Abstract
BACKGROUND:
Acute myeloid leukemia (AML) is characterized by heterogeneity in phenotypic, genotypic, and clinical traits. miRNAs play an important role in pathogenesis and diagnosis of adult AML. Such information is not available about miRNA expression role in pediatric AML.
OBJECTIVE:
We aimed to investigate the expression of miR-370 and miR-375 as new diagnostic biomarkers to discriminate pediatric AML patients and to predict their roles in the disease molecular basis.
METHODS:
The expression of both miR-370 and miR-375 in peripheral blood (PB) of pediatric AML patients was assessed by QPCR; their impact for diagnosis was evaluated by ROC curve and their roles in pediatric AML development were predicted by bioinformatics analysis.
RESULTS:
The expression of miR-370 and miR-375 levels was significantly decreased in pediatric AML patients, suggesting them as tumor suppressor miRNAs as supported by bioinformatics analysis. miR-370 showed better potential and sensitivity toscreen pediatric AML patients and more significant correlation with AML risk than miR-375. This is the first study to report the positive correlation between both miR-370 and miR-375.
CONCLUSION:
miR-370 level in peripheral blood can serve as a potential non-invasive diagnostic biomarker and was significantly correlated with AML risk. We strongly recommend PB miRNAs as diagnostic biomarkers for pediatric AML.
Introduction
Acute myeloid leukemia (AML) is a hematological malignancy of clonal stem cell or progenitors, characterized by abnormal proliferation and lack of differentiation of immature myeloid cells [1, 2]. This abnormal growth is due to numerous genetic mutations and epigenetic dysregulation [3]. AML makes up to
MicroRNAs (miRNAs) are a family of biological molecules including single stranded non-protein coding RNAs
miR-370 is located within the imprinted Delta-like homologue 1- type 3 deiodinase region (DLK1-DIO3) on human chromosome 14q32. The DLK1-DIO3 region was considered a cancer-related genomic region, suggesting the implication of miR-370 in carcinogenesis [15]. miR-370 was reported to be upregulated and to have an oncogenic role for a number of human malignancies, including prostate cancer [16], gastric carcinoma [17], AML [18] and Wilms tumor [19]. However, it was also reported to be downregulated and to have a tumor suppressor role in bladder cancer and cholangiocarcinoma cells [20, 21].
miR-375 is found between the Crystallin Beta A2 (CRYBA2) and Coiled-coil domain containing 108 (CCDC108) genes on human chromosome 2q35 [22]. miR-375 had a dual role in different types of human cancers [23, 24]. The expression of miR-375 may play an important role either as tumor suppressor or oncogenic miRNA [25, 26]. miR-375 acts as a tumor suppressor in gastric cancer [27], colorectal cancer [28], and pancreatic cancer [29]. It may have an inhibitory role during the tumor development [30]. Also, miR-375 promotes migration and invasion in lung cancer cells [25], glioma [30], and AML [11].
Because of the need for a non-invasive biochemical marker to diagnose pediatric AML and due to the presence of fluctuating data about the expression levels of miR-370 and miR-375 in different cancer types, this study aimed to investigate the expression patterns and the diagnostic relevance of miR-370 and miR-375 in newly diagnosed Egyptian pediatric AML patients.
Material and methods
Experimental subjects
The present study was conducted at Children’s Cancer Hospital, Egypt (CCHE-57357). A written informed consent was obtained from the legal guardians of all the patients and controls, after the approval of the Institutional Review Board (CCHE-IRB), according to national law and regulations. The study included 72 patients aged less than 18 years (47 boys and 25 girls; median age, 9 years; range, 0.5–17.9 years) with the diagnosis of de novo AML presented to the CCHE during the period 2013–2017. Patients with acute promyelocytic leukemia, AML with Down syndrome and those with acute myeloid leukemia arising from myelodysplasia and therapy-related myeloid neoplasms were excluded from present study. All specimens were collected from the BioBank, made anonymous according to ethical and legal rules of the hospital. A number of 15 age-sex-matched control group (7 boys and 8 girls; median age, 6 years and 7 months; range, 3–14 years) was randomly selected from healthy bone marrow transplantation donors.
