Abstract
BACKGROUND:
Osimertinib is an epidermal growth factor receptor-tyrosine kinase inhibitor that specifically targets the T790M mutation in cancer.Unfortunately, most non-small cell lung cancer (NSCLC) patients develop osimertinib resistance. Currently, the molecular biomarkers for monitoring osimertinib resistance are not available.
OBJECTIVE:
This study aimed to examine the profile of exosomal miRNA in the plasma of osimertinib-resistant NSCLC patients.
METHODS:
Plasma exosomal miRNA profiles of 8 NSCLC patients were analyzed by next-generation sequencing at osimertinib-sensitive and osimertinib-resistance stage.The expression of dysregulated exosomal miRNAs was validated and confirmed in another cohort of 19 NSCLC patients by qPCR. The relationship between exosomal miRNA upregulation and clinical prognosis, survival analysis was evaluated by Kaplan-Meier curves.
RESULTS:
In osimertinib-resistant NSCLC patients, 10 exosomal miRNAs were significantly dysregulated compared to baseline. Upregulation of all 10 candidate exosomal miRNAs tended to correlate with increased latency to treatment failure and improved overall survival. Among them, 4 exosomal miRNAs, miR-323-3p, miR-1468-3p, miR-5189-5p and miR-6513-5p were essentially upregulated and show the potential to be markers for the discrimination of osimertinib-resistance from osimertinib-sensitive NSCLC patients with high accuracy (
CONCLUSIONS:
Our results highlight the potential role of these exosomal miRNAs as molecular biomarkers for the detection of osimertinib resistance.
Introduction
Non-small cell lung cancer (NSCLC) is the most common type of lung cancer, accounting for approximately 80% of all cases, and has become the leading cause of cancer-related death worldwide [1, 2]. At present, surgery is the recommended treatment method for patients in early stages, while chemotherapy, chemo-immunotherapy, and targeted therapy are preferred for advanced lung cancer [3]. Although multidisciplinary therapy has been used, acquired resistance remains a major problem, and the 5-year survival rate is still poor [1]. Activating mutations in the epidermal growth factor receptor (EGFR) have been reported to be key driver mutations in NSCLC and are found in approximately 10% and 50% of Caucasian and Asian patients, respectively [4, 5]. Although these mutations are initially sensitive to EGFR-tyrosine kinase inhibitors (TKIs) however most patients develop acquired resistance to TKIs within 9 to 15 months. Moreover, 60% of all resistant cases are EGFR-T790M mutation [6, 7, 8]. Recently, the third generation of EGFR-TKIs, such as osimertinib, has been prescribed for T790M-positive cases [9]. Unfortunately, patients developed resistance to osimertinib 6-17 months after treatment [9, 10]. The EGFR C797S mutation accounts for approximately 20% of all osimertinib-resistant NSCLC patients, and the treatment criteria have not been established for this patient population [11]. Hence, it is critical to properly understand the underlying resistance mechanisms to allow for further development of specific treatments for these patients. Importantly, identification of the molecular biomarkers for detecting resistance to third generation EGFR-TKIs is crucial in assisting clinicians in making decisions.
Exosomes are small membrane vesicles with a size of approximately 30–150 nm released from all cell types including tumor cells [12]. Exosomes play an important role in cell-cell communication by transferring and stabilizing a variety of functional biomolecules such as proteins, RNAs, microRNAs (miRNAs), lncRNAs, and DNAs [13, 14]. MiRNAs are endogenous small non-coding RNAs with 20–22 nucleotides in length. A large number of studies have reported the role of miRNAs in cancer progression, such as proliferation, invasion, metastasis, metabolism, and drug resistance [15]. Dysregulation expression of circulating miRNAs was found to be correlated with tumor progression [16], suggesting their potential as cancer biomarkers. Moreover, a large number of studies reported that exosomal miRNAs are involved in lung cancer progression and have, therefore, been proposed as potential diagnostic biomarkers for lung cancer [17, 18, 19, 20, 21, 22, 23]. Tumors are highly heterogeneous and evolve; thus, a tissue biopsy is commonly inadequate for comprehensive tumor characterization.To overcome these limitations, an analysis of circulating cancer-derived exosomes, often referred to as a “liquid biopsy” is making big-time waves in the field of cancer management [24]. Cancer exosomes regulate the biological functions of distant recipient cells through the transportation of specific cargos [12, 13, 14]. The secreted exosomes can be collected from body fluids including blood, CSF, saliva, breast milk, and urine [25]. Importantly, exosome content is unique and often reflects the disease state. Consequently, several studies have demonstrated the discovery of potential molecular biomarkers for lung cancer from circulating exosomes [26, 27, 28]. However, studies have not investigated the exosomal miRNAs profiles of osimertinib-resistant NSCLC patients. Therefore, the present study aimed to elucidate the expression profile of exosomal miRNAs before and after osimertinib resistance in NSCLC patients.
Materials and methods
Ethics statement
This study was approved by the ethics committee of The Human Research Ethics Committee of Ramathibodi Hospital, Mahidol University (IRB no. EC600169). All patients and healthy controls have signed written informed consent.
