Abstract
BACKGROUND:
MicroRNAs constitute promising biomarkers.
OBJECTIVE:
The aim was to investigate diagnostic and prognostic implications of miR-182-5p and miR-205-5p in p16-positive and p16-negative oropharyngeal squamous cell carcinomas (OPSCCs).
METHODS:
Expression of miR-182-5p, miR-205-5p were determined via quantitative real-time-PCR in fresh frozen tissues of 26 p16-positive, 19 p16-negative OPSCCs and 18 HPV-negative oropharyngeal controls. Associations between miRNA-expression, clinicopathological characteristics and prognosis were analyzed.
RESULTS:
Higher miR-182-5p expression was associated with significant inferior disease-specific survival for p16-positive OPSCCs (HR
Higher miR-205-5p expression was associated with an inferior progression-free survival (HR
CONCLUSIONS:
Results indicate that miR-182-5p and miR-205-5p can further stratify patients with p16-positive OPSCC into prognostic groups.
Introduction
An increasing subset of oropharyngeal squamous cell carcinomas (OPSCC) is associated with human papillomavirus (HPV) infections and/or overexpression of the protein p16 [1, 2]. The latter is considered as a surrogate marker to detect high-risk-HPV-infections [3, 4, 5].
Distinction between HPV/p16-positive and HPV/p16-negative OPSCCs was recently introduced to the 4th edition of the World Health Organization classification for malignant tumors [5] and the 8th edition of the Union International Contre le Cancer (UICC) and the American Joint Committee on Cancer (AJCC) cancer staging manual [1, 3]. This is consistent with demographic, molecular, morphological, clinical and prognostic diversities of HPV/p16-positive or -negative OPSCCs [3, 5]. Nevertheless, trials having investigated modified therapeutic strategies for HPV/p16-positive OPSCCs [6, 7] have so far not resulted in altered standard treatment recommendations [8].
Additional biomarkers could provide new insights into the pathobiological basis underlying HPV/p16-positive and HPV/p16-negative OPSCCs, and therefore might also help to establish a more precise and individualized oncological management of patients with OPSCCs [8, 9, 10].
MicroRNAs (miRNAs, miRs) are approximately 22-nucleotide-long, single stranded RNAs [11]. Hybridizing with complementary sequences of messenger RNAs, they can epigenetically influence the expression of multiple genes [9, 11]. Dysregulation of miRNAs was related to pathogenesis, progression, immune and treatment response of tumors [9, 10, 11], but also to viral infections like HPV [12, 13]. Combined with their stable tumor-type specific expression, stability, and reliable detectability in clinical specimens, they were considered as potential biomarkers, as well as therapeutic targets for various cancers [9, 11, 14].
To date, only a few studies have been investigating the association of miRNAs with clinicopathological characteristics and prognosis within a uniform OPSCC cohort, and in addition, considering the tumor’s p16-/HPV-status [15, 16, 17, 18, 19, 20, 21, 22]. None of them focused on miR-182 or miR-205. Both were shown to modulate important carcinogenic genes and pathways in HNSCCs (miR-182-5p [23, 24, 25, 26, 27, 28, 29]; miR-205-5p [30, 31, 32, 33, 34, 35]). In particular both miRNAs were discussed in context with TP53 mutation and dysregulation (miR-182 [28, 36, 37]; miR-205-5p [36, 38]), which is common in p16/HPV-negative OPSCCs, but untypical for p16/HPV-positive OPSCCs [39]. Additionally, both were discussed in context with HPV-infections of cells (miR-182-5p [12, 40]; miR-205-5p [13]). Furthermore, both miRNAs were suggested as miRNAs of potential prognostic relevance in HNSCCs (miR-182-5p [16]; miR-205-5p [36, 41]) and a variety of other cancers (miR-182-5p [42, 43, 44, 45, 46, 47]; miR-205-5p [48, 49, 50, 51, 52, 53]). Even though to the best of our knowledge the potential prognostic implications of both as single markers remained so far unexplored in a selective OPSCC dataset. Therefore, the aim of the present study was to investigate the diagnostic and prognostic implications of miR-182-5p and miR-205-5p in a well characterized cohort of patients with p16-positive and p16-negative OPSCCs treated with curative intent.
