Abstract
Hepatocellular carcinoma (HCC) is the most aggressive type of gastrointestinal tumor, with a high rate of mortality. However, identifying biomarkers for the treatment of HCC remains to be developed. We aimed to determine whether cell division cycle 25C (CDC25C) could be used as a novel diagnostic and therapeutic biomarker in HCC. Expression of CDC25C in HCC was analyzed by using GEPIA (Gene Expression Profiling Interactive Analysis) and UALCAN databases. GEPIA and CBioPortal databases were applied to analyze patients’survival and CDC25C mutations, respectively. PPI (Protein-Protein Interaction) network was further built by STRING (Search Tool for the Retrieval of Interacting Genes) and Metascape Web portals. To the best of our knowledge, the novel observations identified in the present study reveal that the expression of CDC25C in HCC was significantly enhanced when compare to that in normal liver tissues (P < 0.001). A higher CDC25C expression resulted in a remarkably shorter disease free survival as well as overall survival. Moreover, the expression of CDC25C in HCC was related to HCC patients’grade and race, but not gender. The expression levels of CDC25C elevated gradually from stage 1 to 3 but decreased in stage 4. The specific gene mutations V41A, L87 H, N222 K and X309-splice of CDC25C occurred in HCC samples and these unique mutations were not detected in any other tumor tissues. Finally, PPI networks and GO enrichment analysis suggested that CDC25C might be associated with cell cycle and p53 signaling pathway. Taken together, bioinformatics analysis revealed that CDC25C might be a potential diagnostic predictor for HCC.
Introduction
Although hepatocellular carcinoma (HCC) is the sixth common cancer, it is the second cause of cancer deaths in Asian/Pacific islander. 1,2 During the past 2 decades, the great advancement of HCC treatment has been achieved, including surgical resection, ablation, liver transplantation and targeted therapy. 3 However, the 5-year survival rate for HCC has remained unsatisfied. Due to toxicity to the liver, radiotherapy is not used for the treatment of HCC. 4 Although liver transplantation is a potential approach for HCC, its application is limited by a number of factors.The reason for liver transplantation is not only liver donor shortage but liver transplantation have low success rate beyond Milan criteria, and etiology is also important because of high relapse rate in HCV infection. Recurrence of hepatitis C virus (HCV) infection following liver transplantation is a major source of morbidity and mortality. Targeted therapy can block the action of key molecules in the formation and progression of liver cancer. However, only a few drugs are currently available for patients with advanced liver cancer. Therefore, it is significant to investigate the key genes and major signal pathways involving in the development for better treatment of HCC. To date, serum alpha-feto protein (AFP) and PIVKA-II (protein induced by vitamin K absence or antagonist-II) are 2 best used biomarker for HCC in clinical screening, diagnosis and monitoring. 5 -10 AFP is not only important for diagnosis but also is important as a prognostic and predictive factor for anti-VEGF treatment and outcome after transplantation and surgery.The combination use of both biomarkers has significantly improved HCC sensitivity detection despite that sensitivity and specificity are far from satisfactory. 11 -14 In order to improve the clinical diagnosis and treatment of HCC, it is necessary to find new genes that are useful for screening, diagnosis and monitoring of HCC.
Dual specificity phosphatase CDC25 originally discovered in yeast can dephosphorylate phospho-threonine and phospho-tyrosine. 15,16 In humans, CDC25 family is composed of CDC25A, B and C. 17 Several studies have shown the association between CDC25C and human cancers. Ozen et al 18 found that CDC25C was closely related to the occurrence, development and prognosis of prostate cancer. Wang et al 19 further confirmed that overexpression of CDC25C played an important role in the occurrence and development of squamous cell carcinoma of female vulva. Furthermore, Xia Z.et al 20 found that CDC25C predicts poor prognosis in Lung Adenocarcinoma(LUAD)and may function in cell cycle regulation and FAS-mediated apoptosis. Al-Matouq et al 21 also found that CDC25B and CDC25C were increased in mouse and human skin cancers.
