Abstract
BACKGROUND:
CPNEs are significant biomarkers which can affect the progression and prognosis of various tumor diseases. However, the prognosis role of CPNEs in multiple myeloma (MM) is still unclear.
OBJECTIVES:
To investigate the prognosis role of CPNEs in MM.
METHODS:
Seven hundred and thirty-five samples from two independent data sets were involved to analyze the clinical and molecular characteristics, and prognosis role of the expression of CPNE1-9 in MM.
RESULTS:
MM patients with higher expressions of CPNE5 and CPNE9 had longer event-free survival (EFS) and overall survival (OS) compared with CPNE5
CONCLUSIONS:
High expressions of CPNE5 and CPNE9 might be used as positive indicators for MM, and their combination was a better predictor for the survival of MM patients.
Introduction
Multiple myeloma (MM) is a common malignant tumor disease of blood system, which is characterized by abnormal proliferation of clonal plasma cells. It is estimated that there were 176,404 new MM patients in 2020, accounting for 0.9% of all cancer patients, and 117,077 deaths of MM patients, accounting for 1.2% of all cancer deaths [1]. Although the prognosis of MM patients has improved with the improvement of MM diagnosis and treatment, only 45% of patients have an overall survival (OS) of 6 years [2]. It is currently believed that genetic changes, bone marrow microenvironmental effects and epigenetic modifications play an important role in the occurrence and development of MM [3, 4, 5]. Although genetic abnormalities have made great progress in the assessment of MM progression and prognosis, there is still an urgent need to find more effective prognostic biomarkers, which are of great clinical significance for optimizing the personalized treatment plan and improving the clinical efficacy of MM patients. It is also a current research hotspot.
CPNEs are highly conserved intracellular calcium-dependent lipid-binding proteins originally identified by Creutz et al. [6] from the extracts of paramecium tetraurelia, which play a fundamental role in eukaryotic cell biology. The N-terminal of CPNEs contains two C2 domains and a domain similar to the A domain, in which the C2 domain plays a role in membrane transport and the A-like domain mediates the interaction between integrins and extracellular ligands. The HUGO Gene Nomenclature Committee has identified nine homologues of CPNEs in humans, namely CPNE1-9. Jiang et al. [7] found that the down-regulation of CPNE1 can inhibit the proliferation and metastasis potential of osteosarcoma cells and reduce the expression of a variety of tumor-related genes. In non-small cell lung cancer (NSCLC), inhibition of CPNE1 expression can effectively reduce the proliferation and movement of cancer cells [8]. In addition, CPNE3 is also highly expressed in a variety of cancer cells and can regulate ErbB2 signaling pathway-mediated movement and migration of cancer cells [9, 10, 11]. Genetic variants in CPNE5 have been found to be associated with alcohol dependence and obesity [12]. As a protein specifically expressed in the brain, CPNE6 has a high degree of homology with human CPNE1, and is closely related to synaptic plasticity and memory formation [13, 14]. A recent study found that Ca
The CPNEs have been deeply researched in a variety of diseases, especially for the pathogenesis, treatment and prognosis of malignant tumors. However, the prognostic value of CPNEs in MM is still unclear. This study explored the asscociation between the expression levels of CPNEs and survival of MM patients.
Subjects and methods
Patients
In this study, two independent microarray data sets (GSE39754 and GSE24080) were selected to analyze prognosis role of the expression of CPNEs in MM. The GSE39754 data set used Affymetrix Human Exon 1.0 ST Array to analyze the gene expression profiles of 6 healthy donors and 170 newly diagnosed MM patients [19, 20, 21]. The GSE24080 data set collected high-purity bone marrow plasma cell gene expression profiles of 559 newly diagnosed MM patients, and used Affymetrix human genome U133 Plus 2.0 array to analyze the expression level of CPNEs in MM patients [22, 23, 24]. All information related to the patients’ clinical and molecular data, as well as microarray data sets comes from Gene Expression Omnibus (GEO). All experimental design, quality control and data standardization are in compliance with the standard Affymetrix protocol. The study was conducted in accordance with the “Declaration of Helsinki”, and all patients had obtained written informed consent.