Patient characteristics and blood collection
All patients were diagnosed with advanced NSCLC with EGFR mutations and were treated with 1
Plasma exosome isolation
Exosome isolation was conducted using ExoQuick (System Biosciences, Palo Alto, CA, USA). Briefly, 3
Electron microscopy
A formvar-carbon-coated 300 mesh copper grid (Electron Microscopy Sciences, Hatfield, PA, USA) was discharged with UV light for 15 min before use. The exosome suspension (5
Western blot analysis
The exosome pellets were lysed with RIPA lysis buffer containing 50 mM Tris-HCl pH 7.4, 150 mM NaCl, 1 mM EDTA, 1% TritonX-100, 1 mM NaF, 1 mM Na
Small RNA extraction
Total small RNA was extracted from exosome samples using an miRNeasy mini kit (QIAGEN, Hilden, Germany) according to the manufacturer’s instructions. Briefly, 700
miRNA deep sequencing
The cDNA preparation and miRNA expression profiling were performed by QIAGEN (QIAGEN, Hilden, Germany). All small RNA extractions were packed in tightly sealed vial tubes and shipped on dry ice. Briefly, cDNA libraries were reverse transcribed from 1
Prediction miRNA target genes and pathway enrichment analysis
Target genes of dysregulated miRNAs were predicted by an online bioinformatics tool miRSystem (ver.20160513) containing 7 well-known miRNA target gene prediction algorithms: DIANA, miRanda, miRBridge, PicTar, PITA, rna22, and TargetScan [29]. Only the target genes predicted from at least three databases were chosen for further analysis. To identify significant signal transduction pathways of miRNA target genes, the putative genes were analyzed according to the KEGG (Kyoto Encyclopedia of Genes and Genomes) pathway using WebGestalt (WEB-based Gene SeT AnaLysis Toolkit) [30, 31].
Reverse transcription and quantitative real-time PCR
First-strand cDNA synthesis was performed using the miScript II RT Kit (QIAGEN, Hilden, Germany) according to the manufacturer’s instructions. Each 20
Statistical analysis
Exosomal miRNA expression levels were compared between those at baseline and those at the resistance stage of osimertinib using paired t-test. Statistical significance was considered as
Characterization of NSCLC patients
Characterization of NSCLC patients
TTF: Time to treatment failure. *Initial mutation,