In this study, to verify the value of CDC25C for the diagnosis and treatment of HCC, the GEPIA and UALCAN databases analysis were performed. Analogously, we observed here that the expression level of CDC25C in HCC tissues was significantly higher than that in normal liver tissues. 22 The expression level of CDC25C in HCC is closely related to tumor size, differentiation, stages and grades. 23 Furthermore, CBioPortal databases were applied to unveil the specific mutations of CDC25C in HCC patients. PPI network and GO enrichment analysis mined some promising CDC25C relevant genes, further suggested that these genes are significant enriched in cell cycle, signal transduction involved in DNA integrity checkpoint, cell cycle G2/M phase transition and P53 signaling pathway.
Materials and Methods
UALCAN Analysis
CDC25C expression data and clinical data of HCC patients in TCGA were downloaded from UALCAN(http://ualcan.path.uab.edu/analysis.html). 5 In this study, we utilized this online tool to analyze the expression levels of CDC25C between HCC specimen and normal tissues.
Survival Analysis
GEPIA(http://gepia.cancer-pku.cn/) is an interactive web resource and database for analyzing cancer transcriptome and patients’ survival.In this study, we utilized this online tool to analyze patients’survival.Using GEPIA, overall survival(OS) and disease free survival(DFS) were presented and the hazards ratio was calculated based on Cox PH Model.95% confidence interval was added as dotted line.The thresholds for high and low expression level cohorts are 50% respectively.
Correlation Genes Analysis
Protein-protein interaction of CDC25C was conducted in STRING online service.The STRING database (http://string-db.org/) was used to analyze PPI networks.The non-interacting genes were excluded.The top 10 genes with the highest degree of connection to the others were presented.
Pathway and Process Enrichment Analysis
The Metascape database (http://metascape.org/) is an online tool for high-throughput functional analysis of genes.All the genes interacted with CDC25C in STRING were included in the enrichment analysis in Metascape database.Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway, Gene ontology (GO) biological process, Reactome Gene Sets and Canonical Pathways were enriched.
Mutations Analysis
CBioPortal (https://www.cbioportal.org/) is a database with integrated genetic data, including DNA mutations, methylations, gene amplifications and proteinalterations.In our study, the cBioPortal database was used to analyze the association between genetic mutations and the development of HCC.The top 10 geng which are related to CDC25C were analyzed by using cBioPortal database.We performed CDC25C gene mutations analysis across all tumor samples (n = 14) from the TCGA-HCC database.
Results
The Expression of CDC25C in HCC Patients
To verify CDC25C expression levels in HCC tissues and the value to the diagnosis and surveillance of HCC, GEPIA database was applied. As shown in Figure 1A, the expression level of CDC25C was significantly higher in HCC (n = 371) compared with that in normal tissues (n = 50;P < 0.001). Moreover, the relationship between CDC25C expression levels and HCC patients’clinicopathological parameters were further analyzed by UALCAN databases. The result demonstrated that expression of CDC25C was higher in Asian HCC patients than that in Caucasian patients(P < 0.05, Figure 1B). The expression levels of CDC25C increased from grade 1 to grade 4, suggesting CDC25C was remarkably correlated with HCC patients’grade (Figure 1C) (grade1 vs grad e3, P < 0.001;grade 1 vs grade 4, P < 0.001;grade 2 vs grade 3, P < 0.001;grade 2 vs grade 3, P < 0.05). As shown in Figure 1D, there are gradually increased expression levels of CDC25C from stage 1 to stage 3 but obviously declined in stage 4 (stage 1 vs stage 2, P < 0.05;stage 1 vs stage 3, P < 0.001).