Survival analysis of CPNE2, CPNE5 and CPNE9 in GSE24080. (A) There wasn’t any significant difference in EFS between CPNE2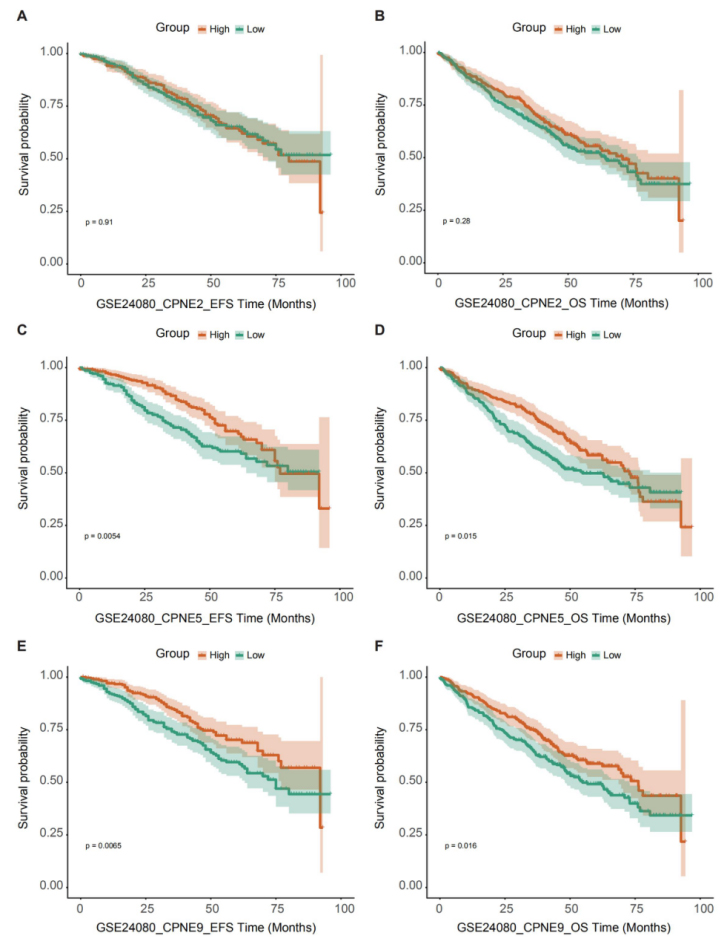
Survival analysis of combined expression of CPNE5 and CPNE9. (A) CPNE5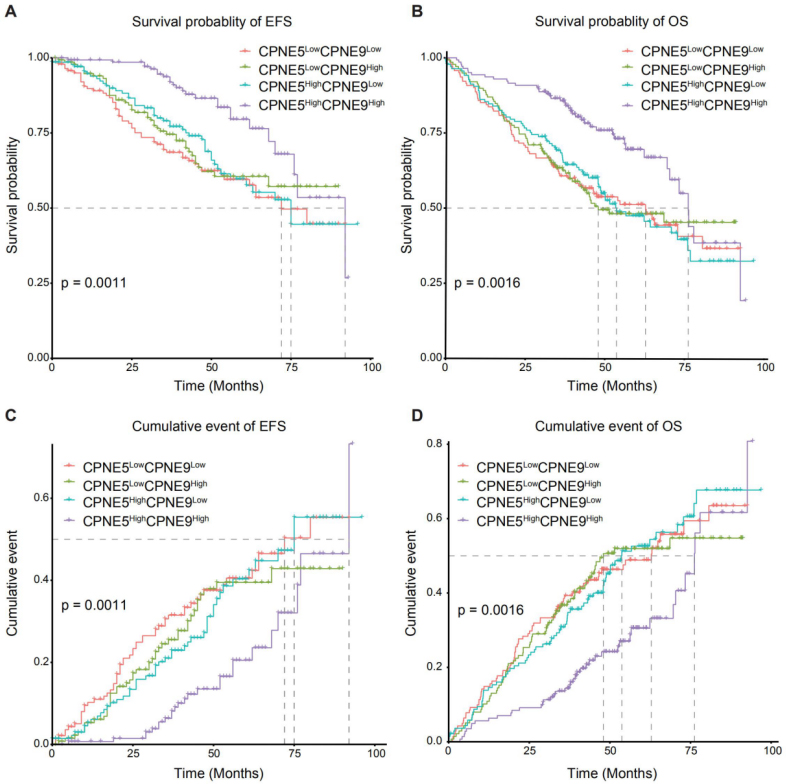
The event-free survival (EFS) and OS were used as the study endpoints. EFS refers to the time from the diagnosis of the disease to the first event such as relapse and death. OS refers to the time from the diagnosis of the disease to the final death of the patient. In GSE39754 (
Results
The expression of CPNE1-9
In this study, GSE39754 was used to explore the expression levels of CPNE1-9 in healthy donors and MM patients. And we found that the expression levels of CPNE1 and CPNE5 in the MM group were significantly higher than that in the healthy group (CPNE1:
Comparison of EFS and OS between the high and low expression levels of CPNE family
Comparison of EFS and OS between the high and low expression levels of CPNE family
EFS event-free survival, OS overall survival.
Patients’ characteristics of 559 MM patients in GSE24080
AGE: age at registration (years); B2M: beta-2 microglobulin (mg/l); CRP: C-reactive protein (mg/l); CREAT: creatinine (mg/dl); LDH: lactate dehydrogenase (U/l); ALB: albumin (35 g/l); HGB: hemoglobin (g/dl); MRI: number of magnetic resonance imaging (MGI)-defined focal lesions (skull, spine, pelvis); no.: number of patients.