The characterization of exosomes isolated from NSCLC patients and healthy controls. (A) The morphology of isolated exosomes was visualized by transmission electron microscopy. (B) The overall size distribution of exosomes derived from NSCLC patients and healthy controls was analyzed using Nanoparticle tracking analysis. (C) Isolated exosomes were probed for traditional exosome markers (Alix, flotillin-1 and CD81) by Western blot analysis.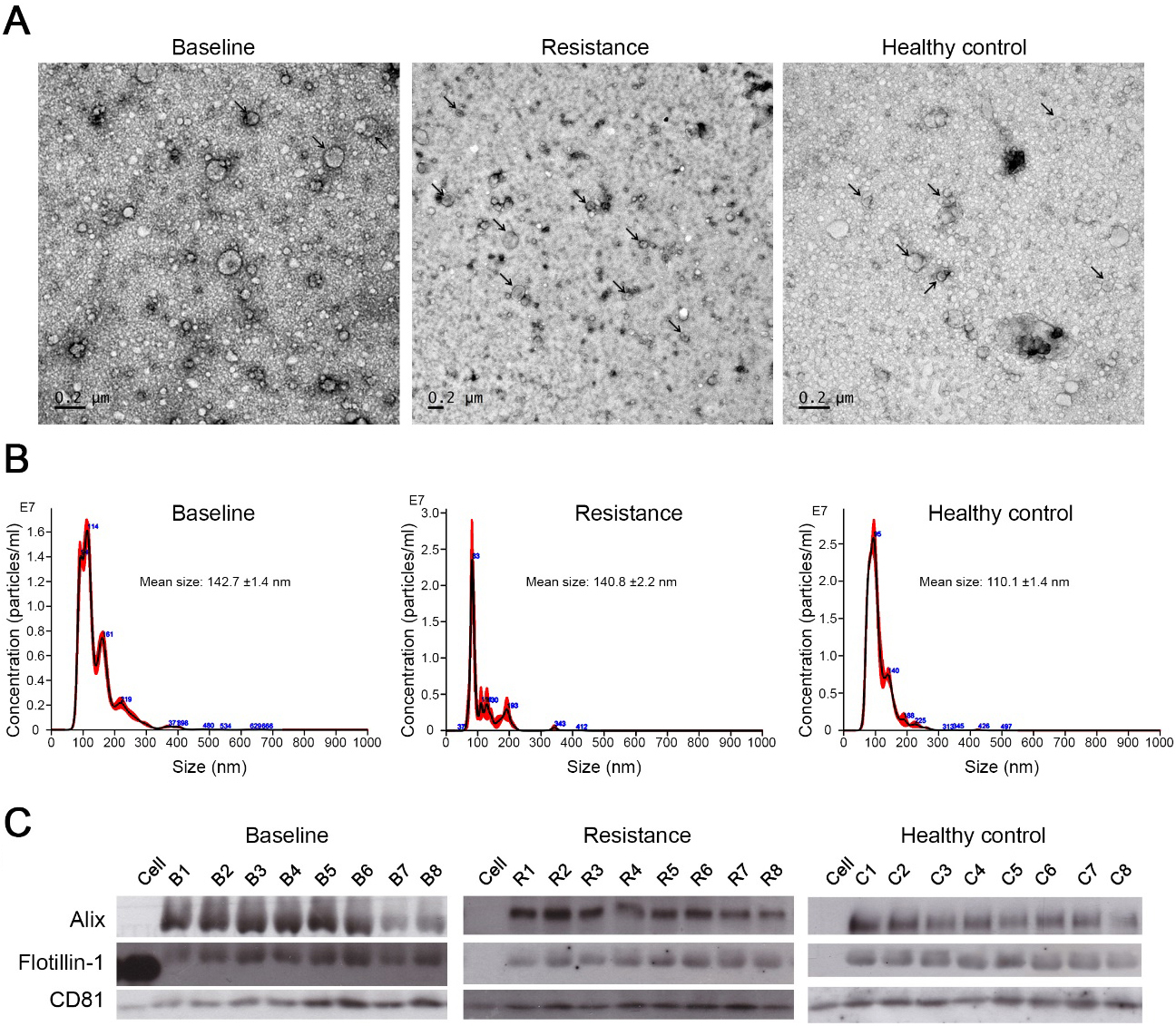
Characterization of exosomes derived from plasma of NSCLC patient
A total of 8 lung cancer patients (3 males and 5 females) were recruited to investigate exosomal miRNA profiles. All patients were treated with TKI and developed TKI-resistance harboring the T790M mutation. As shown in Table 1, the median patient age was 59.5 years. The median TTF was 11.5 months. Among them, 3 patients have EGFR L858R mutation while 5 patients have exon 19 deletion. Plasma samples were collected before osimertinib treatment (baseline) and at the initial point of the osimertinib-resistance (resistance). A total of 8 age- and sex-matched healthy controls were recruited for this study. Plasma exosomes were isolated using ExoQuick as outlined in Materials and Methods. Isolated exosome morphology was visualized using the transmission electron microscopy. In each group, exosomes displayed a spherical vesicular shape with diameters less than 200 nm (Fig. 1A). Exosome size was further confirmed by nanoparticle tracking analysis. The median size of exosomes derived from the plasma of lung cancer patients at baseline and the resistance stage were approximately 142.7
Differentially expressed exosomal miRNAs and functional annotation of dysregulated miRNAs. (A) Differentially expressed plasma exosomal miRNAs from the baseline group vs healthy controls (136 miRNAs, 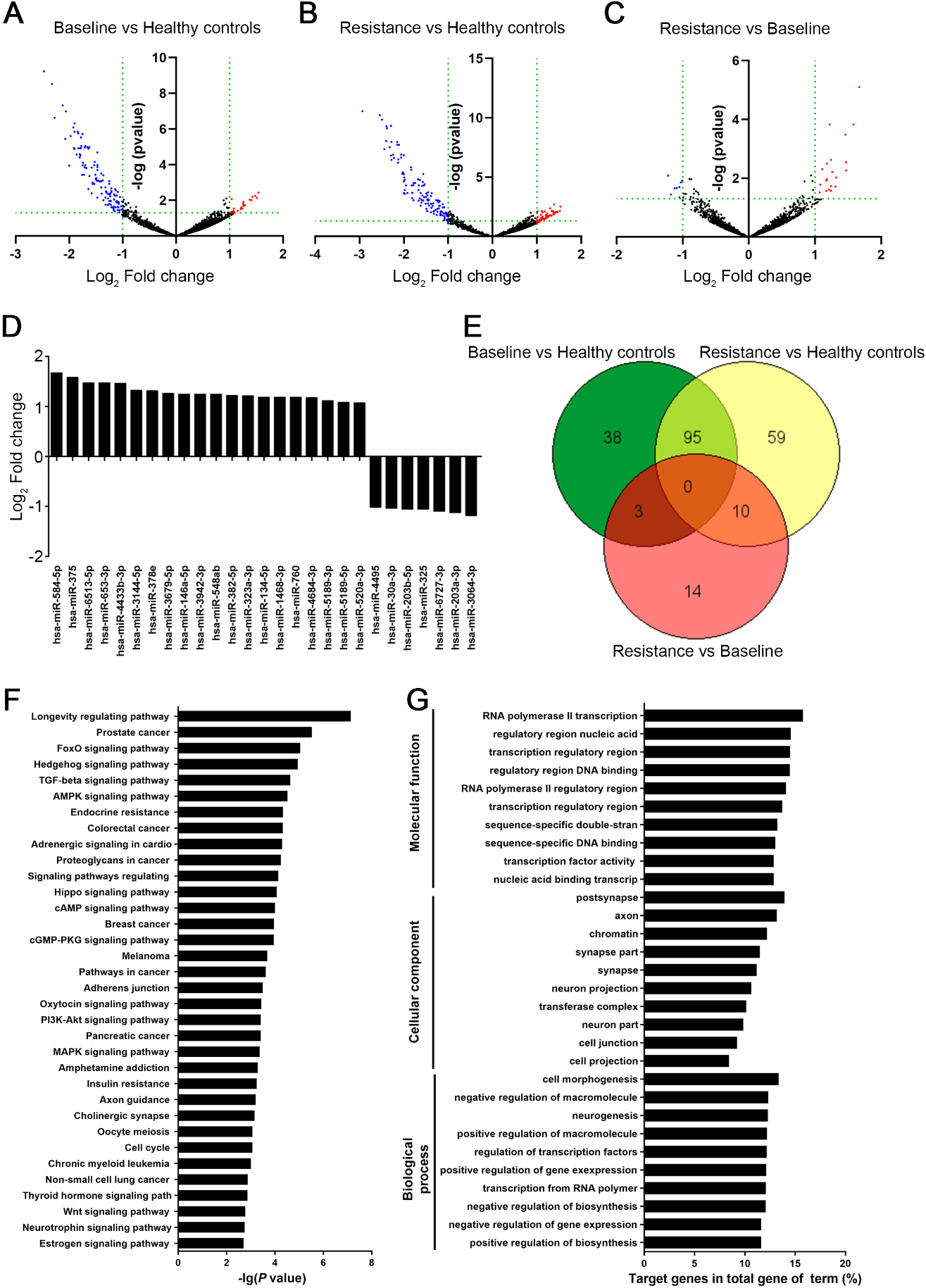
The plasma exosomal miRNA profiles of lung cancer patients before osimertinib treatment (baseline) and after the osimertinib resistance stage, as well as healthy controls, were analyzed by the NGS Illumina platform. On average, 13.1% reads across all samples (21–24 million mapped reads) mapped to the miRNAs in the human GRCh38 genome (Supplement Fig. S1). Among these, 2,409 miRNAs were characterized as known