Abstract
BACKGROUND:
Sphingolipids are bioactive lipids that play a role in cancer development. However, the clinical role of sphingolipid (SPL)-related genes in oral cancer (OC) remains not fully understood.
OBJECTIVE:
This study, aimed to examine the mRNA expression of 14 sphingolipid-related genes in oral cancer patients and their implication with clinicopathological features and prognosis.
METHODS:
qPCR analysis was performed in 50 OC tissues and their matched surgical margins. Next, Kaplan-Meier, Cox regression, and Receiver operating characteristics (ROC) analysis were applied to evaluate the impact of sphingolipid-related genes expression on the prognosis of OC.
RESULTS:
The genes SET, ACER3, SK1 and S1PR5 were predominantly up-regulated, while ABCG2, S1PR1, ABCB1 and SPNS2 were down-regulated in OC patients. Analyzing the Cancer Genome Atlas Head-Neck Squamous Cell Carcinoma (TCGA-HNSC) data, which are predominantly composed of OC samples, these genes displayed a similar profile. In OC patients, high levels of SK1 were associated with lymph node metastasis, extracapsular invasion, desmoplasia, locoregional relapse, and disease status. Low levels of SPNS2 were associated with lymph node metastasis, perineural invasion, and disease status. Furthermore, OC and HNSC patients with higher SK1 expression demonstrated shorter disease-free survival (
CONCLUSION:
Several sphingolipid-related genes are deregulated in OC at the mRNA level and are associated with clinicopathological features and presented potencial for the prediction of poor prognosis in OC patients.
Introduction
Oral squamous cell carcinoma (OSCC) is the most frequent malignant neoplasm of all Head-Neck Squamous Cell Carcinoma (HNSCC) and represents over 90% of the total oral cancer worldwide [1]. The 5-year survival rate of oral cancer was about 68% during 2009–2015 in the United States of America [2]. However, in developing countries oral cancer is frequently detected in advanced stages, what raise the mortality rate. In advanced stage disease, the overall 5-year survival rate is below 40% [2, 3]. A full understanding of the molecular traits in oral cancer could lead to improvements of diagnosis, treatment, and prognosis.
Ceramides (Cer) and sphingosine-1-phosphate (S1P) are bioactive sphingolipids that play critical roles in plasma membrane dynamics and signaling processes. S1P is generated by breaking ceramide into sphingosine, followed by phosphorylation via sphingosine kinases (SphK1 and SphK2). S1P regulates numerous biological functions, such as inflammation, proliferation, cell survival, migration, and angiogenesis in tumor cells [4, 5]. As a consequence, S1P has been gradually established in the latest years as an oncogenic sphingolipid implicated in several cancer types, including lung, colon, breast, and oral cancer [4, 5]. On the other hand, ceramide has demonstrated tumor suppressor properties via mitochondrial-mediated apoptosis [6]. However, ceramide can be converted to ceramide-1-phosphate (C1P) by ceramide kinase (CERK), which can enhance survival, migration, and invasion of tumor cells [7]. In addition, the activity of ceramide and S1P can be altered by other proteins [8, 9]. Previous studies from our group have indicated that S1P activity can be influenced by the action of the SET (SET nuclear proto-oncogene) protein (data not published), which is altered in several types of cancer, including HNSC [10].
The availability of intracellular C1P and S1P may be affected by the activity of sphingomyelin synthase and ceramidase. Sphingomyelin synthases can convert ceramide into sphingomyelin, thus decreasing the levels of ceramide. Sphingomyelin synthase I (SGMS1) has been shown to be inhibited in melanoma, promoting an imbalance in sphingolipid homeostasis and may contribute to melanoma development [11]. Ceramidases cleave ceramide-producing sphingosine, thus increasing the supply of sphingosine for the production of S1P [12]. Ceramidases have been implicated in cancer, for example, alkaline ceramidase 3 supports the growth, survival, and AKT signaling activation in leukemia cells [13].
S1P performs its main biological functions when it is secreted and activates five specific G protein-coupled receptors (S1PRs) located on the cell surface. These S1PRs exhibit tissue-specific expression and may play an oncogene or tumor suppressor role depending on the cellular context and the downstream signaling pathway activated [14, 15]. The ABC family of transporters mediates S1P released from cells. In breast cancer, estradiol can induce the export of S1P mediated by ABCC1 (ATP binding cassette subfamily C member 1) and ABCG2 (ATP Binding Cassette Subfamily G Member 2) transporters [16]. Recently a more specific S1P transporter has been found, the sphingolipid transporter 2 (SPNS2) [17]. In vivo, SPNS2 plays a role in the control of blood and lymph S1P levels [18].
The greatest interest in sphingolipids signaling pathway, especially ceramides and S1P, is that their balance in the intracellular environment is crucial to define the fate of the cell, in other words, whether apoptosis by ceramide will be activated or not by increasing S1P [19]. Therefore, it is essential to study the genes involved in the different stages of metabolism and signaling of ceramide and S1P. Increased levels of SphK1 have been shown in head and neck cancer, in association with worse prognosis [20]. However, in oral cancer, knowledge of other genes that regulate S1P/ceramides is scarce.
The present study aimed to examine the mRNA levels of several sphingolipid-related genes encoding sphingosine kinases, ceramide kinases, ceramidase, sphingomyelin synthase, S1P receptors, S1P transporters, and SET protein in oral cancer tissues, and to analyze their potential association with clinical factors and prognosis in oral cancer (OC) patients.
Materials and methods
Ethics approval
This study was approved by the Human Research Ethics Committee of FCFRP-USP and ICESP/FM-USP (CEP/FCFRP; CAAE: 90532418.6.0000.5403, CAAE: 90532418.6.3002.0065). The tissue samples were obtained from patients during the oncological surgery at ICESP/FM-USP, Brazil, and were immediately stored in the sample bank.
Tissue specimens and patient information
Total RNA isolated from 50 oral cancer (short for oral squamous cell carcinoma) specimens and their corresponding surgical margins were obtained from the tissue bank of the Cancer Institute of São Paulo (ICESP), School of Medicine, University of São Paulo, Brazil. The tissue samples were obtained from patients during the oncological surgery at ICESP and were immediately stored in the sample bank. All patients signed a written consent form. Oral cancer (OC) patients included in these studies were admitted at ICESP from 2017 to 2019 and comprised 42 men and 8 women, and the mean age was 58.9 (age range, 38–90 years).
The exclusion criteria to select the cases were the insufficient amount of biological samples from both matched tumor and surgical margin and low quality of extracted RNA. Consequently, all patients with available samples with high RNA quality were included. For each patient, data concerning the following clinicopathological parameters were gathered: gender, age, smoking, metastasis, capsular extravasation, perineural invasion, inflammatory infiltrate intratumoral, desmoplastic, cancer stage (TNM), adjuvant therapy, locoregional relapse, disease-free interval, and survival status. Cancer stage was classified according to the staging criteria proposed in the 8th edition of the TNM Classification of Malignant Tumors of the Union for International Cancer Control [21].
