Abstract
OBJECTIVE:
This study investigates expressions of circ0001429, miR-205-5p and vascular endothelial growth factor (VEGFA) in bladder cancer tissues and their effects on the proliferation, migration and apoptosis.
METHODS:
Arraystar Human CircRNA chip was applied to analyzing the differential expression of circRNA in four bladder cancer tissues and paired adjacent normal bladder tissues. Real time quantitative PCR was utilized to detect the expression of circ0001429 in tissue specimens. Bioinformatics, RNA immunoprecipitation and luciferase reporter assays were used to verify the relationship among circ0001429, miR-205-5p and VEGFA in bladder cancer. Cell propagation was determined by CCK8 assay and roles of circ0001429 and miR-205-5p in cell migration were verified with transwell migration assay. Flow cytometry and TUNEL staining were conducted to observe the impact on cell apoptosis ability. Xenograft experiment was also performed to validate the influence of circ0001429 on tumor growth in vivo.
RESULTS:
Expressions of circ0001429 and VEGFA were up-regulated, whereas miR-205-5p expression was down-regulated in bladder cancer tissues in comparison with paired adjacent normal bladder tissues. Circ0001429 enhanced the propagation and metastasis abilities of T24 cells and 5637 cells in vitro, but reduced cell apoptosis. In vivo experiments revealed the inhibitor role of sh-circ0001429 in tumor growth and lung metastasis. Circ0001429 sponged miR-205-5p that targeted VEGFA, thereby modulating the protein level of VEGFA. Meanwhile, miR-205-5p restrained the cell viability and mobility and promoted the apoptosis in bladder cancer. Circ0001429 could accelerate cell propagation, migration and invasiveness through increasing VEGFA expression via miR-205-5p.
CONCLUSION:
Circ0001429 and VEGFA were highly expressed in bladder cancer, while miR-205-5p were lowly expressed in bladder cancer. The circ0001429 could target at miR-205-5p to regulate VEGFA and promote the development of bladder cancer.
Introduction
Bladder cancer, with estimated 386,300 new cases and 150,200 deaths per year, is a frequently seen urinary system cancer [30]. Metastases happen more frequently in muscle-invasive bladder cancer patients and always correlate with poor prognosis [2]. Although appreciable advance has been made in surgical techniques and adjuvant chemotherapies, the mortality of metastatic bladder cancer has not been significantly reduced [30]. Therefore, further comprehension of the gene expression may be favorable to the treatment of bladder cancer.
Circular RNA (circRNA), a class of endogenous non-coding RNAs that contain a circular loop, was found to exhibit multiple biological effects [27]. With the rapid development of high-throughput sequencing and bioinformatics analysis, lots of circRNAs have been successfully identified in various cell lines from different species, but their functional significance remains unclear [30]. Latest studies have reported that circRNAs may be involved in neurological disorders [12] and cancers [22]. Recently, circRNAs are demonstrated to act as “miRNA sponges” and can expose a negative regulation on miRNAs [30]. All these findings strongly indicate potential functions of circRNAs in disease pathogenesis by regulating gene expressions [18].
MicroRNAs (miRNAs) are a group of highly conserved small (19–25 nucleotides-long) single-stranded non-coding RNAs. MiRNAs can down-regulate gene expressions in various ranges of biological functions by binding to the 3’-UTR of target genes [28]. Numerous studies have manifested that dysregulation of miRNAs may be an important factor for carcinogenesis and tumorigenesis [28, 4]. Accumulative researches have revealed the down-regulated expression of miRNAs in many human malignancies, including bladder cancer [7, 15]. Notably, miR-205-5p was proved to be suppressor of VEGFA in human glioblastoma cells [29]. Downregulation of miR-205-5p were significantly linked to progression in non-muscle invasive bladder tumors [19]. Thus, miR-205-5p could be new molecular indicators for bladder cancer patients’ diagnostics, monitoring and therapeutics [4].
Vascular endothelial growth factor (VEGF), has been identified as a critical protein in promoting angiogenesis on tumor growth and metastasis [3]. The VEGF gene is located on chromosome 6p21.3 and consists of 9 exons. VEGFA is one of the most potent inducers of angiogenesis, and increased expression of VEGFA has been detected in almost all known tumors, including bladder cancer [24]. VEGFA can activate ERK1/2, PI3K-Akt/PKB pathway, and phospholipase C-
Here, we identified a circular RNA, termed circ000 1429, which was obvious reduced in bladder cancer tissues and cell lines. Additionally, we found that circ0001429 knock-down inhibited growth and metastasis of bladder cancer cells via suppressing VEGFA expression by miR-205-5p expression increase.