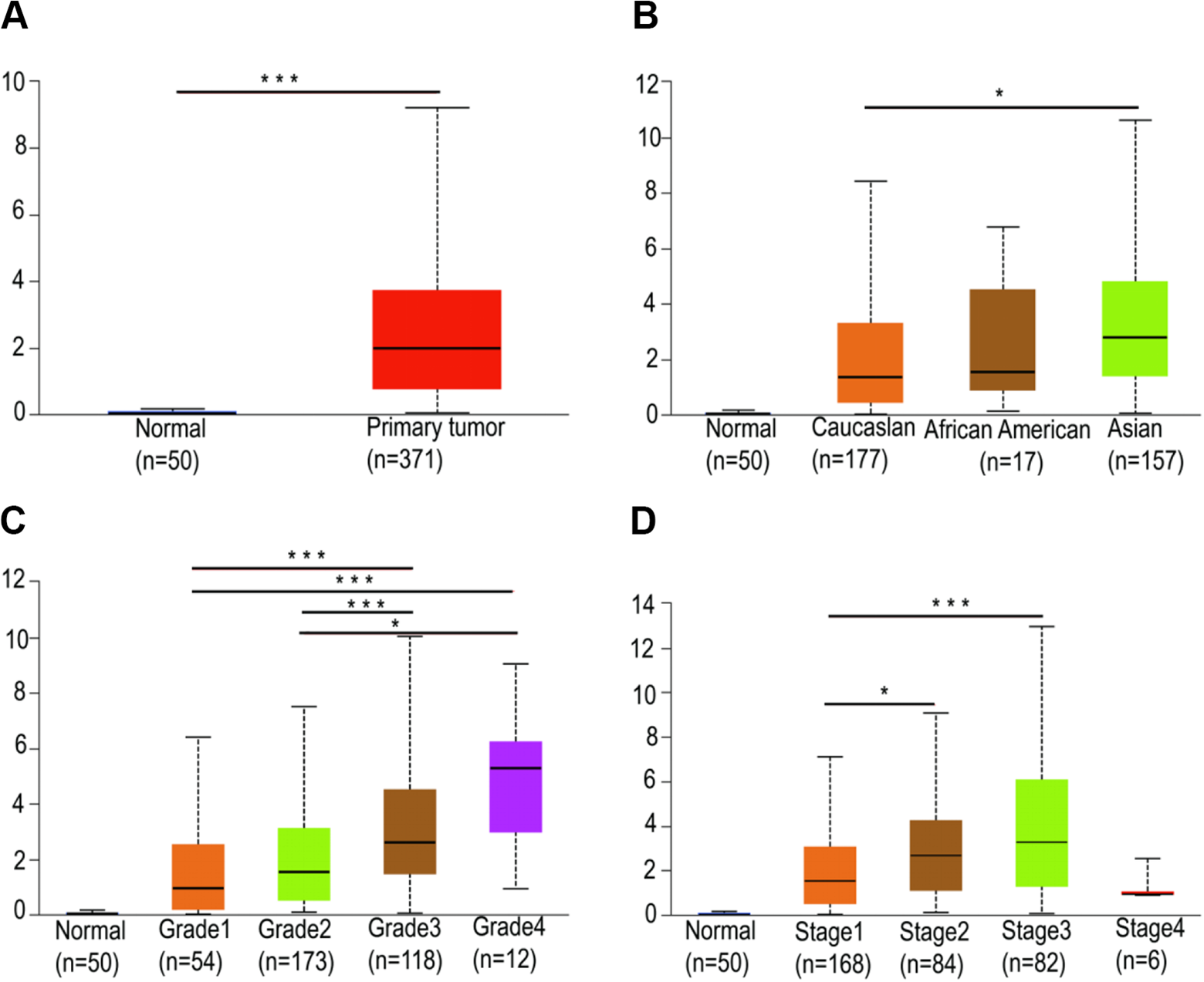
Expression of CDC25C in patients with HCC. (A) The expression of CDC25C in HCC obtained from GEPIA online tool (P < 0.001; tumor vs. normal). (B) Expression of CDC25C in HCC based on race (P < 0.05; Caucasian vs. Asian), and (C) individual cancer grade (P < 0.001; grade 1 vs. grade 3; grade 2 vs. grade 3;grade 1 vs. grade 4; grade 2 vs. grade 4) and (D) individual cancer stage (P < 0.001; stage 1 vs. stage 2; stage 1 vs. stage 3).*, P < 0.05;**, P < 0.01;***, P < 0.001.
Effect of CDC25C Expression Level on Patient Survival
In survival analysis, CDC25C expression of HCC patients were divided into low-expression group and high-expression group (cutoff-high is 50%, cutoff-low is 50%). As shown in Figure 2A and Figure 2B, a lower CDC25C expression level resulted in a significantly higher survival probability (P < 0.001). Survival curves analysis showed that CDC25C was suitable for predicting HCC patients’prognosis.