Quantitative RT-PCR (qRT-PCR)
RNA samples were prepared by the tumor biobank staff using AllPrep
TCGA data
The association between the expression of S1P-related genes and overall survival and disease-free survival in Head-Neck Squamous Cell Carcinoma (HNSC) patients was evaluated using HNSC data from The Cancer Genome Atlas (TCGA). RNA-Seq data of S1P-related genes and survival data of HNSC patients from the TCGA cohort were downloaded using the UCSC Xena Functional Genomics Explorer. Gene expression modulation was calculated as Fold Change of tumors samples (
Bioinformatics and statistical analysis
The heatmap from gene expression data were plotted using ggplot2 package (v3.3.2) R version 4.0.0. The Euclidean distance of clusters was determined using complete-linkage clustering method. Univariate and multivariate Cox proportional hazards regression models were adopted in this study for evaluating the correlation of survival time and disease-free interval with one or more covariates. Cox regression models were implemented using the survival package (v3.2-3) in R. The survival and disease-free curves were drawn by the Kaplan-Meier method and compared applying the log-rank test. The survminer package (v0.4.7) in R was utilized to plot Kaplan-Meier curves. Gene signatures from the expression profile of the S1P-related genes were generated and analyzed using lattice (v0.20-41), gdata (v2.18.0), bitops (v1.0-6) and caTools (v1.17.1) packages in R. The pROC package (v1.16.2) in R was used to perform receiver operating characteristic (ROC) analysis and to calculate the area under the curve (AUC). All bioinformatics analyses were conducted in RStudio (v1.3.959). Gene expression modulation between 2 groups was analyzed by two-tailed Student’s
Results
SPL-related genes expression in oral cancer patient tissues
