Abstract
BACKGROUND:
The prognosis of patients with recurrent and/or metastatic oral squamous cell carcinoma (OSCC) remains poor, and its incidence is especially high in developing countries. Multiple long non-coding RNAs (lncRNAs) are recently identified as crucial oncogenic factors or tumor suppressors. This study aimed to probe into the role of lncRNA small nucleolar RNA host gene 17 (SNHG17) on the progression of OSCC.
METHODS:
The expression level of SNHG17 in OSCC samples was tested using quantitative real-time polymerase chain reaction (qRT-PCR). Human OSCC cell lines CAL-27 and Tca8113 were used in in vitro studies. Cell counting kit-8 (CCK-8) and BrdU assays were used to assess the effect of SNHG17 on OSCC cell proliferation. Flow cytometry was used to study the effect of SNHG17 on OSCC cell apoptosis. Transwell assay was conducted to detect the effect of SNHG17 on migration and invasion. Moreover, luciferase reporter assay was employed to confirm targeting relationship between miR-375 and SNHG17. Additionally, Western blot was used to observe the regulatory function of SNHG17 on PAX6.
RESULTS:
SNHG17 expression in OSCC clinical samples was significantly increased and was correlated with unfavorable pathological indexes. Its overexpression remarkably accelerated proliferation and metastasis of OSCC cells, while reduced apoptosis. Accordingly, knockdown of SNHG17 suppressed the malignant phenotypes of OSCC cells. Overexpression of SNHG17 significantly reduced the expression of miR-375 by sponging it, but enhanced the expression of PAX6.
CONCLUSION:
SNHG17 is a sponge of tumor suppressor miR-375 in OSCC, enhances the expression of PAX6 indirectly, and functions as an oncogenic lncRNA.
Introduction
According to the latest report, in 2019, about 53000 people suffered from carcinoma of oral cavity and pharynx in the United States, with estimated deaths of 10860 [1]. Oral squamous cell carcinoma (OSCC) is one of the most commonly known cancers in the oral cavity. In China, about 3–6 cases/100,000 people are diagnosed with OSCC every year. Despite the progress in treatment strategies, the mortality of OSCC is still high [2, 3]. OSCC is associated with changes in gene expression during tumorigenesis and progression. In recent years, the molecular mechanism of OSCC has been continuously investigated, and some key molecules have become the targets of early diagnosis and treatment of OSCC [4].
lncRNAs, a kind of transcripts with a length of over 200 nucleotides (nt) and no protein coding ability, are proved to be important regulators of gene expression and biological processes [5]. More and more research points out that during the progression of tumors, there were many dysregulated lncRNAs. These lncRNAs can be used as prognostic indicators and potential therapy targets [6]. On the other hand, more and more data show that lncRNAs participate in regulating cancer cell proliferation, apoptosis, and metastasis [7]. For instance, lncRNA PANDAR expression is up-regulated in cervical cancer and lncRNA PANDAR serves as a poor prognostic diagnostic indicator, which can promote the proliferation of cervical cancer cells [8]. The expression of lncRNA NEAT1 not only can be used as a prognostic marker in renal cell carcinoma, but also can facilitate cell proliferation, migration, and invasion [9]. As a member of lncRNAs, lncRNA small nucleolar RNA host gene 17 (SNHG17) is reported to be an unfavorable prognostic factor for colorectal cancer (CRC), which can promote the proliferation of CRC cells through epigenetic silencing of p57 [10]. However, there is no report exploring the function and mechanism of SNHG17 in OSCC.
MiRNAs are small non-coding RNAs involved in gene regulation, which regulate the expression of target proteins at post-transcriptional or translation levels by binding to the 3’-UTR (untranslated region) of the mRNA [11, 12]. Study finds that miRNAs are also abnormally expressed in tumors, and they are involved in cancer biology by controlling the proliferation, metastasis, chemoresistance and so on [13]. In OSCC, miRNAs also play an important role. For example, miR-155 can regulate the OSCC procession by controlling p27Kip1 to regulate the proliferation and apoptosis [14]. MiR-375 is reported to be lowly expressed in hepatocellular carcinoma and significantly correlated with lymphatic metastasis and TNM stage [15]; in OSCC, miR-375 can block cancer progression by targeting PDGF-A [16]. However, the regulatory mechanism of miR-375 in OSCC remains unclear.
Paired box 6 (PAX6) is a member of the PAX family. It contains conserved paired box domain and homeobox domain, which can bind to DNA and function as regulators of gene transcription [17]. In non-small cell lung cancer, high expression of PAX6 is markedly correlated with lymph node metastasis [18]; in colorectal cancer, PAX6 expression is negatively regulated by miR-7 and PAX6 enhances cell proliferation, colony formation, and invasion [19]. Interestingly, study shows that PAX6 is a target gene of miR-375 [20]. This suggests that miR-375/PAX6 is probably an important axis in regulating cancer progression.
