Abstract
BACKGROUND:
Tumor-infiltrating immune cells are indispensable to the progression and prognosis of clear cell renal cell carcinoma (ccRCC).
OBJECTIVE:
The aim of this study was to explore the clinical implications of immune cell infiltrates in ccRCC.
METHODS:
The Cancer Genome Atlas (TCGA) database (
RESULTS:
Survival analyses revealed that 13 genes significantly associated with the overall survival (OS). Furthermore, multivariate Cox analysis identified an immune risk score on the basis of mast cells, natural killer CD56bright cells, T helper 17 (Th17) cells, and Th2 cells. The immune risk score was associated with OS, with hazard ratios of 2.72 (95% CI 2.17–3.40) and 3.24 (95% CI 1.64–6.44) in TCGA and E-MTAB-1980 datasets, respectively. This immune risk score was significantly correlated with some immunotherapy-related biomarkers.
CONCLUSIONS:
We profiled a prognostic signature and established an immune risk score model for ccRCC, which could provide novel predictive markers for patients with ccRCC and an indicator for immunotherapy response measurement.
Introduction
Renal cell carcinoma (RCC) is a heterogeneous group of cancers with distinct clinicopathologic and genetic features. It ranks among the top 10 lethal forms of tumors in both men and women in the United States [1, 2]. Clear cell renal cell carcinoma (ccRCC) – consisting of 75% of all RCC [3, 4] – is the most common subtype. An in-depth analysis of the biological characteristics and identification of individuals’ survival risk are critical for the management and treatment of patients with ccRCC. Thanks to the revolutionary advances in high-throughput biotechnology, a continuous deepening understanding of ccRCC is having a clear impact on clinical treatment and drug development. For example, the Cancer Genome Atlas (TCGA) project comprehensively provided magnanimous high-throughput sequencing to help cancer researchers systematically know the clinicopathologic features and multiomics molecular characteristics of ccRCC [5]. These public high-throughput sequencing data provide additional prognostic information to analyze and visualize the molecular characteristics of cancers.
In addition to the genetic alterations and gene expression, proteomics analyses have unveiled the molecular mechanisms of ccRCC, the tumor immune microenvironment, which consists of multiple immune cells and is of increasing interest. Recently, the rise of immunotherapy has been shown to be promising avenues for the treatment of ccRCC [6, 7, 8, 9]. Because of the heterogeneity and influence of immune cells in tumor clinical outcomes, it is critical to characterize the interaction between cancer cells and immune cell infiltrates [10, 11]. The immune microenvironment also has remarkable clinical significance in ccRCC. Notably, in the TCGA database, ccRCC has the highest T-cell infiltration score among 19 types of cancers [12]. However, the clinical effects of immune cells in ccRCC as well as their molecular functions have not been fully explored. Therefore, it is imperative to understand the characteristics and survival implications of the immune microenvironment in ccRCC.
Consequently, the aim of the present study was to explore whether or not a risk score based on immune cells could be constructed for survival prediction and immune therapy response evaluation. We characterized the immune infiltration characteristics of patients suffering from kidney renal clear cell carcinoma (KIRC) on the basis of the infiltration levels of 24 immune cells. Then, integration of the genomics and clinicopathologic data of immune cell infiltrates provided novel insights. Importantly, an immune risk score was constructed using univariate and multivariate Cox analyses, which have the potential to act as robust prognostic biomarkers. Furthermore, the survival prediction performance was validated in another independent dataset.
Methods
ccRCC datasets and preprocessing
The RNA-sequencing data (Fragments Per Kilobase Million, FPKM values) for the TCGA KIRC cohorts were achieved by using the TCGAbiolinks package in R software [13]. The data were converted into per kilobase million (TPM) values. A total of 539 tumor and 72 nontumor samples were included. The corresponding clinical data and follow-up information were obtained from the TCGA pan-cancer project [14]. The agilent microarray dataset that included 101 patients with ccRCC was downloaded from the ArrayExpress database (accession ID: E-MTAB-1980) [15].
Immune cell infiltration inference
To quantify immune cell infiltrates from gene expression profiles of bulk tumors, we conducted single-sample gene set enrichment analysis (ssGSEA) implementation in the “GSVA” R package [16]. ssGSEA is a computational algorithm that computes an enrichment score for a gene list of interest. It is a rank-based method that calculates the enrichment score using the ranks of genes by their absolute expression in a sample [17]. The normalized RNA-sequencing and microarray datasets mentioned previously were submitted to ssGSEA. We obtained the marker gene set for 24 types of immune cells from the previous report, including B cells, T cells, T helper cells, T central memory, T effector memory, T helper 1 (Th1) cells, Th17 cells, Th2 cells, T follicular helper cells, TReg, CD8 T cells, gamma delta T cells, cytotoxic cells, natural killer (NK) cells, NK CD56dim cells, NK CD56bright cells, dendritic cells (DCs), activated DCs, immature DCs, plasmacytoid DCs, eosinophils, macrophages, mast cells, and neutrophils [18]. The output for each immune cell is a near-Gaussian list of decimals that can be used for further visualization and statistical analysis.