In order to verify the potential role of sphingolipids pathway genes in oral cancer development and progression, we first compared the expression levels of 14 SPL-related genes among 50 OSCC tissues and their paired surgical margin tissues. As shown in Fig. 1, SET, ACER3, SK1, and S1PR5 were the most highly expressed genes in OSCC, with 19, 20, 25, and 29 tumor samples showing up-regulation of at least 2-fold, respectively. On the other hand, ABCG2, S1PR1, ABCB1, and SPNS2 were the most down-regulated genes in OSCC, with 30, 34, 36, and 41 tumor samples demonstrating at least 2-fold reduction, respectively (Fig. 1).
Heatmap showing the expression profiles of 14 Sphingolipid-related genes in 50 OC patient samples. The expression levels of SPL-related genes are showed as Fold Change (Log2) of tumor vs matched surgical margin.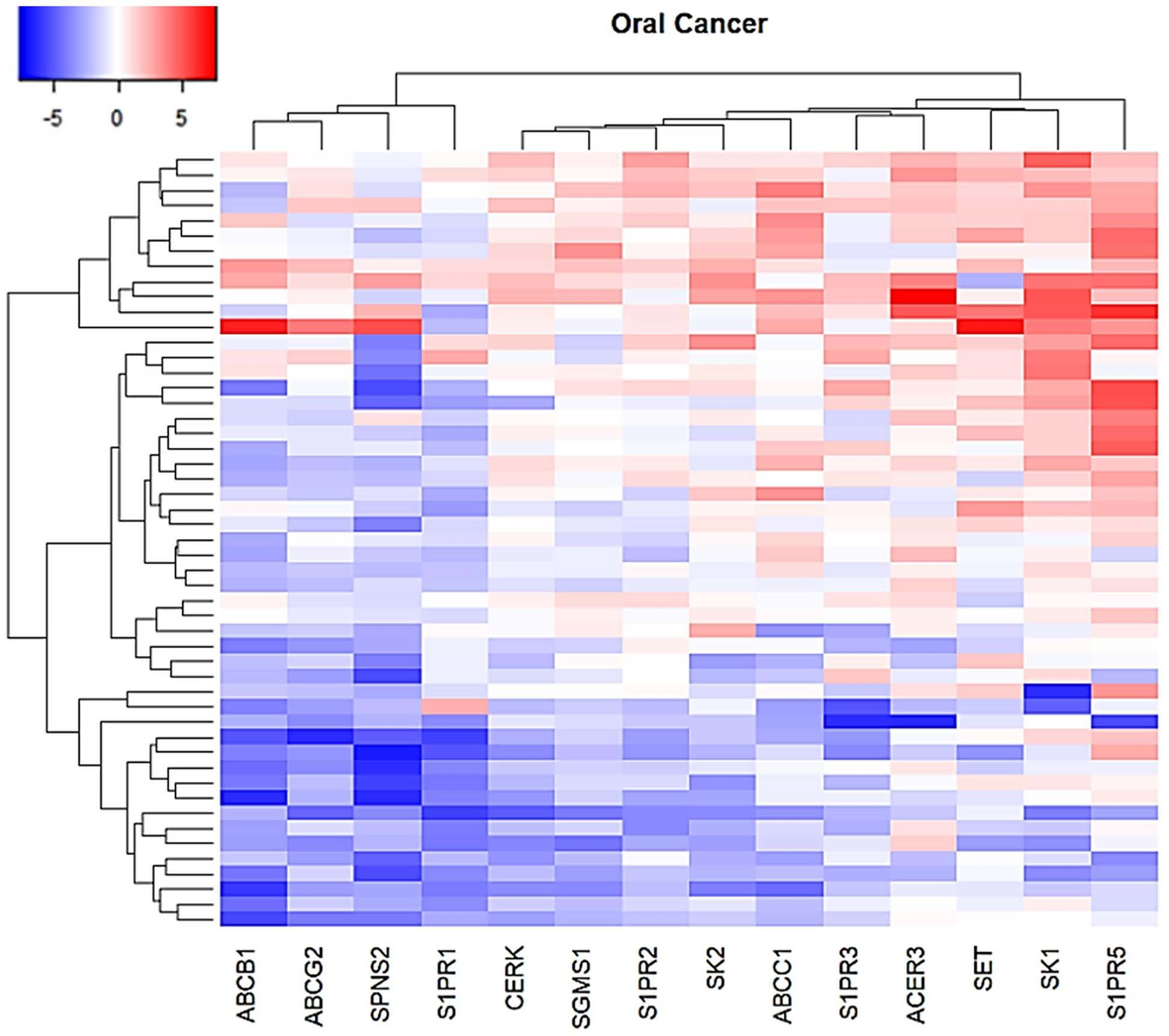
The expression levels of Sphingolipid-related genes in TCGA head and neck samples as compared to solid tissue normal samples. Heatmap showing the expression profiles of 14 genes in 520 tumor tissues. Expression data as Fold Change (Log2) of tumor samples (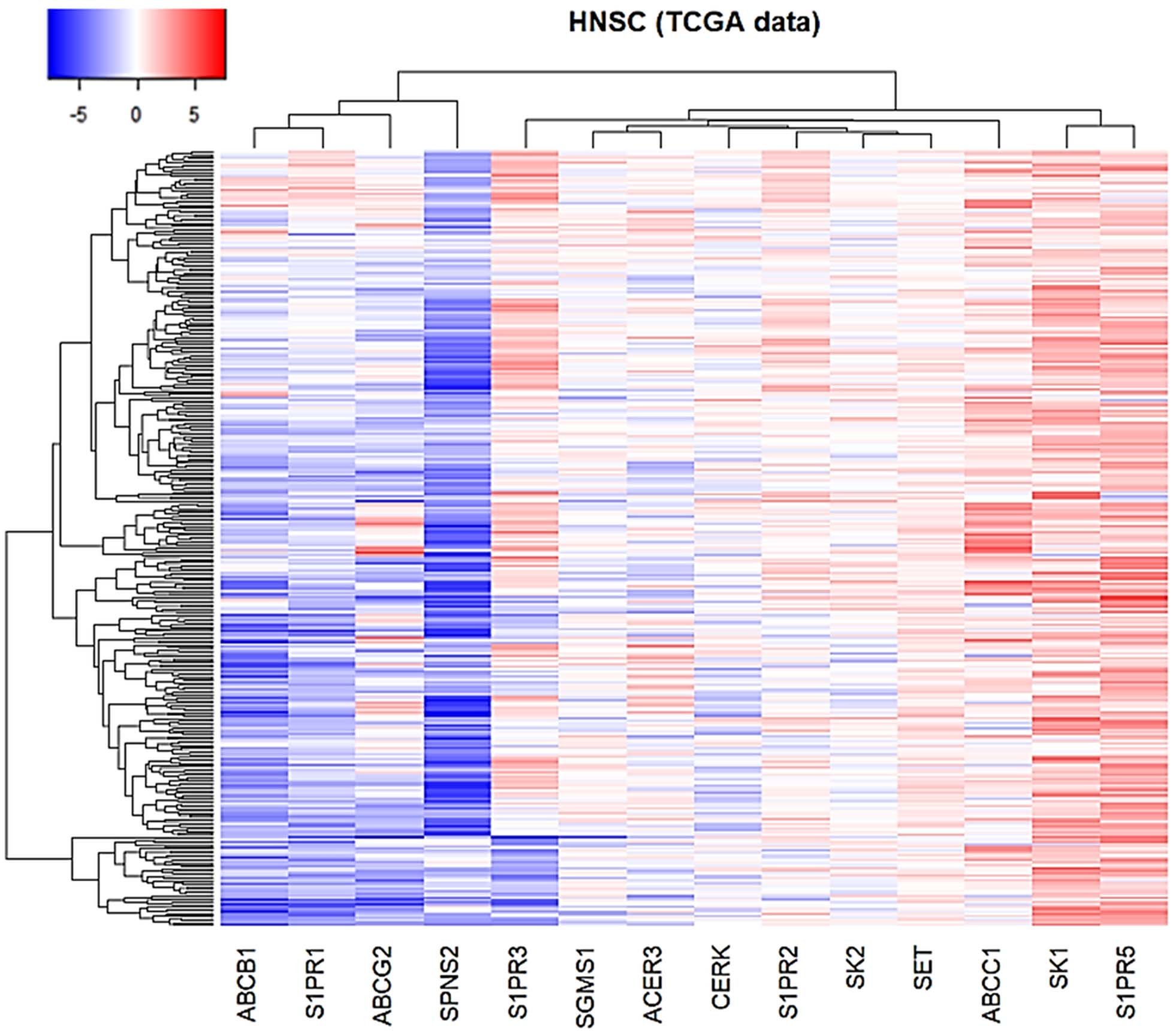
Clinicopathological characteristics of oral cancer patients
Comparison of the Sphingolipid-related genes expressions in OC tissue with different clinicopathological parameters. a. Extracapsular invasion. b. Desmoplasia. c. Lymph node metastasis. d. Perineural invasion. e. Locoregional relapse. f. Disease status.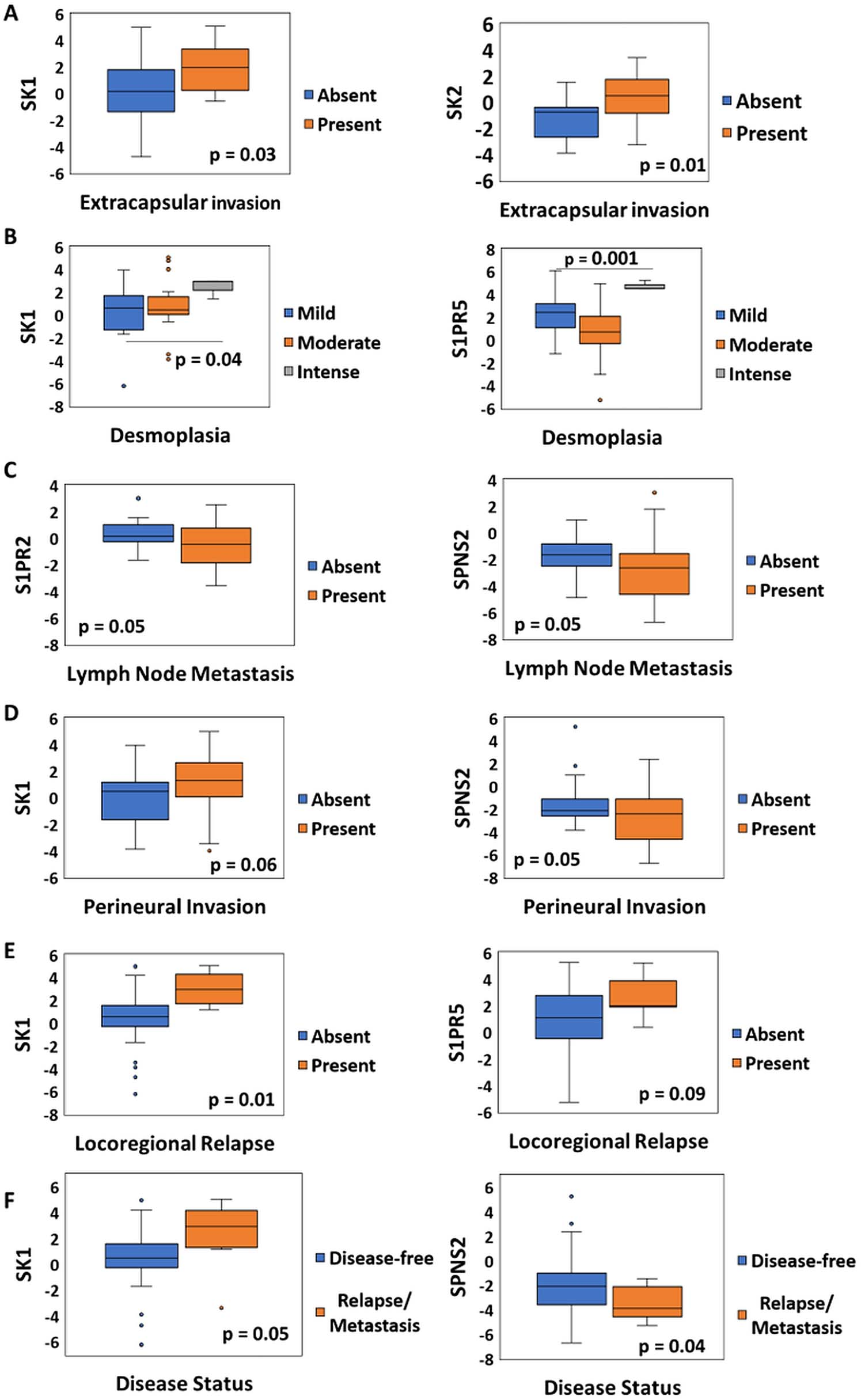
Next, we examined the levels of S1P-related genes expression in HNSC samples (
Realizing that SPL-related genes were differentially expressed in oral cancer tissues, we further attempted to analyze the associations between gene expression profile and clinicopathological parameters. We investigated the relationships of gene expression with all clinicopathological parameters of OC patients (Table 1).