Patients and methods
Patients and specimens
Patients with OPSCC, who received treatment with curative intent at an academic tertiary referral center were included. Time span of inclusion was from 07/2011 to 04/2016, with subsequent systematic follow-up. Cases were identified, and data obtained from the hospital’s cancer database and original medical records. It was analyzed anonymously. The study was conducted according to the criteria of the Declaration of Helsinki and was approved by the institutional review board (Ethics Committee of the University Medical Center Göttingen) according to the national regulations (application number: 9/12/10). Written informed consent for anonymized analysis of their data and tissues was obtained from all patients prior to treatment at the University Medical Center Göttingen.
Due to the aim of assessing oncologic outcomes in the setting of curative treatment the following patients were excluded from analysis: one patient was excluded due to distant metastasis, two patients, who were treated with palliative intent, one patient due to complete rejection of any therapy, and further two patients due to lack of follow-up data of at least twelve months after primary diagnosis. Furthermore, of 34 cases no sufficient material for p16-immunohistochemistry and micro-RNA quantification was available. Subsequently, 45 patients with histopathological proven OPSCC were included to the study. Tumors’ main sublocalizations were assigned according to the predominantly infiltrated region by the surgeon. Control specimens were obtained from 18 non-cancer patients, who underwent tonsillectomy.
Staging, treatment and follow-up
Regarding staging, the 7th edition of classification of the UICC [54] and the AJCC [55] was applied.
The therapeutic strategy was primary radiotherapy in 8.9% (
Patients received a systematic follow-up after treatment for at least five years. After 60 months without recurrence a patient was considered as cured, but follow-up examinations continued on a regular basis.
Tissue analyses
All included malignant tissue was harvested during initial panendoscopy and biopsy or surgery of the primary tumor. Fresh frozen (FF) tissues were snap-frozen in liquid nitrogen and stored at
p16-immunohistochemistry and HPV-typing
For tumor cases without an assessment for p16-status at time of initial diagnosis, p16-immunohistochemistry was performed using the FFPE tissue of the oropharyngeal primaries. Monoclonal p16 antibodies (p16 (JC8): sc-56330, Santa Cruz Biotechnology, Inc., Dallas, TX) (1:50; pH 9) combined with diaminobenzidine and the EnVision Flex
In the non-cancerous control group HPV-genotyping was performed in order to rule out HPV-infections. Therefore, DNA was isolated from FFPE tissues using innuPREP FFPE DNA Kit – IPC16 combined with the InnuPure
RNA extraction, cDNA synthesis, microRNA quantification
Expression levels of hsa-miR-182-5p (UUUGGCAAUGGUAGAACUCACACU), hsa-miR-205-5p (UCCUUCAUUCCACCGGAGUCUG), and the housekeeping miRNA miR-103a-3p (AGCAGCAUUGUACAGGGCUAUGA) were determined using quantitative real-time-PCR (qRT-PCR). The middle section of 21 slices of the FF tissues was stained with hematoxylin and eosin and was considered as representative to estimate the amount of tumor cells (
MiRNA cDNA was synthesized using the Applied Biosystems™ TaqMan™ Advanced miRNA cDNA Synthesis Kit. Finally, the expression of miRNAs was assessed by qRT-PCR with TaqMan™ Fast Advanced Master Mix technology combined with specific miRNA Assays (all Thermo Fischer Scientific Inc., Waltham, MA; Assay-ID: 478253_mir (hsa-miR-103a-3p), 477935_mir (hsa-miR-182-5p), 477967_mir (hsa-miR-205-5p)). All qRT-PCRs were conducted in duplicates. Exclusively, if the discrepancy between the duplicates was less than one cycle, the mean of the threshold cycle (CT) values was included in the further analysis. For other cases, the qRT-PCR was repeated. Among all samples the maximum detected CT value was 35 of 40 total cycles. miRNA expression was normalized to the housekeeping miRNA miR-103a-3p. This miRNA was recommended for normalization by Thermo Fisher Scientific.