Higher expressions of CDC25C was associated with poorer DFS (A) and OS (B) in HCC patients. (P < 0.0001).
PPI Networks and Process Enrichment Analysis of CDC25C
As shown in Figure 3, PPI network analysis indicated that CDC25C interacted with 10 functional genes, including CDK1, CCNB1, CHEK2, CHEK1, PLK1, CCNA2, YWHAZ, TP53, CCNB2 and WEE1. To further determine the potential function CDC25C in HCC, GO enrichment analysis and KEGG patheway analysis of CDC25C were performed using Metascape. The result revealed that those proteins were biologically closely associated with cell cycle and p53 signaling pathway (Figure 4 and Table 1).

PPI network of the top 10 hub genes related to CDC25C.
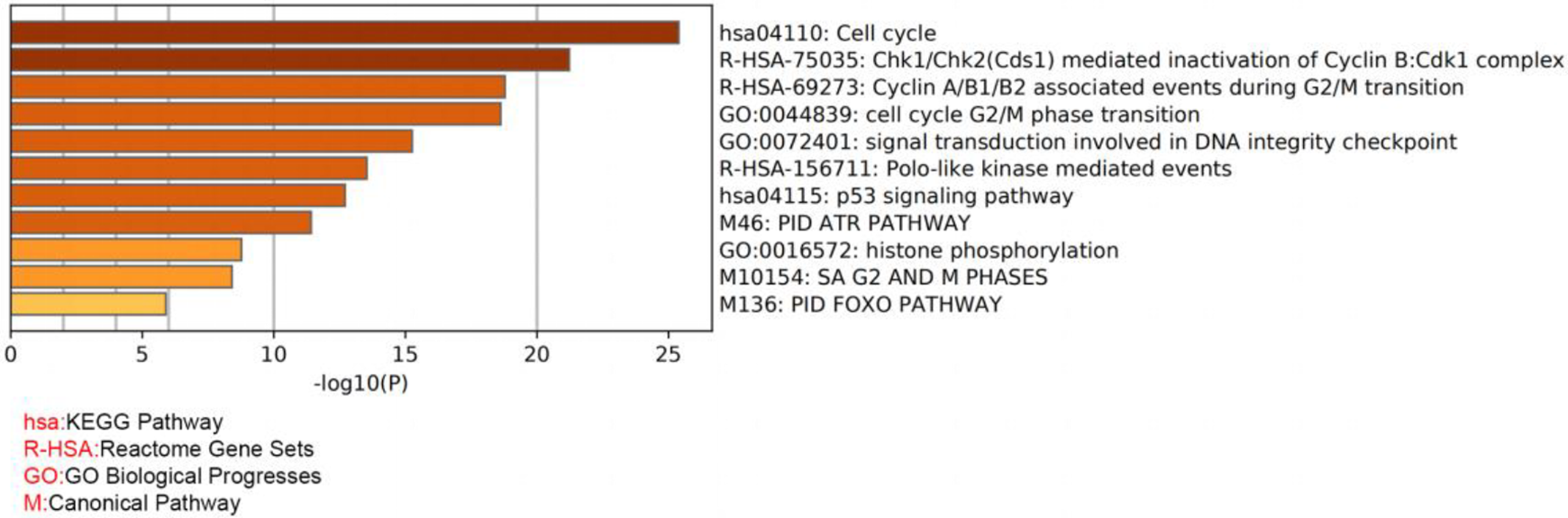
GO enrichment analysis of the top 10 hub genes.
Significantly Enriched GO Terms and KEGG Pathways of CDC25C.