Among them, seven parameters demonstrated association with gene expression. Figure 3 shows the top 2 genes with the greatest relationship to each of these clinicopathological characteristics. Oral tumors with extracapsular invasion showed higher mRNA levels of SK1 (
To further elucidate whether SPL-related genes expression is associated with overall survival (OS) and disease-free survival (DFS) of OC patients, we used Kaplan-Meier method and log-rank test to estimate the survival probability. Survival curves of the enrolled OC cohort were plotted in Figs 4 and 5 and S2-S3. The survival analysis showed that OC patients with high SK1 expression presented poorer outcomes regarding DFS compared to those with lower SK1 expression (
Kaplan-Meier analysis of overall survival (OS) in patients with oral cancer based on Sphingolipid-related genes expression level. Kaplan-Meier curves for evaluating the relationship between OS and expression of a. SK1. b. SK2. c. ACER3. d. S1PR2. e. SPNS2.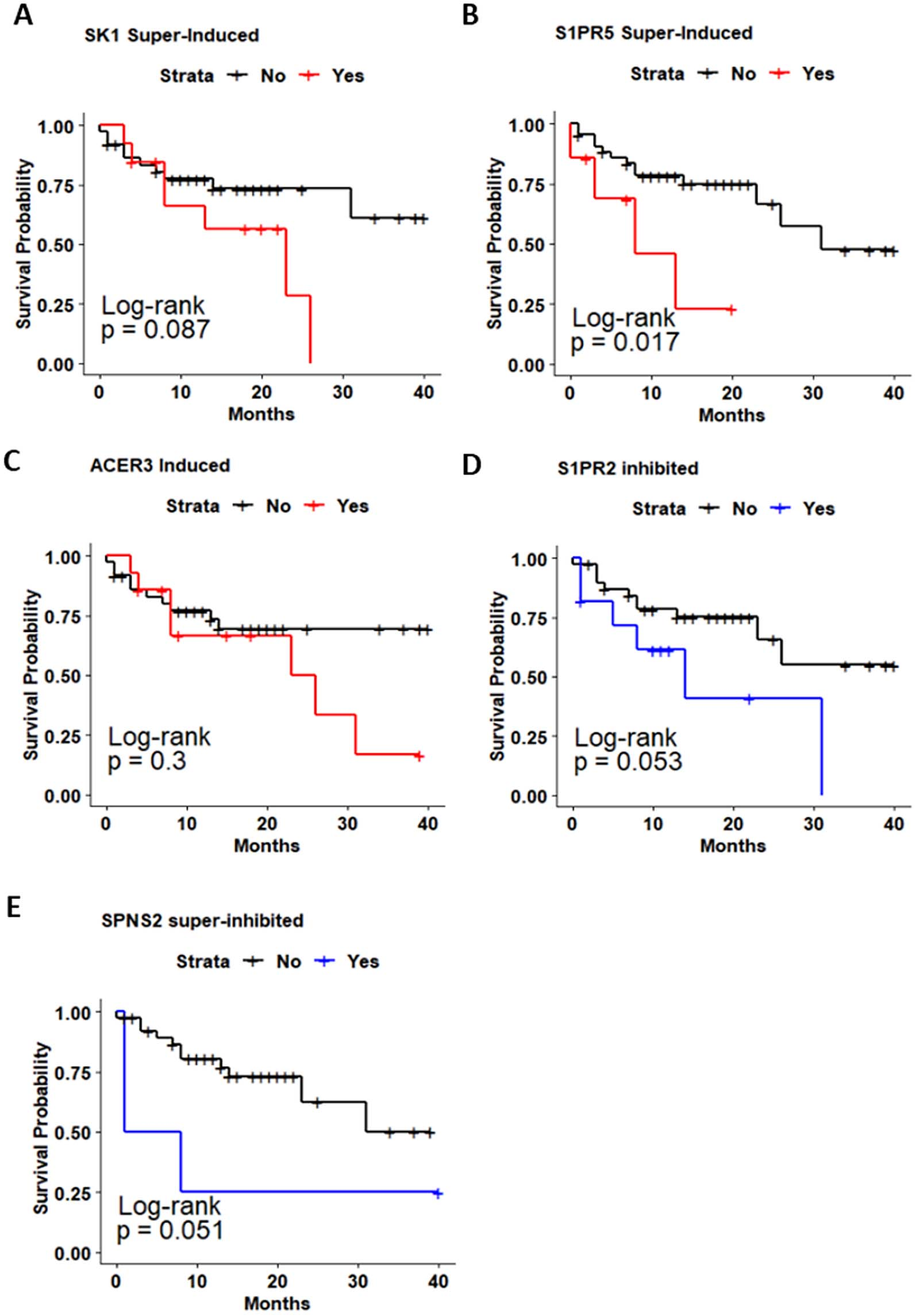
Kaplan-Meier analysis of disease-free survival (DFS) in patients with oral cancer based on Sphingolipid-related genes expression. Kaplan-Meier curves for evaluating the relationship between DFS and expression of a. SK1. b. SK2. c. ACER3. d. S1PR2. e. SPNS2.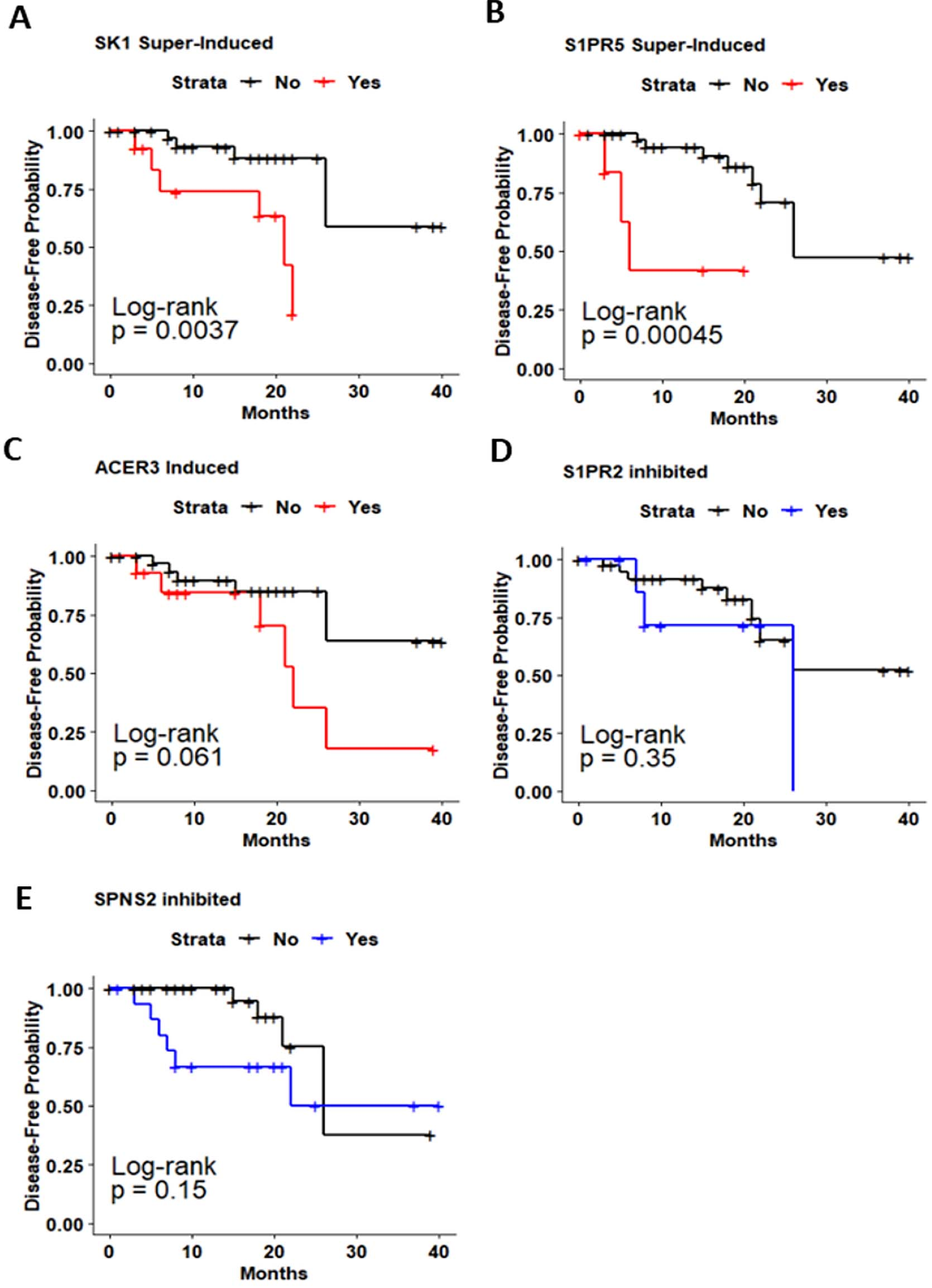