miRNAs, while 5,680 potential miRNAs were unknown. Differential miRNA expression analysis on the mapped miRNAs was analyzed with DESeq2. To remove the background from non-tumor cells, we compared the plasma exosomal miRNA profiles from lung cancer patients with healthy controls. As shown in Fig. 2A, 136 exosomal miRNAs derived from baseline lung cancer plasma were differentially expressed relative to healthy control miRNAs (
Dysregulated miRNAs in plasma exosomes of osimertinib resistance patients are associated with diverse cancer-related signaling pathways
To understand the biological functions of differentially expressed miRNAs in lung cancer patients, we predicted target genes of the 10 lung cancer-dysregulated exosomal miRNAs using the miRSystem database. We found that 1,175 putative targets were regulated by these 10 miRNAs (Supplement Table S4). Pathway analysis demonstrated that many diverse pathways were associated with these dysregulated miRNAs (adjust
Validation of the expression of candidate exosomal miRNAs
To explore the potential of these dysregulated miRNAs as useful biomarkers for osimertinib resistance, we compared the expression of these 10 lung cancer-dysregulated exosomal miRNAs of 19 lung cancer patients at the baseline and the resistance stages (Table 1). This sample group consisted of 6 males and 13 females with median age of 67 years old. The median TTF was 12 months. Among them, 7 patients have EGFR L858R mutation while 12 patients have
Validation of candidate exosomal miRNA expression in osimertinib resistance. The expression of the 10 lung cancer-dysregulated exosomal miRNAs was analyzed by quantitative real-time PCR. The miRNA expression was compared between at baseline and at the resistance stage of 19 paired samples of osimertinib-treated NSCLC patients. The data are represented as mean 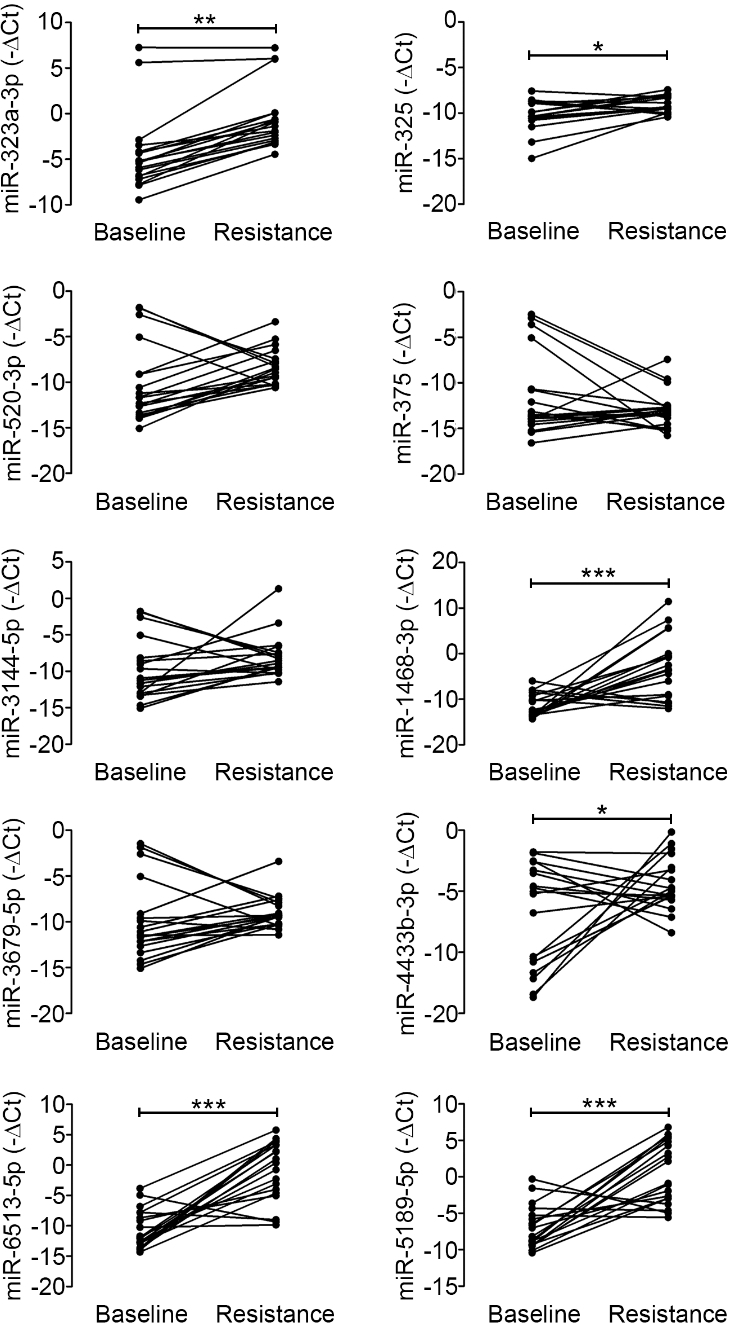
exon 19 deletion. Of these 19 NSCLC patients, all except one had acquired T790M mutation after received 1
