Abstract
BACKGROUND:
Long non-coding RNAs (lncRNAs) were detected extraordinarily expressed in various tumors and could combine with microRNAs (miRNAs) to play important role in tumor cells. This study is to explore the role of lncRNA RP11-909N17.2 in NSCLC and discuss in what way it functions in NSCLC.
METHODS:
120 NSCLC patients were enlisted in this study. Expression levels of lncRNA RP11-909N17.2 and miR-767-3p were detected and the correlation between lncRNA RP11-909N17.2 expression and the clinical data characteristics was analyzed. Prognosis potential of lncRNA RP11-909N17.2 was inferred with Kaplan-Meier and multivariate Cox regression assays. Biological functions of NSCLC cells were accessed by cell counting Kit-8, transwell migration and invasion assay. Mechanism of RP11-909N17.2 action on NSCLC cells was investigated by luciferase activity assay with wide-type or mutation.
RESULTS:
LncRNA RP11-909N17.2 has an ascendant expression while miR-767-3p has descended one in NSCLC tissue specimens and cells. Over-expression of lncRNA RP11-909N17.2 can shorten the overall survival period of NSCLC patients when compared with low expression. Knockdown of lncRNA RP11-909N17.2 suppressed biology function of NSCLC cell including proliferation, migration, and invasion.
CONCLUSION:
LncRNA RP11-909N17.2 can be developed into a prognostic index for NSCLC. LncRNA RP11-909N17.2 plays a promoting role in NSCLC cells possibly by binding miR-767-3p as a sponge.
Introduction
As a common malignant tumor of the respiratory system, lung cancer has led to a high morbidity and mortality rates worldwide [1]. Lung cancer has been subdivided into two subsets pathologically: small cell lung cancer or SCLC [2]; and non-small cell lung cancer or NSCLC (Including adenocarcinoma, squamous cell carcinoma and large cell carcinoma) [3]. Almost one-quarter of all cancer deaths are due to lung cancer. According to data from 2009 through 2015, the 5-year relative survival rate for lung cancer is only 19% for all stages combined [4]. In view of this, it is important to study and find reliable prognostic factors of lung cancer.
long non-coding RNAs (lncRNAs) is a subgroup of non-coding RNAs composed of more than 200 nucleotides. It has been reported that lncRNAs play a critical role in physiological processes and progression of human diseases [5, 6]. Tang et al. identified three lncRNAs, RP11-397D12.4, AC007403.1, and ERICH1-AS1, as potential fingerprints for tumorigenesis prediction [7]. Also, lncRNA TUC338 was found an elevated expression in lung cancer and increased the lung cancer cell activity by MAPK pathway [8]. As reported, lnc-IGFBP4-1 was significantly upregulated in lung cancer tissues compared with paracarinoma tissue [9]. Recently, lncLINC00461 was concluded as a promotion of the NSCLC progression of by the miR-30a-5p/ZEB2 axis, and can be used as a potential treated target for NSCLC [10]. All these reports inspire the interest in further study of the lncRNA regulatory nexus that controls the carcinogenesis of NSCLC.
There are several ways by which lncRNAs function in mammalian cells, such as binding to the microRNA (miRNA) [11]. LncRNAs contain multiple miRNA-binding sites that have been investigated as competing endogenous RNAs (ceRNA) to indirectly modulate mRNAs through binding to miRNAs [12]. Guan et al. studied the function of lnc RNA SNHG20 in breast cancer cells and arrived at a conclusion that SNHG20 reversely regulated miR-495 leading to a boosted ability in biological characteristics of breast cancer cells [13]. lnc RNA RP11-909N17.2 was reported differentially expressed in breast cancer and normal, which was significantly associated with survival time of breast cancer patients [14]. He et al. screened four abnormal expression genes in adenocarcinoma of the lung from the Cancer Genome Atlas (TCGA) database, one of them is RP11-909N17.2 [15]. But the expression level and detail of RP11-909N17.2 function in NSCLC have not been concerned.
This study is aimed to reveal the ability of RP11-909N17.2 as a potential prognosis factor, and its biological effects in NSCLC. And, furtherly we also aimed to explore the regulatory mechanism of RP11-909N17.2-miR-767-3p axis in NSCLC process.