Statistics
For descriptive analysis, the mean value with corresponding standard deviation, median, minimum and maximum or frequencies with corresponding proportions in percentages were calculated for each variable. Clinicopathological parameters were tested for differences between p16-positive and p16-negative patients using Pearson’s Chi Squared Test.
The Kaplan-Meier method [57] was applied for calculation of overall survival (OS), disease-specific survival (DSS), progression-free survival (PFS), recurrence-free survival (RFS) and local control rate (LCR). The date of initial histopathologic diagnosis was defined as the starting point for interval calculations. Recurrence was defined as the same tumor entity occurring not earlier than 3 months after initial diagnosis, otherwise it was considered as residual disease with subsequent completion of treatment. With regard to OS, death from any possible cause was counted as an event and all other observations were considered as censored. Regarding DSS, exclusively death related to the oropharyngeal primary tumor was considered as an event, while all other observations were censored. Calculating the PFS, events were defined as local, regional, or locoregional recurrence, as well as distant metastasis and death from all causes. Loss of follow-up was defined as censored observations. Regarding RFS, local, regional, or locoregional recurrence, distant metastasis and death related to the oropharyngeal primary tumor were counted as events. Correspondingly, death from other causes, as well as loss of follow-up were considered as censored. Referring to the LCR, local recurrence was considered as an event, whereas alive patients without local recurrence or death regardless the reason were accounted as censored. The association of variables with the relative risk of having an event was assessed by univariate Cox regression proportional hazard models. Calculated hazard ratios (HR) are shown with the respective 95% confidence interval (CI). Log-rank test was applied in order to test for differences of risks between comparison groups.
Relative differences of miRNA expression levels between tumors and benign controls were calculated applying the 2
Statistica 13.3 (Dell Inc., Round Rock, Texas) was used to calculate Spearman’s rank correlation, Mann-Whitney-U test, Kruskal-Wallis test, Pearson’s Chi-squared test and Kaplan-Meier estimates. Survival analysis and visualization was conducted using the R statistical platform (R Core Team, R: A Language and Environment for Statistical Computing, R Foundation for Statistical Computing, version 3.5.1 [July 2, 2018]) in combination with the survival and Survminer R packages. Box plots were generated using GraphPad Prism Version 7.0e (Graph Pad Software Inc., San Diego).
miRNA
Results
Patients and therapy
This study included 45 patients diagnosed with OPSCC. Oropharyngeal tissue of 18 patients with a non-malignant diagnosis served as a control group.
Primary tumors originated from tonsillar regions (51.1%,
The distribution of patient, disease, and treatment characteristics within the complete cohort and p16-stratified subgroups is depicted in Table 1. As shown, patients with p16-positive vs. p16-negative OPSCC shared the same baseline characteristics (
Patient and disease characteristics and follow-up data of the complete cohort and the p16-stratified subgroups
Patient and disease characteristics and follow-up data of the complete cohort and the p16-stratified subgroups
Abbreviations: SD, standard deviation; min, minimum; max, maximum; c, clinical; p, pathological; (C)RT, (chemo) radiotherapy; ND, neck dissection; TLM, transoral laser microsurgery; OPSCC, oropharyngeal squamous cell carcinoma;
Median follow up was 26.2 months
miR-182-5p and miR-205-5p expression in tumor versus normal tissues
Based on a literature research [23, 24, 25, 26, 27, 28, 29, 30, 31, 32, 33, 34, 35, 36, 41, 42, 43, 44, 45, 46, 47, 49, 50, 51, 52, 53, 59, 60, 61, 62], miR-182-5p and miR-205-5p were selected for this study. To analyze the potential dysregulation of miR-182-5p and miR-205-5p in malignant tissue, expressions were examined in FF samples of 26 p16-positive and 19 p16-negative primary OPSCCs, as well as 18 HPV-negative control specimens.