Materials and methods
Human tissue specimens
Twenty pairs of bladder cancer tissues and paired adjacent normal bladder tissues were collected from patients undergoing no chemoradiotherapy or other treatment for the tumor therapy at General Hospital of Western Theater Command. All participants were confirmed by exhaustive diagnosis and the tissue samples were independently examined by three well-qualified pathologists. Tissue specimens were frozen in liquid nitrogen instantly after surgical removal, and conserved at below 80
Microarray analysis
Affymetrix microarray platform GPL19978 and microarray data GSE92675 were gained from Gene Expression Omnibus (GEO) database (
Cell culture
Human bladder cancer cell lines T24 and 5637 were obtained from ATCC, SV-HUC-1 and BIU-87 were provided by BNCC. T24, 5637 and BIU-87 were grown in RPMI 1640 (Gibco Laboratoties, Grand Island, NY, USA) supplemented with NaHCO
RNA Extraction and qRT-PCR
Total RNAs extraction was performed using TRIzol Reagent (Invitrogen Corp., Carlsbad, CA), followed by quantification by NanoDrop 2000. 200 ng of total RNA per sample was converted to cDNA using the ReverTra Ace qPCR RT Kit (Toyobo, Japan). Quantitative reverse transcription PCR (qRT-PCR) was implemented using the THUNDERBIRD SYBR
Prime sequences for qRT-PCR
Prime sequences for qRT-PCR
The cells were lysed in RIPA buffer (Beyotime) and quantified with Pierce BCA Protein Assay Kit (Pierce, Rockford, IL, USA) following the conditions suggested by the manufacturer. Protein (100
Cell transfection
Circ0001429, sh-circ0001429 and sh-NC were dissolved and ligated to the multi-cloning sites of the pVax1 expression reporter plasmid (GenePharma, Shanghai, China). MiR-205-5p mimics and miR-205-5p inhibitor were produced by Sangon Biotech (Shanghai, China). 24 h pre-transfection, 1