Materials and methods
Patients and clinical specimens
Tumor specimens and para-tumor normal samples were provided by 120 paired tissues from NSCLC patients receiving surgical treatment in Yidu Central Hospital of Weifang during the study period and were immediately stored in liquid nitrogen for the purpose of further analysis. The patients received no preoperative treatment (chemotherapy or radiotherapy), and all tissues were confirmed to be NSCLC through histopathological examination. Clinical data of the enrolled NSCLC patients were obtained, and key clinical characteristics were recorded in Table 1. The patients were classified into different TNM stage of the tumors following the 7
Correlation of the lnc RP11-909N17.2 expression with clinical characteristics in NSCLC
Correlation of the lnc RP11-909N17.2 expression with clinical characteristics in NSCLC
The study was conducted in accordance with the Declaration of Helsinki (as revised in 2013) and approved by the institutional ethical committee of Yidu Central Hospital of Weifang. Written informed consent was gathered from each patient enrolled in the study.
Four human NSCLC cell lines (H1299, A549, NCI-H650, and HCC827) and an immortalized human bronchial epithelial cell line (BEAS-2B) were gained from Institute of Cell Biology, Chinese Academy of Sciences, Shanghai (Shanghai, China). All cells were cultured in Dulbecco’s Modified Eagle’s medium (DMEM; Gibco, NY, USA) fortified with 10% Fetal Bovine Serum (FBS; Gibco, NY, USA) in the condition of 5% CO
The small interfering RNA (siRNA) sequences targeting RP11-909N17.2 (si-RP11-909N17.2) was synthesized by Genepharm. Co., (Shanghai, China). The miR-767-3p inhibitor (anti-miR-767-3p), inhibitor NC (anti-NC), miR-767-3p mimic (miR-767-3p), and mimic NC (miR-NC) were obtained from RIBOBIO (Guangzhou, China). All oligonucleotides or vectors were respectively transfected into cells using Lipofectamine 3000 (Invitrogen) in accordance with the manufacturer’s procedure.
RNA isolation and real-time quantitative PCR (RT-qPCR) assay
The total RNA was extracted from patients’ tissue specimens and cultured cells with the addition of TRIzol reagent (Thermo Fisher Scientific, Waltham, MA, USA), in line with the manufacturer’s instructions. Then the RNA samples (A260/A280 ratio: between 1.8 and 2.0) were subjected to reverse transcription to synthesize cDNA using the Superscript III transcriptase (Invitrogen, Grand Island, NY) according to the manufacturer’s instructions. Subsequently, the qRT-PCR assay was conducted at a 7500 Real-Time PCR System (Applied Biosystems, USA) with the SYBR
Cell proliferation assay
Transfected cells were prepared to be a single cell suspension with a cell density of 4
Cell migration and invasion assays
24-well Transwell chambers (8 m pore; Corning, NY, USA) were used to estimate the cell migration and invasion of A549 and H1299 cells. For the migration assay, upper chambers were loaded with cell suspension (5
Bioinformatic analysis for RP11-909N17.2-target miRNA
To predict the potential target miRNA of RP11-909N17.2, an online prediction tool, LncBase Predicted v.2 was used (
Dual-luciferase reporter assay
The sequences of RP11-909N17.2 was synthesized and subcloned into luciferase reporter vector (Promega, Shanghai, China), named WT-RP11-909N17.2; The corresponding mutant versions without miR-767-3p binding sites were synthesized and subcloned into luciferase reporter vector (Promega, Shanghai, China), named MUT-RP11-909N17.2. Then H1299 cells were seeded in 24-well plates and co-transfected with WT-RP11-909N17.2 or MUT-RP11-909N17.2 and miR-767-3p or miR-NC with the help of Lipofectamine 2000 (Invitrogen). After 48 hours, the luciferase activity was determined by dual-luciferase reporter assay kit (Promega) according to the manufacturer’s protocol.