To investigate the relationship between exosomal miRNA upregulation and clinical prognosis, survival analysis was performed on all 27 osimertinib-treated NSCLC patients (Table 1). TTF was defined as time since the initiation of osimertinib until stopping the drug due to disease progression or treatment toxicities. OS was defined as time since the initiation of osimertinib until death from any causes. The upregulation of our 4 previously identified miRNAs (hsa-miR-323-3p, hsa-miR-1468-3p, hsa-miR-5189-5p, and hsa-miR-6513-5p) was observed in 17 patients (Supplement Table S5). As shown in Fig. 5A, the median TTF of patients who had upregulation of all these 4 miRNAs was longer compared to the rest of the patients who did not have all 4 miRNAs upregulation (11.9 months vs. 8.8 months, HR, 0.72; 95% CI, 0.32–1.61,
ROC curve of individual exosomal miRNA for distinguishing between osimertinib-sensitive and osimertinib-resistance stages. 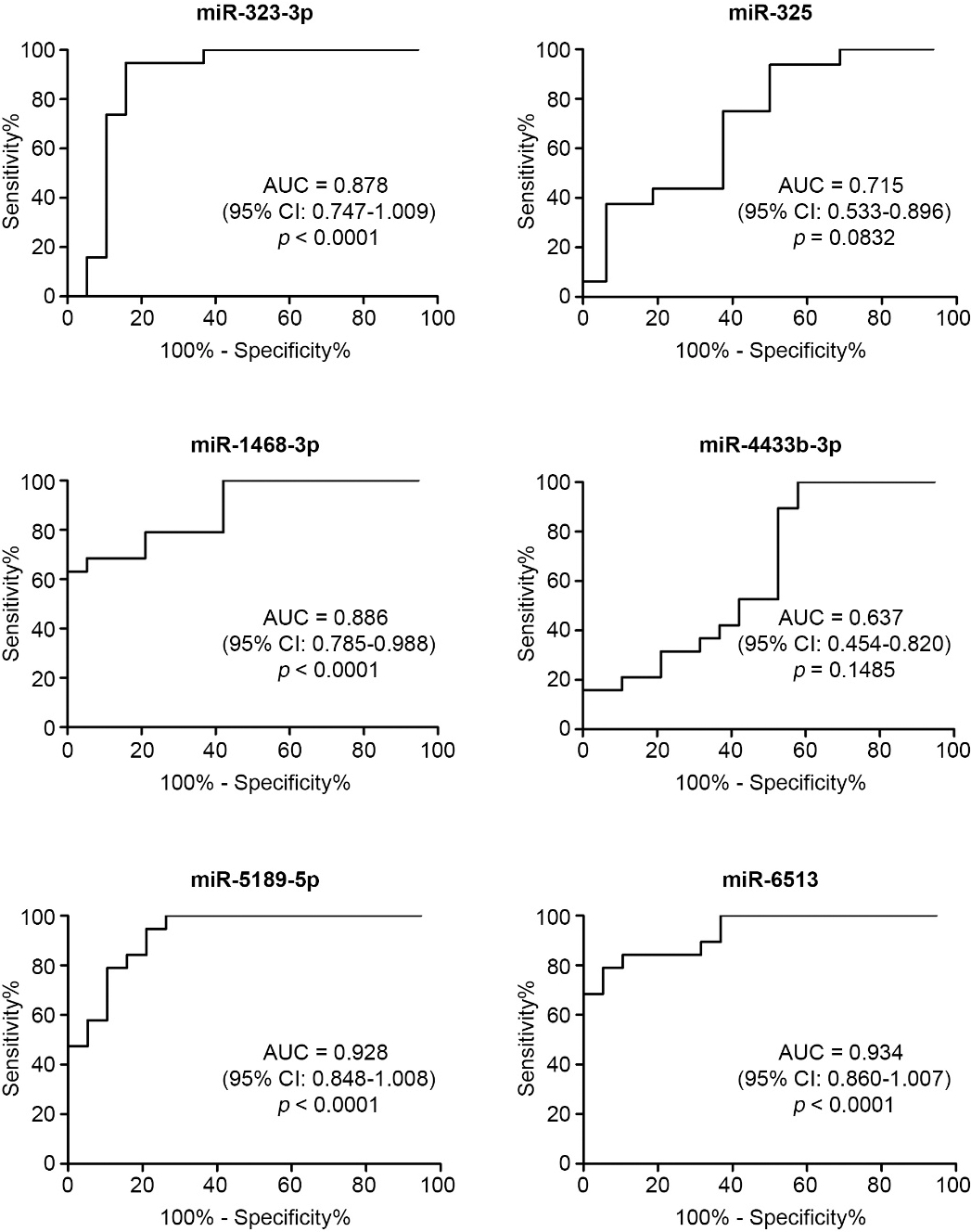
Kaplan-Meier curves according to the upregulation of exosomal miRNAs. Illustrates the TTF curves and OS curves as compared between the patients with and without the 4 upregulated miRNAs (A and B) and the patients who had all and not all 10 miRNAs upregulation (C and D). 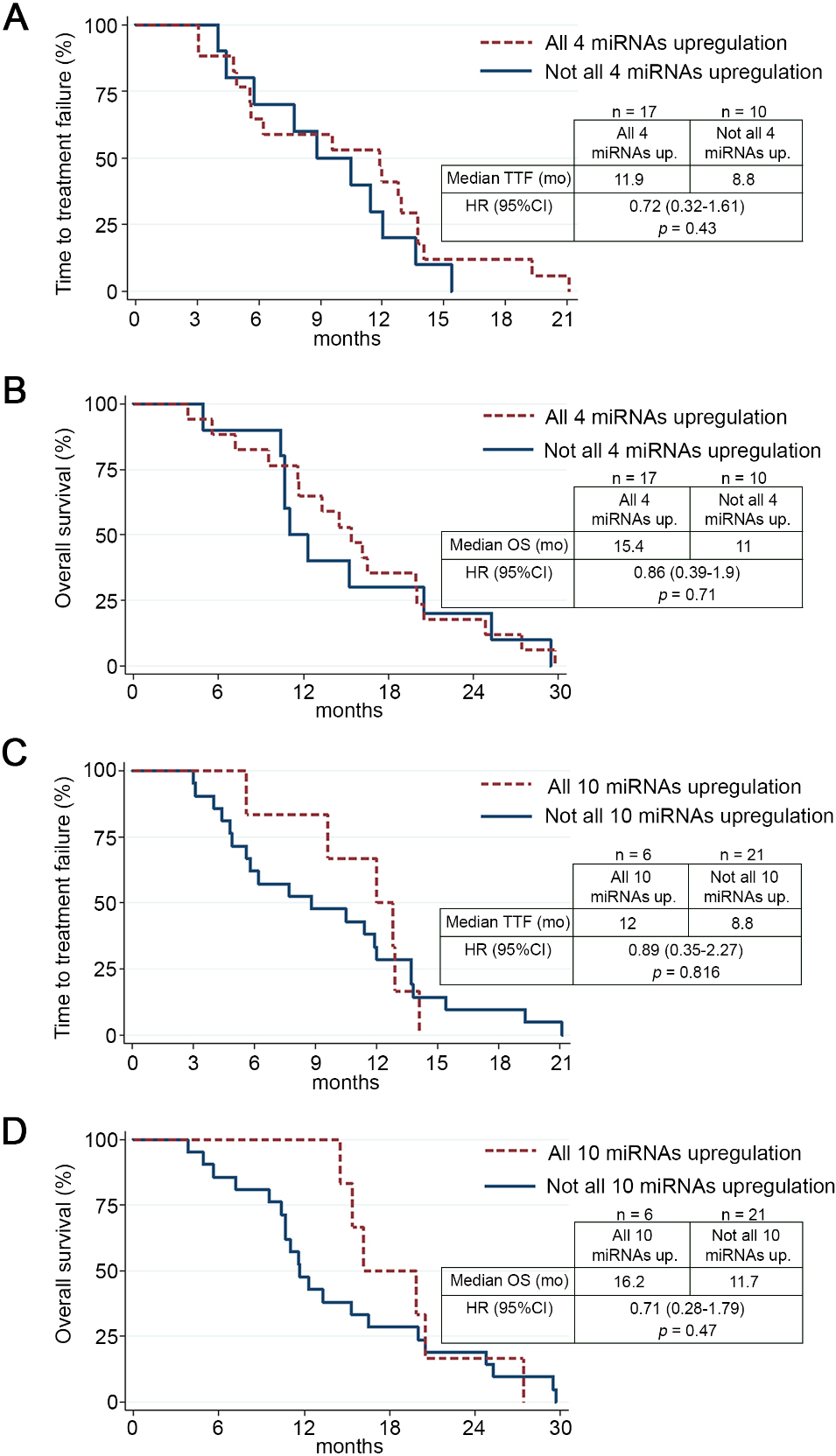
Drug resistance in cancer is a major treatment barrier, particularly for patients who display resistance to targeted drug therapy. In lung cancer, resistance to third generation EGFR-TKIs treatment is frequently observed in NSCLC patients harboring the T790M mutation [9, 10]. The lack of dependable biomarkers for predicting treatment response is a major cause of recurrence and poor prognosis in lung cancer, making it a leading cause of cancer-related death worldwide. In the present study, we aimed to develop a system for predicting resistance to third generation EGFR-TKIs. To this end, we utilized multiple emerging bioinformatic techniques to identify signaling pathways and potential molecular biomarkers for EGFR-TKI resistance. Specifically, using exosomal miRNAs isolated from the plasma of NSCLC patients, we compared miRNAs profiles at baseline (before osimertinib treatment or osimertinib responding) and the resistance stage (osimertinib resistance). Interestingly, using NGS, we demonstrated differential expression of miRNAs in exosomes isolated from osimertinib-sensitive (baseline) and osimertinib-resistant NSCLC patients. As such, we found 10 exosomal miRNAs that were statistically dysregulated in osimertinib-resistant NSCLC patients; 9 