Compared to benign controls, expression of miR-182-5p showed no significant differences within either the complete cohort of tumors, or the p16-stratified subgroups (complete cohort: relative fold change (RFC)
In contrast, miR-205-5p was significantly overexpressed in tumor tissues compared to benign controls in both, the complete cohort as well as the p16-negative subgroup (complete cohort: RFC
Figure 1 illustrates the expression of miR-182-5p and miR-205-5p in the complete cohort as well as the p16-stratified subgroups compared to benign controls, respectively.
miRNAs and clinicopathological characteristics
In order to examine associations between miR-182 -5p and miR-205-5p and clinicopathological characteristics within the complete cohort as well as the p16-stratified subgroups the Mann-Whitney-U-Test, Kruskal-Wallis-Test and Spearman’s rank correlation test were applied.
Associations of miR-182-5p and miR-205-5p with clinicopathological characteristics in the complete cohort as well as in the p16-stratified subgroups
Associations of miR-182-5p and miR-205-5p with clinicopathological characteristics in the complete cohort as well as in the p16-stratified subgroups
Abbreviations: miR, microRNA;
Expression of miR-182-5p in oropharyngeal squamous cell carcinoma (OPSCC) tissue compared to benign control tissue in the complete cohort (A), p16-positive subgroup (B), and p16-negative subgroup (C), respectively. Expression of miR-205-5p in OPSCC tissues compared to benign control tissues in the complete cohort (D), p16-positive subgroup (E), p16-negative subgroup (F), respectively. The relative fold change (RFC) was computed with the 2
As shown in Table 2, among each of the three subgroups no associations were found between the expression levels of miR-182-5p and miR-205-5p with respect to sex, age, T-categorization (T1 vs. T2 vs. T3 vs. T4a/b), local tumor extent (circumscribed T1-2 vs. advanced T3-4), N-categorization (N0 vs. N1 vs. N2 vs. N3), lymph node metastases (N0 vs. N
Patients with a positive history of alcohol consumption exhibited significant higher miR-182-5p levels compared to those patients, who had negated any drinking history (
After dichotomization according to high or low levels of miR-182-5p and miR-205-5p expression based on a median separation, the survival differences between the resulting groups were analyzed within the complete cohort as well as the p16-stratified subgroups, respectively. The corresponding Kaplan-Meier curves are displayed in Fig. 2 (miR-182-5p) and Fig. 3 (miR-205-5p).
Association between high (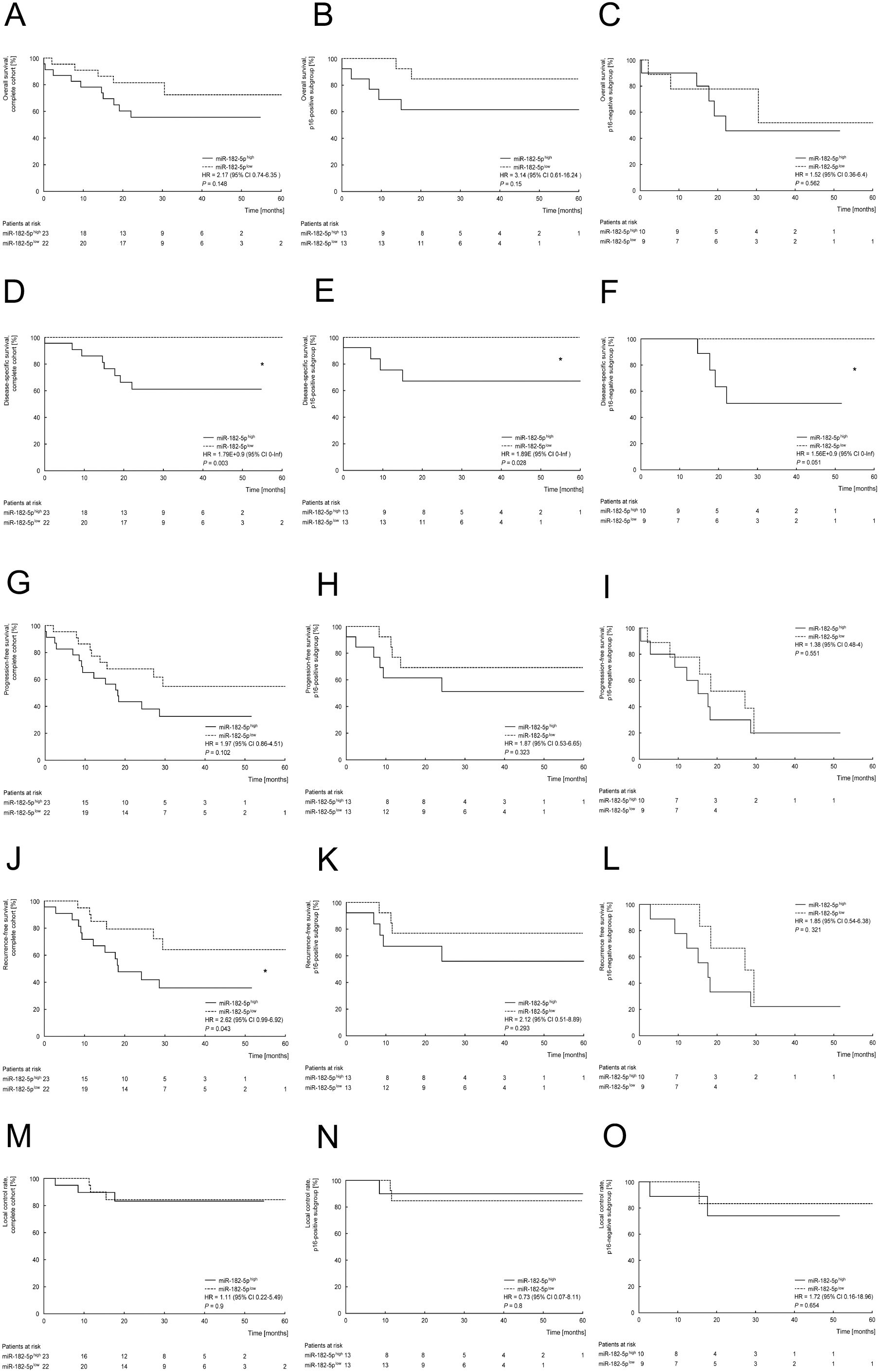