Cell proliferation
Cell proliferation was determined by Cell Counting Kit-8 (CCK8) (Beyotime, Shanghai, China) Assay. T24 and 5637 cells (8000) suspended in RPMI1640 medium (100
Luciferase reporter assay
To create a luciferase reporter construct, 3’-UTR segments of circ0001429 wildtype (WT), circ0001429 mutant (MT), VEGFA wildtype (WT) and VEGFA mutant (MT) were cloned into the pGL3-control (Promega, Madison, WI) vector. T24 and 5637 cells were co-transfected with luciferase reporter constructs containing the wild-type or mutant 3’-UTR firefly luciferase reporters, pRL-TK, and miR-552 inhibitor (Sangon, Shanghai) or control (Sangon, Shanghai) using Lipofectamine™ 2000. Luciferase activities were detected 48 h after the transfection by the Dual-luciferase Reporter Assay System (Promega, WI, USA).
RNA immunoprecipitation (RIP)
Magna RIP™ RNA-binding protein immunoprecipitation kit (Millipore, Billerica, MA) was used for RIP in accordance with the instructions. Anti-Argonaute-2 antibody (ab186733, Abcam) was applied to RIP assay. Bound RNA was then eluted from the beads by directly adding Trizol (Invitrogen) to the beads, followed by RNA extraction and RT-real time PCR as described previously.
Apoptosis assay
Cell apoptosis was evaluated by Annexin V-FITC Apoptosis Detection Kit (BD Science, Bedford, MA, USA). Cells after transfected 48 h were resuspended in binding buffer containing annexin V-FITC and propidium iodide (PI) according to manufacturer’s directions. Stained cells were measured by FACS Calibur using FACS Diva software (BD, USA). Each sample was run in triplicate.
Transwell migration assays
The migration assays were carried out using transwell chamber following the manufacturer’s manual (BD Science). Cells (5