Clinical significance of immune cell infiltrates
The Mann-Whitney
The expression landscape of immune cells in clear cell renal cell carcinoma (ccRCC) and nontumor tissues. A. The heatmap analysis was performed by R software to observe the expression profiles of immune cell infiltrates in ccRCC and nontumor tissues; B. The box plot shows the difference of immune cells in tumor and nontumor tissues.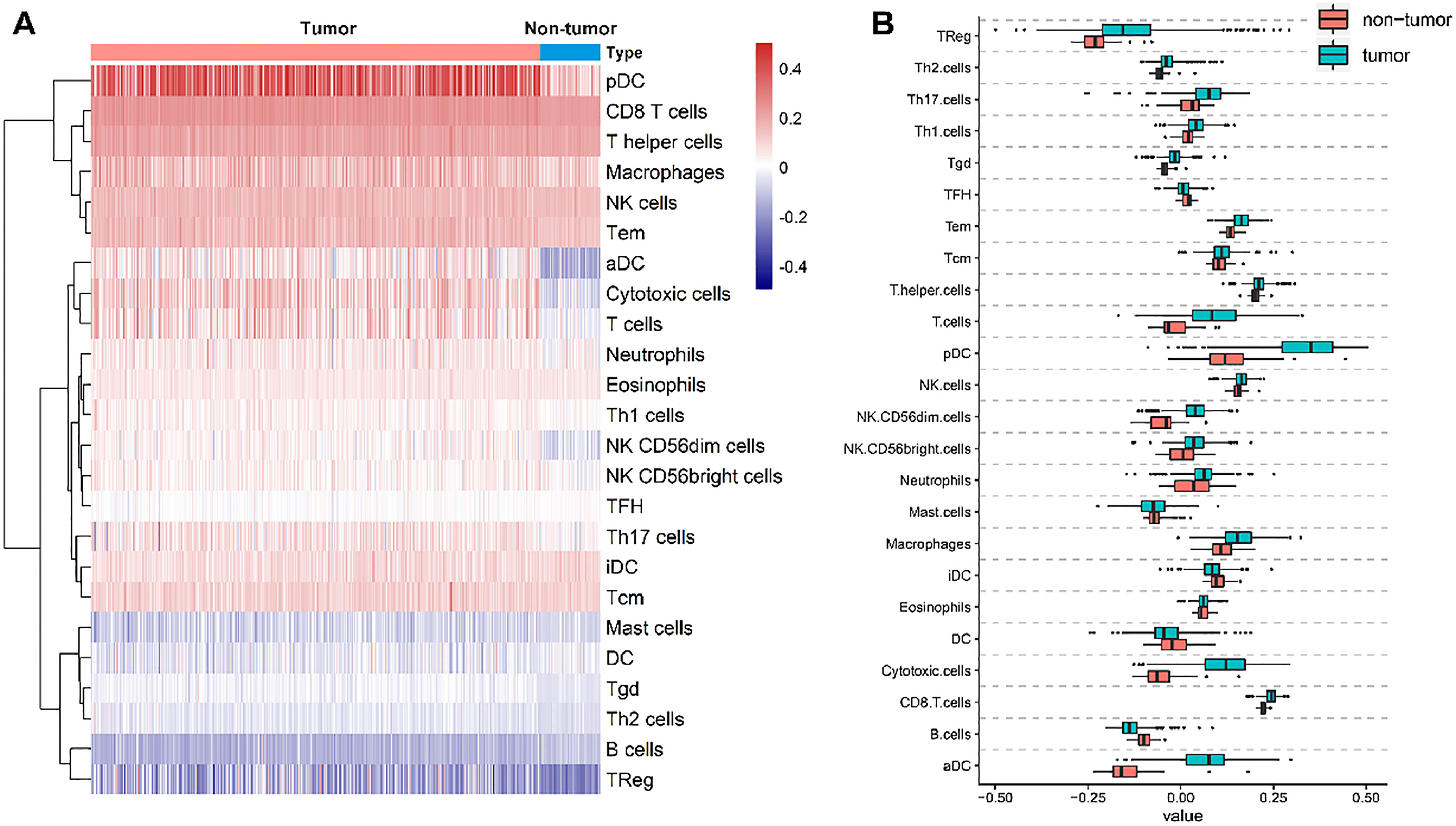
The characteristics of the tumor immune microenvironment and prognostic value of immune cells. A. Most immune cells correlated with each other, and these immune cells were mainly grouped into four types on the basis of the correlation; B. The prognostic value of immune cells. The size of each dot represents the survival impact of each immune cell. Favor for overall survival with 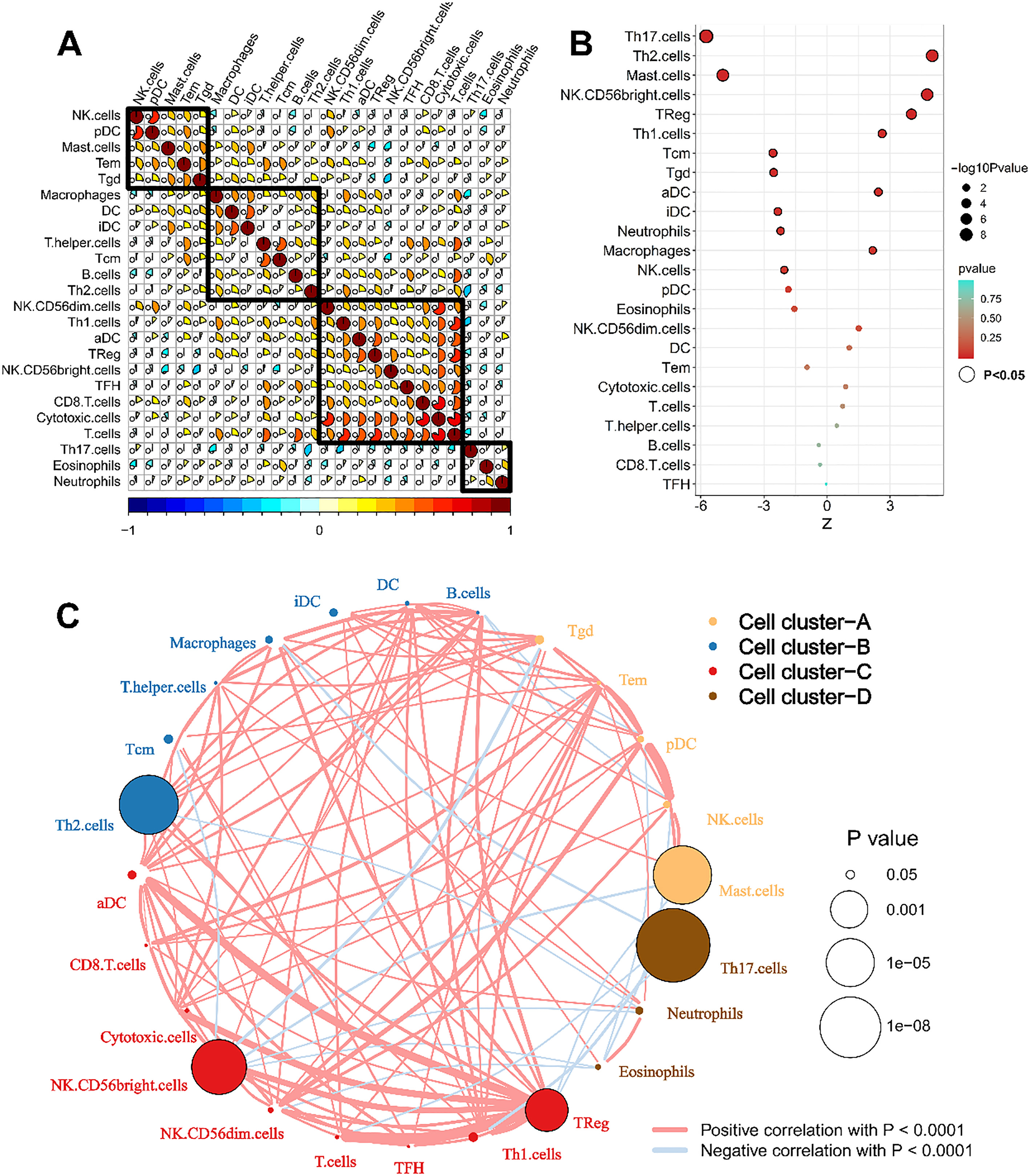
The prediction of perturbed molecular pathways be- tween high and low immune cell infiltration was analyzed using GSEA. GSEA was conducted to recognize the perturbations of different gene sets, which represent functionalities. Here, the collective gene ontology terms and Kyoto Encyclopedia of Genes and Genomes pathways were selected as gene sets for analysis. The gene sets were downloaded from the MSigDB database of Broad Institute. A GSEA algorithm with 1000 permutations was carried out.