Association between high (
High expression levels of miR-182-5p were associated with a significant inferior DSS and RFS within the complete cohort (DSS: HR
With regard to miR-205-5p, patients with higher expression levels demonstrated a significant inferior PFS, RFS, as well as LCR within the complete cohort (PFS: HR
Association between high (
Abbreviations: 95% CI
Table 3 summarizes the analysis of miR-182-5p and miR-205-5p for prognostic stratification of the complete cohort as well as the p16-stratified subgroups.
In order to identify potential prognostic relevant cofounders, the association of clinicopathological characteristics and survival endpoints were analyzed. Therefore, patients were stratified according to clinicopathological characteristics, that were also analyzed for difference of miRNA expression and their association with prognosis. Results are illustrated in Supplementary Table S1. Referring to the complete cohort, lower histopathological differentiation (G1-2 vs. G3) was significantly associated with an inferior OS (HR
In the p16-positive subgroup, nodal involvement (N
In the p16-negative subgroup an advanced T-category (c/pT1-2 vs. c/pT3-4a/b) was significantly associated with worse PFS and RFS (PFS: HR
Finally, comparing all these results (association of clinicopathological characteristics with prognosis; Supplementary Table S1) with the univariate analyses of the miRNAs (association of miRNA expression with prognosis; Table 3), there was no overlap between a significant association of any of the clinicopathological characteristics, or one of the miRNAs with one of the prognostic endpoints in the same group, respectively. Therefore, the significant associations observed in the univariate analysis between miR-182-5p or miR-205-5p and prognosis were considered as independent of any of the tested clinicopathological characteristics and thus, no further multivariate testing was performed.
Discussion
An increasing subset of OPSCC is associated with HPV-infections [1, 63, 64, 65]. Corresponding to the observed prognostic superiority of patients with p16-/HPV-positive disease, trials investigated altered therapeutic treatment strategies for “HPV associated” OPSCCs [6, 7]. Nevertheless, to which extent the HPV/p16-status can be applied for appropriate patient selection is still widely debated [8, 66, 67].
Additional biomarkers could help in order to refine the prognostic value of HPV/p16 and provide more detailed information about the molecular [15, 16, 20] and clinical [68] diversity between HPV/p16-positive and HPV/p16-negative OPSCCs [8, 9, 10].
