Abstract
BACKGROUND:
Gastric cancer is the third leading cause of cancer-related deaths worldwide.
OBJECTIVE:
The present study aims to identify key long non-coding RNAs (lncRNAs) and their potential roles in the pathogenesis of gastric adenocarcinoma.
METHODS:
The lncRNA and mRNA expression profile between gastric adenocarcinoma and adjacent non-tumor tissues were obtained from The Cancer Genome Atlas (TCGA). Differentially expressed lncRNAs (DElncRNAs) and mRNAs (DEmRNAs) between gastric adenocarcinoma and adjacent non-tumor tissues were identified after bioinformatics analysis. DElncRNA-DEmRNA co-expression network and DElncRNA-nearby DEmRNA interaction network were constructed, respectively. Functional annotation for DEmRNAs interacted with DElncRNAs was performed. Receiver operating characteristic (ROC) analysis of selected DElncRNAs was conducted.
RESULTS:
Based on TCGA, the mRNA and lncRNA expression profiles of 375 gastric adenocarcinoma and 32 adjacent non-tumor tissues were downloaded. A total of 1502 DEmRNAs and 928 DElncRNAs between gastric adenocarcinoma and adjacent non-tumor tissues were identified. HOXC-AS3 might involve with gastric adenocarcinoma by regulating a set of HOX genes (HOXC8, HOXC9, HOXC10, HOXC11, HOXC12 and HOXC13) with cis-effect. AC115619.1-APOA4/APOB and AP006216.2-APOA1/APOA4 integrations might play roles in gastric adenocarcinoma through regulating pathways of Fat digestion and absorption and Vitamin digestion and absorption. Six lncRNAs including (HOTAIR, C20orf166-AS1, PGM5-AS1, HOXC-AS3, HOXC-AS2 and AC012531.1) have excellent diagnostic value for gastric adenocarcinoma.
CONCLUSIONS:
This study identifies key lncRNAs in gastric adenocarcinoma which provides clues for exploring the pathogenesis and developing potential biomarkers for gastric adenocarcinoma.
Keywords
Introduction
Gastric cancer is one of the most common types of cancers that is the third leading cause of cancer-related death worldwide [1]. The incidence rate of gastric cancer is particularly high in East Asia and South America [2]. Gastric adenocarcinoma (GAC) is the most common histological type of gastric cancer. Due to delayed diagnosis, the five- year survival rate of patients with gastric cancer is low [1]. Although the strongest risk factor for gastric cancer,
Patient characteristic
Patient characteristic
Top 20 up- and down-regulated DElncRNAs between gastric adenocarcinoma and adjacent non-tumor tissues
DElncRNA, differentially expressed long non-coding RNA between gastric adenocarcinoma and adjacent non-tumor tissues. log2 fold-change, log2-transformed fold-change of DElncRNA expression in gastric adenocarcinoma tissues relative to adjacent non-tumor tissues. FDR, false discovery rate. Regulation, the trend of DElncRNA expression in gastric adenocarcinoma tissues compared with adjacent non-tumor tissues.
Long non-coding RNA (lncRNA) is a new class of non-coding RNA with transcripts more than 200 bp in length, which has been receiving increased attention [3]. Through cis- or transacting mechanisms, lncRNAs could regulate gene expression at transcriptional, epigenetic, and translative levels [4, 5]. Accumulated evidences indicated that abnormal expressed lncRNAs play roles in the occurrence and progression of tumors [6, 7]. Moreover, several gastric cancer-related lncRNAs including MEG3, DANCR, SNHG7, HOXA11-AS, HOTAIR and HOXC-AS3 have been identified [8, 9, 10, 11, 12, 13], which highlighted the importance of lncRNAs in gastric cancer.
In the present study, we identify differentially expressed lncRNAs (DElncRNAs) and mRNA (DEmRNAs) between GAC and adjacent non-tumor tissues based on The Cancer Genome Atlas (TCGA) and bioinformatic analysis. Furthermore, functional annotation of DEmRNAs interacted with DElncRNAs contributes to exploring function of DElncRNAs in GAC. In addition, several lncRNAs indicate excellent diagnostic value for GAC. Our study provides clues for better understanding of pathogenesis and developing novel biomarkers for GAC.