The diagnosis of AML was performed by morphological, cytochemical, immunophenotyping, cytogenetic and molecular testing. The patients were stratified according to the cytogenetics and molecular markers into
All the patients received same chemotherapy regimen adopted from the COG-AAML1031 protocol. According to patient’s response post induction I, they were subsequently divided into 2 groups: patients with known high-risk disease features and those with evidence of residual AML (MRD
Detection of microRNA expression by RT-QPCR
RNA extraction and cDNA synthesis
Total RNA was extracted from 2 mL PB samples on EDTA using a Direct-zol
Real time quantitative PCR
Detection of miRNA-370 and miR-375 expression was performed by real-time quantitative PCR. Each 15-
Statistical analysis
Descriptive statistics for the patients’ demographics was represented as frequencies and percentages for categorical variables. Quantitative data were statistically represented in terms of median (IQR). Comparison between different groups in the present study was performed using Mann-Whitney Test for comparing two non-parametric groups. Kruskall-Wallis Test was used when comparing more than two non-parametric groups. Comparison between different groups in the present study was performed using Chi-Square Test. Correlation between various variables was done using Spearman rank correlation coefficient (R). Receiver Operating Characteristic (ROC) Curve was used to generate the Cut-off value, area under the curve (AUC), sensitivity, specificity for miRNAs. Odd ratios (ORs) and 95% confidence intervals (CIs) were calculated by logistic regression to assess the relative risk conferred by our markers. All statistical analyses were performed using statistical software SPSS (Statistical Package for Social Science) statistical program version 21.0. Graphs were built using SPSS statistical program version [21.0] and Microsoft Excel program version 2016.
Relative expression of a. miR-370 and b. miR-375 in pediatric AML patients vs controls, 
To predict the possible role of miR-370-5p, miR-370-3p and miR-375 in the AML development, bioinformatics target prediction and functional annotation enrichment analysis were performed. First, microRNA target prediction analysis was done using miRWalk 2.0 server (mirwalk.uni-hd.de/). MiRWalk is a comprehensive database from which both predicted and experimentally validated miRNA-targets could be obtained. In our analysis, we selected the target prediction algorithms: miRWalk, Targetscan v7.0, miRanda, and RNA22 to obtain the common predicted targets between at least two target prediction algorithms with cut-off
The functional enrichment analysis for the predicted and validated targets was performed using Enrichr server, (comprehensive gene set enrichment analysis web server; amp.pharm.mssm.edu/Enrichr/). The most essential enriched pathways (KEGG pathways), which were extracted from Enrichr with
Results
Expression level of miR-370 and miR375 in pediatric AML patients
We analysed the expression level of miRNA-370 and miRNA-375 from 72 pediatric AML patients at the time of diagnosis using QRT-PCR. miR-370 was significantly downregulated in patients compared to normal controls (patients
Correlations of miR-370 and miR-375 with AML clinical parameters
Correlation analysis was performed to assess the association between miR-370 and miR-375 expression levels and different laboratoryand clinical parameters (Table 1). There was a significant correlation between miR-370 level and the TLC of the patients (
Correlation of miR-370 and miR-375 level with clinical characteristics of pediatric myeloid leukemia patients
Correlation of miR-370 and miR-375 level with clinical characteristics of pediatric myeloid leukemia patients
TLC: Total leukocyte count, Hb: Hemoglobin, FAB: French-American-British.
ROC-Curve for both miR-370 and miR-375 of pediatric AML patients
AUC: Area under the ROC curve. CI: Confidence interval of the estimated crude odds ratio.