Development and validation of immune risk score
Then, 13 survival-associated immune cells in univariate analysis were submitted to multivariate Cox analysis. Finally, four immune cells that showed significance in the multivariate cox analysis were used to develop the immune score. An immune risk score was developed using the linear combination of immune cells weighted by the regression coefficients. On the basis of the results of multivariate Cox analysis, the immune risk score for each patient was calculated. Receiver operating characteristic (ROC) curve was drawn using the “survivalROC” package in R software to assess the survival prediction value of the immune risk score for 1, 3, and 5 years. Subgroup analyses were performed to estimate the prognostic significance of the immune risk score in different subtypes of cancers.
Clinicopathological characteristics of ccRCC patients included in the study
Clinicopathological characteristics of ccRCC patients included in the study
Unsupervised consensus clustering method identifies patients suffering from clear cell renal cell carcinoma (ccRCC) with distinct immune cell infiltrates. A. Heatmap of the consensus clustering matrix for two groups. Patients could be separated into two groups with distinct immune cell profiles; B. The Kaplan-Meier overall survival (OS) curves for the 515 patients with ccRCC stratified by cluster; C. Upregulated and downregulated Kyoto Encyclopedia of Genes and Genomes (KEGG) pathways when C2 subtype compared with C1 subtype; D. Upregulated and downregulated gene ontology (GO) terms when C2 subtype compared with C1 subtype.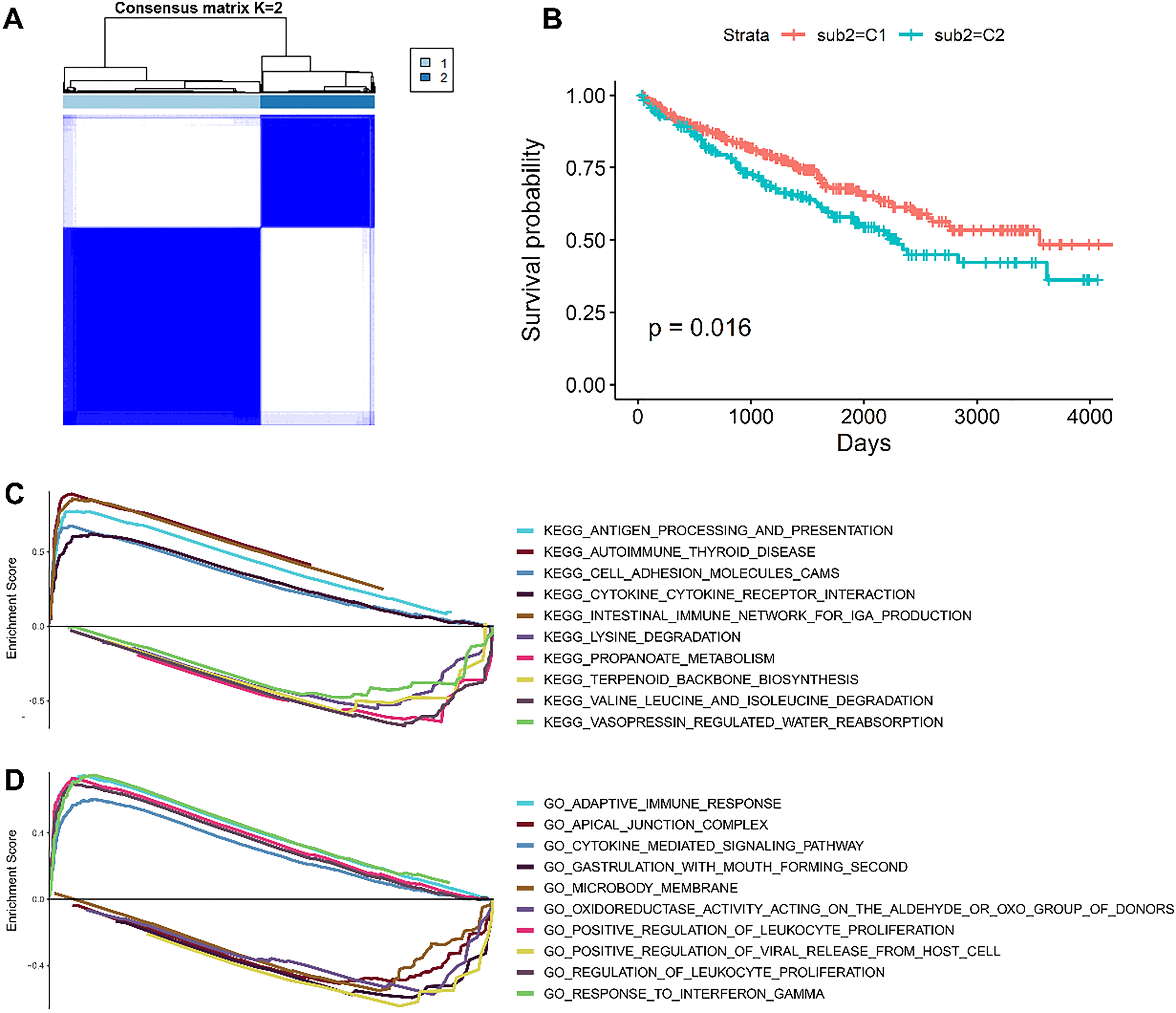
The heatmap of immune cell infiltrates and progression-related clinical parameters.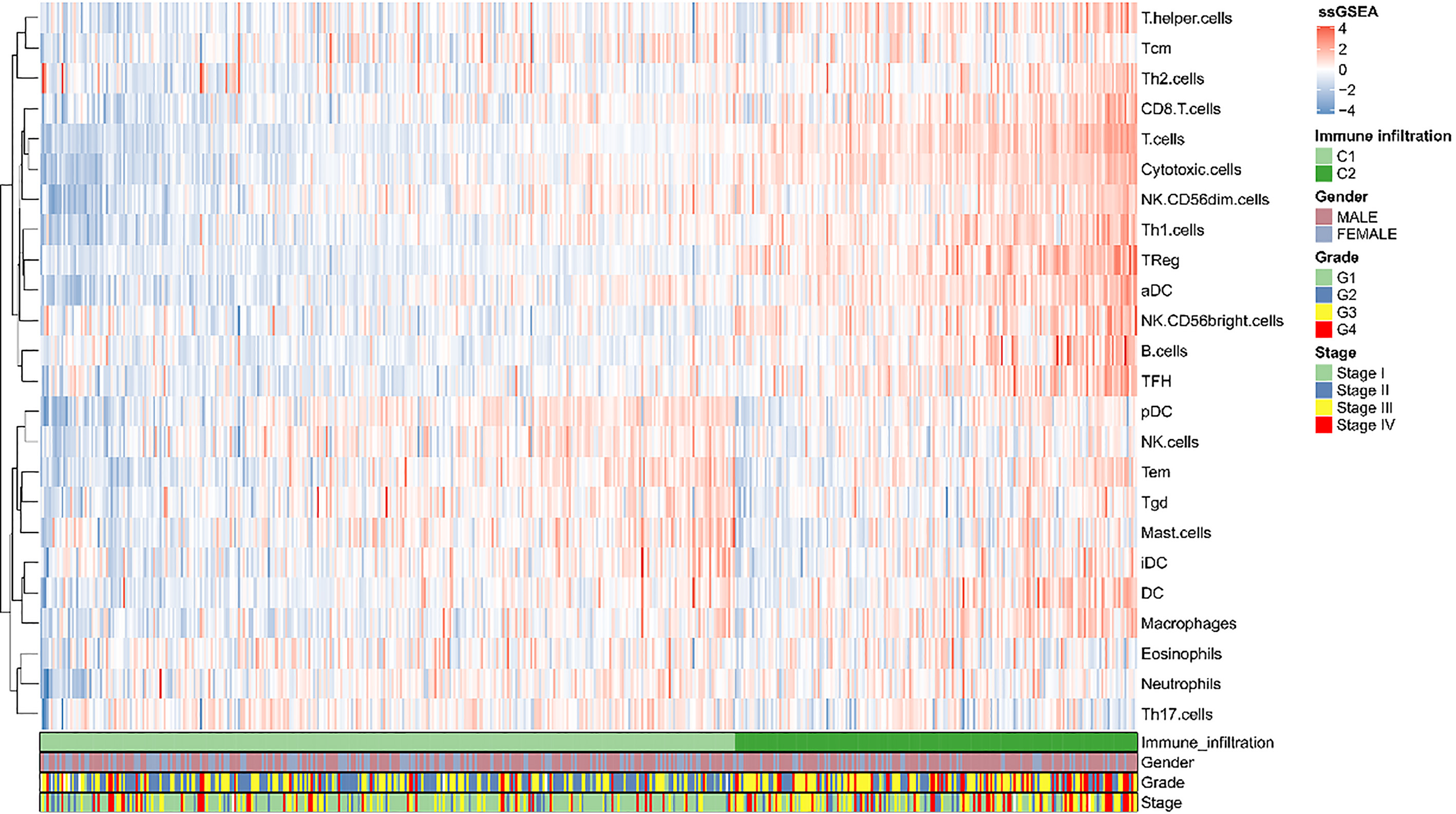
The distribution of the immune risk score, survival status along with survival times of patients with clear cell renal cell carcinoma (ccRCC), and heatmaps of four included immune cells.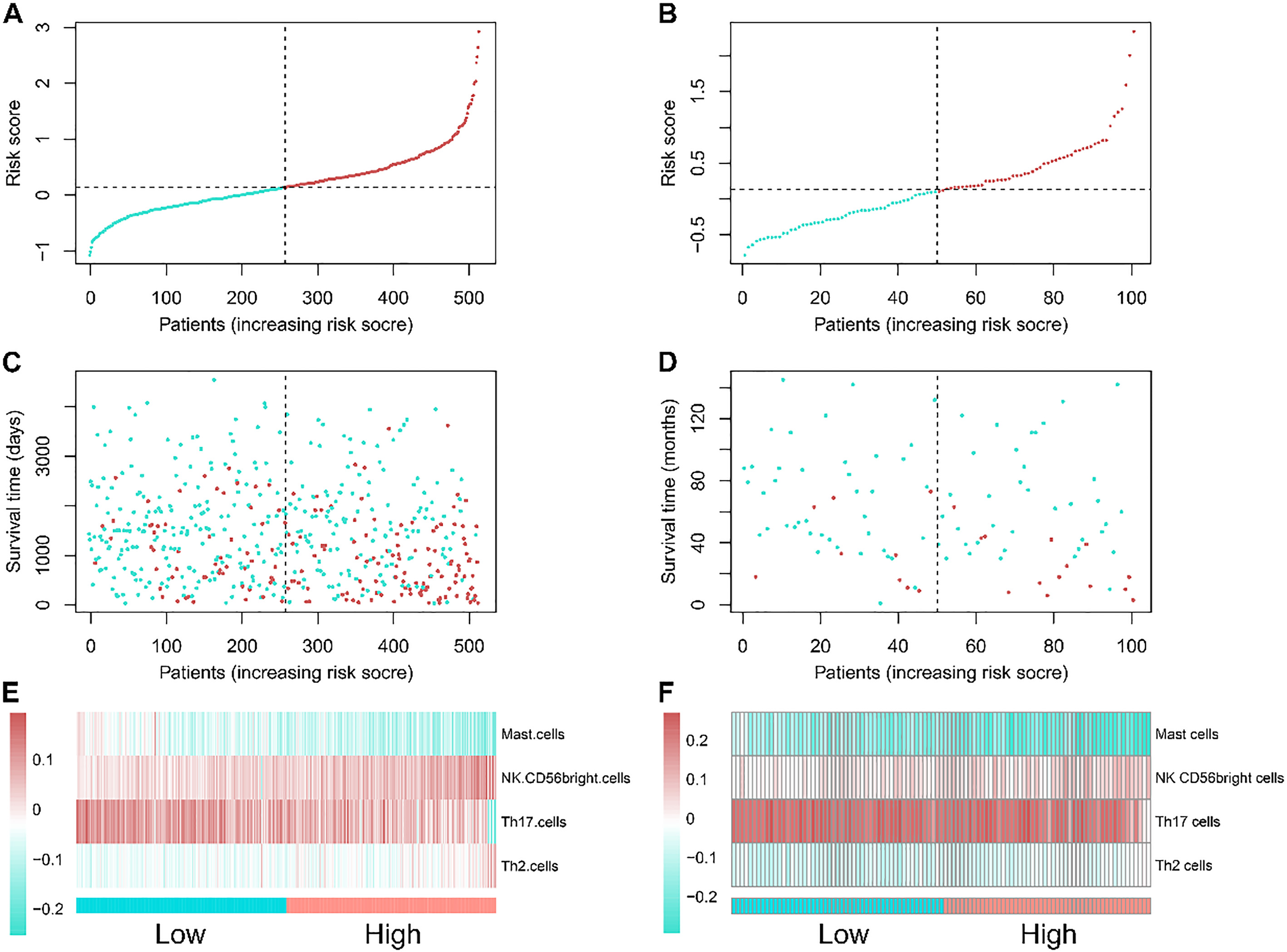
The survival prediction performance of the immune risk score. Receiver operating characteristic (ROC) analysis of sensitivity and specificity of the immune risk score in predicting the overall survival (OS) of patients with clear cell renal cell carcinoma (ccRCC) in the training cohort (A) and validation cohort (C). The hazard ratios of the immune risk score in different subgroups in the training cohort (B) and validation cohort (D).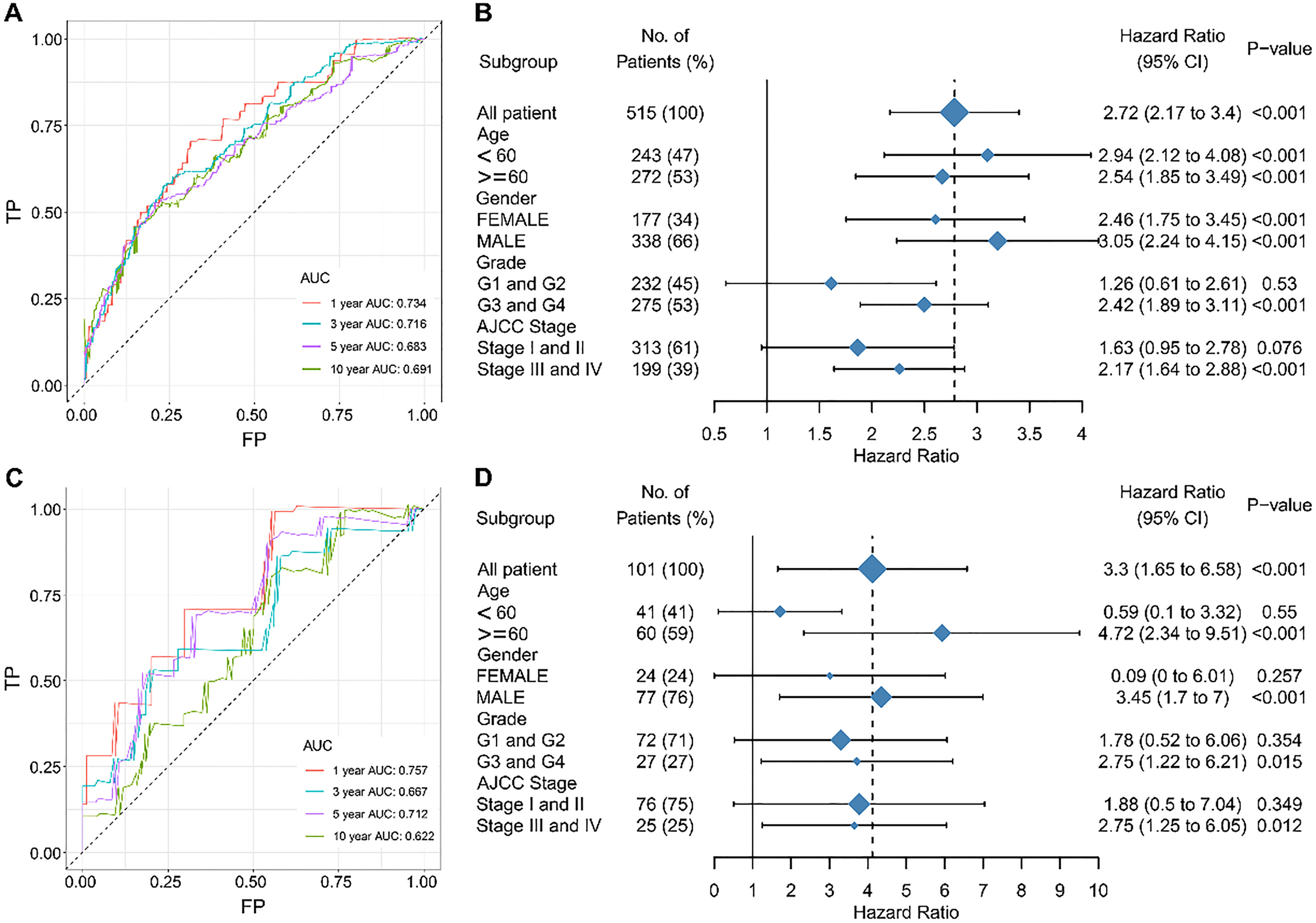