mRNA and lncRNA expression profiles of GAC in TCGA
The Cancer Genome Atlas (TCGA) is a central bank for multi-dimensional data of various cancers at DNA, RNA and protein levels. In this study, the clinical data of patients with GAC were downloaded from TCGA data portal (
Identification of DElncRNAs and DEmRNAs between GAC and adjacent non-tumor tissues
DElncRNAs and DEmRNAs between GAC and adjacent non-tumor tissues were calculated via R package DESeq2 (
DElncRNA-DEmRNA co-expression analysis
The pairwise Pearson correlation coefficients between DElncRNAs and DEmRNAs were calculated. DElncRNA-DEmRNA pairs with
DElncRNA-nearby DEmRNA interaction analysis
LncRNAs were reported to regulate genes that were transcribed near them, consistent with activity in cis [14]. To identify the nearby DEmRNAs of DElncRNAs with cis-regulatory effects, DEmRNAs transcribed within a 10kb/100kb window up-or down-stream of DElncRNAs between patients with GAC and adjacent non-tumor tissues were searched. The DElncRNA-nearby DEmRNAs interaction network was constructed by using Cytoscape software (
Functional annotation
Functional annotation including Gene ontology(GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) molecular pathway enrichment analysis was performed for DEmRNAs co-expressed with DElncRNAs and nearby DEmRNAs of DElncRNAs using DAVID 6.8 (
Hierarchical clustering analysis of DElncRNAs and DEmRNAs between gastric adenocarcinoma and adjacent non-tumor tissues. A). DElncRNAs; B). DEmRNAs GAC, gastric adenocarcinoma. Row and column represented DElncRNAs/DEmRNAs and tissue samples, respectively. The color scale indicated the expression of DElncRNAs and DEmRNAs. Red and green color indicated up- and down-regulation, respectively.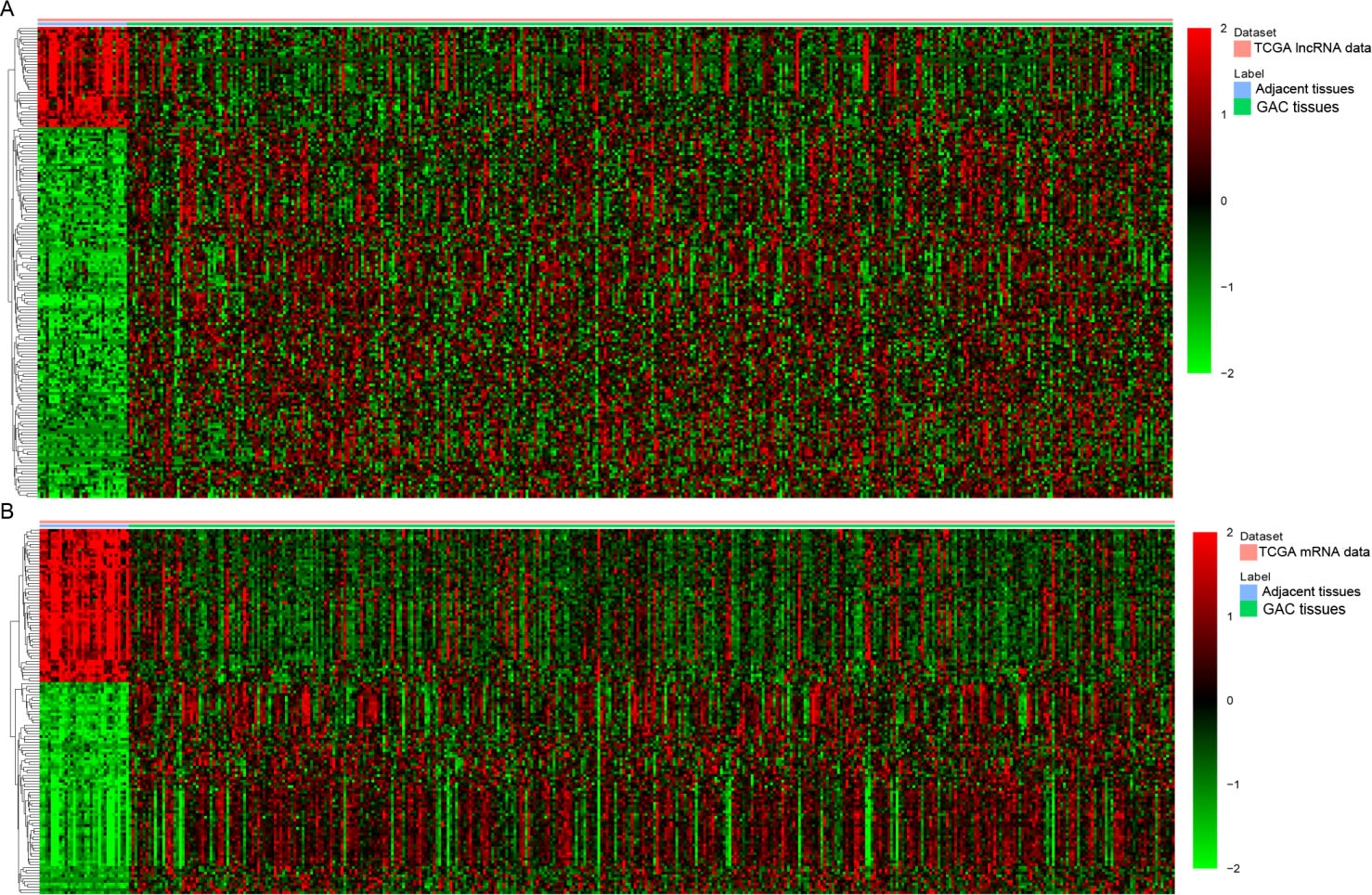
DElncRNA-DEmRNA co-expression network. Red and green ellipses represent up- and down-regulated DEmRNAs between gastric adenocarcinoma and adjacent non-tumor tissues, respectively. Orange