The expression and correlation of circ0001429, miR-205-5p and VEGFA. (A, B) Volcano, heat map and hierarchical clustering analysis of circRNAs which were differentially expressed between bladder cancer and normal bladder tissues. Each column represents the expression profile of a tissue sample (four bladder cancer and normal bladder samples, respectively), and each row corresponds to a circRNA (log fold change (FC) 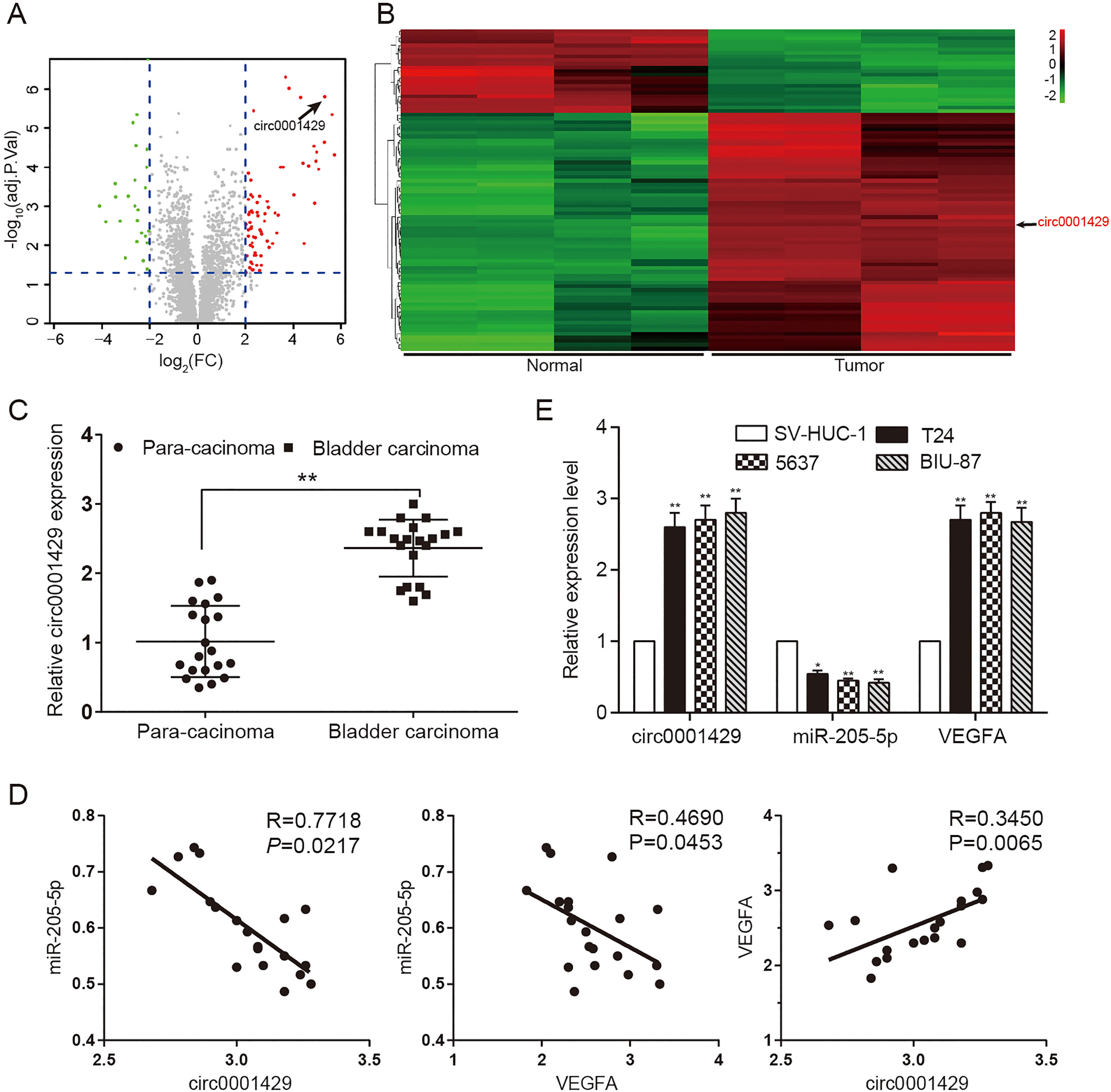
The 12 Balb/c male mice (4–6 weeks old) used for this study was divided equally into 2 groups. All mice were purchased from Shanghai Medical Experimental Animal Care Commission (Shanghai, China). To establish a tumor model, T24 cells stably transfected with sh-NC or sh-circ0001429 were subcutaneously injected into Balb/c mice. Besides, tumors volume and tumor weight were measured in every 3 days. The tumor growth curve was calculated and plotted following the formula (
TUNEL staining and histologic examination
Apoptosis via DNA fragmentation was detected by using the terminal deoxynucleotidyl transferase-mediated dUTP nick end labeling (TUNEL) technique with a situ apoptosis detection kit (Neurotacs II, R&D, USA), according to the manufacturer’s recommendations. The cells were lightly counterstained using DAPI to enable microscopical counting of TUNEL
Statistical analysis
The GraphPad Prism version 6.0 (GraphPad Software Inc.) was used for statistical analysis. The differences between groups were compared by means of unpaired student’s
Results
Circ0001429 was overexpressed in bladder cancer
The up-regulated and down-regulated mRNAs were respectively filtered using
circ0001429 affected bladder cancer cells growth. (A) The expression of circ0001429 in T24 and 5637 cells treated with circ0001429 or sh-circ0001429 were detected by real-time PCR. The relative levels of circ0001429 were normalized to the value measured in the mock treatment. Data were expressed with mean 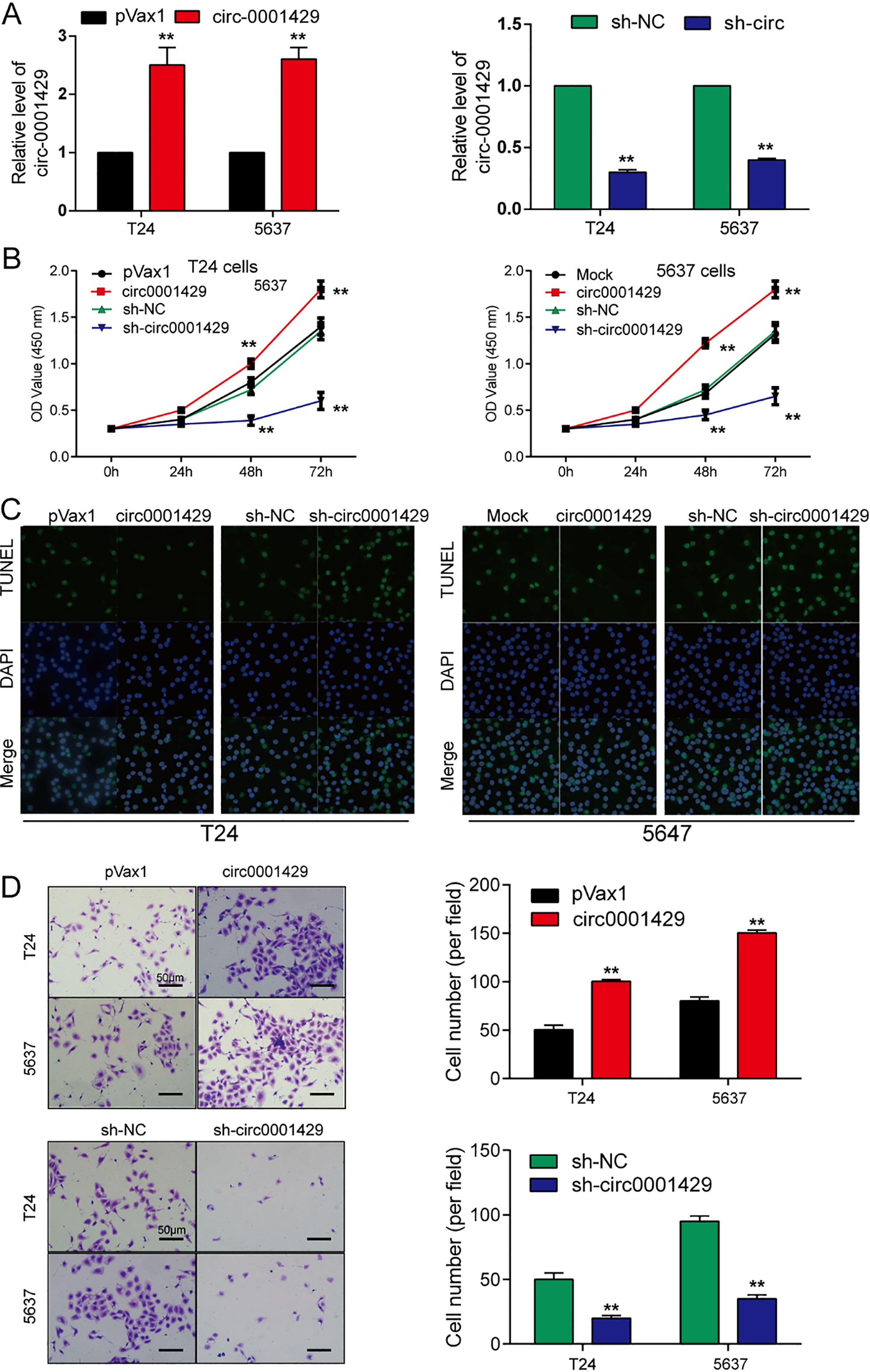