We found that there were binding sites between miR-375 and SNHG17, and miR-375 and 3’UTR of PAX6. In view of this, we made a hypothesis that SNHG17 could regulate the malignant phenotypes of OSCC by targeting miR-375/PAX6 axis. This study provided a theoretical basis for elucidating the molecular mechanism of OSCC progression and potential therapeutic targets for OSCC patients.
Materials and methods
Clinical samples
This study was ratified by the Ethics Committee of Stomatological Hospital of Nanchang University, and each patient signed informed consent. Forty pairs of tissue samples (cancer tissues/adjacent normal tissues) of OSCC patients were collected. Tissues were obtained from the tongue (17 cases), gingiva (10 cases), buccal mucosa (7 cases), and floor of the mouth (6 cases). Tissue specimens were immediately stored in liquid nitrogen at
Cell culture and transfection
Normal human oral keratinocyte (NHOK) and four OSCC cell lines (including Fadu, SCC-25, CAL-27 and Tca8113) were purchased from the American Type Culture Collection (Manassas, VA, USA). The above cells were cultured in Dulbecco’s Modified Eagle’s Medium (DMEM, Invitrogen, Carlsbad, CA, US) supplemented with 10% fetal bovine serum (FBS) and 1% penicillin/streptomycin (Invitrogen, Carlsbad, CA, US). The culture plates were placed in an incubator containing 5% CO
Quantitative real-time polymerase chain reaction (qRT-PCR)