miRNAs were upregulated and 1 miRNA was downregulated. Among these 10 miRNAs, we further confirmed the dysregulation of 6 exosomal miRNAs in another lung cancer cohort. Through further ROC analysis, we found 4 exosomal miRNAs (hsa-miR-323-3p, hsa-miR-1468-3p, hsa-miR-5189-5p, and hsa-miR-6513-5p) that could be used to differentiate osimertinib-resistance from osimertinib-sensitive NSCLC patients. The target genes of these dysregulated exosomal miRNAs are associated with several signaling pathways reported to be involved with various oncogenic processes. To our knowledge, this is the first report showing four potential biomarkers of osimertinib-resistance in NSCLC patients.
Tissue biopsy is still standard procedure necessary for cancer diagnostic and prognostic. However, since tumors are highly heterogeneous, a tissue biopsy is inadequate to comprehensively characterize a tumor and its invasiveness. To overcome these limitations, analysis of circulating cancer-derived exosomes has been proposed as a diagnostic and prognostic tool [24]. Cancer exosomes can regulate the biological functions of distant recipient cells through the transportation of specific cargos [12, 13, 14]. Here, we purified extracellular vesicles (EVs) from the plasma of NSCLC patients, before and after osimertinib-resistance, and healthy controls. The mean diameter of the isolated EVs from each group ranged from 100 to 150 nm, which is similar to previously reported exosome sizes in the plasma of NSCLC patients [32]. Furthermore, these EVs were positive for well-known exosomal markers, Alix, Flotillin-1, and CD81 [33], suggesting that the isolated EVs were indeed exosomes. Exosomal content is unique and often reflects the disease state. In addition, study on the circulating exosomes could overcome the treatment limitation from tumor heterogeneity [34]. Previously, several studies have isolated potential molecular biomarkers from circulating exosomes in lung cancer [25, 26, 28]. Furthermore, the exosomal phospholipid membrane can protect encapsulated RNA, including miRNAs, from degradation by ribonuclease [27]. Thus, exosomal miRNAs represent potential biomarkers for studying cancer progression and drug resistance.