Expression level of RP11-909N17.2 and miR-767-3p in NSCLC tissue samples and cells. (A) RP11-909N17.2 is upregulated in NSCLC tissue samples and para-tumor normal samples; (B) miR-767-3p is downregulated in NSCLC tissue samples and para-tumor normal samples; (C) The correlation between miRNA-767-3p and RP11-909N17.2 expression in NSCLC tissues was analyzed by Spearman correlation analysis; (D) Expression level of RP11-909N17.2 in NSCLC cell lines (H1299, A549, NCI-H650, and HCC827); (E) Expression of miR-767-3p in NSCLC cell lines (H1299, A549, NCI-H650, and HCC827). 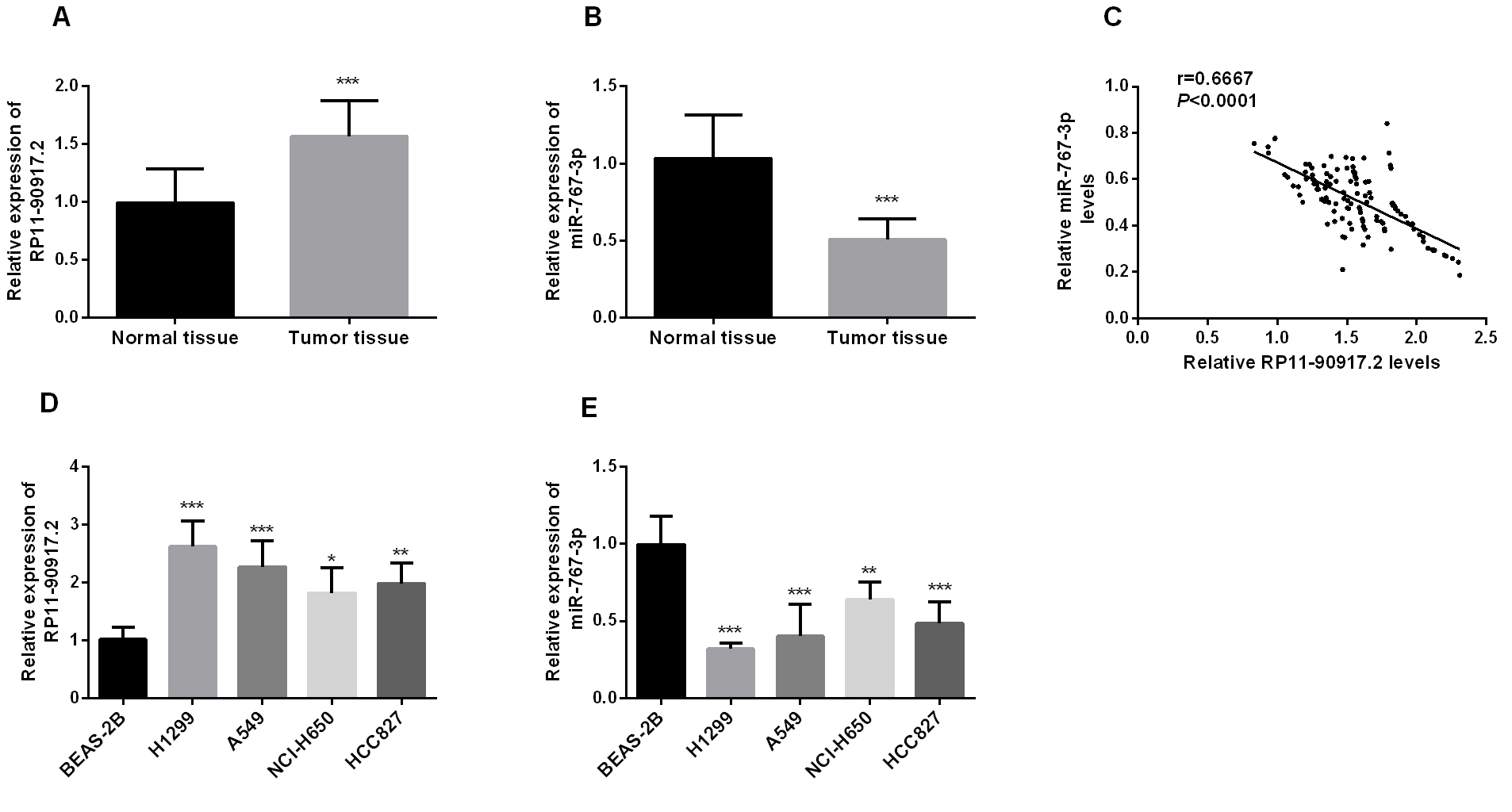
Magna RIP
Statistical analysis
Data are given as means
Results
Correlation between expression of lncRNA RP11-909N17.2 and clinical data of patients with NSCLC
The enrolled 120 patients with NSCLC were grouped into low lnc RP11-909N17.2 expression (
RP11-909N17.2 is upregulated in NSCLC tissues and cell lines but miR-767-3p is downregulated
RT-qPCR was performed to evaluate the expression levels of RP11-909N17.2 and miR-767-3p in NSCLC samples and different cell lines. As shown in Fig. 1A, RP11-909N17.2 was significantly upregulated (About 1.5-fold) in NSCLC tissue specimens compared with adjacent normal tissues (