In order to confirm the association of SPL-related genes expression with OC prognosis, we performed a survival analysis in the HNSC-TCGA dataset. Kaplan-Meier curves of the HNSC cohort are shown in Figs 6, 7 and S4. HNSC patients with elevated SK1 expression presented worse prognosis regarding DFS (
Due to the relationship of SPL-related genes expression with the survival ratio of OC patients, we subsequently performed Cox proportional hazard regression analysis to investigate whether gene expression could act as an independent prognostic factor for OC patients. First, univariate regression analysis of clinicopathological parameters (Table 2) indicated that age
Univariate Cox proportional hazard regression of potential clinicopathological prognostic factors in oral cancer patients
HR: hazard ratio, CI: confidence interval.
Kaplan-Meier analysis of overall survival (OS) in patients with head and neck cancer (HNSC-TCGA) based on Sphingolipid-related genes expression. Kaplan-Meier curves for evaluating the relationship between OS and expression of a. SK1. b. SK2. c. S1PR1. d. S1PR2. e. S1PR3. f. ACER3. g. SPNS2. h. ABCC1. i. ABCB1. j. SET.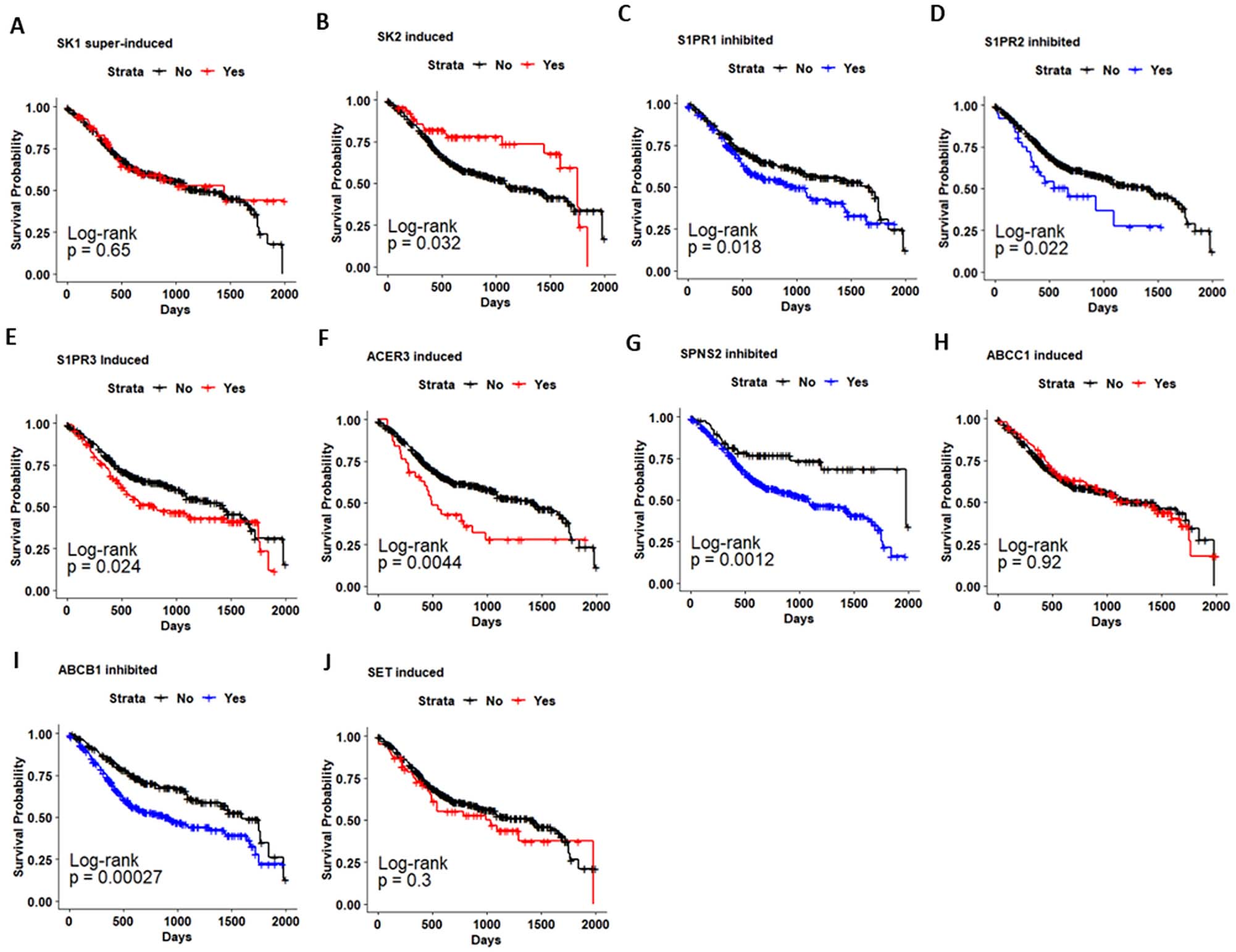
Multivariate Cox proportional hazard regression of potential gene expression prognostic factors in oral cancer patients
HR: hazard ratio, CI: confidence interval.