In the present study, we identify 10 exosomal miRNAs that were statistically dysregulated in osimertinib-resistant NSCLC patients. The target genes of these dysregulated exosomal miRNAs are associated with several oncogenic signaling pathways, such as the Hedgehog, Hippo, PI3K/AKT, and MAPK signaling pathways. Previous studies have correlated hyperactivation of Hedgehog signaling with EGFR-TKI resistance in NSCLC cell lines [35, 36]. In addition, MET amplification can activate the PI3K/AKT signaling pathway, thereby providing a bypass mechanism of EGFR-TKI [37, 38]. Similarly, exosomes shed by gefitinib-resistant NSCLC cells can transfer drug resistance to sensitive target cells through activating the PI3K/AKT signaling pathway [39]. Furthermore, the present study found that miR-323a-3p, miR-325, miR-1468-3p, miR-4433b-3p, miR-5189-5p, and miR-6513-5p were significantly dysregulated in osimertinib-resistant NSCLC patients. Among them, 4 of these miRNAs have been previously associated with lung cancer. Upregulation of miR-323a-3p has been seen in lung tissue of stage I NSCLC patients, while the downregulation of miR-325-3p was associated with poor prognosis for NSCLC patients [40, 41]. Moreover, in NSCLC patients, plasma exosomal miR-4433b-3p was dramatically upregulated in progressive disease after anti-PD-1 treatment compared to pre-treatment [42]. Likewise, miR-1468 expression was positively correlated with the recurrence-free survival of lung adenocarcinoma patients and, thus, has been suggested as a potential prognosis biomarker [43]. Currently, there are no reports on the roles of miR-5189-5p and miR-6513-5p in cancer progression. However, using the KEGG pathway analysis, our study found that the predicted target genes of miR-5189-5p were mostly associated with the hippo signaling pathway. Moreover, overactivation of YAP, a downstream molecule of the hippo signaling pathway, plays a crucial role in EGFR-TKI resistance of lung adenocarcinoma cells [44]. Similarly, we found that the predicted target genes of miR-6513-5p were associated with the EGFR-TKI resistance pathway. These results indicate that these two new miRNAs, miR-5189-5p and miR-6513-5p, may regulate EGFR-TKI resistance. However, further studies are required to fully evaluate the roles of these miRNAs on osimertinib sensitivity.