Down-regulation of circ0001429 inhibited the growth and metastasis of bladder cancer xenograft. (A, B, C) Hypodermic injection of T24 cells stably transfected with sh-circ0001429 or sh-NC control into BALB/c nude mice established subcutaneous xenograft tumors (2 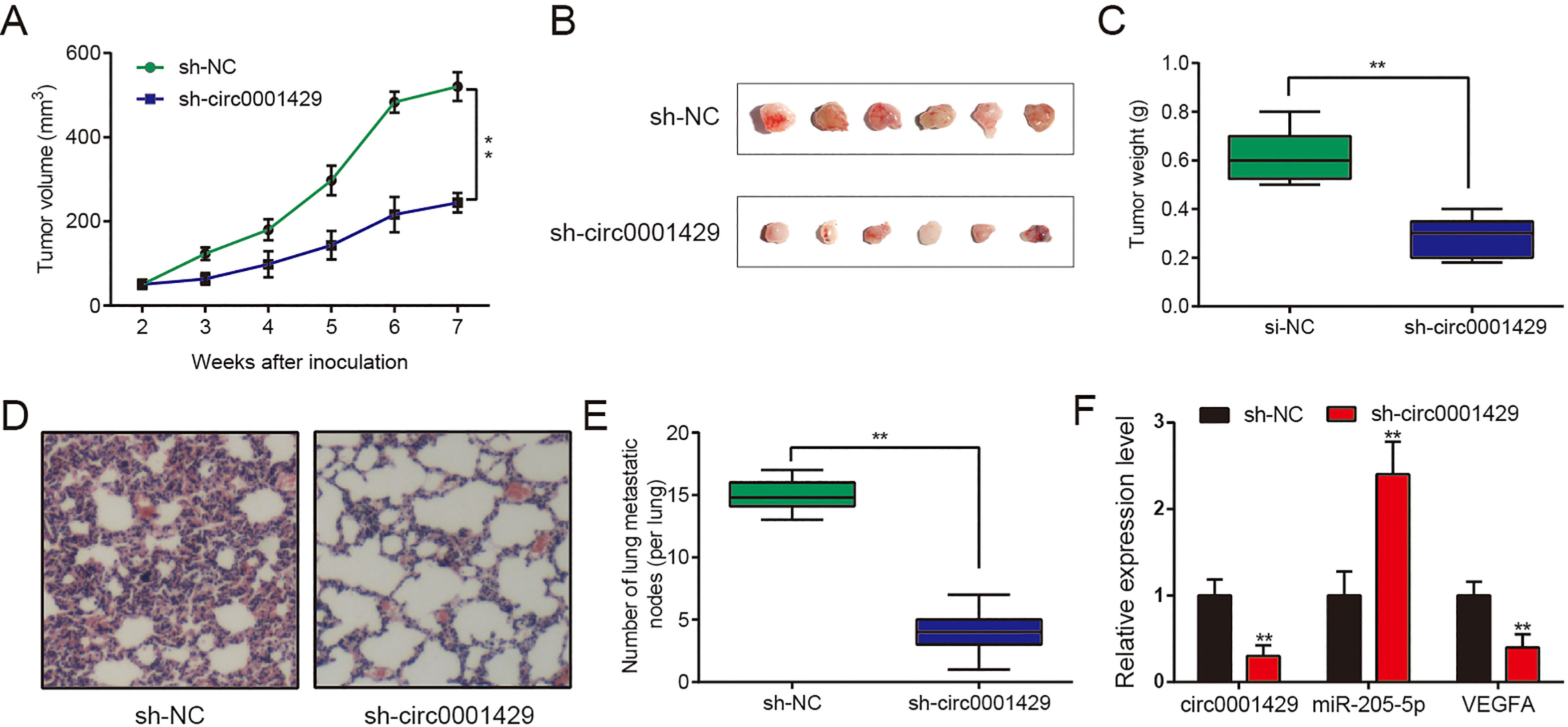
To explore the effect of circ0001429 in bladder cancer cells, T24 and 5637 were transfected with circ0001429, circMock, sh-circ0001429 and sh-NC. The change of circ0001429 expression post-transfec-tion was validated using qRT-PCR (Fig. 2A,
miR-205-5p bound to circ0001429 and VEGFA. (A) Base pairing complement suggested the putative miR-205-5p binding position at 3’-UTR of circ0001429. (B) Luciferase Report assay of circ0001429 3’-UTR treated with miR-205-5p inhibitor or miR-205-5p NC showed that circ0001429 3’-UTR bind with miR-205-5p. Data are mean 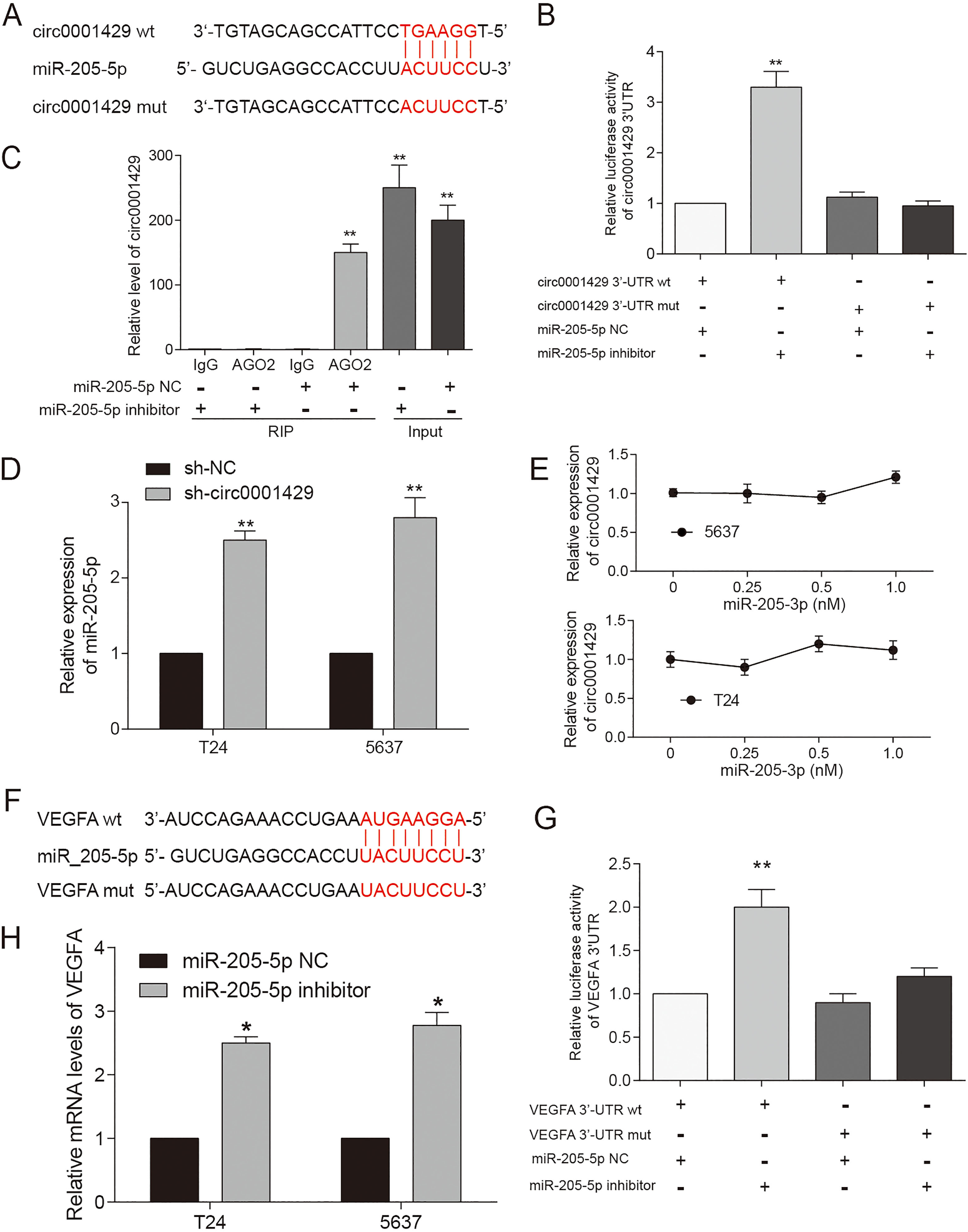
To verify the positive effect of circ0001429, a xenograft mouse model was established. T24 cells stably transfected with si-NC or sh-circ0001429 were subcutaneously injected into nude mice. The tumor sizes (Fig. 3A and B) and tumor weight (Fig. 3C) of sh-circ0001429 group were significantly lower than sh-NC group, indicating that down-regulation of circ000 1429 obviously inhibited xenograft tumor growth. In addition, tumor metastasis were found in the lungs according to the HE staining (Fig. 3D and E), and we found that sh-circ0001429 significantly reduce the number of metastases nodules compared with si-NC group (Fig. 3D and E). The relative expressions of circ0001429, miR-205-5p and VEGFA mRNA in xenograft tumor suggested that circ0001429 and VEGFA were down-regulated, while miR-205-5p was up-regulated (Fig. 3F).