Total RNA was extracted from tissues and cells by TRIzol (Invitrogen, Carlsbad, CA) method. Total RNA was then reversely transcribed with First Strand cDNA Synthesis Kit (Thermo Fisher Scientific Inc., Rockford, IL, USA). qRT-PCR was performed using SYBR Green Premix Ex Taq II (TaKaRa, Dalian, China) in line with the manufacturer’s protocol. GAPDH and U6 were used as internal references, and the primer sequences were shown in Table 1. Relative expression was calculated according to 2
Primer sequence of qRT-PCR
Primer sequence of qRT-PCR
Cells in logarithmic growth phase were trypsinized with 0.25% trypsin to make a single cell suspension. After counting the cells, about 2,000 cells were inoculated into 96-well plates per well and cultured in 37
BrdU assay
The transfected cells were seeded on cover slides, placed in a 24-well plate, and cultured overnight. BrdU kit (10
Flow cytometry
Cells were trypsinized with trypsin and collected by centrifugation. After washing the cells with PBS twice, 400
Transwell assay
Transwell assay was used to detect cell migration and invasion. 2
Western blot analysis
The cells were washed with PBS and lysed with RIPA lysis buffer (Thermo Science, Rockford, IL, USA) containing protease inhibitors. The supernatant was collected after high-speed centrifugation, and the protein was quantified by BCA method. The supernatant was heated in a water bath to denature the protein. SDS-PAGE was used to separate the protein samples, and then the protein was transferred to nitrocellulose (NC) membrane (Millipore, MA, USA), followed by the block with skimmed milk for 30 min at 37