The relationships between the immune risk score and known immune checkpoint genes. (A) The associations among immune risk score, programmed cell death 1 (PD-1), programmed cell death ligand 1 (PD-L1), and cytotoxic T-lymphocyte-associated protein 4 (CTLA-4). The immune risk score is significantly correlated with the expression of PD-1 (B) and CTLA-4 (D). However, no significance of correlation is observed between the immune risk score and PD-L1 (C).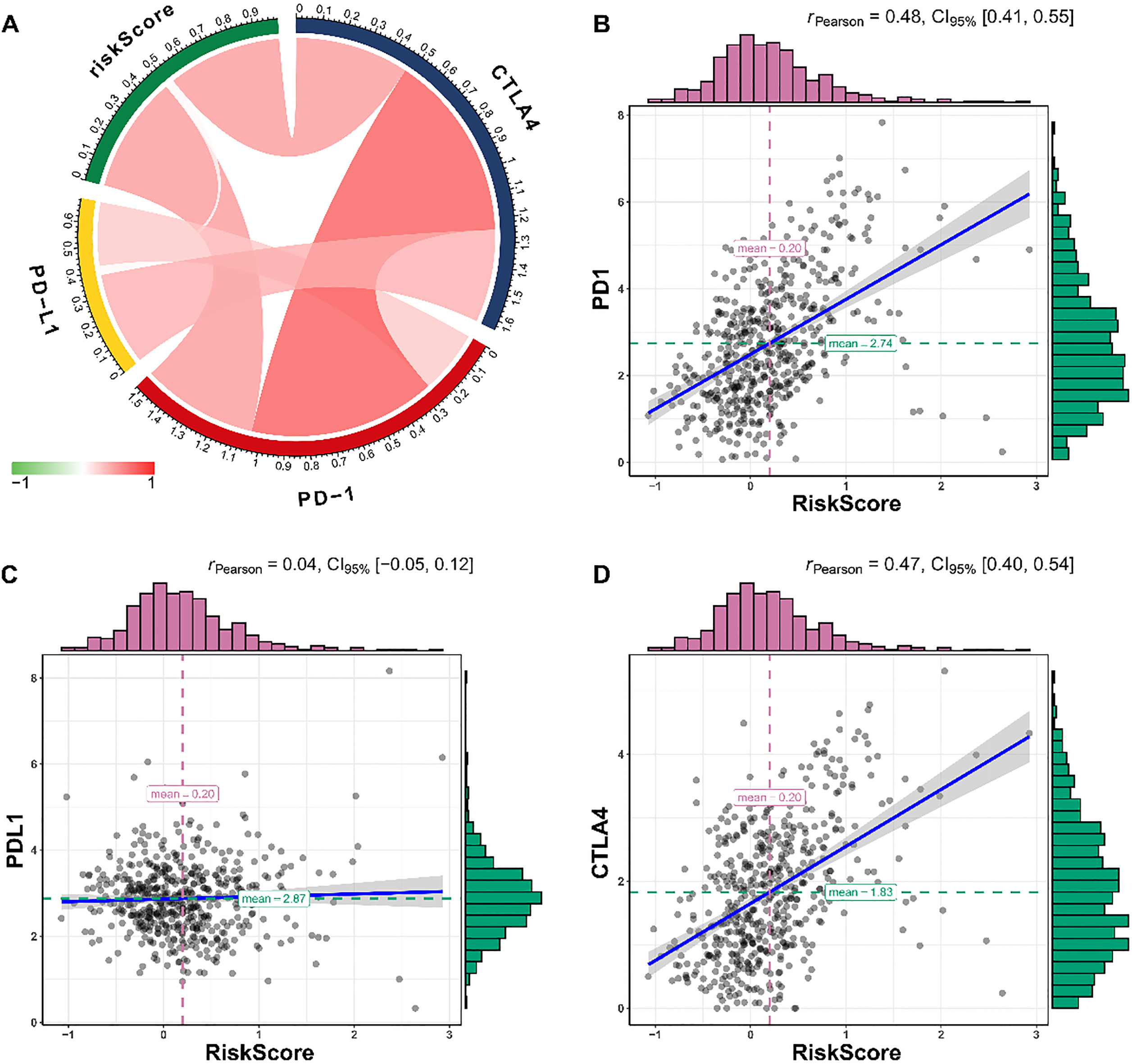
Immune cell infiltration in ccRCC
Using ssGSEA, we identified 24 types of immune cells in 539 tumor and 72 nontumor samples (Fig. 1A). Most immune cells showed differential expression between ccRCC and nontumor samples (Fig. 1B). After removing patients with ccRCC who had a follow-up time of less than 30 days, a total of 515 and 101 patients from TCGA and E-MTAB-1980 datasets, respectively, were used (Table 1). According to the cophenetic correlation among these immune cells, we found that these genes are mainly grouped into four clusters (Fig. 2A). To identify key immune cells that could explain the