Abstract
BACKGROUND:
Immunosuppressive receptor LILRB1 regulates tumors progression by transducing immune inhibitory signals via intracellular immunoreceptor tyrosine-based inhibitory motifs. However, its role in Hepatocellular Carcinoma (HCC) remains vague.
OBJECTIVE:
This study is aimed to disclose the association between LILRB1 and HCC.
METHODS:
Immunoblotting and qRT-PCR were employed to evaluate the level of LILRB1 in hepatocarcinoma cells. LILRB1-positive cells in tissue array were measured using immunohistochemistry staining. The relation among LILRB1, SHP1 and SHP2 and survival rates were analyzed using Gene Expression Profiling Interactive Analysis (GEPIA) and Oncomine database.
RESULTS:
LILRB1 was robustly reduced in hepatocarcinoma cells compared to normal cells. Clinically, LILRB1 was significantly higher in 49 of 75 (65%) paired paracarcinoma tissues than that in paired HCC samples. 48 of 75 (64%) HCC subjects in tissue microarray showed low level of LILRB1, compared to 25 of 75 (33%) in paired-adjacent tissues. Oncomine database and GEPIA analysis confirmed that LILRB1 was lower in HCC than normal tissues. Additionally, lowLILRB1 had a significant association with clinicopathological characteristics and Disease Free Survival, but no association with Overall Survival in HCC patients. Mechanismly, positive correlation between LILRB1 and SHP1, but not SHP2 was observed in HCC.
CONCLUSIONS:
LILRB1 possibly plays an antitumor effect in hepatocarcinoma cells by integrating SHP1, providing evidence that LILRB1 might be involved in the pathologic progression of HCC.
Introduction
Hepatocellular carcinoma (HCC) is the fifth most common malignancy worldwide, with a high degree of malignancy and a poor prognosis [1]. Currently, immunotherapy has been developed as an effective therapeutic option for HCC. It has been proved that the complex microenvironment of HCC can activate immunosuppressive receptors [2]. Binding of activated immunosuppressive receptors to its ligands results in a blunt immune response and tolerance of HCC [3]. However, immune checkpoint inhibitors are able to remotivate the immune system through intervening the combination of immunosuppressive receptors and its ligand, resulting in the inhibition of cancer progress [4]. Immune checkpoint inhibitor Opdivo (nivolumab) which targets programmed death-1 (PD-1), has been approved for the treatment of advanced liver cancer patients [5]. Unfortunately, However, a subset of subjects with advanced HCC treated with immune checkpoint inhibitors failed to respond to therapy and presents adaptive resistance to immune checkpoint inhibitors [6]. Therefore, we urgently need to expose new effective immunosuppressive receptors and illuminate their underlying mechanisms in HCC, as this will build a solid foundation for the development of new immune checkpoint inhibitors and offer novel concepts for improving the present situation of patients with advanced HCC
Leukocyte immunoglobulinn (Ig)-like receptor subfamily B (LILRBs) acting as a group of type I transmembrane glycoproteins are mostly expressed on the surface of immune cells such as macrophages, T cells, B cells, and NK cells [7, 8]. Once activated by its’ receptors, LILRBs transduce intracellular inhibitory signals, leading to an immune response silence [9]. LILRB1, also known as ILT2, CD85J and LIR1, is composed of extracellular Ig-like domains, transmembrane domain and intracellular immunoreceptor tyrosine-based inhibitory motifs (ITIMs) [10]. After binding with its specific ligands such as human Leukocyte Antigen-G (HLA-G) [11], UL18 [12], S100A8 and S100A9 [13], the tyrosine residues in the ITIM motif are phosphorylated by Src kinase, resulting in the activation of LILRB1 [14]. It has been demonstrated that the interaction of HLA-G with LILRB1 promotes the expansion of myeloid-derived suppressor cells (MDSCs), which causes the inhibition of the immunocompetence activity and cytotoxicity of T cells [15]. The binding of HLA-G to LILRB1 also inhibits the polarization and cytotoxicity of NK-cells [16]. Besides, LILRB1 also competitively inhibits the combination of HLA I to CD8, leading to the decrease of CD8
Activated LILRB1 negatively regulates immune cells mainly by activating intracellular ITIMs, which subsequently recruit protein tyrosine phosphatase non-receptor type 6 (PTPN6/SHP-1), PTPN11/SHP-2, or Src homology 2 domain-containing inositol phosphatase (SHIP) [21]. In B-cell lymphoma, LILRB1 curbs the intracellular downstream signal including AKT, mTOR, STAT, c-Raf and GSK3-
Herein, we demonstrated that LILRB1 was downregulated in HCC in vitro and in human specimens with HCC, and a low level of LILRB1 was closely related to patients’ age, and pathologic grades, tumor diameter, and clinical stages. Further, the expression of LILRB1 was positively correlated with SHP1 but not SHP2 in different liver subtypes and low LILRB1 had a poor DFS. Our data suggested LILRB1 was also expressed in solid liver tumor cells and might act as a tumor suppressor for hepatocellular carcinogenesis involvement with protein tyrosine phosphatase SHP1, providing a new idea for the study of immunosuppressive receptors in solid liver tumors.
