Abstract
INTRODUCTION:
Osteosarcoma (OS), aggressive neoplasms of the bone, is the most common primary bone cancer in children. MiR-196a usually low expressed in several tumors and its functions in osteosarcoma still unclear.
MATERIALS AND METHODS:
Quantitative real-time polymerase chain reaction (qRT-PCR) was employed to assess the expression of miR-196a and the HOXA5. Cell metastasis and epithelial-mesenchymal transition (EMT) abilities were assessed using Transwell and western blot. The dual luciferase reporter assay was carried out to verify whether miR-196a directly targeted the 3’-untranslated region (UTR) of HOXA5 mRNA.
RESULTS:
MiR-196a was overexpressed and HOXA5 was low expressed in osteosarcoma versus the non-tumor tissues and normal cell lines. Upregulation of miR-196a or downregulation of HOXA5 was associated with worse outcome of osteosarcoma patients. MiR-196a enhanced cell migration, invasion and EMT by regulating the expression of HOXA5 through directly targeting the 3’-UTR of its mRNA in osteosarcoma. HOXA5 partially reversed roles of miR-196a on metastasis and EMT in osteosarcoma.
CONCLUSIONS:
MiR-196a promoted cell metastasis and EMT by targeting the 3’-UTR of HOXA5 mRNA in osteosarcoma. The newly identified miR-196a/HOXA5 axis provides novel insight into the pathogenesis of osteosarcoma.
Introduction
Osteosarcoma originates from the long bones’ metaphysis of young adults and adolescents, and is a leading cause of cancer death in adolescents [1, 2]. Although the current treatment of osteosarcoma has improved, the survival rate has not improved in the past three decades [3, 4]. Thus, it is important to find potential osteosarcoma therapeutic targets.
MicroRNAs (miRNAs), a quantity of short non-coding RNAs with a length of 19–28 nucleotides, could inhibit the function of target genes through degradation of the mRNA or suppressing its translation in post-transcriptional regulation [5, 6]. MiR-196a can be used as a prognostic biomarker for esophageal squamous cell carcinoma and ER
The homeobox A5 (HOXA5) is a part of A cluster on chromosome 7 that encodes a DNA-binding transcription factor [13]. HOXA5 played a role in inhibiting cell proliferation in gastric cancer [14]. HOXA5 impaired pathological angiogenesis and was associated with tumor progression [15]. HOXA5 depletion suppressed osteogenic differentiation and proliferation of stem cells from the apical papilla [16]. HOXA5 inhibited tumor initiation and progression of breast cancer through E-cadherin and CD24 [17]. Moreover, HOXA5 enhanced cell apoptosis of osteosarcoma via p53 and p38
Materials and Methods
Patients and tumor samples
A total of 47 osteosarcoma patients who were hospitalized in Jinan City People’s Hospital from 2016 to 2018 were collected, and pairs of osteosarcoma and corresponding paracancerous tissues were obtained through surgical operation. No patients had accepted chemotherapy or radiotherapy before surgery. The fresh tissues were immediately frozen in liquid nitrogen after surgery, and stored at
Cell culture and treatment
An osteoblast cell line hFOB 1.19 and human osteosarcoma cell lines U2OS, Saos-2 and MG63 were obtained from American Type Culture Collection (ATCC; Rockville, MD, USA). All the cells were cultured in Dulbecco’s modified eagle medium (DMEM) (Gibco, Rockville, MD, USA) containing 10% Fetal Bovine Serum (FBS) (Gibco, Rockville, MD, USA) and incubated at 37
Cell transfection
MiR-196a mimic or miR-196a inhibitor (Gene Pharma, Shanghai, China) was used to increase or down-regulate the intracellular levels of miR-196a. U2OS cells with a density of 70% were seeded into 6-well plates. The transfection was performed using Lipofectamine 2000 (Invitrogen, Carlsbad, CA, USA), which was diluted in Opti-MEM medium. Next, the mixture was added to the cells and the cells were incubated at 37
RNA Extraction and quantitative real-time polymerase chain reaction (qRT-PCR)
Total RNA or miRNA were extracted using the TRIzol reagent (Invitrogen, Carlsbad, CA, USA) or the miRCURY RNA Isolation Kit (Exiqon, USA), and then quantified by using the NanoDrop spectrophotometer (NanoDrop, Thermo Fisher Scientifc, Waltham, MA, USA). The first cDNA chain was synthesized using a High-Capacity cDNA Reverse Transcription Kit (Applied Biosystems, Foster City, CA, USA). The SYBR PrimeScript miRNA RT-PCR kit (Takara Bio, Tokyo, Japan) or the SYBR PrimeScript miRNA RT-PCR kit (Takara Bio, Inc., Otsu, Japan) was employed to calculate the expression of HOXA5 or miR-196a. The glyceraldehyde 3-phosphate dehydrogenase (GAPDH) and U6 are used as the internal reference. The primers were: miR-196a F: 5’-CCGGGACTGTTGAGTGAAGT-3’, R: 5’-AACTGCCATGAATCAGGTGG-3’; U6 F: 5’-CTCGCTTCGGCAGCACA-3’, R: 5’-AACGCTTCA CGAATTTGCGT-3’; HOXA5: F: 5’-ATGCGCAAGCT GCACATAAG-3’; R: 5’-CGGGTCAGGTAACGGTTG AA-3’; GAPDH F: 5’-TCAACGACCACTTTGTCAAG CTCA-3’, R: 5’-GCTGGTGGTCCAGGGGTCTTACT-3’.