Up-regulation of SNHG17 expression in OSCC. A. The expression of SNHG17 in normal paracancerous tissues and OSCC tissues was detected by qRT-PCR. 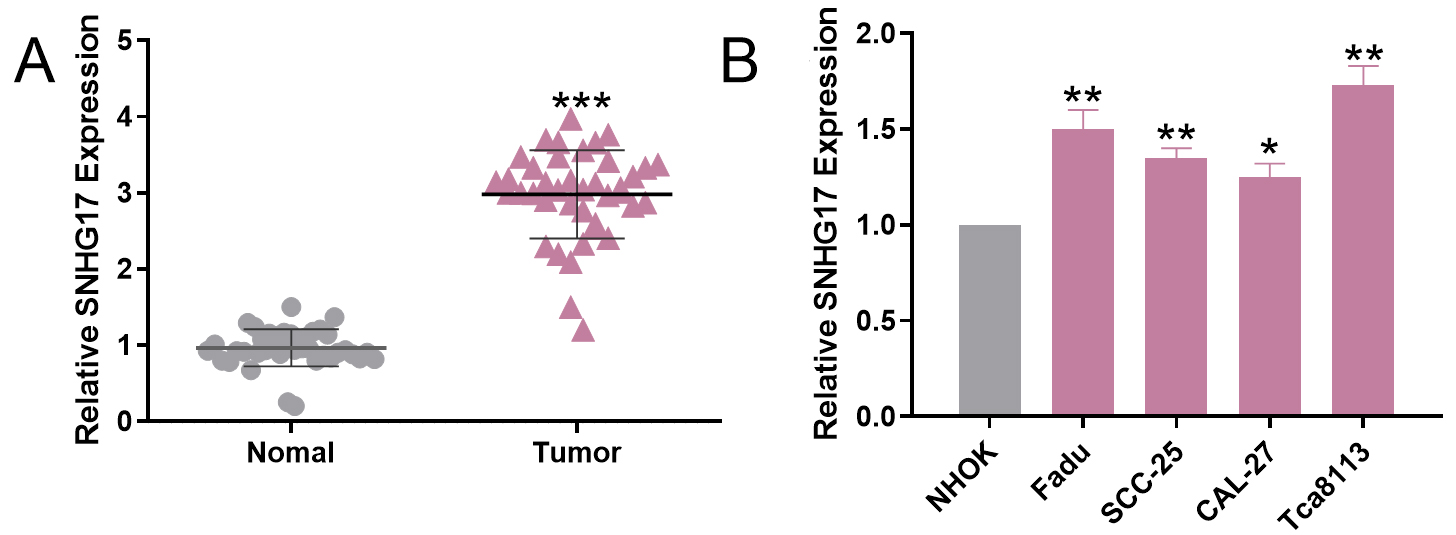
All luciferase reporter vectors (SNHG17-WT, SNHG17-mut, PAX6-wt and PAX6-mut) were constructed by Promega (Promega, Madison, WI, USA). Cells (4.5
Statistical analysis
SPSS 17.0 software (SPSS Inc., Chicago, IL, USA) was used to perform statistical analysis. The measurement data were expressed by mean
Results
SNHG17 expression was up-regulated in OSCC and associated with unfavorable pathological parameters
To investigate the expression of SNHG17, we tested the expression of SNHG17 in OSCC tissues and normal tissues adjacent to cancer by qRT-PCR. As shown, the expression of SNHG17 was significantly increased in OSCC tissues (
Correlation between SNHG17 and pathological parameters in OSCC
Correlation between SNHG17 and pathological parameters in OSCC
SNHG17 promoted the proliferation, migration, and invasion of OSCC cells. A. Models of overexpression and low expression of SNHG17 were established and the expression of SNHG17 was detected by qRT-PCR. 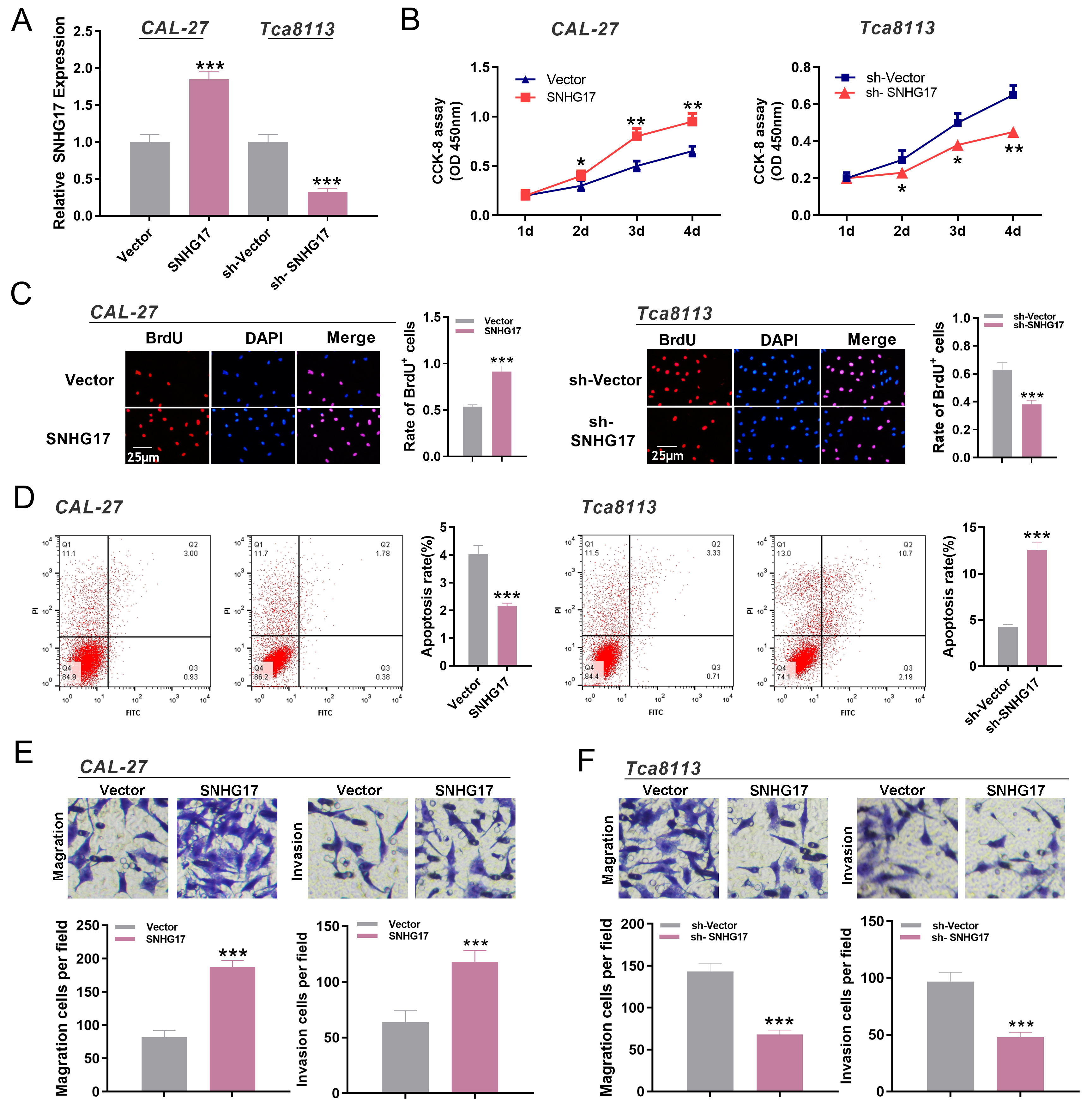