Specific Mutations of CDC25C Genes in HCC Patients
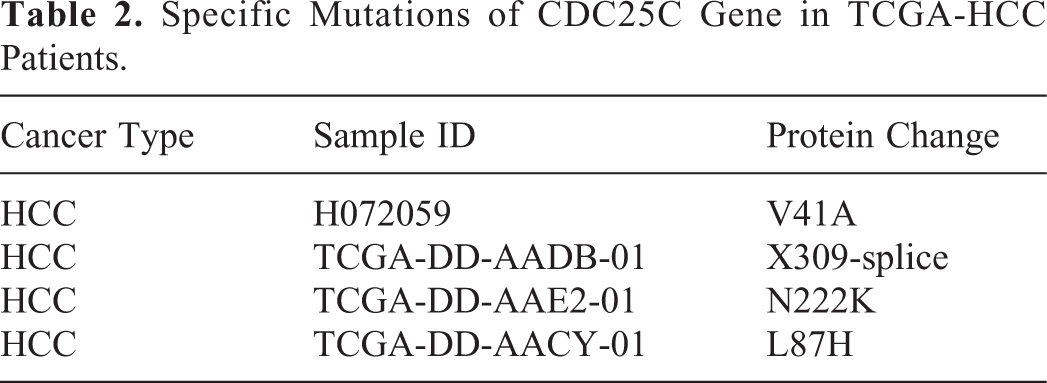
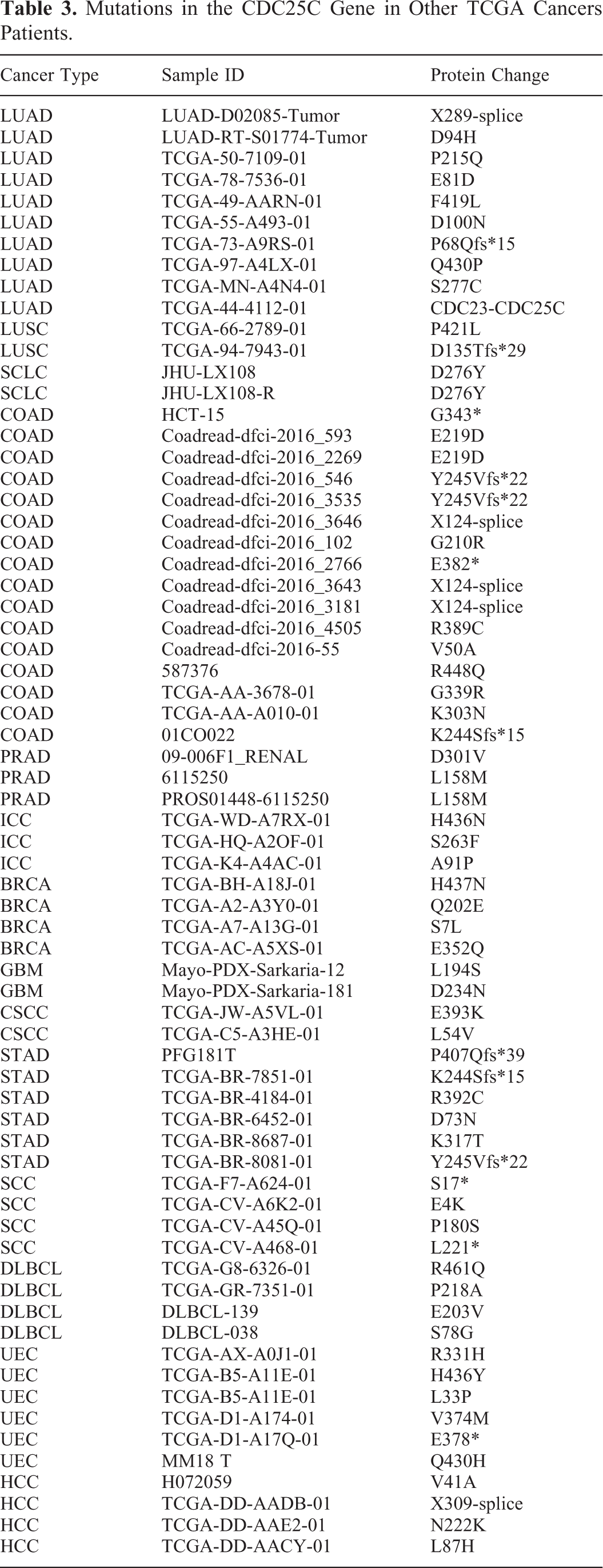
In order to analyze the mutations of CDC25C gene in HCC patients’ tissues, the CBioPortal database analysis was employed. As shown in Tables 2 and 3, we performed CDC25C gene mutations analysis across all tumor samples (n = 14) from the TCGA-HCC database (https://www.cbioportal. org/). These particular mutations of CDC25C in HCC patients might contribute greatly to HCC clinical diagnosis and monitoring. Moreover, the mutations of CDC25C and its interacted genes (CDK1, CCNB1, CHEK2, CHEK1, PLK1, CCNA2, YWHAZ, TP53, CCNB2 and WEE1) were analyzed by means of the cBioPortal dataset. The alteration statuses of 10 key genes were analyzed. The frequency of alteration of 10 hub genes were shown in Figure 5A and Figure 5B. Intriguingly, there were 4 specific mutations V41A, L87 H, N222 K and X309-splice (Table 2, Figure 5C and Figure 5D) in the HCC samples that were not present in any other tumor samples. Furthermore, the OS (Figure 5E) and DFS (Figure 5F) were significant better in no alterations group than that in alterations group (P < 0.05). Survival curves analysis showed that CDC25C gene mutation may affect the prognosis of HCC patients.