Upregulation of miR-196a predicted poor prognosis of osteosarcoma. (A) MiR-196a was upregulated in osteosarcoma tissues versus the corresponding paracancerous tissues. (B) Overexpression of miR-196a was associated with poor overall survival of osteosarcoma patients. (C) The expression of miR-196a was upregulated in U2OS, Saos-2 and MG63 cells versus hFOB 1.19 cells. (D) The transfection efficiency of transfecting the miR-196a mimic and the miR-196a inhibitor in U2OS cells.
MiR-196a enhanced the metastasis and the EMT of U2OS cells. (A) and (B) The miR-196a mimic caused an increase in the migratory and invasive behavior of the cells, whereas inhibition of the miR-196a reduced cell migration and invasion (200 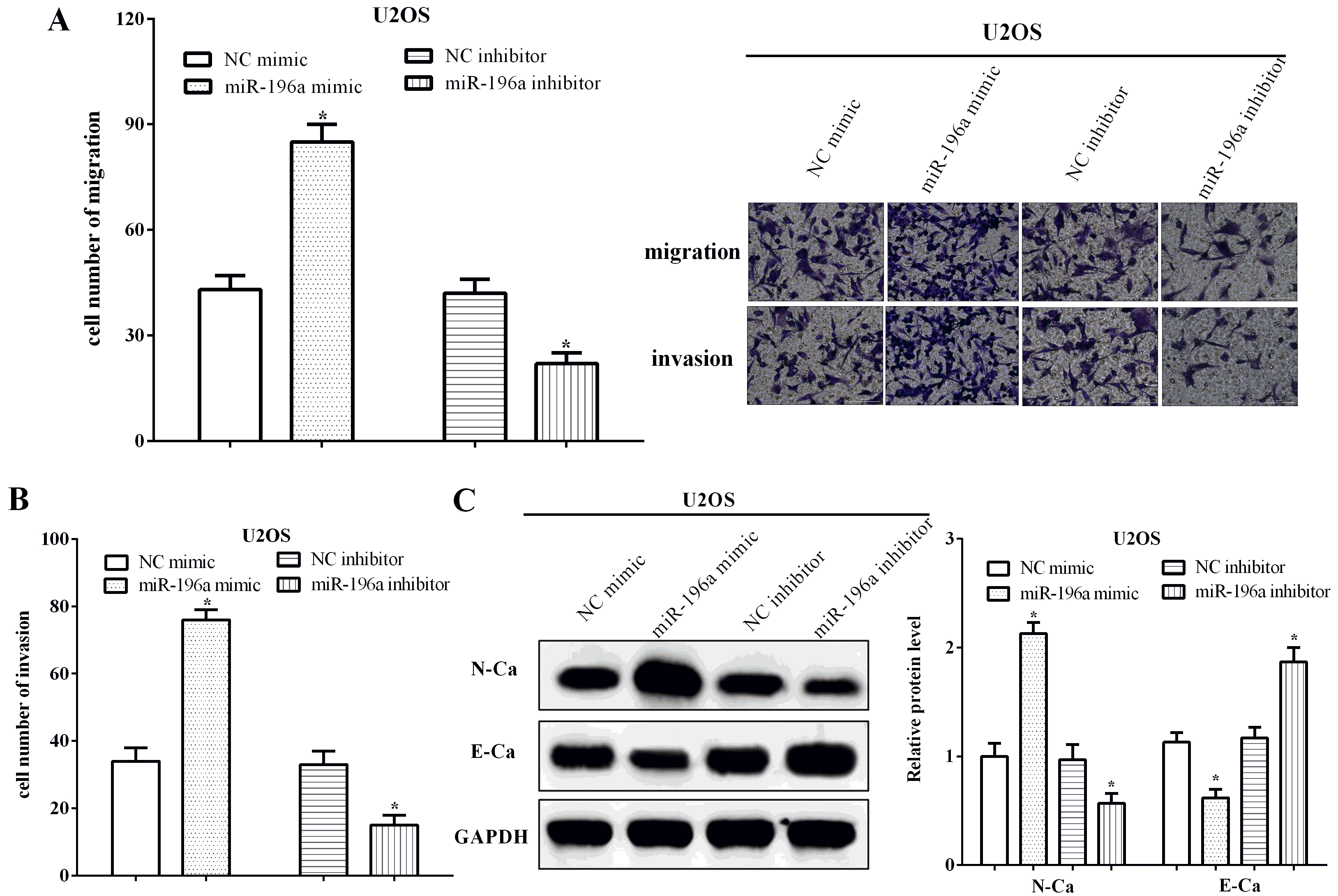
Transwell assays without or with Matrigel were utilized to investigate the migration and invasion abilities of osteosarcoma cells. Before the experiment, transwell chambers were placed in a 24-well plate, and formed upper and lower chamber. 200
Western Blotting
Radioimmunoprecipitation assay (RIPA) buffer containing protease inhibitors (SigmaAldrich, St. Louis, MO, USA) was applied to lyse the cells and extracted proteins. After centrifugation at 12,000 rpm for 15 min, the concentration of total protein was assessed by using the bicinchoninic acid (BCA) Protein Assay Kit (Thermo Fisher Scientific, Waltham, MA, USA). The proteins were separated using 10% sodium dodecyl sulphate-polyacrylamide gel electrophoresis (SDS-PAGE), and followed transferred onto polyvinylidene fluoride (PVDF) membranes (Millipore, Billerica, MA, USA).
After blocking by incubating 5% skimmed milk at room temperature for 5 h, the membranes were subsequently incubated with the primary antibodies. The primary antibodies were against HOXA5 or GAPDH (Cell Signaling Technology, Danvers, MA, USA). Followed, the membranes were incubated with the HRP-conjugated secondary antibody (Cell Signaling Technology, Danvers, MA, USA) (1:5000). Western electrochemiluminescence (ECL) Substrate (Bio-Rad, Hercules, CA, USA) was applied to carry out the visualization.