Circ0001429 sponged miR-205-5p, which targeted to VEGFA
Studies have shown that the main function of circRNA is to act as a miRNA ‘sponge’ and depress the functional miRNA [25]. Therefore, the miRNA ‘sponge’ function of circ0001429 and its role in downstream regulation was verified. Bioinformatics analysis predicted the potential target was miR-205-5p, which shared complementary binding sites with circ0001429. MiR-205-5p contains six paired nucleotides with circ0001429 (Fig. 4A). Besides, VEGFA was predicted as a candidate target of miR-205-5p (Fig. 4F). Luciferase reporter assay (Fig. 4B) and RIP (Fig. 4C) results confirmed that circ0001429 sponged miR-205-5p expression. In addition, miR-205-5p in T24 and 5637 cells transfected with si-circ0001429 were significantly up-regulated compared to si-NC treated group (Fig. 4D,
miR-205-5p inhibited bladder cancer progression
In T24 and 5637 cells, ectopically expressed miR-205-5p mimics significantly reduced the protein level of VEGFA and overexpression of VEGFA rescued this change (Fig. 5A). The propagation (Fig. 5B) and migration (Fig. 5C) rate of T24 and 5637 cells transfected miR-205-5p mimics were significantly down-regulated, and were rescued by overexpression of VEGFA. Apoptosis assay suggested that miR-205-5p mimics promoted the apoptosis of T24 and 5637 cells (Fig. 5D). Taken together, miR-205-5p inhibited propagation and migration of bladder cancer cells and promoted cell apoptosis, which was rescued by overexpression of VEGFA.