Specific Mutations of CDC25C Gene in TCGA-HCC Patients.

Mutations in the CDC25C Gene in Other TCGA Cancers Patients.

(A) A visual summary of Genetic alterations (data from HCC in TCGA) shows the genetic alteration of 10 key genes which were altered in 184 (49%) of 377 HCC patients. (B) The total alteration frequency of 10 key genes is illustrated. (C) the genetic alteration of CDC25C genes which were altered in 7 (2%) of 377 HCC patients. (D) The particular mutations (V41A, L87 H, N222 K, X309-splice) of CDC25C in HCC patients. (E) Overall survival Kaplan Meier estimate. (F) Progression free survival Kaplan Meier estimate.
Prediction of Relevance Genes to CDC25C in HCC
The correlation between CDC25C and its 10 interacted genes were selected to further analyzed in HCC patients. We found that the 6 genes(CCNB1, CCNB2, CDK1, CHEK1, CHEK2, PLK1) were considered to be relevant genes and the scatter plots were shown in Figure 6.
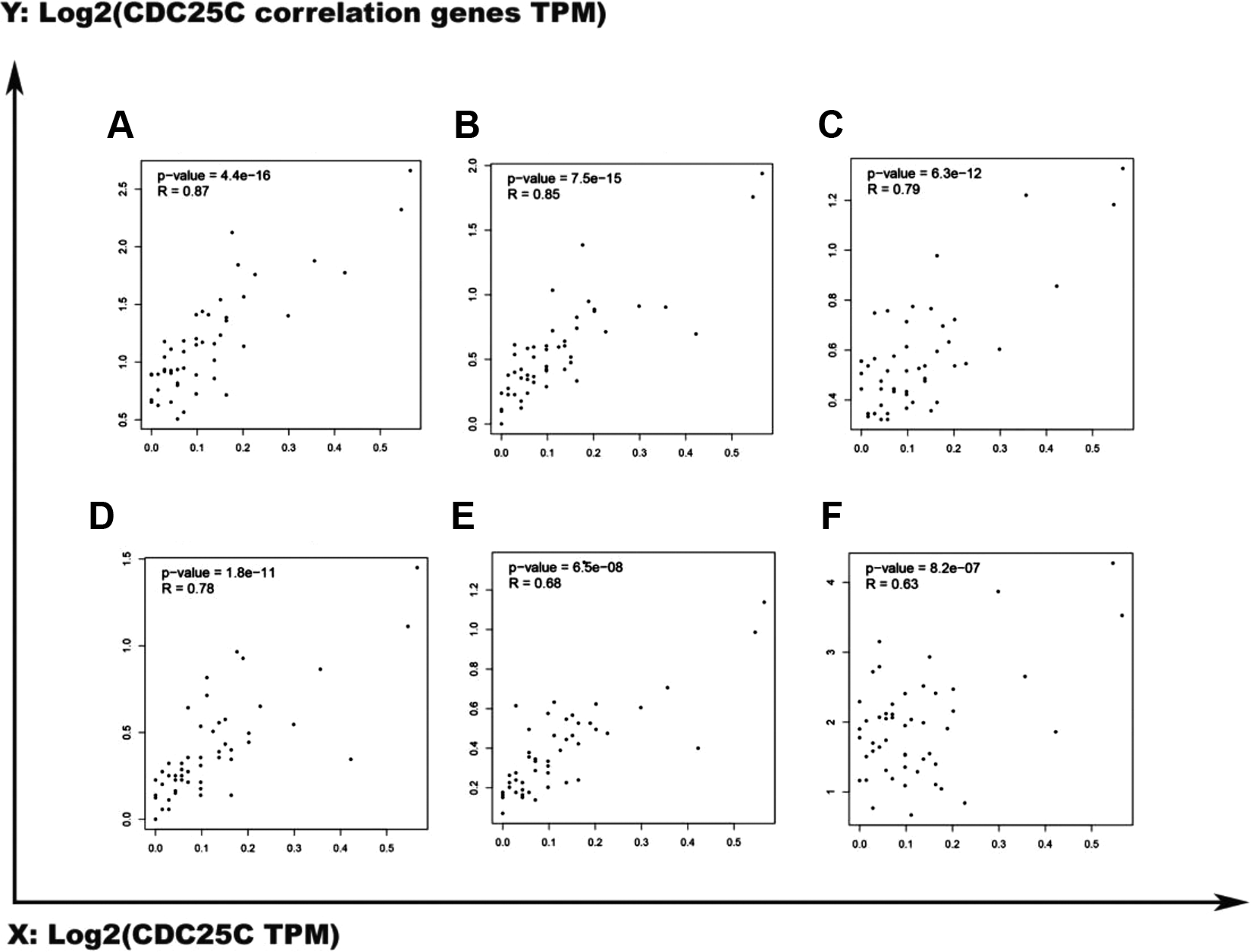
The plots of 6 pair-wise correlation genes. (A) CDC25C and CCNB1, (B) CDC25C and CDK1, (C) CDC25C and CHEK1, (D) CDC25C and CCNB2, (E) CDC25C and PLK1, (F) CDC25C and CHEK2.
Discussion
Bioinformatics methods can provide us with gene expression levels and predict potential therapeutic targets. 