Kaplan-Meier analysis of disease-free survival (DFS) in patients with head and neck cancer (HNSC-TCGA) based on Sphingolipid-related genes expression. Kaplan-Meier curves for evaluating the relationship between DFS and expression of a. SK1. b. SK2. c. S1PR1. d. S1PR2. e. S1PR3. f. ACER3. g. SPNS2. h. ABCC1. i. ABCB1. j. SET.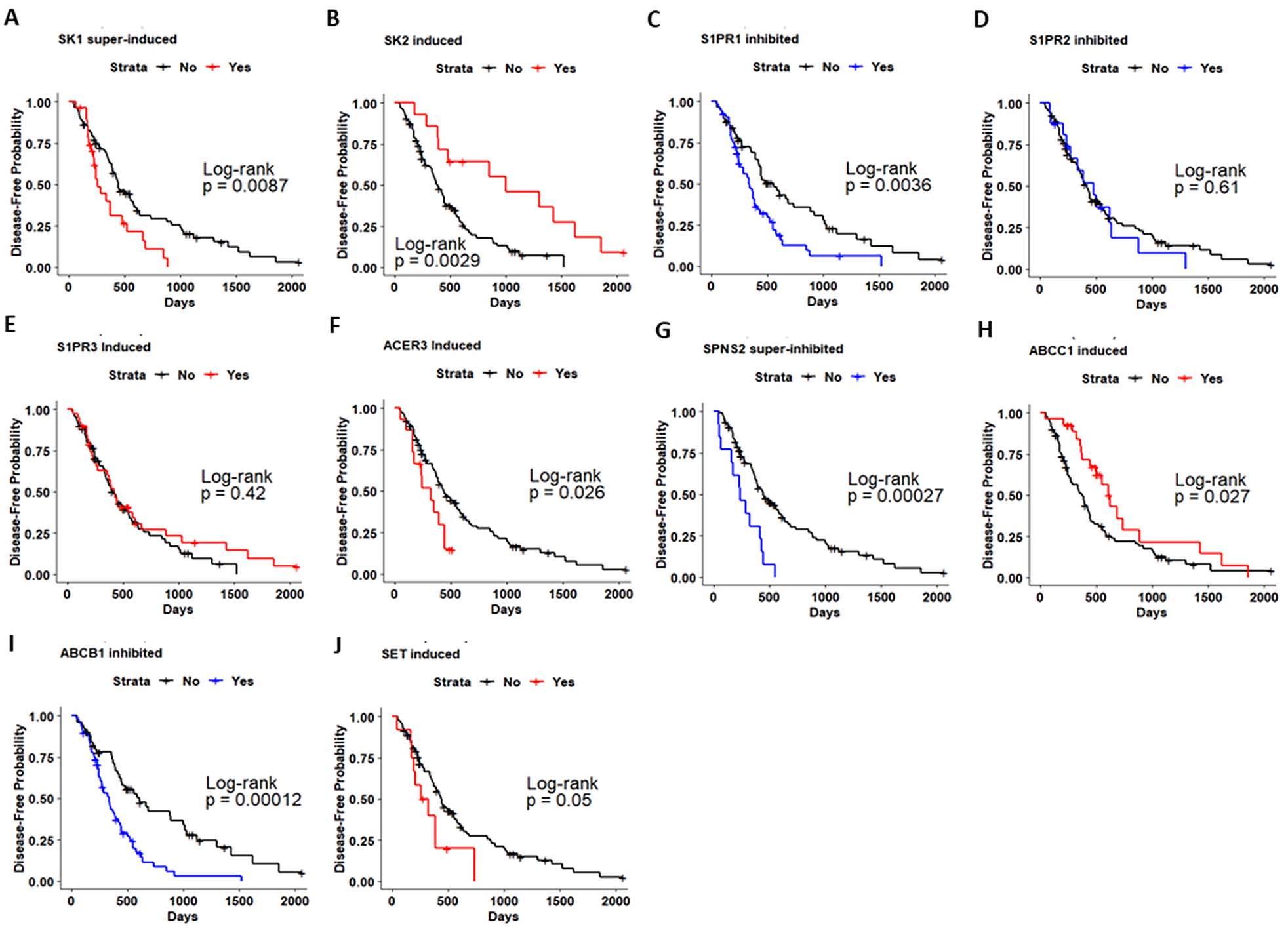
Next, we performed multivariate regression analysis, where clinicopathological factors associated with survival in the univariate analysis were included in the multivariate model along with each gene investigated (Table 3). Multivariate analysis indicated that high S1PR5 expression was associated with OS (HR
Multivariate Cox proportional hazard regression of potential gene expression prognostic factors in the HNSC-TCGA cohort
HR: hazard ratio, CI: confidence interval.
Multivariate Cox proportional hazard regression of mRNA signatures in oral cancer and HNSC patients
HR: hazard ratio, CI: confidence interval.
Following, multivariate analysis was also conducted on the HNSC-TCGA dataset (Table 4). High levels of SK1 (HR
To further explore the relationship between the genes and survival in OC, we generated gene signatures (combinations of genes) and analyzed their relationships with overall survival. Through the combination of 14 genes, 16383 mRNA signatures were generated, and some demonstrated a relationship with OS in both the OC and HNSC cohorts. Multivariate Cox regression analysis indicated that the mRNA signature I (SPNS2, ABCG2 and ABCB1), II (SK2 and S1PR5), III (S1PR3, S1PR5 and SK1) and IV (S1PR3, S1PR5 and ACER3) were prognostic factors for lower OS of patients from our OC cohort and HNSC-TCGA (Table 5).
Kaplan-Meier curves showed that OC and HNSC patients with mRNA signature I (
Kaplan-Meier analysis estimates the overall survival of oral cancer (a–d) and HNSC patients (e–h) using the Sphingolipid-mRNA signatures.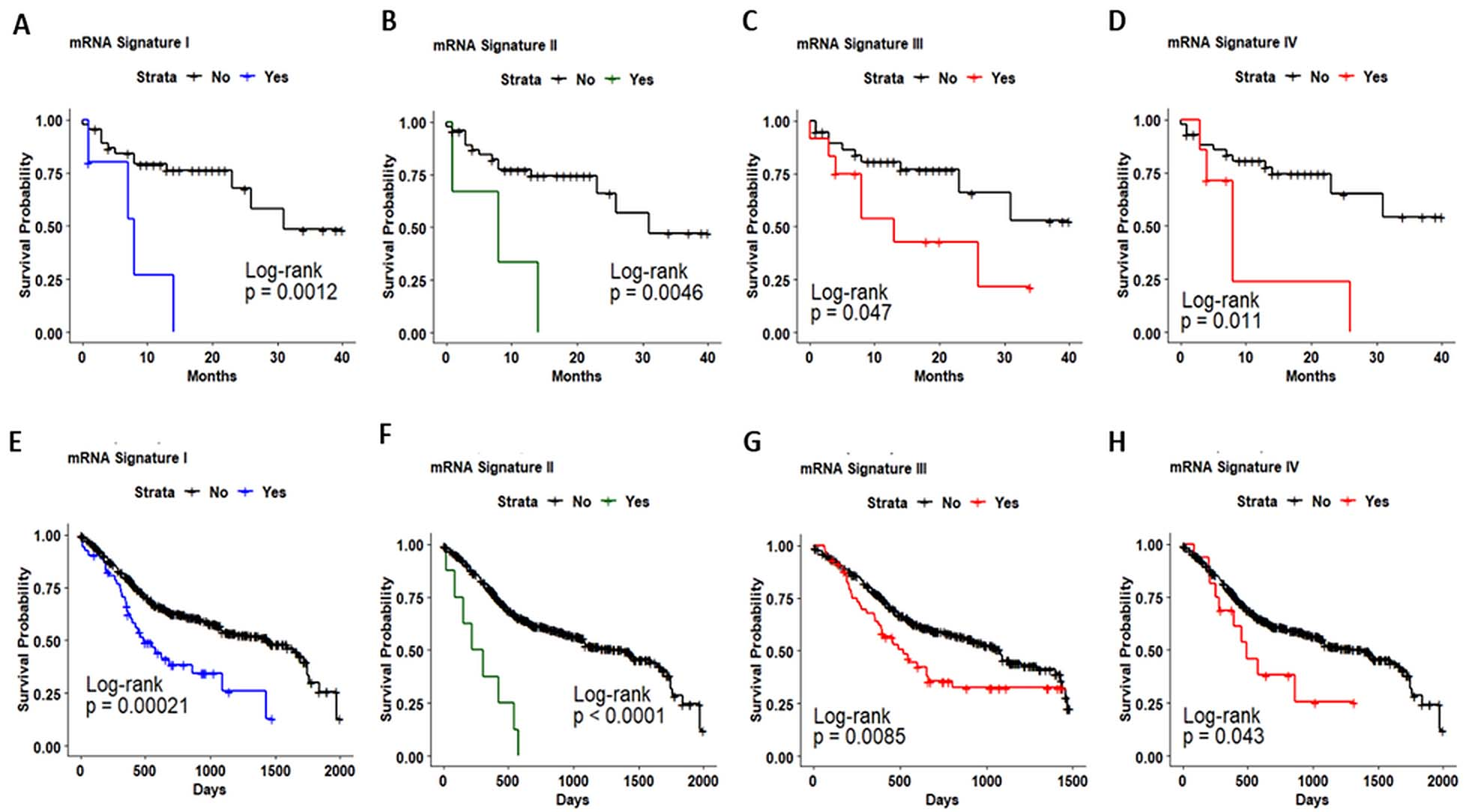