and blue rectangles represent up- and down-regulated DElncRNAs between gastric adenocarcinoma and adjacent non-tumor tissues, respectively. Edges indicate DElncRNA-DEmRNA co-expression. DEmRNAs, differentially expressed mRNAs. DElncRNAs, differentially expressed lncRNAs.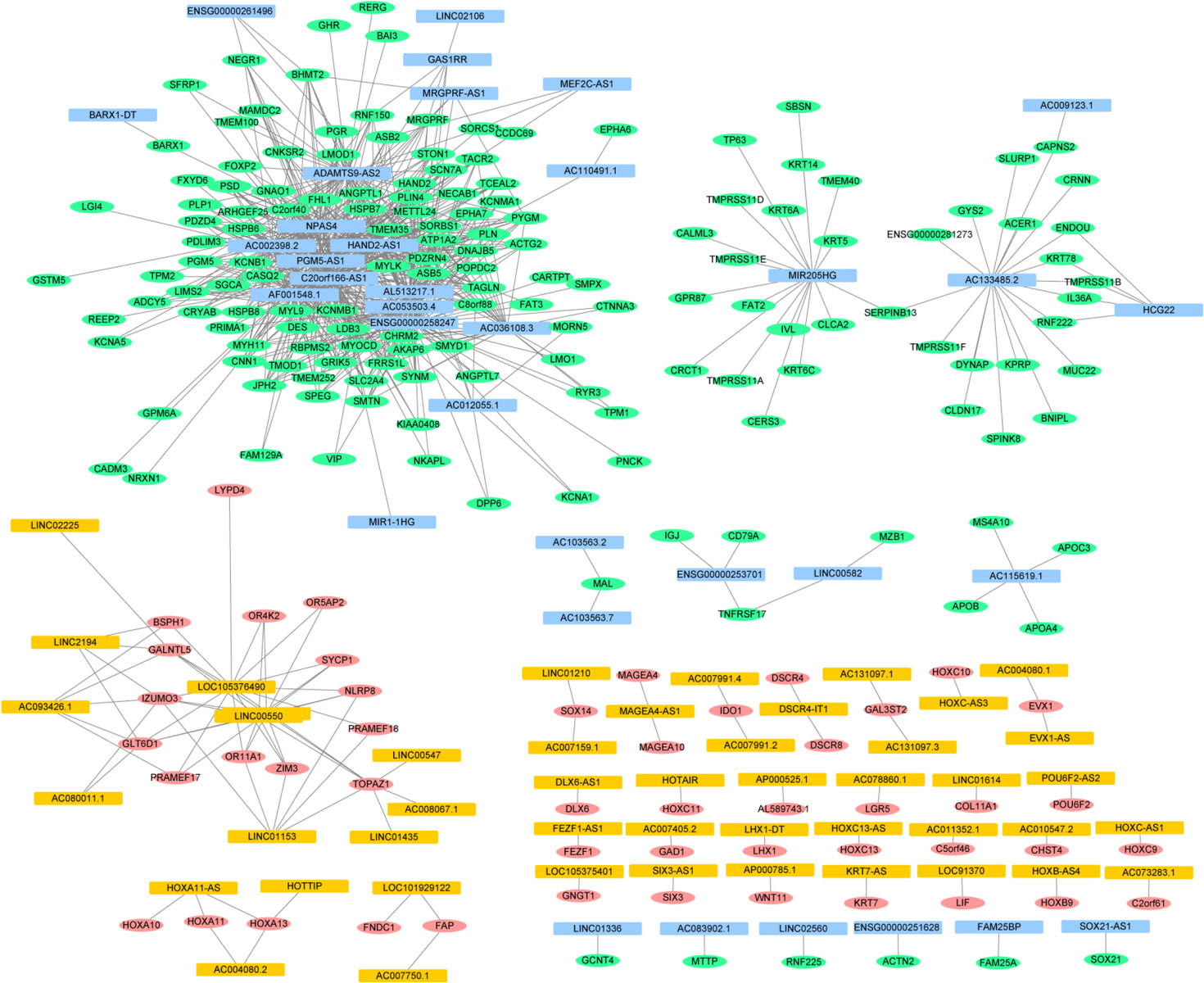
Pathway of fat digestion and absorption. The red rectangles represented the particles which regulated by DEmRNAs between gastric adenocarcinoma and adjacent non-tumor tissues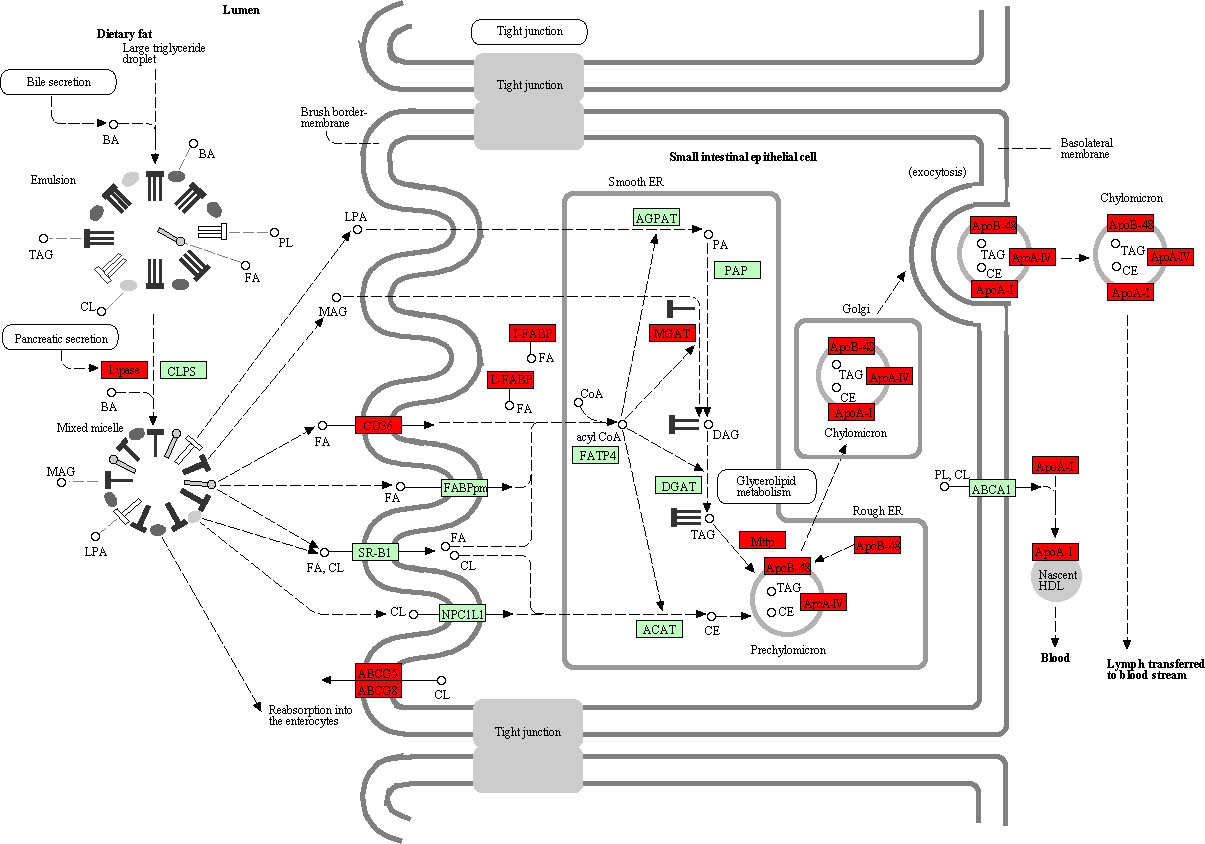
Pathway of vitamin digestion and absorption. The red rectangles represented the particles which regulated by DEmRNAs between gastric adenocarcinoma and adjacent tissues.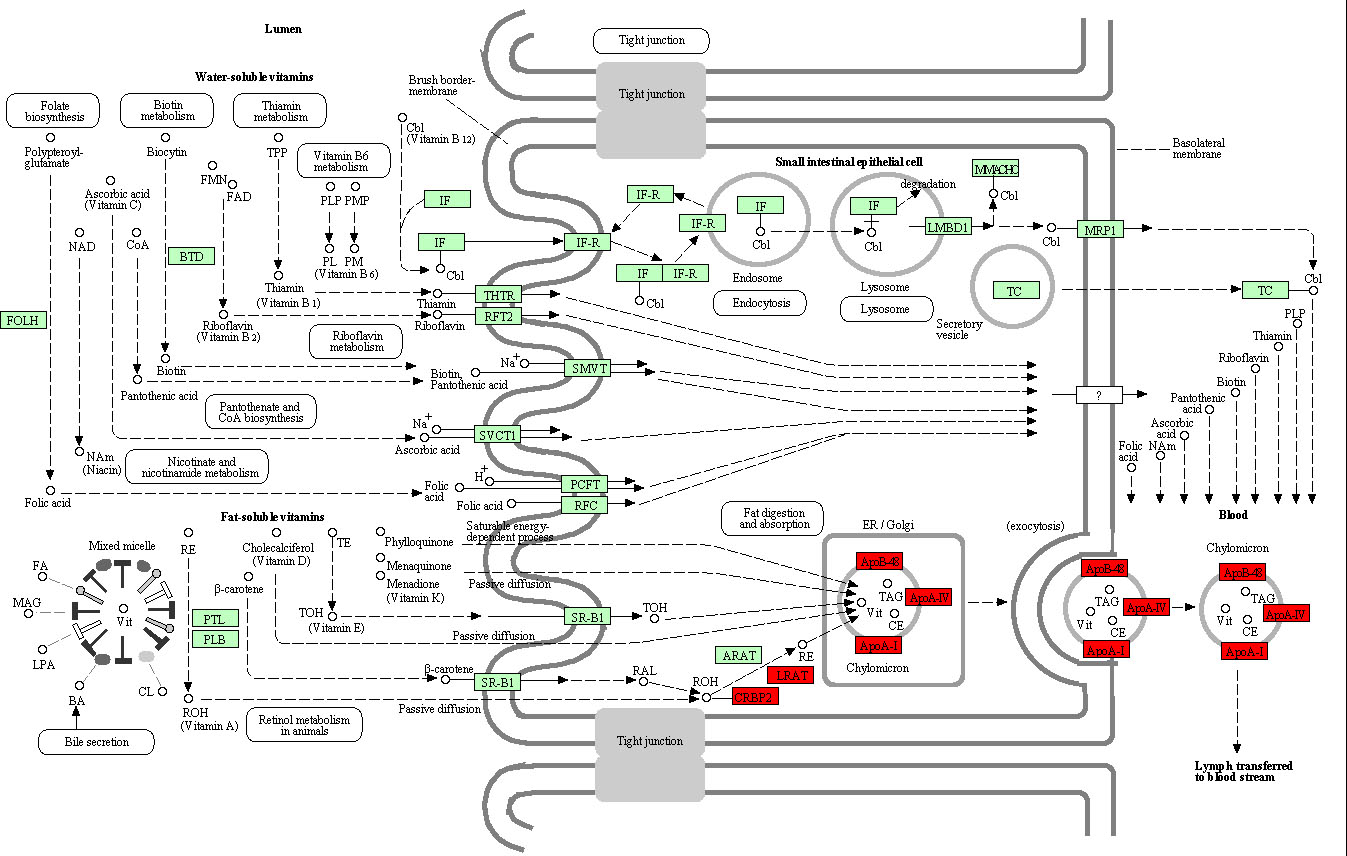
In order to access the diagnostic value of DElncRNAs for GAC, the “pROC” package (
Results
DEmRNAs and DElncRNAs between GAC and adjacent non-tumor tissues
The clinical data of 375 patients with GAC and the mRNA and lncRNA expression profiles of 375 GAC tumor tissues and 32 adjacent non-tumor tissues were downloaded from TCGA. Detailed information of these 375 patients with GAC was displayed in Table 1. A total of 928 DElncRNAs (684 up-regulated and 244 down-regulated lncRNAs) and 1502 DEmRNAs (685 up-regulated and 817 down-regulated mRNAs) between GAC and adjacent non-tumor tissues were identified with FDR
Top 20 up- and down-regulated DEmRNAs between gastric adenocarcinoma and adjacent non-tumor tissues
Top 20 up- and down-regulated DEmRNAs between gastric adenocarcinoma and adjacent non-tumor tissues
DEmRNAs, differentially expressed mRNAs between gastric adenocarcinoma and adjacent non-tumor tissues. log2 fold-change, log2-transformed fold-change of gene mRNA expression in gastric adenocarcinoma tissues relative to adjacent non-tumor tissues. FDR, false discovery rate. Regulation, the trend of gene mRNA expression in gastric adenocarcinoma tissues compared with adjacent non-tumor tissues.