MiRNAs were described as potential important players in carcinogenesis, immune, and treatment response [9, 10, 11] that are also dysregulated in the course of HPV-infections [12, 13]. Combined with their relatively stable tumor-type specific expression, as well as stability and reliable detectability in clinical specimens, they were suggested as potential biomarkers and therapeutic targets for various cancers [9, 11].
A limited number of studies investigated miRNAs’ associations with clinicopathological characteristics and prognosis within a uniform OPSCC cohort with in addition considering the tumor’s p16- or HPV-status [15, 16, 17, 18, 19, 20, 21, 22]. The current study analyzed the diagnostic and prognostic implications of miR-182-5p and miR-205-5p in a well characterized, p16-stratified cohort of 45 patients with OPSCCs treated with curative intent.
miR-182
miR-182 was considered to act oncogenic but also tumor-suppressive in HNSCC. For example higher expression of this miRNA was indicated to enhance cell-cycle progression, colony formation, invasiveness and proliferative capacity e.g. by directly suppressing genes like RASA1 and SPRED1, leading to an activation of the Ras-MEK-ERK pathway [27]. The proliferation of tumor cells could also be affected by miR-182-5p through targeting genes like CAMK2N1 [25]. Another way miR-182 could mediate cell migration and the evasion of invasion control and thereby influence the evolution and progression of HNSCCs is by suppress the gene of the Metastasis Suppressor Protein 1 [24]. Nevertheless, enhanced proliferative, migratory, or invasive ability of HNSCC cells, which was associated with miR-182 expression could be reverted by an induced expression of the genes
In the present study miR-182-5p was not significantly dysregulated between OPSCCs and benign tissues, irrespective of the p16-status. Unaltered tissue levels of miR-182-5p are in accordance with another study focusing on OPSCCs [16]. In contrast, miR-182 was described to be downregulated in tissue and cell lines of laryngeal cancer (LSCC) [26]. Whereas, an upregulation was observed in cell lines [24, 25, 27] and specimens deriving from oral SCC (OSCC) [24, 25, 27, 69], as well as pooled HNSCC datasets [23, 28, 36, 70]. Interestingly, Ganci et al. observed a significant upregulation of miR-182-5p in TP53-mutated only, and not in the TP53 wild-type HNSCC [36]. Consistent with this, Wang et al. described an upregulation of miR-182 predominantly in p53-overexpressing HNSCC tumor tissue [28]. Therefore, this potential dependency of miR-182-5p expression and tumor’s TP53/p53-status, suggests a promising approach for further investigations of the differences of miR-182-5p dysregulation and its role in cancer of the head and neck region.
Furthermore, in vitro and in vivo experiments indicated, that the oncoprotein E7 of high-risk HPV is able to cause an upregulation of miR-182 in tumor, as well as adjacent normal cells via activating the TGF-
In contrast to the current study, associations between miR-182-5p with poor differentiation, advanced T-category, and lymph node metastases were observed within an OSCC dataset [24], indicating that miR-182-5p might serve as a marker for a more aggressive diseases. These aspects are consistent with the above-mentioned oncogenic potential of miR-182 [23, 24, 25, 27, 28], its association with an enhanced proliferation, cell-cycle progression, invasiveness, colony formation, migration, and reduced apoptosis in vitro [25, 27, 28], as well as an extended tumor volume, and growth rate in vivo [25, 28]. Moreover, it is in accordance with the herein observed association between higher levels of miR-182-5p with an inferior DSS and RFS in the complete cohort, an unfavorable DSS within the p16-positive subgroup, and a similar trend in the p16-negative subgroup. This is in line with a meta-analysis focusing on different types of cancer [46] and the inverse association between miR-182 with prognosis, e.g. within patients with glioma [44], prostate [43], hepatocellular [42], and colorectal [45, 47] cancer. Nevertheless, it contradicts the positive prognostic association found in, e.g. patients with NSCLC [59].