ROC curves of Sphingolipid-mRNA signatures in oral cancer and head and neck cancer. ROC analysis of sensitivity and specificity using the four selected mRNA signatures in predicting the overall survival in OC (a–d) and HNSC patients (e–h).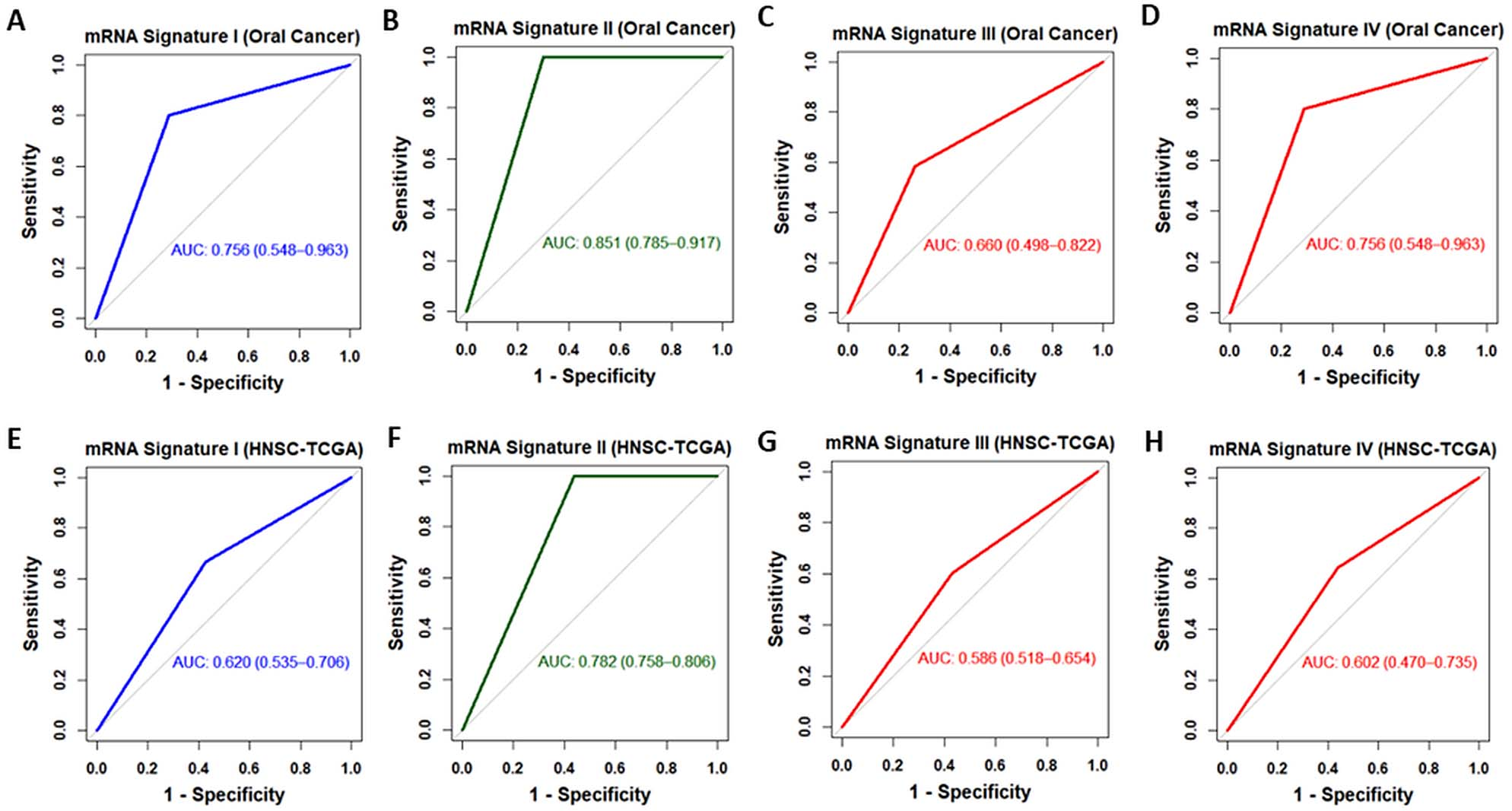
Furthermore, ROC curves were constructed by assessing the sensitivities and specificities of the identified prognostic mRNA signatures to predict survival in OC. ROC analysis indicated that mRNA signatures I and II had good predictive power (AUC
Previous studies have revealed that sphingolipids (SPL) are related to cancer development and progression in numerous tumor types [22, 23, 24]. However, the role of SK1 is the most studied, while other genes have not been entirely addressed, especially in oral carcinoma. In the present study, we thoroughly examined 14 genes of the SPL metabolism and signaling to determine their association with clinicopathological parameters and prognosis of two independent cohorts of oral cancer (OC) and head and neck cancer patients (HNSC). Through analyzing the mRNA expression of these 14 genes in a cohort of OC (
In contrast, ABCG2, S1PR1, ABCB1 and SPNS2 were the most down-regulated genes in oral tumors. Next, comparing these findings with the level of S1P-related genes expression in 520 HNSC-TCGA samples, we found a very consistent expression profile between OC and HNSC cohorts. SK1 and S1PR5 were highly expressed in HNSC tissue. While ABCG2, S1PR1, ABCB1 and SPNS2 were the most repressed genes in HNSC tissue. Moreover, in HNSC tissues, the genes S1PR3 and ABCC1 were predominantly induced. The up-regulation of SK1 and SET in OC has been reported before, but as far as we know, this is the first study describing that S1PR5 is up-regulated and S1PR1 and SPNS2 are down-regulated at mRNA level in OC tissues and HNSC.