To check the feasibility of miR-370 and miR-375 as diagnostic markers, ROC curve analysis was performed. The ROC curve showed AUC of 0.966 for miR-370, and 0.858 for miR-375 (Table 2). The miR-370 level at 4.191 and miR-375 level at 9.284 were the clear cut-off value to screen pediatric AML patients from controls. Based on this cut-off value, the sensitivity and specificity of miR-370 for distinguishing AML was 96.9% and 93.3%, respectively (Fig 2a) and the values for miR-375 were 75.4% and 80.00%, respectively (Fig. 2b). In addition, at specificity 100%, miR-370 at cut-off 7.398 showed sensitivity of 72.3%, while miR-375 redundant showed 50% sensitivity at cut off 10.324.
ROC curve analysis of a. miR-370 and b. miR-375 expression in pediatric AML patients.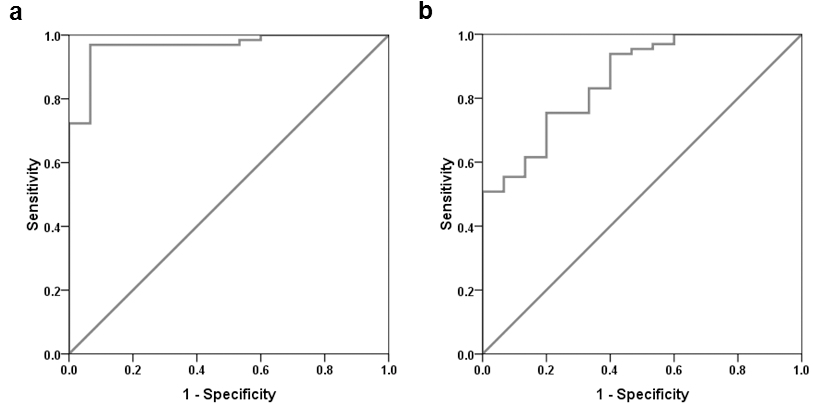
To assess the relative risk conferred by miR-370 and miR-375 for pediatric AML, logistic regression analysis model was performed (Table 3). It revealed that miR-370 is significantly associated with increased risk for AML (OR
Logistic Regression Results for both miR-370 and miR-375 of pediatric myeloid leukemia patients
Logistic Regression Results for both miR-370 and miR-375 of pediatric myeloid leukemia patients
OR: Odds ratio; CI: Confidence interval.
The predicted genes and pathways targeted by miRNAs in AML. a. Overlapped target genes of all miRNAs. b. The enriched pathways of the miRNAs’ targets, 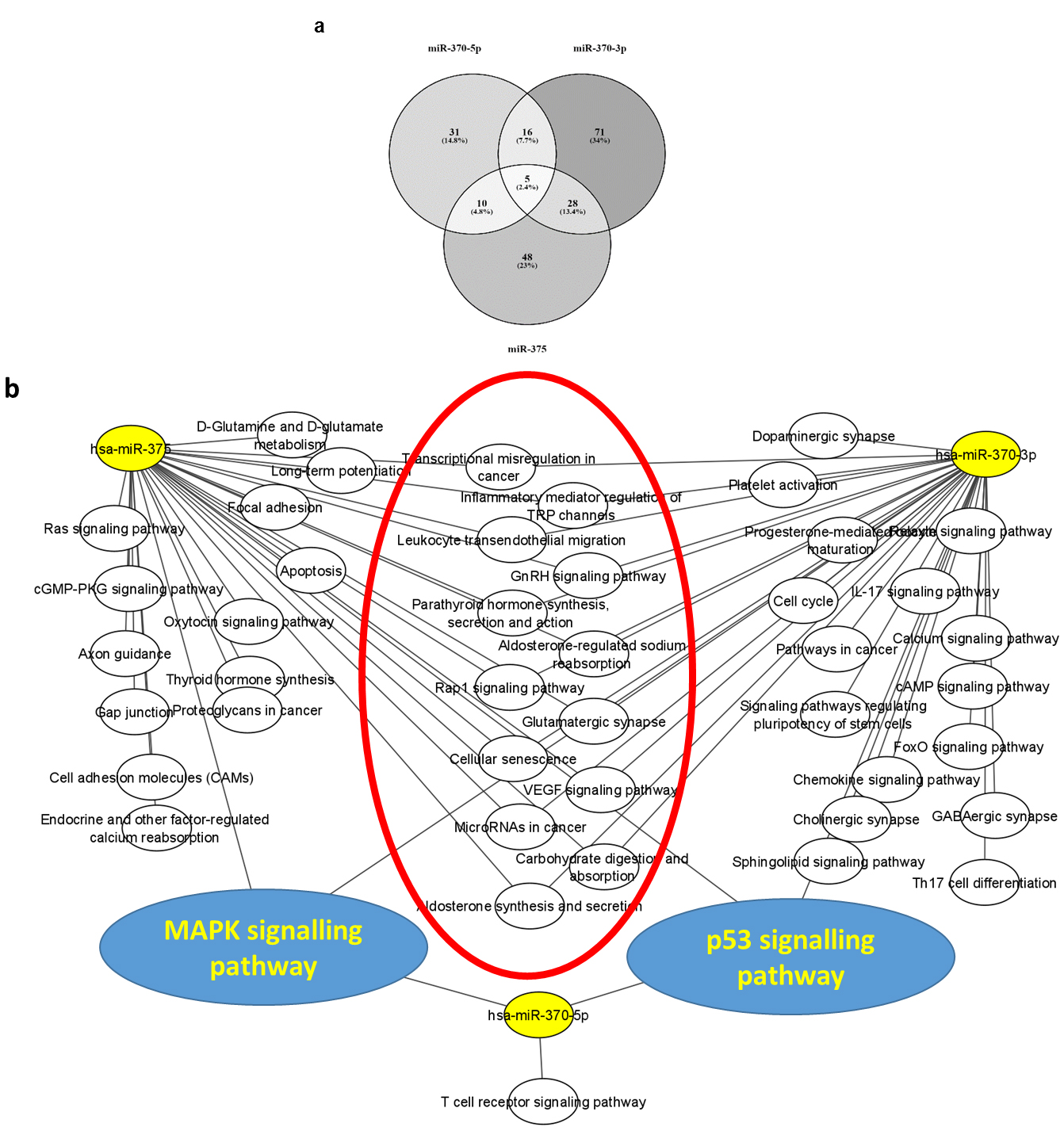
The previous data suggested that downregulation of miR-370 and miR-375 expression may have an important role in AML development, which drove us to predict their possible functionally enriched pathways in AML (Fig. 3 and Supplementary Table 1). After overlapping the miR-370-3p, miR-370-5p and miR-375 miRwalk targets to the differentially upregulated genes in AML (adj