In summary, the results of the present study indicate, that although miR-182-5p was not dysregulated in OPSCCs compared to benign tissues, higher expression levels in OPSCC at initial diagnosis may help to identify patients with an inferior prognosis. Consequently, they could be excluded from, e.g. de-escalation trials for p16-positive OPSCCs.
miR-205
Within HNSCC miR-205 could act tumor-suppressive via targeting genes like Tissue Inhibitor of Metallopro- teinases-2 leading to a reduced activation of Pro-matrix Metalloproteinase-2 and finally in a reduced migration and invasion of cancer cells [30]. Furthermore miR-205 was negatively associated with the expression of proliferative markers like the dihydrofolate reductase and the proliferating cell nuclear antigen as well as the apoptotic regulator Bcl-2. In vivo, the induced expression of miR-205 was further associated with an increase in apoptotic cells in xenograft tumors [31]. Nevertheless, miR-205 could also positively influence the progression of HNSCCs, e.g. through inducing genomic instability by targeting the DNA repair proteins of BRCA1 and RAD17 [35] or leading to stemmness associated characteristics and an enhanced tumor growth via targeting COMMD1 [32]. Also through the repression of CDK2AP1 and the induction of proliferation and invasion, miR-205 could potentially contribute the progression of HNSCCs [33]. Last miR-205 was also discussed to negatively affect the response to radiotherapy by regulating the well-known tumor suppressor PTEN, leading to an activation of AKT in HNSCC [34].
In the current study, both, p16-positive and p16-negative OPSCCs exhibited at least a trend towards an upregulation of miR-205-5p compared to controls, indicating miR-205-5p elicits oncogenic functions in OPSCCs. These results are consistent with the previously described upregulation of miR-205 in tissue of laryngeal [33, 73, 74], oral [41, 69], nasopharyngeal carcinomas [75], as well as in tumor samples [62, 76, 77], sera [78], fine needle aspirations of lymph node metastases [79], and diverse cell lines [80, 81] of HNSCC. Interestingly, Ganci et al. identified an upregulation of miR-205-5p, as already described above for miR-182-5p, exclusively in TP53-mutated, but not TP53-wild-type HNSCC [36]. This corresponds to the observed upregulation of miR-205-5p in HNSCC cell lines, that overexpressed mutant p53 [38]. However, miR-205 was also considered as a marker for squamous epithelium irrespective of the tissue’s dignity [82], consistent with the unaltered levels of miR-205 observed, e.g. in OSCC [82, 83, 84]. Furthermore also downregulation of miR-205 was reported, e.g. in tissues of LSCCs [31], OSCCs [30], pooled HNSCC datasets [85]. Summarized, these results indicate a dysregulation of miR-205-5p possibly occuring in diverse HNSCCs, including p16-positive and p16-negative OPSCCs at time of initial diagnosis. However, in context with the reported ability of miR-205 to influence oncogenic, as well as tumour-suppressive genes and pathways, its potential role and therefore its up- or downregulation appears target- and tumor-specific.
Moreover, in vitro studies demonstrated an association between expression of HPV16’s oncoproteins E6 and E7 with depletion of miR-205 [13]. Although, in line with the present study, no altered tissue levels of miR-205 were reported between HPV/p16-positive vs. negative HNSCCs by others [15, 16, 62, 71, 72].
However, within the present study miR-205-5p was not significantly associated with any of other clinicopathological features within the p16-stratified subgroups and therefore was (as miR-182-5p) considered as independent from them. Nevertheless, for instance, in pooled HNSCC and LSCC datasets, high miR-205 expression levels were previously associated with absence of features that were considered indicative for more aggressive and moreover metastatic disease [31, 62]. In contrast, in a cohort including OSCCs, as well as OPSCCs an upregulation of miR-205 was significantly associated with lymph node metastases and a trend towards association with higher pathological stage [60], supporting the consideration of miR-205 as a marker for more advanced and aggressive tumors.
This is also consistent with the above described oncogenic potential of miR-205 [32, 34, 35], its association with a malign cellular phenotype [32, 34, 35], its upregulation in tumor tissues, and its association with an inferior prognosis within the current study’s complete cohort (PFS, RFS, LCR), as well as p16-positive subgroup (PFS, LCR). Fittingly, miR-205-5p expression alone, as well as in a pattern with other miRNAs, was identified as a predictor for lower RFS of HNSCC patients independent from other clinicopathological characteristics [36]. Furthermore, an inverse association between miR-205 and prognosis was reported for patients with OSCC [41], as well as outside the HN-region, e.g. for patients with endometrial [53], ovarian [51], non-small cell lung [48], esophageal SC [52] carcinomas. In contrast, higher miR-205 levels were associated with a beneficial prognosis of patients with, e.g. colon [49], prostate [50], breast [53] cancer.