Comparison of 5-year survival curves of NSCLC patients with high and low expression of RP11-909N17.2 by Kaplan-Meier method. Survival of the high expression group showed a shorter overall survival rate than the low expression group (log-rank test 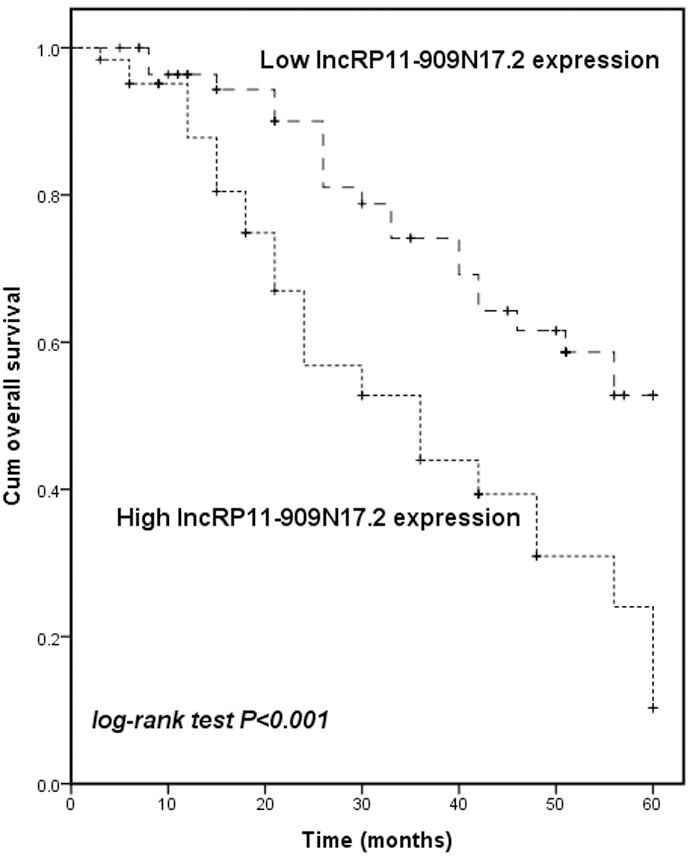
Over a 5-year follow-up period, the Kaplan-Meier curve was drawn to present the relationship of RP11-909N17.2 expression and prognosis in NSCLC patients. As shown in Fig. 2, patients with a raised RP11-909N17.2 expression showed a shorter overall survival time than those with low one (log-rank test,
Multivariate Cox analysis of clinical characteristics in relation to overall survival
Multivariate Cox analysis of clinical characteristics in relation to overall survival
H1299 and A549 cells were cultured to study the function of RP11-909N17.2 in vitro. By RT-qPCR, the result showed that RP11-909N17.2 si-RNA significantly suppressed the expression level of RP11-909N17.2 (Fig. 3A and B). The knockdown of RP11-909N17.2 also led to a significant decline in OD values which means the proliferation level of cells was attenuated (Fig. 3C and D). From the result of transwell assay, the cell number presented a significant decrease both for migration and invasion if RP11-909N17.2 si-RNA was involved, which means knockdown of RP11-909N17.2 can inhibit migration, invasion of NSCLC cells (Fig. 3E–H).