heterogeneous immune profiles in distinct clinical outcomes, univariate Cox analysis was performed. The univariate Cox regression analysis indicated that 13 immune cell infiltration levels were correlated to the OS of patients with ccRCC (Fig. 2B). Moreover, the immune cell interaction network displays a comprehensive landscape of cell clusters and their effects on the OS of patients with ccRCC (Fig. 2C).
Immune cell infiltrates’ status for patients with ccRCC
Using the infiltration profile of immune cells, consensus clustering of 515 patients with ccRCC found that high- and low-infiltration patients were distinguished significantly (Fig. 3A). The survival analysis showed that high immune cell infiltration was associated with an improved OS (
Construction of immune risk score in ccRCC
Considering that patients with different immune cell infiltration levels showed distinct clinical outcomes, prognosis-associated immune cells were further submitted to develop the risk signature. A prognostic model consisting of four immune cells (mast, NK CD56bright, Th17, and Th2 cells) was identified to predict OS. The immune risk score formula based on the immune cell level and regression coefficients of the four immune cells was developed as follows: Risk score
Association between the immune risk score and patients’ OS
The four-immune-cell signature was useful in separating patients into different survival statuses. Furthermore, the risk score was closely related to the OS of patients, leveraging the risk score as a continuous variable in both the training and validation cohorts (training cohort: Hazard ration (HR)
The area under the curve (AUC) values were used to estimate the performance of the four-immune-cell signature in predicting survival. In the training cohort, the AUC values were 0.734, 0.716, 0.683, and 0.691 in 1, 3, 5, and 10 years, respectively. These results suggested that the immune risk score has high sensitivity and specificity and has excessive potential to become a survival prediction indicator in clinical applications. More importantly, in the test cohort, the AUC values were 0.757, 0.667, 0.712, and 0.622 in 1, 3, 5, and 10 years, respectively. These results further validated that the novel prognostic signature could effectively determine the prognosis of ccRCC. Although the results of subgroup analysis were heterogeneous, the immune risk score acted as an effective prognostic biomarker in the training and validating cohorts (Fig. 6).