By examining the functional enrichment analysis results of our miRNAs’ targets upregulated in AML, 45 KEGG signalling pathways are significantly enriched (
Discussion
Cancer is one of the most important public health problems in the world [32], and it remains one of the major health concerns of the 21
In the present study, a significant decrement in miR-370 level was detected and this was in agreement with previous studies [33, 35]. Lin et al. [33] highlighted that the expression levels of miR-370 in serum and bone marrow were both decreased in pediatric AML patients, and confirmed that low expression plays an important role in both diagnosis and prognosis. Saied et al. [34] reported that the expression of miR-370 in AML peripheral blood mononuclear cells (PBMCs) was decreased and that might point to its potential role as a tumor suppressor miRNA Also, Zhang et al. [35] reported the downregulation of miR-370 in both leukemic cell lines and primary leukemic cells from patients with de novo AML, and confirmed that miR-370 could slow down the cell cycle progression of AML cells.
We have also found that miR-370 showed significant correlation with TLC
The ROC curve proposed miR-370 as a significant diagnostic biomarker for pediatric AML (AUC
In the current study, a significant decrement in miR-375 level was shown and that was in agreement with a previous study [24], which reported that the expression of miR-375 significantly decreased in leukemic cell lines and primary AML blasts. Interestingly, Zhi et al. [11] found that miR-375 in serum was one of the upregulated miRNAs in pediatric AML. Also, Wang et al. [36] showed that the upregulated miR-375 in bone marrow of pediatric AML played a role as a prognostic marker and could be a factor for AML development. We assume that the sample type may be a reason for these expression differences.