miR-205-5p inhibited bladder cancer progression. (A) Western Blot of VEGFA protein in T24 and 5637 cells with miR-205-5p mimics, NC, miR-205-5p mimics 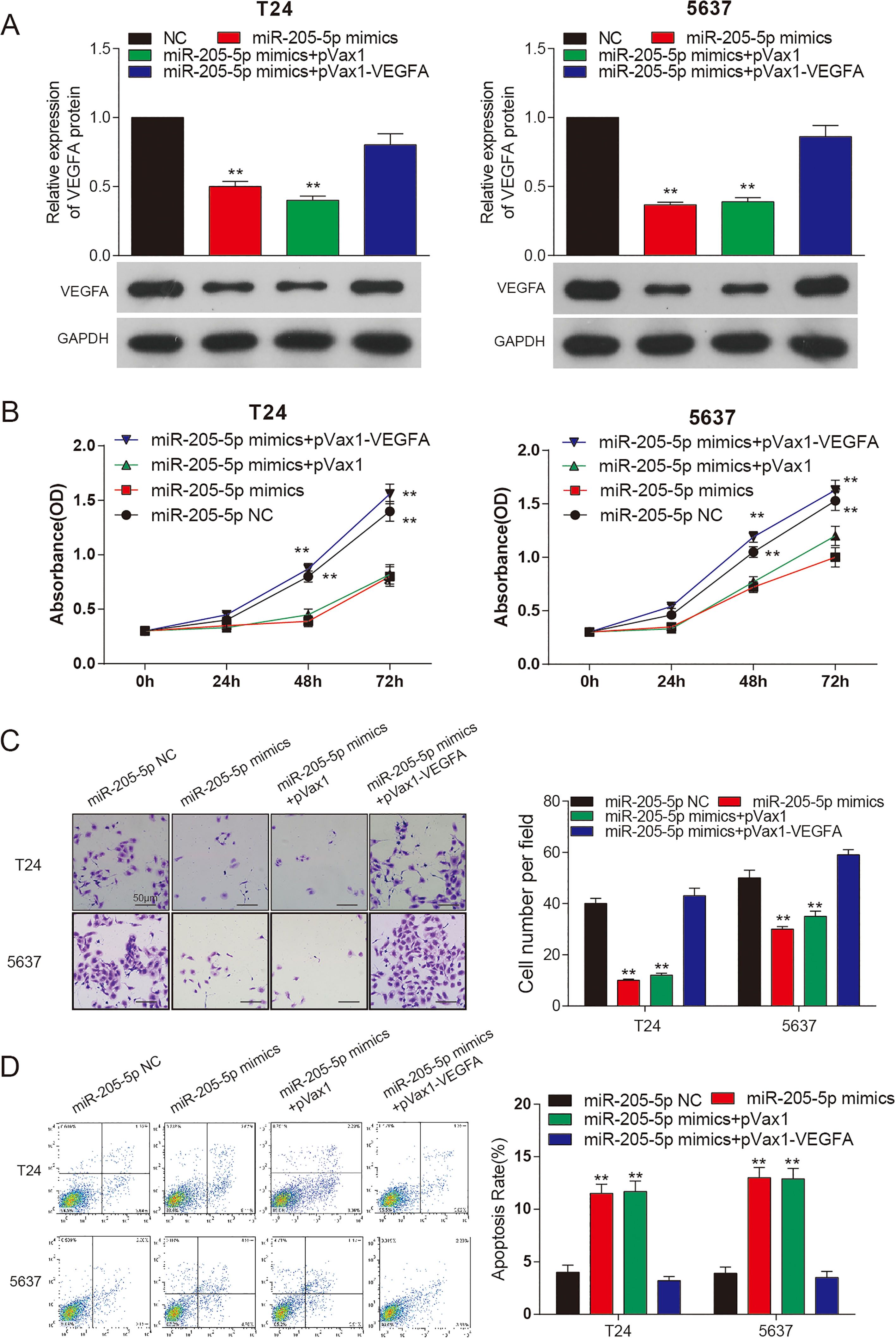
As shown in Fig. 6A, ectopically expressed circ000 1429 improved the luciferase activity with the wild-type 3’UTR of VEGFA, while the expression of miR-205-5p attenuated this positive effects. These results showed that circ0001429 could promote the expression of VEGFA, which could be relieved by miR-205-5p. Western Blot assay further proved this conclusion (Fig. 6B). The miR-205-5p reduced the propagation of T24 cells, while circ0001429 rescued this effect (Fig. 6C). Similarly, circ0001429 rescued the apoptosis of T24 cells co-transfected miR-205-5p mimics (Fig. 6D).
Overexpression of circ0001429 promoted VEGFA expression through targeting miR-205-5p. (A) Luciferase Report assay of VEGFA 3’-UTR treated with miR-205-5p mimics, miR-205-5p-NC or circ0001429. Data are mean 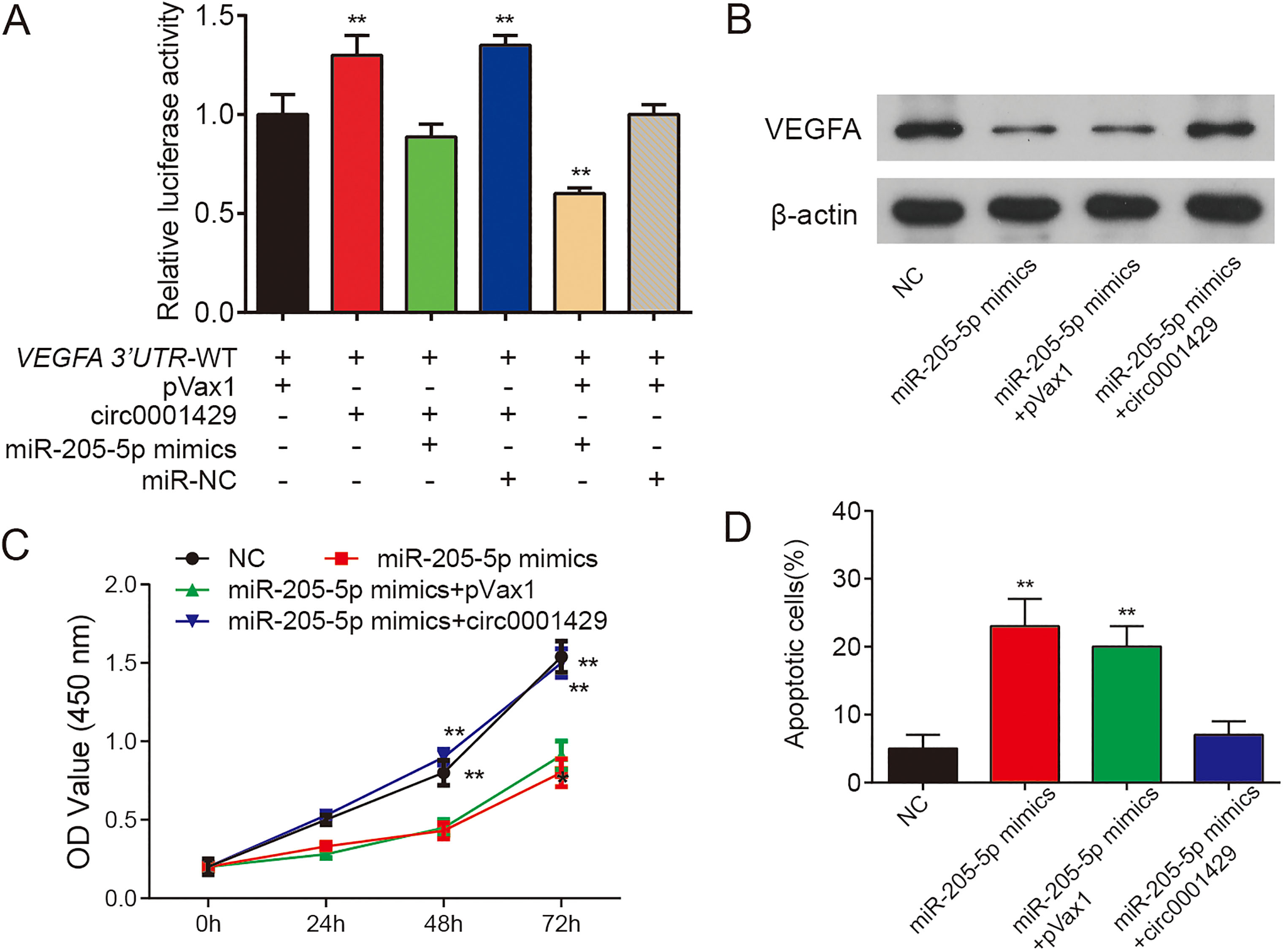
Here, we first identified that circ0001429 was up-regulated in bladder cancer tissues and cells and it promoted the propagation and metastasis of bladder cancer cells in vitro and in vivo. Then we found that circ0001429 could efficiently sponge miR-205-5p, which was also verified by dual luciferase reporter assay and RIP. Finally, we proved that circ0001429 down-regulated VEGFA protein level through sponging miR-205-5p to promote bladder cancer cells growth and metastasis.