Proliferation, migration and invasion of H1299 and A549 cells was suppressed after downregulation of RP11-909N17.2 compared to untreated cells. H1299 and A549 cells were transfected with si-NC or si-RP11-909N17.2 (A) and (B) Expression level of RP11-909N17.2 was determined in H1299 cells and A549 cells by RT-qPCR; (C) and (D) Proliferative capacity of H1299 and A549 cells were measured by CCK-8; (E) and (F) Migratory ability of H1299 and A549 cells were measured by transwell assay; (G) and (H) invasive ability of H1299 and A549 cells were measured by transwell assay. 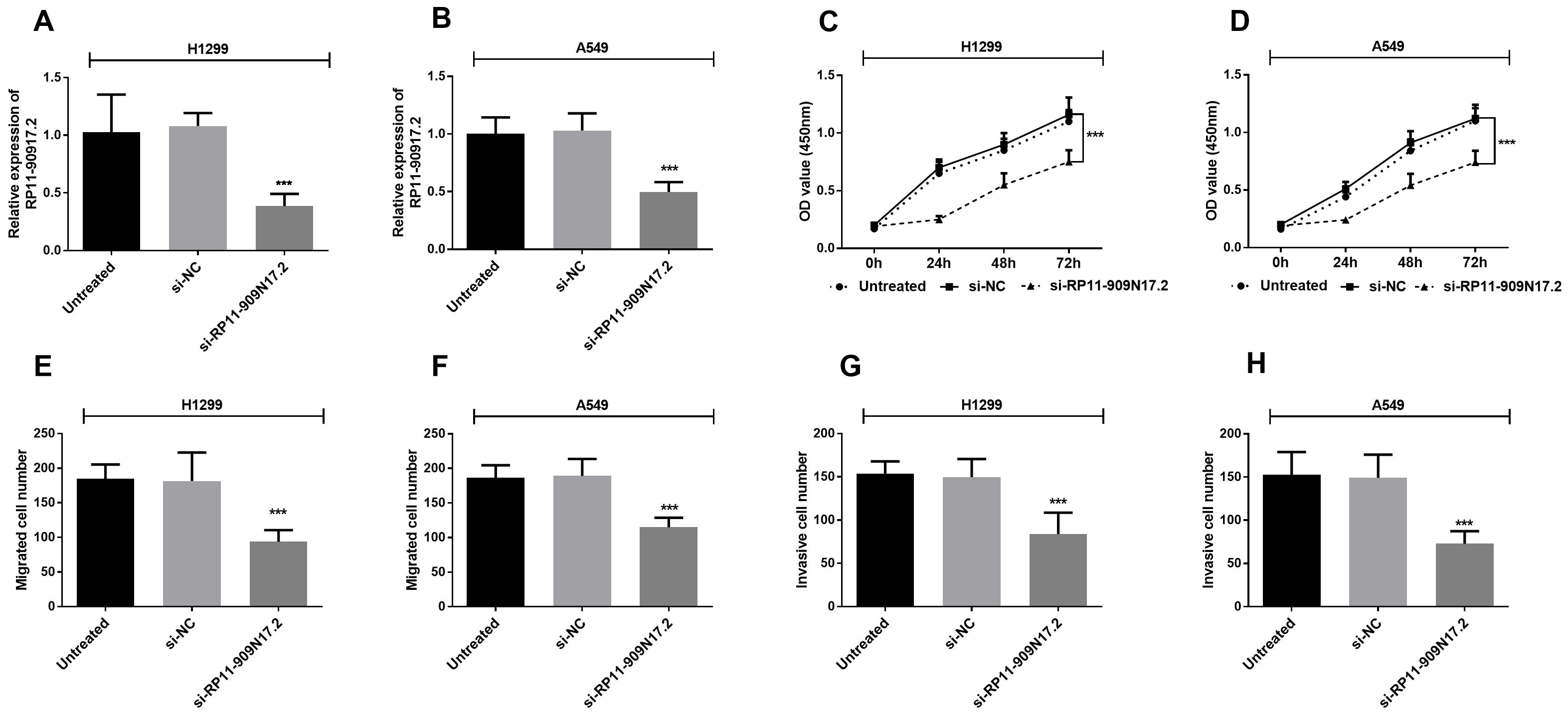
RP11-909N17.2 interacted with miR-767-3p. (A) The binding site between RP11-909N17.2 and miR-767-3p is shown; (B) Luciferase activity was examined in H1299 cell cotransfected with WT-RP11-909N17.2, or MUT-RP11-909N17.2 and miR-767-3p mimic or mimic NC. (C) H1299 cells were transfected with miR-767-3p mimics or negative control, followed by the measurement of RP11-909N17.2 enrichment with anti-Ago2 by RT-qPCR, and anti-IgG used as control. 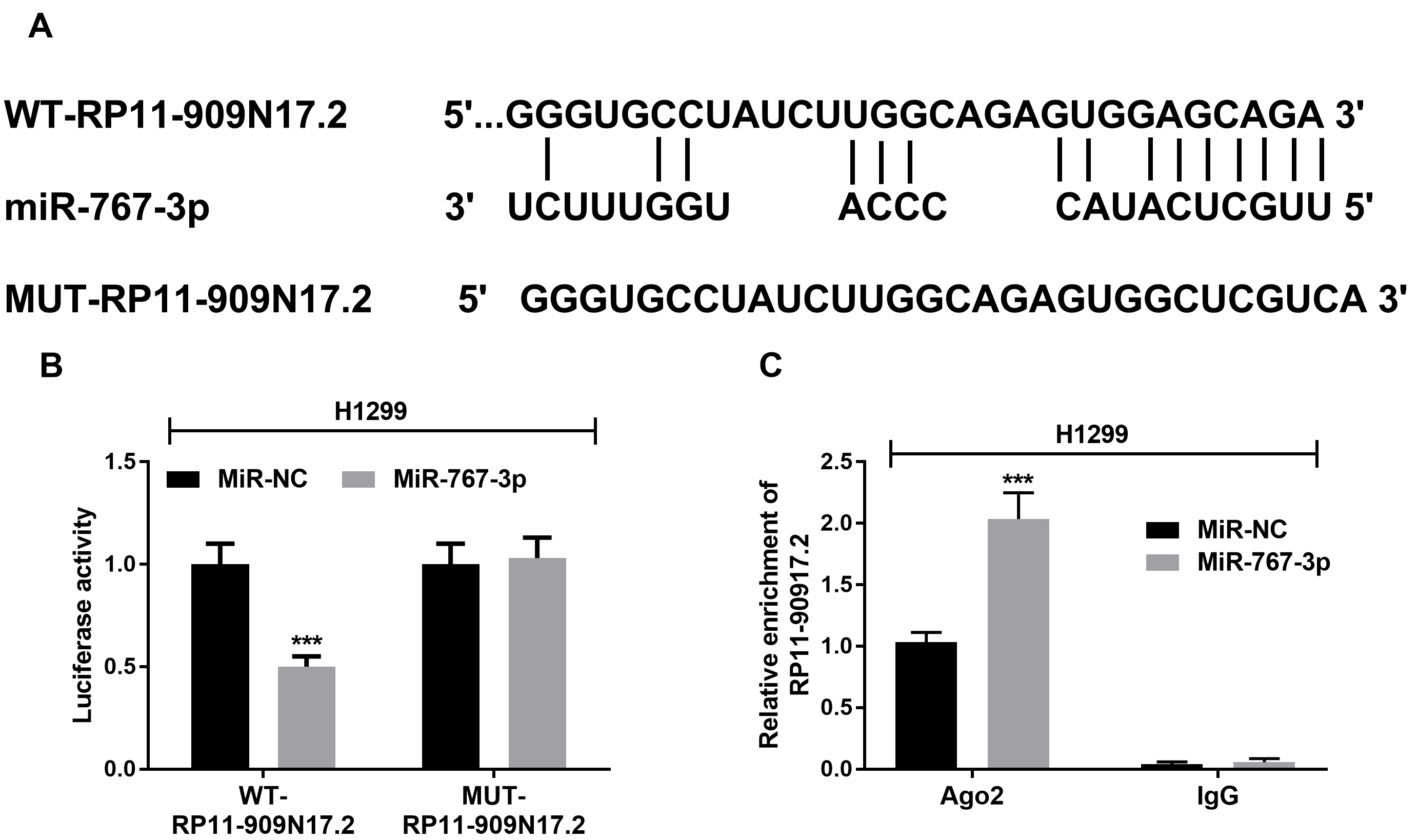