We also found that miR-375 showed no significant correlation with TLC, and that was in agreement with what has been reported before [36]. In addition, in our study there was no correlation between FAB subtypes, cytogenetic risks groups, response end of induction I and PB miR-375 levels. Though, miR-375 has been previously shown to be significantly associated with both FAB subtypes M7 and the unfavorable cytogenetic risks [36].
The ROC curve showed thatthe value of detection of single marker for identifying patients was significantly high for miR-375. However, miR-375 has no significant association with AML risk at pediatrics as shown by the logistic regression analysis. This was in contrast with previous results, which proved that miR-375 expression can also predict poor clinical outcome as a prognostic indicator in AML [24].
To the best of our knowledge, this is the first study to present a significant correlation between the expression levels of both miR-370 and miR-375 in pediatric AML patients (R
The primer assays used in this study assess the levels of the pre-miRNA, therefore to predict the tumor suppressor roles of these downregulated miRNAs, we performed the target and functional enrichment analysis on miR-370-3p, miR-370-5p, and miR-375. The three miRNAs target 5 common genes; PRKX, ASPH, ZMAT3, SPATA6 and PLXNA4. Four of these target genes are shown by the results of previous studies to be associated with increased cell proliferation, invasiveness, poor prognosis and age & sex related signalling pathways in AML [31, 37, 38]. Moreover, the enrichment analysis of the miRNAs targets for KEGG pathways reflected relevance to AML in several aspects such as cell cycle regulation, metabolic, and immune pathways, as well as apoptosis and transcriptional misregulation in cancer [31]. These results are in agreement with the involvement of miR-370 and miR-375 as tumor suppressor miRNAs through induction of aging, and inhibition of the cell growth in AML cells [34, 35], as well as, reduction of the colony number and formation in leukemic cell lines and primary AML blasts [24]. It is noteworthy that p53 and MAPK signalling pathways enriched for the miRNA targets. Both pathways are known to be essential for pediatric AML development and progression [39, 40]. These results suggest that restoring the expression of miR-375 and miR-375 may improve the treatment outcomes in AML pediatric patients through regulation of these pathways. However, we have demonstrated by using the logistic regression analysis the association between the downregulation of only miR-370 and higher risk of pediatric AML, which may correlate with its higher potential for pediatric AML diagnosis. We suggest more experimental and mechanistic validations for these miRNAs to investigate their clinical implications.
Conclusion
miR-370 and miR-375 are significantly downregulated in PB of pediatric AML. This expression profile may point to their potential roles as tumor suppressors in AML pediatric patients; a role that was supported by bioinformatics analysis. More interestingly, despite their significant roles as diagnostic markers we demonstrated higher potential of PB miR-370 for pediatric AML diagnosis. Our findings also suggest for the first time the positive correlation between both miR-370 and miR-375 expressions. Taken together, miR-370 is a promising non-invasive diagnostic and a possible prognostic biomarker for pediatric AML that merits further investigations.
Author contributions
M.M.A carried out the experimental work, analysed the data, and prepared the first draft of the manuscript. R.H.M. performed bioinformatics analysis and provided statistical analysis, data interpretation and writing assistance. S.M. and D.A.Y supervised the experimental work and provided manuscript writing assistance. W.M.E and A.A.S designed and supervised the work. W.M.E critically read and approved the final version of the manuscript. All authors have read, revised and approved the final manuscript.
Supplementary data
The supplementary files are available to download from http://dx.doi.org/10.3233/CBM-210360.
sj-xlsx-1-cbm-10.3233_CBM-210360.xlsx - Supplemental material
Supplemental material, sj-xlsx-1-cbm-10.3233_CBM-210360.xlsx
Footnotes
Conflict of interest
The authors declare no conflict of interest and that this manuscript is not published or submitted elsewhere.