In the early 1990’s, endogenous circRNAs was first hinted from research of the DCC transcript in human cells [9]. Latterly, with the development of next-generation sequencing, mounting circRNAs were identified from various animal genomes, and many of these are abundant and stable [1]. CircRNAs which may have certain potential functions in regulation of gene expression, was dysregulated in laryngeal cancer [13], colorectal cancer [1], esophageal squamous cell carcinoma [5], hepacellular carcinoma [17] and bladder cancer [34]. CircRNAs arise from diverse genomic locations, and most of them are formed by circularization of exons [20]. In current research, we gained a circular RNA, termed circ0001429 which was dramatically up-regulated in bladder cancer. The role of circ0001429 in bladder cancer is not yet studied.
CircRNAs, containing certain miRNA response elements (MREs) is an efficient miRNA sponges [11]. For instance, CDR1as contains over 70 binding sites of miR-7 and significantly suppresses the activity of miR-7 [20]. It was reported that circRNA-MYLK could promote cell propagation and invasion in bladder cancer [33]. Down-regulated circPTK2 expression could markedly inhibited cell migration and propagation [32]. However, circRNAs can also inhibited migration and propagation of cancers. According to Li et al., circHIPK3 inhibited migration, invasion, and angiogenesis of human invasive bladder cancer T24T and UMUC3 cells by targeting miR558, which was contrast to our result [30]. Our research indicated that circ0001429 promoted growth and migration of T24 and 5637 cells by sponging miR-205-5p, which was not reported by previous studies. These findings suggested that circ0001429 was involved in complex regulatory networks in bladder cancer.
MiRNAs could either retard translation or induce degradation of the target mRNA by incomplete or complete base pairing to 3’UTR of mRNA [8]. However, mounting evidences indicated that miRNAs can also promote the transcription of mRNA via binding to the promoter region of target genes [23]. Nunez Lopez et al. illustrated that miR-205-5p was upregulated in a broad range of head and neck squamous cell carcinoma subtypes [31]. Switlik et al. reported that miR-205-5p was upregulated in non-small cell lung cancer (NSCLC) tissues, which was counter to our research [28]. Decreased expression of miR-205-5p was observed in Hepatocellular carcinoma (HCC) cells including HCCLM3, MHCC97-H, MHCC97-L, Huh-7, Bel-7402 and SMMC-7721 cells compared with the normal liver cell line, LO2 [16]. Downregulation of miR-205-5p was significantly linked to progression in non-muscle invasive bladder tumors [19]. However, Thorsten reported that there was no difference of the miR-205-5p expression between nonmalignant and muscle-invasive bladder cancer (MIBC) samples [26]. Interestingly, our results suggested miR-205-5p was markedly down-regulated in bladder cancer tissues and cells. Then, over-expression of miR-205-5p significantly inhibited bladder cancer cell growth and mobility in vitro.
VEGF regulated tumor invasion and metastasis th- rough multiple signaling cascades, including ERK1/2, PI3K-Akt/PKB pathway, and phospholipase C-
Nonetheless, some limitations also existed in this report which is worth mentioning. For instance, we identified circ0001429 as “microRNA sponges” which down-regulated VEGFA expression via miR-205-5p, but downstream pathway of VEGFA was not explored. Moreover, although we found that circ0001429 and miR-205-5p regulated protein level of VEGFA, the protein degradation should be considered.
In conclusion, we showed that circ0001429 was up-regulated in human bladder cancer, and it could efficiently sponge miR-205-5p to inhibit VEGFA expression. We also demonstrated that down-regulation of circ0001429 expression obviously inhibited growth and metastasis of bladder cancer cells via miR-205-5p/VEGFA axis. Our findings provided novel evidences that circRNAs acted as “microRNA sponges” and also provided a potential therapeutic target for bladder cancer therapy.
Footnotes
Conflict of interest
The authors declare that they have no conflicts of interest with the contents of this article.