RP11-909N17.2 silence suppresses the proliferation, migration and invasion in NSCLC H1299 cells by interacting with miR-767-3p. H1299 cells were transfected with si-NC or si-RP11-909N17.2 separately, or co-transfected with si-RP11-909N17.2+anti-miR-NC or si-RP11-909N17.2+anti-miR-767-3p separately. (A) Expression of miR-767-3p was measured using RT-qPCR; (B) Cell proliferation was measured by CCK-8; (C) Cell migration was analyzed using transwell; (D) Cell invasion was analyzed using transwell. 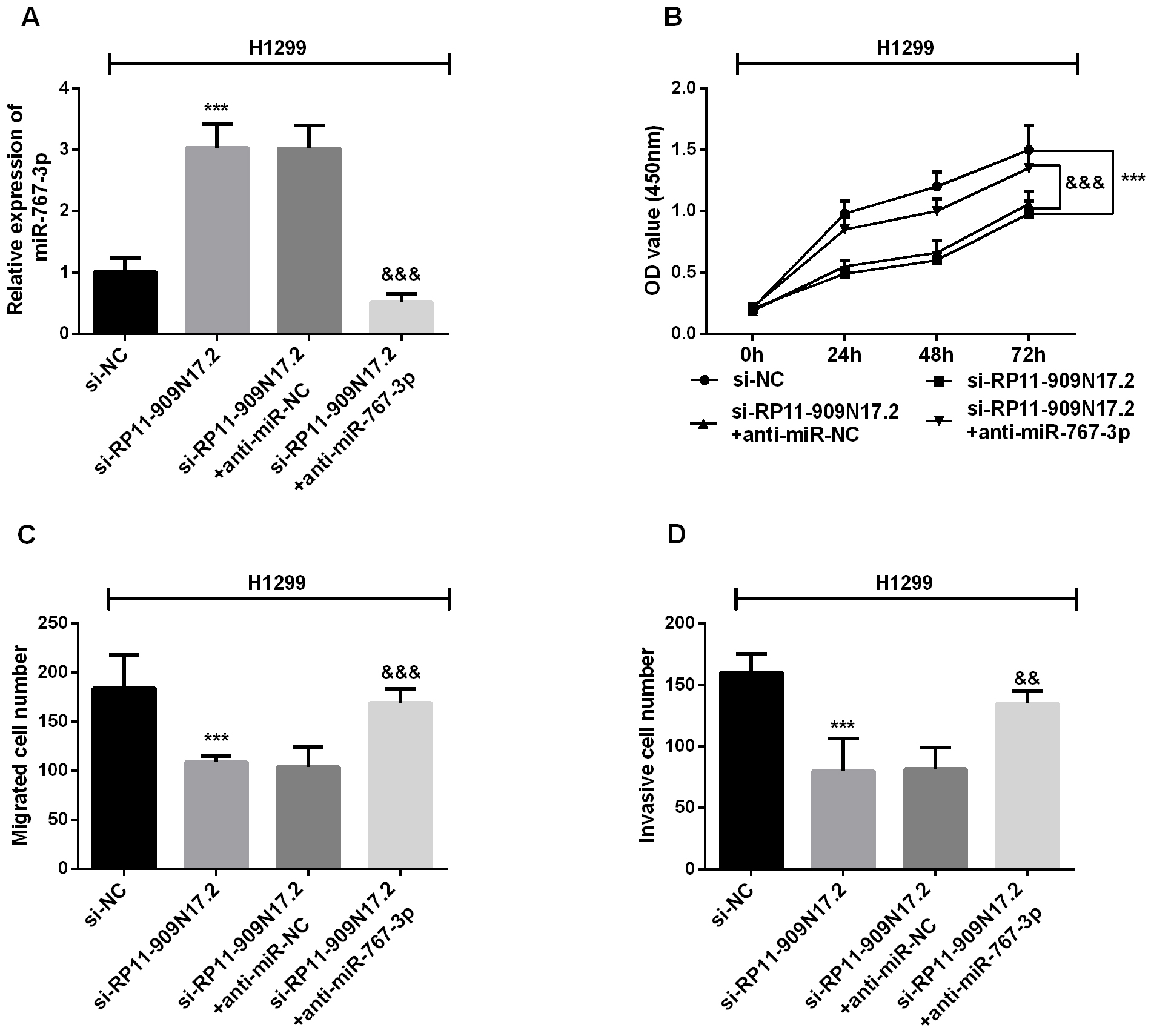
To explore the function of RP11-909N17.2 and miRNA-767-3p on NSCLC, H1299 and A549 cell lines, in which RP11-909N17.2 has an especially high expression, were selected. Among the miRNAs predicted fom LncBase Predicted v.2, miRNA-767-3p, which is a potential tumor suppressor in lung adenocarcinoma, was one of the target miRNAs of RP11-909N17.2. The binding sites of RP11-909N17.2 and miRNA-767-3p were predicted as Fig. 4A. Subsequently, a mutated RP11-909N17.2 construct was obtained by mutating the binding sites on RP11-909N17.2. According to the luciferase assay, the luciferase activity of the mutated-RP11-909N17.2 transfected cells was not affected by the expression of miRNA-767-3p, while the luciferase activity of the wildtype-transfected cells was significantly inhibited by the increased expression of miRNA-767-3p (Fig. 4B). To further validate this interaction, we conducted