Nearby DEmRNAs that co-expressed with DElncRNAs
DElncRNA, differentially expressed long non-coding RNAs between gastric adenocarcinoma and adjacent non-tumor tissues. DEmRNAs, differentially expressed mRNAs between gastric adenocarcinoma and adjacent non-tumor tissues. Chr, chromosome location of DElncRNA. Start, start site of DElncRNA location. End, end site of DElncRNA location.
A total of 795 DElncRNA-DEmRNA co-expression pairs including 82 DElncRNAs (47 up-regulated and 35 down-regulated DElncRNAs) and 206 DEmRNAs (48 up-regulated and 158 down-regulated DEmRNAs) were obtained with
DElncRNA-nearby DEmRNA interaction
A total of 336 DElncRNA-nearby DEmRNA pairs (Supplemental Fig. S2) were obtained, which were consisted of 221 DElncRNAs (135 up-regulated and 86 down-regulated DElncRNAs) and 246 DEmRNAs (119 up-regulated and 126 down-regulated DElncRNAs). Four up-regulated lncRNAs including HOXC13-AS (degree
The intersection of DElncRNA-nearby DEmRNA pairs and DElncRNA-DEmRNA co-expression pairs consisted of 46 DElncRNA-DEmRNA pairs, in which DEmRNAs were not only co-expressed with DElncRNAs but also the nearby DEmRNAs of DElncRNAs (Table 4).
Top 30 KEGG pathways for DEmRNAs co-expressed with DElncRNAs and nearby DEmRNAs of DElncRNAs
Top 30 KEGG pathways for DEmRNAs co-expressed with DElncRNAs and nearby DEmRNAs of DElncRNAs
DEmRNAs, differentially expressed mRNAs between gastric adenocarcinoma and adjacent non-tumor tissues. DElncRNAs, differentially expressed lncRNAs between gastric adenocarcinoma and adjacent non-tumor tissues. KEGG, Kyoto Encyclopedia of Genes and Genomes. Count, the number of DEmRNAs that enriched in this KEGG pathway. Genes, gene symbol of DEmRNAs that enriched in this KEGG pathway.
The top 15 most significantly enriched GO terms, including “biological process”, “molecular function”, and “cellular component” for DEmRNAs co-expressed with DElncRNAs, as well as nearby DEmRNAs of DElncRNAs, were displayed in Supplemental Table S1 and Supplemental Table S2, respectively. Based on the functional annotation of DEmRNAs co-expressed with DElncRNAs, Digestion (GO: 0007586,
According to the KEGG enrichment analysis (Table 5), Metabolism of xenobiotics by cytochrome P450 (hsa00980,
ROC curves of selected DElncRNAs between gastric adenocarcinoma and adjacent non-tumor tissues. The ROC curves were used to show the diagnostic ability of these selected DElncRNAs in gastric adenocarcinoma with sensitivity and specificity. The 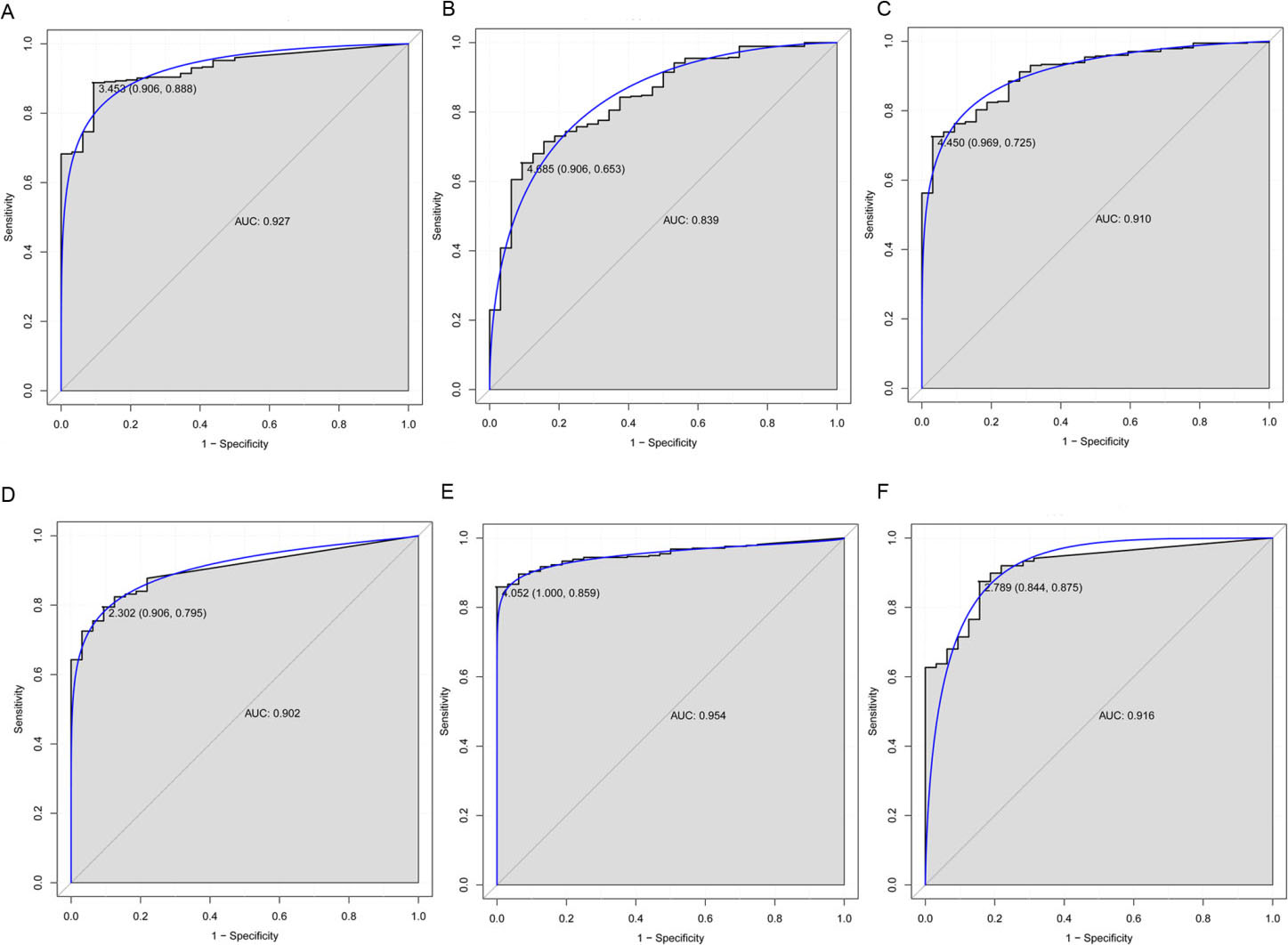
ROC curve analyses were conducted to assess the diagnostic value of selected DElncRNAs for GAC. The AUC of every selected DElncRNAs including HOTAIR (0.927), C20orf166-AS1 (0.839), PGM5-AS1 (0.910), HOXC-AS3 (0.902), HOXC-AS2 (0.954) and AC012531.1 (0.916) was more than 0.8 (Fig. 5), which suggested that all these six DElncRNAs had excellent diagnostic value for GAC.