Luciferase reporter assay
To evaluate the interaction of miR-196a and its response elements in the 3’-UTR of the HOXA5 mRNA, miR-196a mimic or inhibitor together with the psiCHECK2 vector including the sequence of the 3’-UTR HOXA5 mRNA were co-transfected into U2OS cells. The psiCHECK2 vector alone or the vector containing the mutant miR-196a response element (MRE) on the 3’-UTR of HOXA5 (MRE-tandem-mut) were used as the negative controls. The relative luciferase activity was determined using the dual luciferase assay kit (Promega, Madison, WI, USA). The firefly luciferase activity was used for normalization.
Statistical analysis
Statistical analysis was performed using GraphPad Prism 7 Software (La Jolla, CA, USA). Data are presented as mean
MiR-196a regulates the expression of HOXA5 by binding to the 3’-UTR of its mRNA. (A) HOXA5 was predicted as a target of miR-196a by TargetScan. (B) Luciferase activity was reduced in cells containing a wild type HOXA5 3’-UTR psiCHECK2 plasmid that were transfected with a miR-196a mimic. A transfected mutated miR-196a had no effect. (C) HOXA5 transcripts were decreased in U2OS cells transfected with a miR-196a mimic, but increased in cells treated with a miR-196a inhibitor. WT, wild type; MUT, mutant.
Downregulation of HOXA5 predicted poor prognosis of osteosarcoma patients. (A) The expression of HOXA5 was lower in osteosarcoma tissue than the normal tissues. (B) HOXA5 was low expressed in U2OS cells, Saos-2 and MG63 cells than hFOB 1.19 cells. (C) Downregulation of HOXA5 was associated with poor overall survival of osteosarcoma patients.