24 A large number of clinical data showed that the death rate of liver cancer is very high.One of the best ways to reduce mortality is to detect accurately and treat successfully. Identifying novel key genes associated with the development and the progression of HCC is crucial for its diagnosis and treatment. In this study, our data showed that CDC25C expression was higher in HCC patients than that in control normal tissues, and correlated with HCC patients’ grade. The results demonstrated that CDC25C expression enhanced gradually from stage 1 to stage 3 but decreased in stage 4. A higher CDC25C expression resulted in a significant shorter DFS and OS in HCC patients, suggesting that CDC25C may play an important role in the prognosis of HCC. The CBioPortal tool have unveiled 4 specific mutations V41A, L87 H, N222 K and X309-splice unique presented in the HCC. These characteristic mutations of CDC25C in HCC patients might facilitate to HCC clinical diagnosis and monitoring. To determine the probably pathogenic mechanism of CDC25C in HCC, PPI network and GO enrichment analysis were further applied. Ten CDC25C interacted proteins: CDK1, CCNB1, CHEK2, CHEK1, PLK1, CCNA2, YWHAZ, TP53, CCNB2 and WEE1 were identified. GO enrichment analysis suggested these genes are enriched in cell cycle, CHK1/CHK2(cds1) mediated inactivation of cyclin B: CDK1 complex, cyclin A/B1/B2 associated events during G2/M transition, FOXO pathway, p53 signaling pathway and PID-ATR pathway.