RIP assay in H1299 cells, and observed that RP11-909N17.2 was specifically enriched by miR-767-3p overexpression with anti-Ago2 by qRT–PCR analysis (Fig. 4C). This suggested that miRNA-767-3p may bind to the predicted sites and then inhibit the expression of RP11-909N17.2.
Due to the correlation between RP11-909N17.2 and miR-767-3p, the RP11-909N17.2/miR-767-3p axis was considered possible in the regulation of NSCLC cellular function. Then H1299 cells were transfected or co-transfected with si-NC, si-RP11-909N17.2, si-RP11-909N17.2+anti-miR-NC, or si- RP11-909N17.2+anti-miR-767-3p. The qRT-PCR results showed that miR-767-3p expression was significantly increased if RP11-909N17.2 was silenced, but this increase was counteracted after miR-767-3p si-RNA involved in (Fig. 5A). Then, CCK-8 assay demonstrated that H1299 cell proliferation was promoted by RP11-909N17.2 knockdown (Fig. 5B). But, when miR-767-3p inhibitor was co-transfected, the inhibition caused by overexpression of RP11-909N17.2 can be recovered, which implied an interaction between RP11-909N17.2 and miR-767-3p. Consistent with the CCK-8 assay, transwell assay revealed that the inhibitory effect of si-RP11-909N17.2 on H1299 cell growth and invasion can be partially restored by miR-767-3p (Fig. 5C and D).
