of 29 tumors and normal tissue from TCGA and Genotype-Tissue Expression (GTEx, Fig. S4A and B). We discovered that in most solid tumors, the expression level of TROAP in normal tissue was lower than in tumor tissue, while in AML, we found that the expression level of TROAP was higher in normal bone marrow than in tumor tissue. (Fig. S4A,
Relationship between the expression level of TROAP and drug sensitivity
To further investigate whether TROAP expression level is related to drug sensitivity, we compared the IC50 values of compounds in the high and low expression groups of TROAP in GDSC 20 AML cell lines. We found significant differences in IC50 between high-TROAP and low-TROAP among 5 drugs including 5-Fluorouracil, LGK974, Irinotecan, Luminespib and Camptothecin (Fig. S7A). 5-Fluorouracil, Camptothecin and Irinotecan are antitumor chemotherapeutic agents used in the treatment of many types of cancer including AML [43]. We found that among the three drugs (5-Fluorouracil, Camptothecin, Irinotecan), the lower the expression of TROAP, the lower the IC50, and the higher the sensitivity to the drugs (Fig. S7B–D; 5-Fluorouracil,
Discussion
An array of adhesion proteins, including L-selectin and trophinin, mediate human embryo implantation [46]. As a cytoplasmic protein, TROAP forms a complex with trophinin and bystin intracellular domain to mediate the interaction between trophoblast cell and endometria cell. After the elementary attachment of the maternal epithelial cells to embryonic cells, an intensive adhesion is induced, and a series of morphological changes are found in the embryo implantation site. The cellular process of embryo implantation may be related to the cell process of malignant tumor cells in invasion and metastasis, which infers that the expression of TROAP gene may be associated with malignant tumors [24, 25, 47].
In previous studies, high TROAP expression has been reported in some malignant solid tumors such as gastric cancer [26], colorectal cancer [29], prostate cancer [30], epithelial ovarian carcinoma [28] and liver cancer [48]. In some solid cancers, like gastric cancer, colorectal cancer, prostate cancer [26, 29, 30], TROAP gene was upregulated compared to the normal counterparts. Besides, TROAP gene promoted cell proliferation, G1 to S cell cycle transition and the ability of migration and invasion of tumor cells, which predicted poor survival significantly. Moreover, in the epithelial ovarian carcinoma, the expression product of TROAP is helpful to predict the stage of malignant tumor. More TROAP genes were expressed in later stage of epithelial ovarian carcinoma [28]. However, some report showed that downregulated TROAP plays a critical role in human hepatocellular carcinoma through upregulation of tumor cell growth and migration [31], which suggested that TROAP is a prognostic biomarker for these hepatocellular cancer patients. These conclusions proved that the prognostic results of TROAP were different in different cancers. In most solid cancers, TROAP gene predicts a poor prognosis in survival. So far, we seldom have known about the relationship between TROAP and none-solid tumors. Importantly, the prognostic relationship between this gene and CN-AML, an intermediate-risk category, has not been researched recently. We analyzed the gene expression of TROAP from CN-AML patients in 4 independent datasets to prove that the correlation of TROAP gene expression and prognosis of CN-AML patients and this gene is an independent risk factor of CN-AML. In our survival analysis research, we showed that the CN-AML patients with high expression of TROAP had better prognosis than those with low TROAP expression. High TROAP expression also predicts good prognosis in AML patients received chemotherapy. The possible mechanism for the better prognosis of high TROAP expression AML patients may be related to the LSC. Lower TROAP expression was observed in LSC
CD34
In the multivariate analysis, we discovered that age and the DNMT3A mutation were related to EFS and OS in CN-AML patients, and that the FLT3 mutation was associated with EFS in CN-AML. Besides, TET2 and NRAS mutations were related to OS in CN-AML patients. Those results showed consistency with previous findings that DNMT3A mutations are associated with shorter EFS and OS [55, 56]. Multivariate analysis showed that high expression of TROAP in CN-AML patients was regarded to be associated with better OS. Furthermore, we found that the correlation between TROAP expression levels and the prognosis of CN-AML patients was better than other factors. These results indicated that TROAP was an independent predictor of clinical prognosis in patients with CN-AML.
However, we have not researched the specific function of TROAP in the pathogenesis-signaling pathway of CN-AML. This study only investigated the expression of individual genes, and future studies combining other different types of biomarkers could predict the prognosis of patients with CN-AML better. Meanwhile, we did not apply data of TROAP expression and clinical pathologic variables to build a predictive model because of the limited number of samples. In our future work, we will investigate the potential clinical prognostic value of TROAP expression in a larger population for constructing a proper predictive model for the prognosis of patients with CN-AML.
Conclusions
The present study indicated that high expression of TROAP gene was a better prognostic factor for CN-AML patients and AML patients received chemotherapy. TROAP could serve as an independent predictor of CN-AML patients and act as a potential biomarker for the prognosis of CN-AML patients and AML patients. The expression level of TROAP is closely related to drug sensitivity.
List of abbreviations
AML: acute myeloid leukemia; CN-AML: cytogenetically normal acute myeloid leukemia; TROAP: Trophinin-associated protein; TCGA: The Cancer Genome Atlas; GEO: Gene Expression Omnibus; EFS: event-free survival; OS: overall survival; LSC: leukemia stem cells; BM-blast: bone marrow blast; PB-blast: peripheral blood blast; WBC: white blood cell; HR: hazard ratio; CI: confidence interval.
Ethics approval and consent to participate
All experimental protocols were approved by a Ethics Committee of Peking University Third Hospital and did not violate the Declaration of Helsinki. All persons gave their informed consent prior to their inclusion in the study.
Consent for publication
Not applicable.
Availability of data and materials
Not applicable.
Competing interests
The authors declare no competing interests.
Funding
This work was funded by National Natural Science Foundation of China (81800195), interdisciplinary medicine Seed Fund of Peking University (BMU2018MB004), Natural Science Foundation of Beijing Municipality (7132183 and 7182178), China Health Promotion Foundation (CHPF-zlkysx-001), and Key Clinical Projects of Peking University Third Hospital (BYSYZD2019026).
Authors’ contributions
HMJ and WLZ made contributions to conception of the project. HX, JH, CJY, and WLZ make contributions to interpretation or analysis of data. HX, JH, JCY, NXL, YLZ, PY, JW, SXL, WZ, GHD, WLZ and HMJ contribute to preparation of the manuscript and revision for important intellectual content. HMJ and WLZ make contributions to supervision of the project. All authors read and approved the final manuscript.
Supplementary data
The supplementary files are available to download from http://dx.doi.org/10.3233/CBM-210042.
sj-pdf-1-cbm-10.3233_CBM-210042.pdf - Supplemental material
Supplemental material, sj-pdf-1-cbm-10.3233_CBM-210042.pdf
sj-docx-1-cbm-10.3233_CBM-210042.docx - Supplemental material
Supplemental material, sj-docx-1-cbm-10.3233_CBM-210042.docx
Footnotes
Acknowledgments
The authors thank TCGA database and datasets GSE12417, GSE22778, GSE71014, GSE76004, GSE37642, GSE42519 from GEO database, GDSC database, DepMap Portal database and GTEx database.