Association of immune risk score with immunotherapy-related signature
Immunotherapy is a revolutionary approach for cancer therapy, especially the immune checkpoint therapy. Programmed cell death 1 (PD-1), programmed cell death ligand 1 (PD-L1), and cytotoxic T-lymphocyte-associated protein 4 (CTLA-4) are the most effective targets. Pearson’s correlation analysis revealed that the immune risk score was significantly correlated with the PD-1 (Pearson’s correlation coefficient
Discussion
Recent studies have advanced our interpretation of the deep molecular machinery and characteristics of KIRC. Given the strong evidential basis for the efficacy of immune checkpoint inhibitors in the treatment of KIRC, we explored the immune landscape of KIRC and developed a useful immune-cell-based classifier. The use of the prognostic signature served as significant survival prediction performance.
The heterogeneity of the immune microenvironment actively reflects the prognosis and treatment response of patients. Previous studies have proposed some prognostic signatures on the basis of the tumor immune microenvironment [20, 21, 22]. Notably, Ghatalia et al. comprehensively analyzed the immune microenvironment and found that the postoperative recurrence of ccRCC was significantly correlated with lower T-cell infiltrate, lower adaptive immune response, and higher neutrophil gene expression [23]. These findings provided an ideal immune surveillance method postnephrectomy. However, previous study mainly focused on the clinical significance of particular types of immune cell. Here, we integrated four types of immune cell to achieve maximum value. Furthermore, another independent cohort validated our results. These findings hinted that the immune score could be a useful survival prediction indicator. These findings provided a novel indicator for ccRCC patients’ survival prediction. Previous excellent study compared established molecular classifications of ccRCC with the immune profiles, and found that molecular subgroups have different TME profiles [24]. Based on previous studies, we mainly focused on the prognostic value of immune cells and then taking advantage of their value of clinical application. In the present study, we also found that patients are of great heterogeneity in immune cell infiltration levels. As expected, patients with high immune cell infiltration could benefit from OS. Owing to the significant difference in prognosis between patients with different immune statuses, we attempted to provide a more accurate individualized signature for prognosis surveillance.
Because of the dramatic heterogeneity and plasticity of the tumor immune microenvironment, there still remains an especially critical challenge to depict the immune cell infiltrates and immune cell interaction [11]. The spirit of science urged cancer researchers to develop many computational methods to infer immune cell infiltrates – for example, CIBERSORT, a versatile computational method for quantifying 22 tumor-infiltrating immune cells from bulk tissue gene expression profiles [25]. Li et al. also developed TIMER, a computational approach to estimate the six immune cell infiltrates using immune-specific markers and immune cell expression signatures [26, 27]. Here, we used ssGSEA to quantify the immune cell infiltrates. Currently, ssGSEA is simple and can easily be adjusted. Furthermore, the gene signature enrichment approach is rank based and is, therefore, suitable for cross-platform evaluation.
In this study, we proposed an immune score on the basis of four-immune-cell infiltration, including mast, NK CD56bright, Th17, and Th2 cells. Among these immune cells, the coefficients of NK CD56bright and Th2 cells were positive, indicating a correlation between higher infiltration level and shorter OS, whereas higher levels of mast and Th17 cell infiltrates correlated with longer OS. Ghatalia et al. [23] estimated the relationship between the recurrence of ccRCC and immune cell infiltrates and found a consistent trend of our results using the University of Alabama at Birmingham dataset (
Effective survival monitoring factors could be useful in clinical decision-making and follow-up. Owing to the great heterogeneity of tumors, a combination of multiple molecular factors that could reflect the characteristics of the tumor could improve the accuracy of survival prediction. Similarly, several previous studies have also provided survival prediction indicators on the basis of different types of molecular events [31, 32, 33]. The immune risk score developed based on immune cell infiltrates is an alternative for immune microenvironment evaluation. More importantly, the immune risk score exhibited moderate prognosis prediction performance, and the performance was validated by another independent cohort. In the past decade, immune checkpoint therapy has achieved noteworthy benefits in various types of cancers [34]. These results suggested that the immune risk score could not be used for accurate survival prediction, but it can also have the potential to be a guide for precision cancer immune checkpoint therapy.
Limitations of our study should be noted. First, the retrospective nature of the study limited its application, although we have used different datasets to validate its performance. Second, the great heterogeneity of immune microenvironment is difficult to be assessed accurately. Although we leveraged different types of immune cells to reduce certain bias, some bias may still remain. As several clinical trials are investigating the efficiency of immunotherapy for ccRCC, illustrating the feature of immune cell infiltration is indispensable. We found that the immune cells have moderate clinical significance, and the immune risk score based on four types of immune cells is an alternative tool for survival prediction and treatment guidance in ccRCC. However, prospective clinical trials are warranted to validate our findings.
Footnotes
Acknowledgments
The results shown in this study are partly based upon the data generated by the TCGA Research Network: http://www.cancer.gov/tcga. The current study was supported by the funds of Natural Science Foundation of Guangxi, China (2018 GXNSFAA281175, 2017GXNSFAA198107), Medical Excellence Award Funded by the Creative Research Development Grant from the First Affiliated Hospital of Guangxi Medical University, Open fund of Guangxi Colleges and Universities Key Laboratory of Biological Molecular Medicine Research (GXBMR201601) and Guangxi Zhuang Autonomous Region Health and Family Planning Commission Self-financed Scientific Research Project (Z20170556).