Discussion
Although function of most lncRNAs remains largely unknown, lncRNAs have been indicated to involve with the pathogenesis of GAC. The current study identified key DElncRNAs in GAC and further explored their potential roles in GAC by functional annotation of DEmRNAs interacted with them.
Two gastric cancer-related DElncRNAs including HOX transcript antisense intergenic RNA (HOTAIR) and HOXC-AS3, were identified in the present study. HOTAIR is a well-studied lncRNA that regulates gene expression through mediating the modulation of chromatin structure [15]. Up-regulated HOTAIR has been observed in various cancers including colorectal cancer, hepatocellular carcinoma, nasopharyngeal carcinoma and gastric cancer [16, 17, 18, 19]. Furthermore, HOTAIR was demonstrated to play a role in carcinogenesis and progression of gastric cancer by involving with inhibition of apoptosis and promoting invasiveness [12]. HOXC-AS3 was reported to involve with tumorigenesis of gastric cancer, and served as a potential diagnosis and therapy for gastric cancer, which could facilitate cell proliferation and migration of gastric cancer through binding to Y-box-binding protein 1 (YBX1) [13].
In addition, several lncRNAs associated with cancers including HCG22, LINC00355, LINC02471, C20 orf166-AS1 and PGM5-AS1 were found to be aberrantly expressed between GAC and adjacent non-tumor tissues, as well. Decreased HCG22 (HLA complex group 22) expression has been observed in pro-state cancer, head and neck squamous cell carcinoma and oral squamous cell carcinoma, which was associated with poor survival of patients [20, 21, 22]. Aberrantly expressed LINC00355 was found in bladder cancer and involved with tumor stage, lymphatic metastasis, and distant metastasis of colon adenocarcinoma [23, 24]. Up-regulated LINC02471 expression was observed in papillary thyroid carcinoma tissues, which had a potential prognostic value for papillary thyroid carcinoma [25]. The expression of C20orf166-AS1 was down-regulated in prostate cancer tissues compared to normal tissues [26]. PGM5-AS1 were significantly associated with the incidence and development of melanoma and high expression of PGM5-AS1 were associated with poor overall survival for patients with in colorectal cancer [27, 28]. To our knowledge, these five cancer-related lncRNAs (HCG22, LINC00355, LINC02471, C20orf166-AS1 and PGM5-AS1) were firstly found to be differentially expressed between GAC and adjacent non-tumor tissues in this study. And we speculated that these five lncRNAs might involve with GAC, and their precise roles in GAC need further research.