SNHG17 targeted miR-375. A. The downstream target of SNHG17 was predicted with StarBase database, and miR-375 was identified as a potential target of SNHG17. B. The luciferase reporter assay proved the targeting relationship between SNHG17 and miR-375. 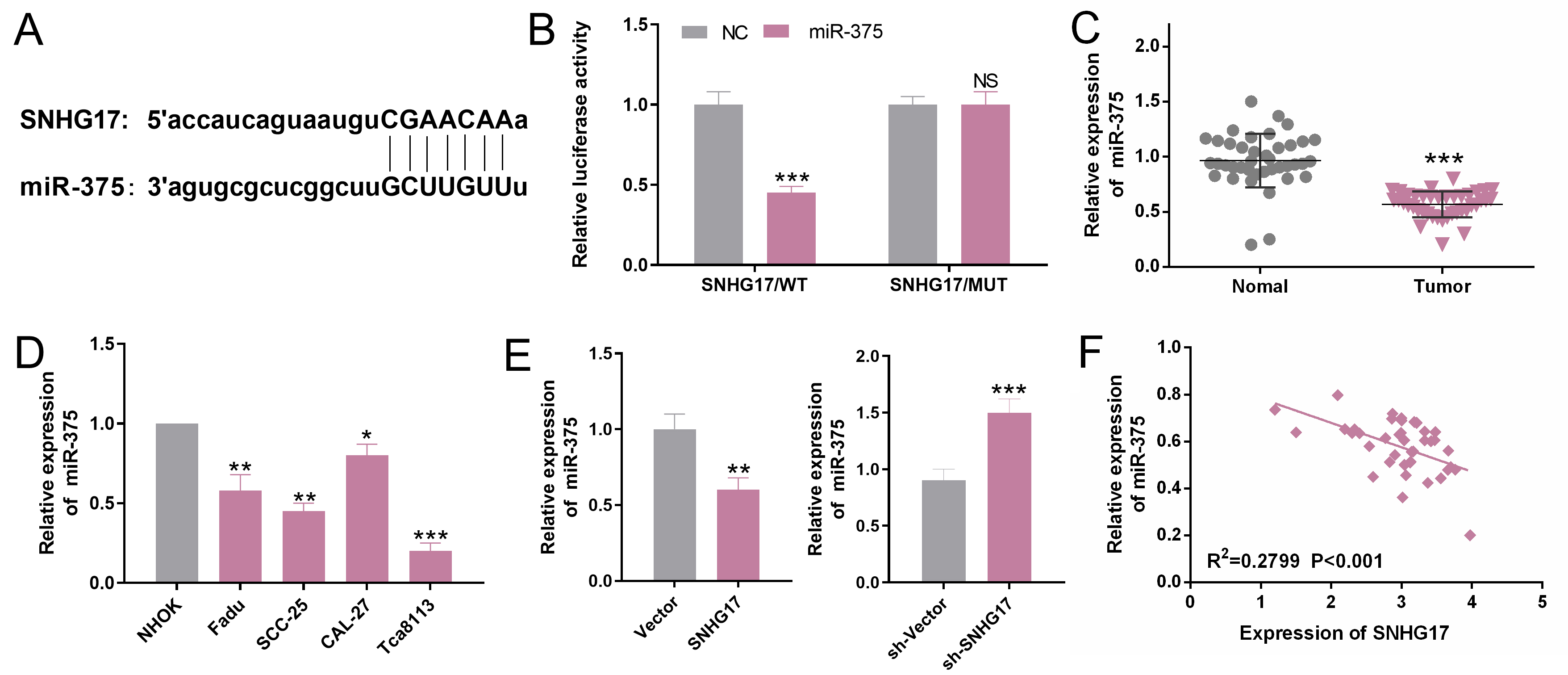
MiR-375 inhibited the proliferation, migration, and invasion of OSCC cells. A. The over-expression and inhibition models of miR-375 were established and the expression of miR-375 was detected by qRT-PCR. 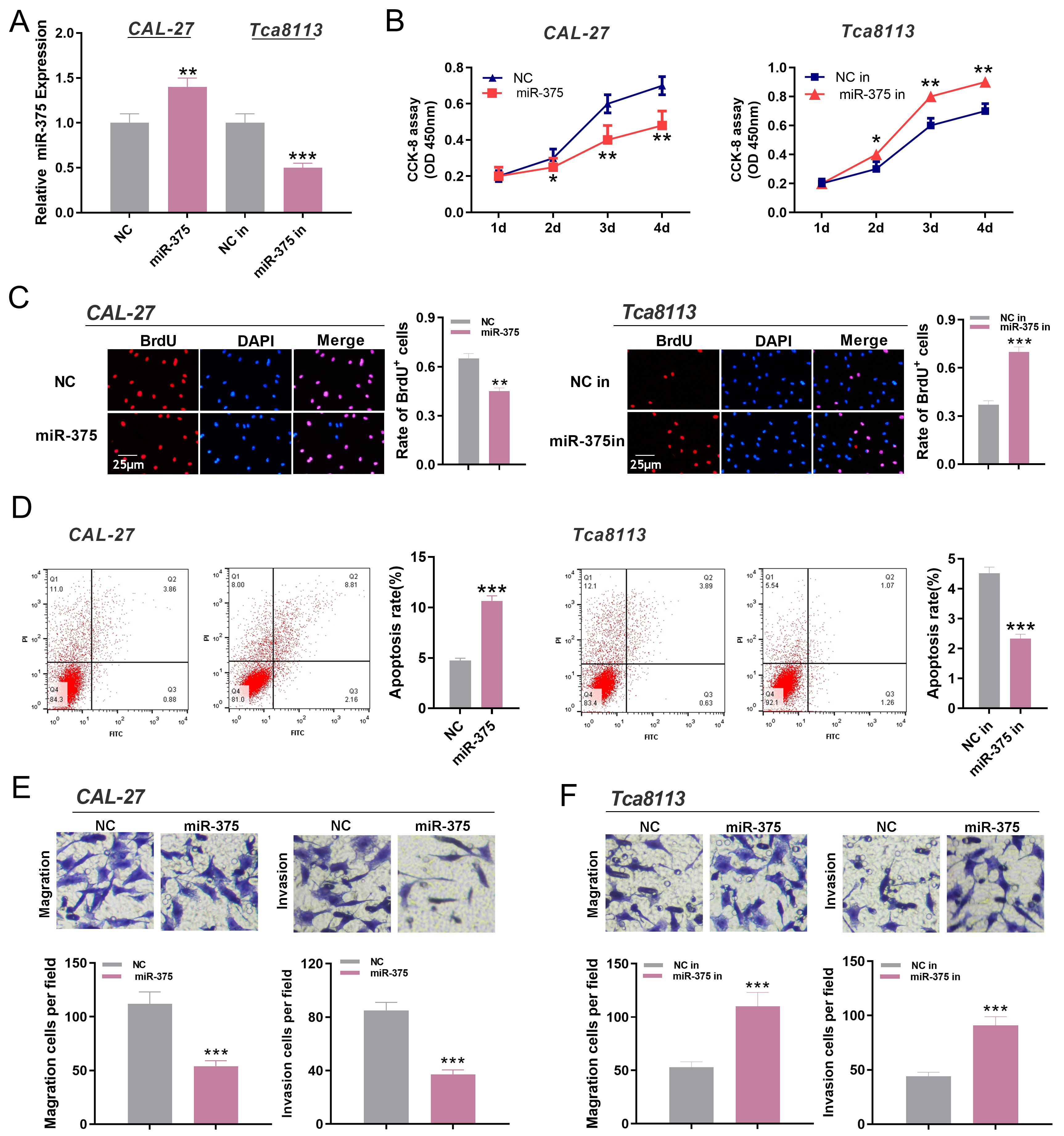
MiR-375 could target PAX6. A. The expression of PAX6 in normal tissues and OSCC tissues was detected by qRT-PCR. 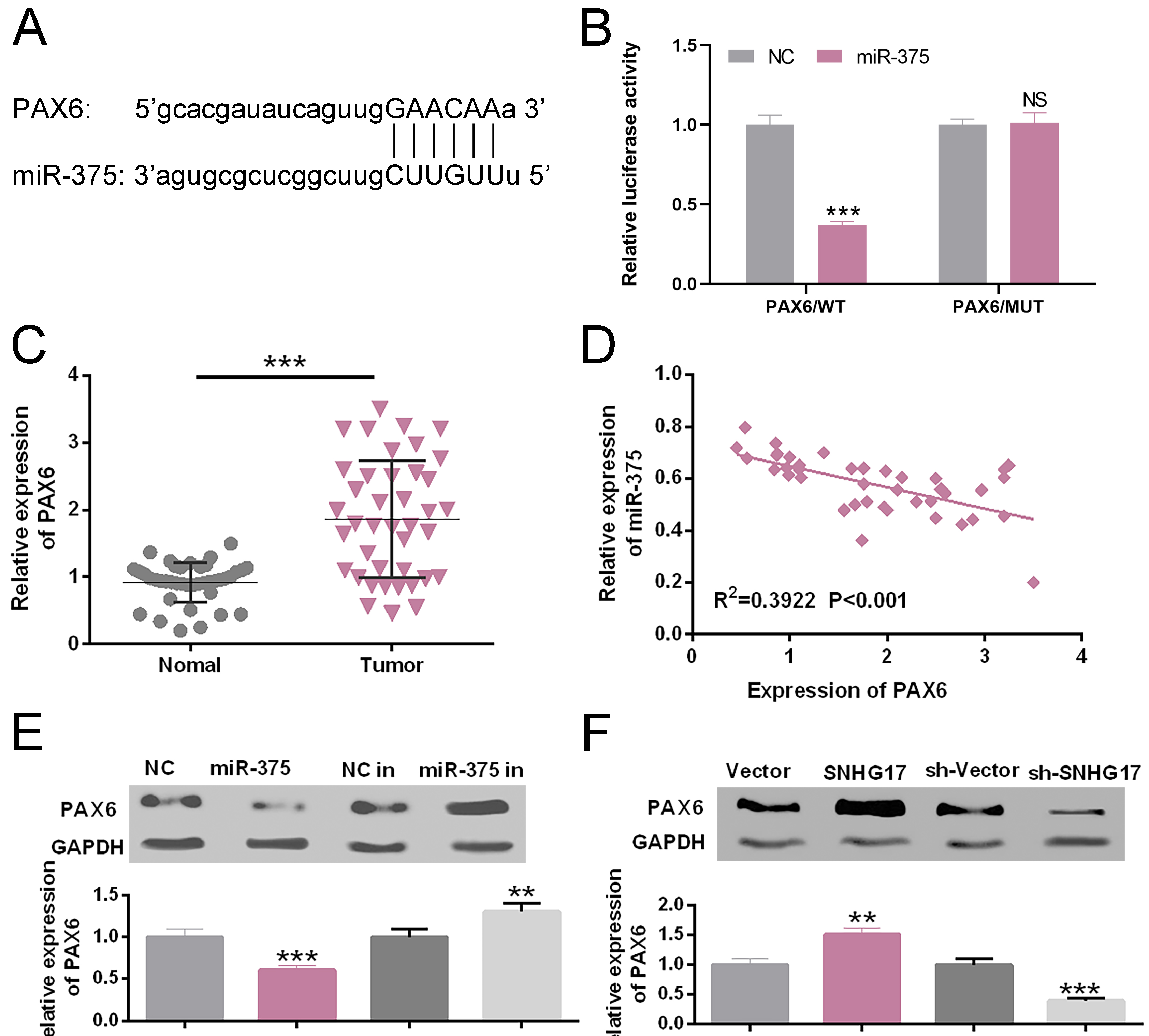
SNHG17/miR-375/PAX6 axis regulated the proliferation, migration, and invasion of OSCC cells. A–C. The expression levels of SNHG17, miR-375, and PAX6 in CAL-27 cells were detected by qRT-PCR, 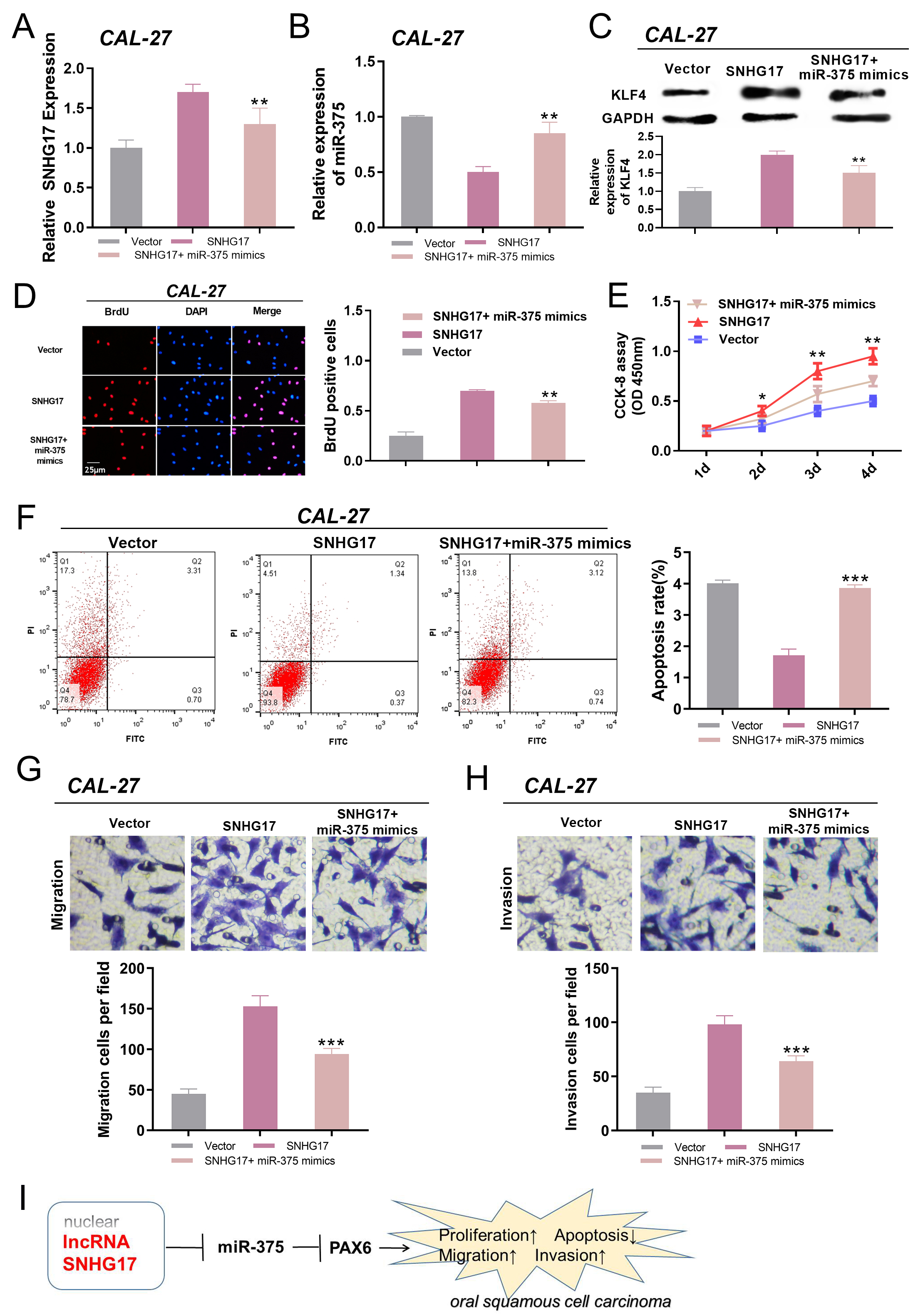
To investigate the effect of SNHG17 on OSCC cells, we established models of over-expression and low-expression of SNHG17 using CAL-27 and Tca8113 cell lines, respectively (
SNHG17 targeted miR-375
Accumulating studies show that lncRNAs play as competing endogenous RNAs (ceRNAs) in targeting the expression of miRNAs. To explore the mechanism of SNHG17 in regulating OSCC cells, we predicted its downstream target through StarBase database, and found that miR-375 was a potential downstream target of SNHG17 (Fig. 3A). Dual-luciferase reporter assay showed that the luciferase activity of the luciferase reporter containing SNHG17-wt was significantly inhibited by miR-375 mimics (