Univariate and multivariate cox regression analyses of EFS and OS
EFS: event-free survival; OS: overall survival; CR: complete remission; HR: hazard ratio; CI: confidence interval; ALB: Albumin, 35 g/l; B2M: Beta-2 microglobulin, mg/l; HGB: Haemoglobin, g/dl; LDH: Lactate dehydrogenase, U/l.
We analyzed the association between expression levels of CPNEs and survival of MM patients in GSE24080. According to the median expression level of each CPNE member, all patients were divided into high and low expression groups. EFS and OS were compared between the two groups in Table 1, the OS and EFS between high and low CPNE5/9 expression were significantly different (OS:
Clinical and molecular characteristics
The clinical and molecular characteristics were shown in Table 2. Compared with the CPNE5
Univariate and multivariate analysis of EFS and OS
To further assess the prognostic significance of CPNE expression, univariate and multivariate COX proportional hazard regression models were constructed (Table 3). Univariate analysis showed that ALB, B2M, HGB, LDH, CPNE5 and CPNE9 were closely related to EFS and OS (all
Co-expression of CPNE1-9
Correlations between the expression levels of CPNE members were analyzed in our study. As shown in Fig. S2, the expressions of CPNE members were not significantly associated with each other (all
Survival analysis of combined expression of CPNE5 and CPNE9
The effect of the combination of CPNE5 and CPNE9 on prognosis was further explored in MM, the results showed that patients with CPNE5
Discussion
In our study, a total of 729 patients were enrolled to explore the prognosis role of CPNE1-9 in MM. High expression of CPNE5 and CPNE9 were proved to be positive prognosis indicators. CPNE5 was an independent favorable predictor for EFS and OS, and CPNE9 was an independent positive indicator for EFS. Combination of high expressions of CPNE5 and CPNE9 might be a better predictor for the survival of MM patients.
CPNE5 mainly exists in cytoplasm of human cells and can be detected in the heart, stomach, spleen, lymph nodes and testis [25]. In esophageal squamous cell carcinoma, the expression of CPNE5 in cancer tissues is lower than that in adjacent tissues, and patients with low CPNE5 expression have shorter OS. Moreover, low expression of CPNE5 may promote the local growth of esophageal cancer and increase the resistance of chemotherapy drugs [26]. These findings suggest that CPNE5 is closely related to tumorigenesis and may affect the prognosis of cancer patients. In our study, MM patients with high expression of CPNE5 had longer OS and EFS. And the clinical and molecular characteristics analysis found that LDH, ALB and the expressions of CDK4 and FGFR3 in the CPNE5
A study by Liu et al. [31] found that CPNE9 is specifically expressed in pancreatic tumor tissues, which indicates that CPNE9 is related to cancer progression. Moreover, CPNE9 was the top five of the prognostic 14-gene signatures that constructed the prognostic model with a high predictive ability for glioblastoma [32]. This suggests that CPNE9 has important clinical significance for the prognosis assessment of glioblastoma patients. In our study, MM patients with high expression of CPNE9 had longer OS and EFS. At the same time, the CPNE9
Multivariate Cox regression analysis showed that LDH and CPNE5 were independent prognostic factors for EFS and OS. ALB, B2M and CPNE2 were independent prognostic factors for OS, and CPNE9 was an independent prognostic factor for EFS. These results further confirmed that high expressions of CPNE5 and CPNE9 were positive prognostic factors in MM patients. The co-expression analysis of CPNE1-9 in MM showed that there was no obvious expression correlation between CPNE family members, indicating that there was no mutual regulation mechanism between CPNE5 and CPNE9. In addition, the survival analysis of the combined expression of CPNE5 and CPNE9 found that the survival probabilities and cumulative events of CPNE5
Conclusion
In summary, we have shown that high expressions of CPNE5 and CPNE9 are positive prognostic factors in MM patients, especially for the combination of CPNE5 and CPNE9, and they may be potential targets for MM therapy. But the specific molecular mechanisms of CPNE5 and CPNE9 in the development of MM are still unclear. Further studies with larger sample size are needed to validate our findings, and the pathogenesis of CPNE5 and CPNE9 in MM also needs additional exploration.
Footnotes
Supplementary data
The supplementary files are available to download from http://dx.doi.org/10.3233/CBM-203108.
Acknowledgments
This work was supported by grants from Xinjiang Joint Fund of National Natural Science Foundation of China (U1903117) and the National Natural Science Foundation of China (81500118).
The results shown here are in whole or part based upon data generated by GEO database:
Conflict of interest
The authors declare that they have no conflict of interest.
Author contributions
Lin Fu, Chaozeng Si were responsible for the conception. Chaozeng Si, Tiansheng Zeng, Pei Zhu, Yan Liu, Longzhen Cui, Wenhui Huang participated in the interpretation or analysis of data. Pei Zhu, Tingting Qian, Tiansheng Zeng, Cong Deng were responsible for preparation of the manuscript, and participated in drafting and revision of the manuscript. Tiansheng Zeng, Cong Deng, Lin Fu was responsible for supervision of the manuscript.