We then profoundly investigated the relationship of SPL-related genes expression with the clinicopathological traits and survival of OC patients. We found that SK1 and SPN2 were the genes that correlated with more clinicopathological features. High levels of SK1 were associated with capsular extravasation, intense desmoplasia, locoregional recurrence, and disease status. The escape of cells from the lymph node metastasis (extracapsular extension) can facilitate the spread of tumor cells, increase the chances of locoregional recurrence, and predict a poor prognosis in patients with some types of head and neck cancer [25, 26].
Desmoplasia is a fibro-inflammatory process that can interfere with cancer treatment. In the liver, SK1 can induce fibrosis by increasing angiogenesis via the activation of the S1PR1 and S1PR3 receptors [27]. Reducing or blocking desmoplasia may be a strategy to improve sensitivity to chemotherapy by increasing blood perfusion and drug distribution within tumor tissue [28]. In turn, reduced SPNS2 levels in OC patients were associated with lymph node metastasis, perineural invasion, and disease status. Lymph node metastasis also had an association with low levels of CERK and S1PR2 in OC cohort. The metastatic lymph node burden is the most important prognostic factor in OC and is associated with shorter survival [29, 30]. The repression of S1PR2 is related to the invasion of esophageal squamous cell carcinoma and it may have an indirect role in the formation of lymph node metastasis [31]. Inhibition of S1PR2 is also implicated in the increased thyroid cancer cell migration via Rho and ROCK signaling [32]. No relationship between SPNS2 and metastasis in lymph nodes has been observed yet, but it has been shown that SPNS2 repression can increase cell migration via the accumulation of intracellular S1P [33], which can increase the potential for metastasis.
Furthermore, we found that SPL-related genes expression was correlated with the survival of OC patients. The overexpression of SK1 and S1PR5 was correlated with poor OS and DFS in OC patients. The high expression of SK1 is related to a poor prognosis in head and neck carcinoma [34], what endorses our study. There are no studies relating to the expression of S1PR5 to survival in OC. However, high S1PR5 expression is associated with lower survival in primary glioblastoma patients [35].
Moreover, we found that strong suppression of S1PR2 and SPNS2 was related with poor OS in OC patients. In diffuse large B-cell lymphoma (DLBCL), the down-regulation of S1PR2 was associated with lower survival in 3 patients cohorts, which was coherent with proapoptotic properties of S1PR2 in DLBCL [36]. In turn, SPNS2 was decreased at the mRNA level in advanced lung cancer, and the SPNS2 inhibition was involved in enhanced lung cancer cell survival [33].
We also performed survival analysis in HNSC-TCGA dataset, in order to validate our data from the OC patients regarding the expression of SPL-related genes. In agreement with that observed in the OC cohort, HNSC patients with high levels of SK1 and low levels of S1PR2 and SPNS2 had worse prognosis for survival. In addition, HNSC patients with reduced expression of SK2, S1PR1 and ABCB1 and elevated expression of S1PR3, ACER3, ABCC1, and SET also presented unfavorable prognosis. SK2 inhibition was not frequent in HNSC patients as it was in OC patients. SK2 inhibition was a rare event; however, it showed a strong correlation with lower survival. The role of SK2 in tumor development is contradictory. In some studies, SK2 acts by suppressing cell growth and in some other studies, it acts to promote cell proliferation. There seems to be an inverse relationship between the levels of SK2 and the formation of xenograft tumor in nude mice, attributing to SK2 two potential antagonistic roles: oncogenic (at a low level in cells) and tumor suppressor (at a high level) [37]. Endorsing this, it has been shown in breast cancer that patients with low SK2 expression have a worse prognosis [38].
As well as observed in HNSC patients, the inhibition of S1PR1 is also related to shorter survival in patients with glioblastoma. Besides, the silencing of S1PR1 in glioma cells promoted cell proliferation [39]. Recently a potential tumor suppressor role of S1PR1 has been demonstrated in breast cancer cells and that higher levels of S1PR1 improve the prognosis of breast cancer patients [38].
Usually the high expression of ABCB1 is related to cancer development and poor prognosis in cancer patients. However, in our OC cohort and HNSC-TCGA cohort, ABCB1 gene was significantly down-regulated, and in HNSC patients this inhibition was related to poor survival. Warta et al. analyzing 40 samples of head and neck carcinoma identified ABCB1 inhibition and its relationship with lower OS and DFS, in that study they also showed the association between up-regulation of ABCC1 and DFS [40].
The up-regulation of ACER3 was more common in OC than in HNSC-TCGA patients. In OC patients, ACER3 may have a relationship with DFS but it did not reach statistical significance (
In previous studies, our group have found that SET is overexpressed at protein level in HNSCC tissues [10]. In the present study, SET mRNA was moderately induced in OC and HNSC cohorts, and we found that HNSC patients with higher levels of SET showed shorter disease-free survival. Although the SET protein has not directly linked to S1P signaling, its function as a PP2A inhibitor and regulator of several proteins such as Akt [10], NF-kB [42] and ERK [43] suggests it Besides, our ongoing studies suggest (data not shown) that SET can interfere with S1P signaling in oral cancer.
Performing multivariate Cox regression analyses in OC and HNSC dataset, we found that SPL-related genes might be potential independent prognostic factors for OC, in particular, the genes SK1, SK2, S1PR1-3, SPNS2 and ABCB1. To further validate our findings, we created gene signatures formed of the combination of the 14 genes examined. Kaplan Meier analysis and Cox regression model showed 4 signatures associated with lower overall survival in OC and HNSC, and their potential to be prognostic factors. Moreover, ROC curves indicated that the mRNA signatures identified in the present study have effectiveness in predicting survival in OC.
Taken together, the results of the present study demonstrated that several sphingolipid-related genes are deregulated in OC and HNSC, and their expression profile are associated with patient survival and may prove to be useful prognostic markers and therapeutic targets for OC and HNSC. To the best of our knowledge this study showed for the first time that S1PR1 and SPSN2 are strongly down-regulated in OC and HNSC at the mRNA level and might be biomarkers for worse OC prognosis.
Author contributions
Conception: Gabriel da Silva (G.S) and Andréia Machado Leopoldino (A.M.L) were responsible for the conception of the article.
Interpretation or analysis of data: G.S., A.M.L, Leandro Luongo de Matos (L.L.M) and Marco Kulcsar (M.K) analyzed the data.
Preparation of the manuscript: G.S. and A.M.L wrote the manuscript.
Revision for important intellectual content: Luiz Paulo Kowalsk (L.P.K) and A.M.L. revised the manuscript.
Supervision: A.M.L was responsible for the supervision.
Supplementary data
The supplementary files are available to download from http://dx.doi.org/10.3233/CBM-203100.
sj-docx-1-cbm-10.3233_CBM-203100.docx - Supplemental material
Supplemental material, sj-docx-1-cbm-10.3233_CBM-203100.docx
Footnotes
Acknowledgments
The authors thank the patients and the technical support by the staff working in the Biobank of tumor samples at the Instituto do Câncer do Estado de São Paulo, Medical School, University of São Paulo, Brazil.
Research was supported by Fundação de Amparo à Pesquisa do Estado de São Paulo (research project 2016/19103-2, 13/08135-2; scholarship: 2018/17480-9), Conselho Nacional de Desenvolvimento Científico e Tecnológico (CNPq) and Coordenação de Aperfeiçoamento de Pessoal de Nível Superior (CAPES).