Summarized, the comparison of miR-205-5p expression in tumor vs. control tissues supports that higher miR-205 levels could help to detect metastatic HNSCC cells within lymph nodes [86], or residual tumor cells at the resection margin [77]. Furthermore, results indicate that higher miR-205-5p levels could help in identifying patients with p16-positive OPSCCs and an inferior prognosis.
Strengths and limitations
In accordance with the 8th edition of the AJCC/UICC cancer staging manual [1, 3] and the 2017 (4th) edition of World Health Organization classification of malignant tumors [5], the need for distinction between HPV/p16-positive and HPV/p16-negative OPSCCs was addressed in the current study. Therefore, the strength of the study lies in the p16-stratified analyses of miRNAs based on FF tissue of the local primaries, strict clinical and laboratory inclusion criteria, and detailed clinical data, which was added by results of further pathological assessment when necessary. This way we were able to provide information to both, potential diagnostic and prognostic implications of miR-182-5p and miR-205-5p.
Limitations of this study are the small sample size, the paucity of comparable analyses in literature, and the absence of an adequate validation cohort. Potential causes for variability and sometimes conflicting results of miRNA analyses include for instance distributions of ethnicity, lifestyle factors like tobacco or alcohol consumption, and other clinicopathological characteristics like p16-/HPV-status within the study populations. Also, basic methodical differences like detection methods (microarray vs. qPCR vs. ISH etc.), type (FFPE vs. FF vs. cell lines vs. saliva etc.) and origin of the investigated biomaterials (pooled datasets vs. specific sites), as well as control groups (non-cancer patients’ specimen vs. adjacent normal tissue from cancer patients) may have contributed to partly diverging results.
Prospective validation in extended cohorts, the examination of miR-182-5p and miR-205 in the context of recurrent or metastatic disease, as well as analysis in different biomaterial (e.g. non-invasive acquirable saliva samples) could further emphasize the translational relevance of the current study’s results. Our findings could serve as a basis for further research on miRNA-based biomarkers in p16-positive and p16-negative OPSCCs.
Conclusion
In conclusion, miR-182-5p and miR-205-5p further stratified patients with p16-positive OPSCC into prognostic groups. These findings may have implications for the development of a more precise and individualized oncological management.
Author contributions
Conception: Bernhard G. Weiss, Mahalia Zoe Anczykowski, Julia Kitz, Mark Jakob.
Interpretation or analysis of data: Bernhard G. Weiss, Mahalia Zoe Anczykowski, Friedrich Ihler, Mattis Bertlich, Jennifer L. Spiegel, Frank Haubner, Martin Canis, Stefan Küffer, Julia Hess, Kristian Unger, Julia Kitz, Mark Jakob.
Preparation of the manuscript: Bernhard G. Weiss, Mahalia Zoe Anczykowski, Jennifer L. Spiegel.
Revision for important intellectual content: Bernhard G. Weiss, Mahalia Zoe Anczykowski, Friedrich Ihler, Mattis Bertlich, Jennifer L. Spiegel, Frank Haubner, Martin Canis, Stefan Küffer, Julia Hess, Kristian Unger, Julia Kitz, Mark Jakob.
Supervision: Bernhard G. Weiss, Mahalia Zoe Anczykowski, Julia Kitz, Mark Jakob.
Supplementary data
The supplementary files are available to download from http://dx.doi.org/10.3233/CBM-203149.
sj-docx-1-cbm-10.3233_CBM-203149.docx - Supplemental material
Supplemental material, sj-docx-1-cbm-10.3233_CBM-203149.docx
sj-pdf-1-cbm-10.3233_CBM-203149.pdf - Supplemental material
Supplemental material, sj-pdf-1-cbm-10.3233_CBM-203149.pdf