Considering lncRNAs have been indicated to involve with cancer by regulating gene expression with cis- and trans-regulatory mechanisms [29], DElncRNA -DEmRNA co-expression network, DElncRNA-nearby DEmRNA interaction network and functional annotation of DEmRNAs integrated with DElncRNAs were used to explore function of key DElncRNAs in GAC. After these bioinformatic analysis, six HOX genes including HOXC8, HOXC9, HOXC10, HOXC11,HOXC12 and HOXC13 were found to be nearby targets of a GAC-related lncRNA HOXC-AS3. Moreover, HOXC10 was co-expressed with HOXC-AS3 as well. All of these six genes were members of HOX genes and accumulated evidences have indicated that HOX genes played important roles in gastrointestinal cancer [30, 31, 32, 33]. HOXC9 promoted the metastasis and stem cell-like phenotype of gastric cancer cells [33]. HOXC10 promoted cell proliferation and metastasis of gastric cancer through MAPK pathway and NF-
Functional annotation indicated that Fat digestion and absorption (hsa04975) and Vitamin digestion and absorption (hsa04977) were two pathways enriched for both nearby DEmRNAs of DElncRNAs and DEmRNAs co-expressed with DElncRNAs, which highlighted that aberrantly expressed lncRNAs in GAC might involve with GAC by regulating expression of genes enriched in these two pathways. Three genes including APOA1, APOA4 and APOB were enriched for both of the two pathways. APOA1 and APOA4 encode apolipoprotein A1 (ApoA1) and apolipoprotein A4 (ApoA4), respectively, which are major constituents of high-density lipoprotein. While APOB encodes apolipoprotein B (APOB) that is the main apolipoprotein of chylomicrons and low-density lipoproteins. Recently, study indicated that ApoB/ApoA1 ratio could act as an independent prognostic factor in gastric cancer [34]. Moreover, both APOA4 and APOB were co-expressed with a same lncRNA, AC115619.1. APOB was a nearby target of AC115619.1, and APOA1 and APOA4 were nearby targets of AP006216.2. Therefore, AC115619.1-APOA4/APOB and AP006216.2-APOA1/APOA4 integrations were speculated to play vital roles in GAC by mediating Fat digestion and absorption and Vitamin digestion and absorption.
In addition, another two lncRNAs including HOXC-AS2 and AC012531.1 have great diagnostic value for GAC, which may serve as potential biomarkers for GAC. ETV6 (ets translocation variant gene 6) encodes a transcriptional repressor, and involves in various cancers-related translocations, such as acute myeloid leukemia and gastrointestinal stromal tumor [35, 36]. In pediatric acute myeloid leukemia, HOXC-AS2 was found to be a translocation partner of ETV6 and ETV6-HOXC-AS2 fusion may result in a loss of function of ETV6 or HOXC-AS2 [37]. We make a hypothesis that ETV6-HOXC-AS2 fusion might play a role in gastric cancer as well. AC012531.1 is a novel lncRNA that has never been reported, and further research is needed to explore its biological function in GAC.
Taken together, the present study identified key lncRNAs in GAC and further explored their potential roles in GAC by functional annotation of their integrated DEmRNAs. Five lncRNAs (HCG22, LINC003 55, LINC02471, C20orf166-AS1 and PGM5-AS1) associated with other cancers are potential regulators of GAC, as well. Six lncRNAs including (HOTAIR, C20orf166-AS1, PGM5-AS1, HOXC-AS3, HOXC-AS2 and AC012531.1) may serve as potential biomarkers for GAC. A GAC-related lncRNA, HOXC-AS3 might involve with GAC by regulating a set of HOX genes (HOXC8, HOXC9, HOXC10, HOXC11, HOXC 12 and HOXC13) with cis-effect. AC115619.1-APOA4 /APOB and AP006216.2-APOA1/APOA4 integrations are speculated to play roles in GAC through regulating Fat digestion and absorption and Vitamin digestion and absorption. This present study provides new clues for exploring the mechanism and developing diagnostic and therapeutic strategies for GAC. Further experiments are needed to confirm our conclusion.
Footnotes
Supplementary data
The supplementary files are available to download from