MiR-375 inhibited the proliferation, migration, and invasion of OSCC cells
To investigate the functions of miR-375 on OSCC cells, we used CAL-27 and Tca8113 cell lines to establish the overexpression and inhibition models of miR-375 respectively (Fig. 4A). CCK-8 assay and BrdU assay showed that transfection of miR-375 mimics inhibited the proliferation of OSCC cells compared with NC group (
PAX6 expression was negatively regulated by miR-375, but positively regulated by SNHG17
PAX6 was reported to be a downstream target of miR-375 [19]. To test whether PAX6 was targeted by miR-375, TargetScan 7.2 and StarBase were used to predict the potential targeted binding sites. The 3’-UTR of PAX6 contained a site complementary to the seed region of miR-375 (Fig. 5A). The luciferase reporter assay revealed that the miR-375 mimic had no significant effect on the luciferase activity of PAX6-mut. In contrast, miR-375 mimic significantly decreased the luciferase activity of PAX6-wt (
SNHG17/miR-375/PAX6 axis regulated the proliferation, migration, and invasion of OSCC cells
From above analysis, we could conclude that SNHG17/miR-375/PAX6 axis might play a role in OSCC progression. To confirm this, we transfected miR-375 mimics into CAL27 cells with SNHG17 overexpression. The expressions of SNHG17, miR-375, and PAX6 were detected by qRT-PCR. The results showed that overexpression of SNHG17 up-regulated the expression of PAX6 (
Discussion
In recent years, previous studies prove that the imbalance of lncRNAs figures prominently in tumorigenesis and cancer progression. These abnormally expressed lncRNAs are found to regulate the proliferation, apoptosis, chemotherapy resistance, radiotherapy resistance and so on [5, 21]. LncRNA also plays a significant regulatory role in the progression of OSCC [22]. For example, the expression of lncRNA ANRIL is up-regulated in OSCC tissues and cells, and it is able to promote the proliferation and suppress the apoptosis of OSCC cells by regulating TGF-
Recently, the abnormal expression of miRNAs in tumors has attracted wide attention, and they are involved in many biological processes through gene regulation [26]. For instance, miR-375, acting as tumor repressors, can significantly promote the expression of E-cadherin and inhibit the expression of N-cadherin in cervical cancer, so as to reduce the invasive ability of SiHa cells [27]. Through regulating the expressions of SEC23A and YAP1, miR-375 can inhibit the apoptosis of prostate cancer cells, thereby inducing the chemoresistance of docetaxel [28]. In this work, we demonstrated that miR-375 expression was abnormally down-regulated in OSCC, and it suppressed the malignant phenotypes of OSCC cells, which further confirmed the previous research results [29, 30]. So, we intended to further investigate its molecular mechanisms.