Interestingly, four exosomal miRNAs (miR-323-3p, miR-1468-3p, miR-5189-3p and miR-6513-5p) showed potential for detecting osimertinib resistance. However, the upregulations of these 4 exosomal miRNAs were not statistically significant correlate with the TTF and OS of osimertinib-treated NSCLC patients. This may be due to small sample size. Therefore, further validation of these exosomal miRNAs in larger cohort is needed to verify their usefulness as biomarkers. However, to further evaluate the potential of the 10 lung cancer-dysregulated exosomal miRNAs as biomarkers, we attempted to correlate miRNA upregulation with patient survivals. We also demonstrated that the miRNAs tended to correlate with longer TTF and OS of osimertinib-treated patients. The reason may be because of 3 (miR-325, miR-375, and miR-520-3p) of the 10 exosomal candidate miRNAs were identified as tumor suppressor genes. For example, miR-325 has been shown to inhibit hepatocellular carcinoma progression [45]. Whereas, miR-520-3p has been correlated with inhibition of colorectal cancer cell migration and NSCLC proliferation [46, 47]. Furthermore, low expression of miR-375 was observed in osteosarcoma patients and associated with brain metastasis in NSCLC patients [48, 49]. This expulsion of tumor suppressor genes in exosomes is commonly seen in cancer cells. For instance, ovarian cancer cells eliminate miR-6126, which inhibits integrin-
In summary, the present study was a pilot study that examined plasma exosomal miRNA profiles in NSCLC patients before and after resistance to osimertinib treatment develops. We showed the tendency of upregulation of 10 lung cancer-dysregulated exosomal miRNAs to correlate with longer TTF and OS of osimertinib-treated patients. Among these 10 miRNAs, 6 were significantly upregulated in osimertinib-resistant compared with osimertinib-sensitive NSCLC patients. Subsequently, by ROC analysis, we identified four miRNAs (miR-323-3p, miR-1468-3p, miR-5189-3p, and miR-6513-5p) that have the potential to discriminate between osimertinib-resistant and osimertinib-sensitive NSCLC patients with high accuracy. Our results highlight the potential role of these exosomal miRNAs as molecular biomarkers for the detection of osimertinib resistance. Therefore, we propose that tissue biopsies can be combined with an analysis of plasma exosomal miRNA expression levels to improve diagnostic and prognostic reliability. However, further validation with larger cohorts is needed to further characterize these plasma exosomal miRNAs as predictive biomarkers of osimertinib resistance. Moreover, the mechanistic roles of these exosomal miRNAs during drug resistance should be further investigated.
Author contributions
Conception: Keatdamrong Janpipatkul, Thanyanan Reungwetwattana, Arthit Chairoungdua
Interpretation or analysis of data: Keatdamrong Janpipatkul, Narumol Trachu, Piyakarn Watcharenwong, Wittaya Panvongsa, Wittawin Worakitchanon, Chanatip Metheetrairut, Songporn Oranratnachai
Preparation of the manuscript: Keatdamrong Janpipatkul, Arthit Chairoungdua
Revision for important intellectual content: Keatdamrong Janpipatkul, Arthit Chairoungdua
Supervision: Thanyanan Reungwetwattana, Arthit Chairoungdua
Supplementary data
The supplementary files are available to download from http://dx.doi.org/10.3233/CBM-203075.
sj-docx-1-cbm-10.3233_CBM-203075.docx - Supplemental material
Supplemental material, sj-docx-1-cbm-10.3233_CBM-203075.docx
sj-pdf-1-cbm-10.3233_CBM-203075.pdf - Supplemental material
Supplemental material, sj-pdf-1-cbm-10.3233_CBM-203075.pdf
sj-xlsx-1-cbm-10.3233_CBM-203075.xlsx - Supplemental material
Supplemental material, sj-xlsx-1-cbm-10.3233_CBM-203075.xlsx
sj-xlsx-2-cbm-10.3233_CBM-203075.xlsx - Supplemental material
Supplemental material, sj-xlsx-2-cbm-10.3233_CBM-203075.xlsx
sj-xlsx-3-cbm-10.3233_CBM-203075.xlsx - Supplemental material
Supplemental material, sj-xlsx-3-cbm-10.3233_CBM-203075.xlsx
sj-xlsx-4-cbm-10.3233_CBM-203075.xlsx - Supplemental material
Supplemental material, sj-xlsx-4-cbm-10.3233_CBM-203075.xlsx
Footnotes
Acknowledgments
This project was supported by Mahidol University and Thailand Research Fund (RSA6280043 to AC) and Thailand Research Fund through the International Research Network (IRN58W0004 to AC and KJ). This research project was partially supported by the CIF Grant, Faculty of Science, Mahidol University. We are grateful to Prof. Dr. Wasun Chantratita and all the staff from the Center for Medical Genomics, Faculty of Medicine Ramathibodi Hospital for their helpfulness for next generation sequencing analysis.