As we known, CDC25C plays an important regulatory role in the G2/M phase transformation of the cell cycle. Lindqvist 25 conducted an experimental study verified that CDC25C could activate the Cyclin B/CDC2 complex through dephosphorylation and facilitate the smooth transition from G2 phase to M phase during mitosis.The cell cycle pathway is one of the most important cellular signaling pathways that regulates both cell division and apoptosis. DNA damage readily leads to dysregulation of the cell cycle, which is an essential step in the initiation and development of human malignancies. 26 -28 It has been reported that cell cycle arrest has been confirmed to be an effective approach in controlling tumor growth. 29,30 There were more cells in the G2 /M phase, whereas cells in the G1 or S phase were reduced. The expression of CDC25C might contribute to the HCC cell cycle by regulating the G2/M phase. By inhibiting the expression level of the proteins CDC25C, HCC cells proliferation and the G2/M phase cell cycle arrest can significantly inhibited. It has been reported that checkpoint is an important regulatory node of the cell cycle, 31 -33 and cell can only passing a checkpoint test can cell enter the next cell cycle. 34
It is widely believed that the CDK1/Cyclin B complex is a key regulatory factor of checkpoint in the G2/M phase. 35 The complex is synthesized in large quantities when the cell passes through the G2/M phase. 36 Therefore, CDK1 /Cyclin B complex is decreased with G2/M cycle arrest. The study showed that the cancer cell arrest at the G2/M phase when the content of the CDK1/Cyclin B complex decresed. It is widely believed that increasing the phosphorylation of CDC25C, can lead to inhibition of the formation CDK1/Cyclin B complex and cell cycle arrest at the G2/M phase. 37,38 The protein kinase CDC25C, which belongs to the dual-specificity CDC25 phosphatase family, dephosphorylates the 2 inhibitory residues of CDK1 (Tyr15 and Thr14) to activate the CDK1-Cyclin B complex at the G2 phase. 39 Therefore, CDC25C might contribute to the HCC cell cycle by activating the CDK1-Cyclin B complex at the G2 phase. In turn, by inhibiting the expression level of CDC25C, HCC cells proliferation and the G2/M phase cell cycle arrest can be significantly inhibited.
Zou Y, et al found that CDK1, CCNB1 and CCNB2 are potential prognostic biomarkers and associated with immune cell infiltration in HCC. 40 The genes may be utilized to predict the reaction of immunotherapy. Combining inhibitors of these genes with immunotherapy may improve the survival time of HCC patients. mRNA expression of CDK1, CCNB1 and CCNB2 was up-regulated in various tumor tissues including HCC. Higher expression of these genes was associated with poorer prognosis in HCC patients. Notably, expression levels of CDK1, CCNB1 and CCNB2 were positively correlated with infiltrating levels of CD4+ T cells, CD8+ T cells and dendritic cells in HCC. Malki A, Elbayaa RY, et al found that CCNB1, CCNB2 and CHEK1 were involved in p53 signaling pathway. 41 Abundant studies found that upregulating CCNB1, CCNB2 and CHEK1 could regulate p53 signaling pathway, thereby promoting the progression of HCC. 42,43 Liu S, et al 44 found that mRNA levels of 3 p53 markers including CCNB1, CCNB2 and CHEK1 were significantly upregulated in 20 HCC tumor tissues versus nontumor adjacent tissues by bioinformatics analysis and experimental verification, and their mRNA expression levels were all negatively correlated with HCC patients’survival rates. In this study, GO and KEGG analyses revealed that CDC25C was involved in pathways including p53 signaling pathway, which might be associated with invasive characteristics of HCC. According to the report, CDC25C can suppress p53-induced growth arrest, 45 and deregulation of p53 signaling pathway might to be principal mechanisms in liver tumors.
In summary, this study has novelly identified that the elevated expression of CDC25C in HCC patients when compared to that in normal tissue.The results from this study may push forward the mechanism underlying CDC25C progression, and provide the high prognostic value of HCC. However, further studies are needed to intensively disclose the molecular mechanism and implication of CDC25C in HCC tumorigenesis and therapy.
Footnotes
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Ethics Statement
The data we utilized were all drawn from TCGA dataset, which were not obtained from our own clinical samples. The current study was approved and consented by the ethics committee of The Second School of Clinical Medicine & Jingzhou Central Hospital of Yangtze University.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by grants from the National Natural Science Foundation of China (31700736), China Scholarship Council (201908420102), Hubei Medical Youth Tip-Top Talent (to XW Wang), Leading Talent Program of Yangtze Talent Project (to XW Wang) and the College Students Innovative Entrepreneurial Training Program in Yangtze University (2018184, 2019372).