Accumulating evidence found that the regulatory network of lncRNA-microRNA-mRNA (mechanism of “ceRNA”) plays a significant role in cancer progression [31, 32]. For example, lncRNA RBM5-AS1 regulates the expression level of miR-1285-3p as a ceRNA, and eventually promotes OSCC progression by positively regulating YAP1 expression [33]. Also lncRNA OIP5-AS1 can boost OSCC progression by regulating miR-338-3p/NRP1 axis [34]. Inspired by this, we predicted the downstream miRNAs of SNHG17 through bioinformatics analysis, and confirmed that miR-375 was one of its potential targets by luciferase reporter assays. Subsequently, we validated that miR-375 expression was negatively regulated by SNHG17, and miR-375 mimics could significantly reverse the promotion of malignant phenotype of OSCC cells induced by SNHG17. According to these findings and results, we proposed that SNHG17 might function as a ceRNA for miR-375 and promote the proliferation, migration, and invasion of OSCC cells.
Bioinformatics and luciferase reporter assay indicated that miR-375 could bind to the 3’UTR of PAX6. PAX6 is a transcription factor encoded by a gene located on chromosome 11p13 in humans, and plays an important role in central nervous system, the development of human eyes and embryogenesis [35, 36]. But, it is identified as a cancer-related gene in recent years. For example, overexpression of PAX6 promotes phenotypes of cancer stem cells in tumors such as lung cancer and glioma [37, 38]. In oral tumors such as tongue squamous cell carcinoma, PAX6 expression is regulated by miR-140-5p and involved in the metastasis of TSCC cells [39]. In addition, studies authenticate that the downregulation of PAX6 expression inhibits the proliferation and invasion of non-small cell lung cancer cells [40]. Up-regulation of PAX6 expression can promote epithelial-mesenchymal transition and metastasis of non-small cell lung cancer cells via activating PI3K/Akt signaling pathway, and enhance the resistance of cancer cells to cisplatin [41]. In OSCC, evidence from genomic and epigenetic data unearth that abnormal PAX6 expression is associated with shorter survival time of OSCC patients [42]. However, the function of PAX6 in OSCC and its upstream regulatory mechanism have not been reported. The current study for the first time explored the regulatory mechanism of PAX6 in OSCC. We demonstrated that the expression of PAX6 in OSCC tissues was upregulated and negatively correlated with the expression of miR-375. Further experiments showed that overexpression of SNHG17 or inhibition of miR-375 could significantly promote the expression of PAX6, while knockdown of SNHG17 or up-regulation of miR-375 expression could inhibit it. Our data explained the SNHG17 might regulate the proliferation, migration, invasion and apoptosis of OSCC cells through miR-375/PAX6 axis, and promote the molecular regulatory mechanism of OSCC. However, further experiments are needed to confirm the oncogenic effect of PAX6 in OSCC.
There are some limitations in this study. First of all, additional clinical data are needed to investigate the clinical and prognostic significance of SNHG17 expression in OSCC patients. Secondly, although two OSCC cell lines were used to repeat the functional study in this study, the functional role of SNHG17 in OSCC needs to be further verified in other OSCC cell lines and animals so that the function of SNHG17 in OSCC will be comprehensively verified.
In summary, the results of this study suggest that SNHG17 has an oncogenic function in OSCC and is significantly associated with poor pathological characteristics of OSCC patients. Functionally, we confirm that SNHG17 promotes OSCC cell proliferation, migration, and invasion through the miR-375/PAX6 axis. This study extends the understanding of the progression of OSCC and offers a new theoretical basis for the diagnosis and treatment of OSCC.
Footnotes
Acknowledgments
This study was supported by funding from Natural Science Foundation of Jiangxi province (No. 20192BAB205055).
Conflict of interest
The authors declare that they have no competing interests.