Knockdown of HOXA5 in U2OS cells lacking miR-196a enhances the EMT phenotype. (A) Transfection However, our results were consistent with the findings of cervical cancer, and we discovered that HOXA5of the silencing vector sh-HOXA5 in to U2OS cells with reduced miR-196a levels causes a decrease in HOXA5 mRNA. (B) Knockdown of both HOXA5 and miR-196 leads to increased cell migration and invasion (200 
Upregulation of miR-196a upregulation predicted poor prognosis of osteosarcoma
The expression of miR-196a was calculated using qRT-PCR in 47 pairs of osteosarcoma tissue samples and corresponding paracancerous tissue samples. As expected, miR-196a was upregulated in osteosarcoma tissues in comparison with the corresponding paracancerous tissues (
Moreover, the expressions of miR-196a were calculated in osteoblast hFOB 1.19 cells and human osteosarcoma cell lines U2OS, Saos-2 and MG63. As expected, the expression of miR-196a was upregulated in U2OS (
MiR-196a enhanced cell metastasis and EMT of U2OS cells
The effect of the miR-196 on osteosarcoma U2OS migration and invasion was determined in cells transfected with a miR-196 mimic and inhibitor and nonspecific controls using a Transwell assay. Expression of the transfected mimic and inhibitor was confirmed by measuring levels of miR-196a (Fig. 1D). Results show that both migration and invasion increased in U2OS cells transfected with miR-196a (
MiR-196a regulated the expression of HOXA5 by directly binding to the 3’-UTR of mRNA
TargetScan was conducted to predict the potential target genes of miR-196a, and HOXA5 was discovered as a target of miR-196a. In order to verify whether miR-196a directly targeted the 3’-UTR of HOXA5 mRNA, the conjectural binding sequences were mutated from ACUACCU to UGAUGGA, and followed the luciferase reporter assay were performed (Fig. 3A). Compared with the NC mimic, the miR-196a mimic inhibited the luciferase activity of cells transfected wild type 3’-UTR of HOXA5 mRNA (
Downregulation of HOXA5 predicted poor prognosis of osteosarcoma patients
qRT-PCR assay indicated that HOXA5 was low expressed in osteosarcoma tissues compared with the normal tissues (
HOXA5 partially restored functions of miR-196a
In order to investigate the functions of HOXA5 in miR-196a inhibitor-transfected cells, sh-HOXA5 plasmid was re-transfected into miR-196a silenced U2OS cells, and the transfection efficiency was measured by qRT-PCR (
Discussion
MiR-196a was overexpressed in thyroid carcinoma and acts as an independent unfavorable prognostic factor [20]. Similarly, miR-196a functioned as a potential prognostic marker to delay lymph node metastasis in early-stage tongue squamous cell carcinoma [21]. Consistent with all the findings, we discovered that miR-196a was upregulated in osteosarcoma tissues and cell lines, and upregulation of miR-196a was correlated with poor outcome of osteosarcoma patients. MiR-196a enhanced cell proliferation and metastasis in laryngeal cancer and in liver cancer [22, 23]. Similarly, miR-196a promoted cell growth and migration via I
HOXA5 can serve as a prognostic factor for non-small cell lung cancer and suppress cell proliferation and invasion [27, 28]. Consistent with all the findings, we discovered that HOXA5 was downregulated in osteosarcoma tissues and cell lines compared with paracancerous tissues and normal cell lines. Downregulation of HOXA5 predicted a worse prognosis for osteosarcoma patients. HOXA5 inhibited cervical cancer cell growth and induced cell apoptosis [29]. In esophageal squamous cell cancer, in contrary, inhibition of HOXA5 inhibited the proliferation, migration and invasion [30]. However, our results are consistent with the findings in cervical cancer, in that, increased HOXA5 made the cells more normal by reducing particular cellular traits associated with transformed cells.
Conclusions
MiR-196a enhanced cell migration, invasion and the EMT by reducing the expression of HOXA5 through targeting the 3’-UTR of its mRNA in osteosarcoma. Our findings provided insight into the biology of osteosarcoma processes and suggested that miR-196a can be used as a new molecular target for the treatment of osteosarcoma.
Footnotes
Conflict of interest
The authors declared no conflict of interest.