Discussion
As the most common type of cancer affecting the lungs, most patients with NSCLC usually lead to poor treatment outcomes [18]. Thus, the novel prognosis factors are key points for assessment of treatment and can lead more therapy options for NSCLC patients. For years, many studies have revealed that exosomes play a key role in many physiological processes of cancer cells [19, 20]. miRNAs and lncRNAs are important constituents of exosomes in cell signaling, and usually considered as cancer-specific biomarkers [21, 22, 23]. Recently, mounting evidence has revealed that lncRNAs play essential roles in cancer, either as tumor suppressors or oncogenes [24, 25]. Wu et al. reported that lnc-RAB1A-2 has a crucial functional role in lung cancer procession and may be a potential prognostic marker for lung cancer [26], based on the study of Zhang et al. who screening out 5563 novel lncRNAs including lnc-RAB1A-2 in mouse [27].
RP11-909N17.2, one kind of antisense LncRNA, was located in Chromosome 8. He et al. have reported that RP11-909N17.2 was identified as a high-risk candidate among the related genes in lung cancer [15]. In line with that, this study found RP11-909N17.2 have an elevated expression level analyzed by RT-qPCR both in NSCLC tissues and cell lines, when compared with normal ones. Subsequent, this study explored the potentiality of RP11-909N17.2 as prognostic biomarker in NSCLC by Kaplan-Meier curve. The log-rank test, using
LncRNAs are often reported to influence the biological functions of various cancer cells. For instance, SIK1-LNC has been reported to suppress the proliferation, migration, and invasion lung cancer cells [28]; lnc-ABCA12-3 was discovered to promotes cell biological function in esophageal squamous cell [29]. To clarify the function of RP11-909N17.2 in NSCLC cells, H1299 and A549 were transfected with si-NC and si-RP11-909N17.2 and a series of cell biological function identification were performed. The transfection efficiency was confirmed by the expression of RP11-909N17.2 using RT-qPCR. Significantly, the proliferation of NSCLC cells was repressed by RP11-909N17.2 knockdown obtained from CCK-8 assay. Meanwhile, suppression was also found in migration and invasion of cells transfected with RP11-909N17.2 si-RNA demonstrated from the result of transwell assay. Thus, inhibition of RP11-909N17.2 has the potential ability to suppress the biological functions of NSCLC cell, including proliferation, migration, and invasion.
Furthermore, the mechanism of RP11-909N17.2 action on cell function was studied. It is common knowledge that lncRNAs can function as miRNA sponges, thus decreasing their ability to target messenger RNAs and influencing their expression. Previously, RP11-909N17.2 was found to sponge miR-767-3p, thereby downregulating miRNA-767-3p expression level in hepatocellular carcinoma (HCC) cell lines [17]. In this study, expression of miRNA-767-3p was analyzed both in NSCLC tumor tissues and cell lines. And a significant downregulation of miRNA-767-3p expression was also found in NSCLC tumor tissues and cells. So, we speculate RP11-909N17.2 may bind to miRNA-767-3p to achieve the results of regulating cell biological function. To confirm, luciferase activity was measured in different co-transfected NSCLSC cells, and found the WT-RP11-909N17.2 markedly inhibited the luciferase activity driven by miRNA-767-3p plasmids. Our study revealed that miRNA-767-3p was a target of RP11-909N17.2, and there involved a reciprocal regulation of RP11-909N17.2 and miRNA-767-3p in NSCLC tissues and cells. Subsequently, the rescue assay supported this relationship between RP11-909N17.2 and miRNA-767-3p. A reduction of cell proliferative capacity was shown when RP11-909N17.2 was knockdown, but the reduction can be partially restored if miRNA-767-3p was inhibited. Similar reduction and recovery can be found in cell migration and invasion. Mutual corroboration between expression assay and cell function can give the information that there exist interactions between RP11-909N17.2 and miRNA-767-3p. Combined with the research of Liu et al. [17], we conclude that RP11-909N17.2 can act as a sponge adsorbing miRNA-767-3p, thereby promoting cell proliferation, migration and invasion of NSCLC cells.
In brief, our study proposed that RP11-909N17.2 is upregulated in NSCLC tissues and cells while miRNA-767-3p is downrgulated. Over-expression RP11-909N17.2, acting as miRNA-767-3p sponge, has the potential to promotes NSCLC cell proliferation, migration, and invasion by binding to miRNA-767-3p. The survival time of NSCLC patients with high RP11-909N17.2 expression is relatively shorter compared than those with low expression. Therefore, it may shed light on new ideas to predicting prognosis and molecular therapy of NSCLC.