Materials and methods
Tissue microarray and Immunohistochemistry
The tissue microarray containing 75 pairs of liver carcinoma tissue and the paired para-carcinoma tissue was purchased from OUTDO Biotechnology co., LTD (Shanghai, China). For immunohistochemistry (IHC) assay, slides were heated in an oven for 30 minutes at 60
Cell culture and treatment
The human immortalized normal hepatocyte cell line-LO2 and three HCC cell lines including HepG2, Huh7, and MHCC97L were purchased from Shanghai cell bank, Chinese Academy of Sciences. These cells were cultured with complete Dulbecco’s modified Eagle’s medium (DMEM; 11320-033, GIBCO, USA) supplemented with 10% fetal bovine serum (10091-148, GIBCO, USA), 100 U/ml penicillin and 100
Data collection, correlation and survival analysis
The expressions of LILRB1 mRNA and Shp1 mRNA in normal liver tissues or precancerous tissue were collected from Oncomine database (Wurmbach Liver Statistics) and analyzed by GEPIA version 2. In Wurmbach Liver Statistics database, 10 normal, 30 liver cancer precursor and 35 HCC patients were enrolled in our correlation analysis between LILRB1 and pathological types. Expression value of LILRB1 which was indicated as the value of Log2 median-centered intensity in these specimens were used to assess the Pearson correlation of LILRB1 and SHP1 or SHP2 expression by SPSS software in different liver tissues. GEPIA was used to analyze the Overall Survival and Disease Free Survival.
Western blotting
Total protein extracts were prepared using RIPA protein lysate (P0013B, Beyotime, China). 40
Co-immunoprecipitation
The expression of LILRB1 in liver cancer cells and tissue array. (A) LILRB1 protein levels in LO2, MHCC97L, Huh7 and HepG2 cells as determined by Western blotting. (B) The expression of LILRB1 mRNA in LO2, MHCC97L, Huh7 and HepG2 cells as analyzed by qRT-pCR. (C) Representative IHC images of LILRB1 expression in the microarray that contained 75 pairs of HCC and paired adjacent liver tissues. Brown color indicates positive staining of LILRB1. Multiple images were taken and representative one was presented. Scale bar: 200 and 50 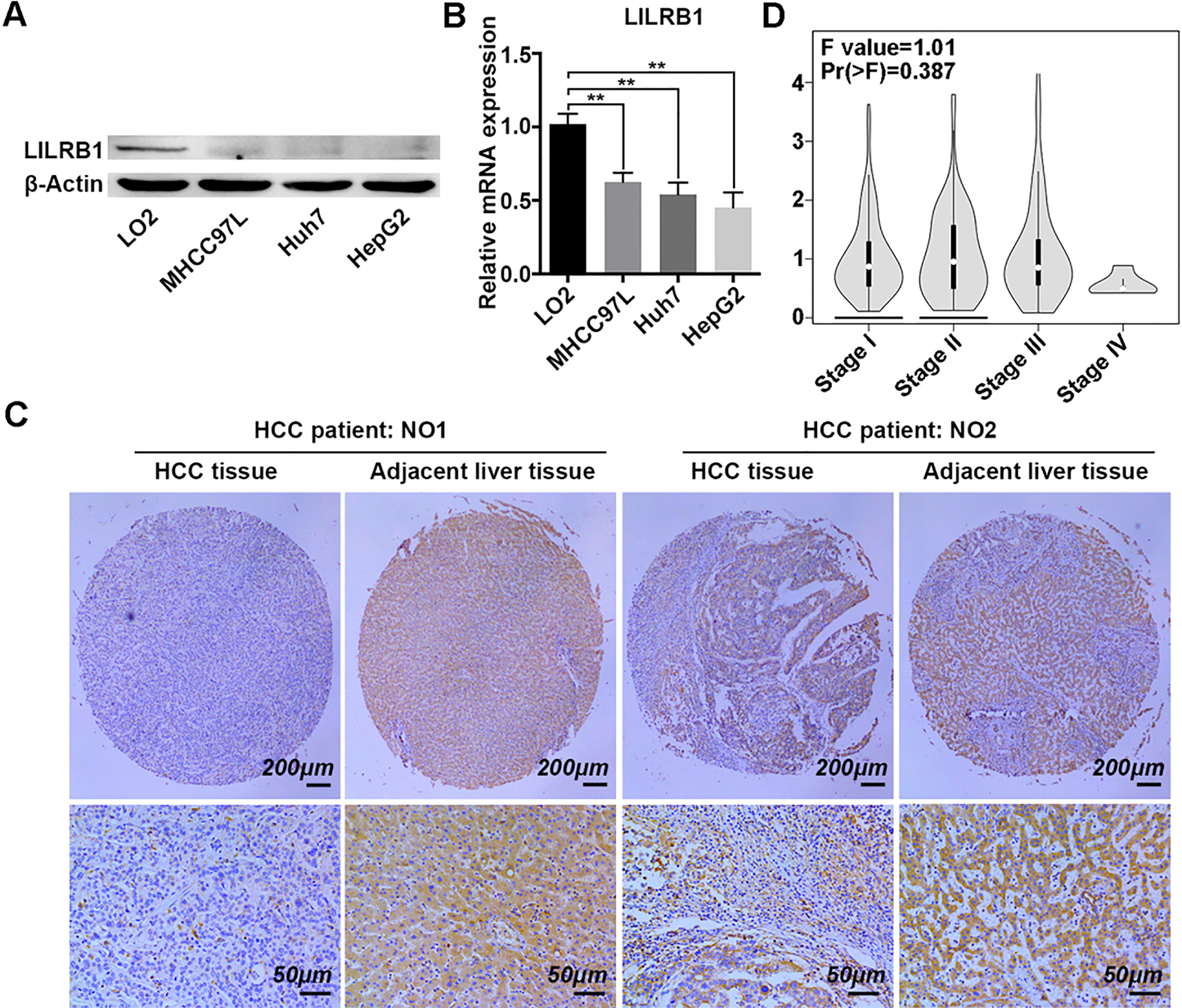
Total protein lysates extracted from LO2 cells were incubated with antibodies against SHP1 (ab227503, abcam, Cambridge, UK) and isotype control IgG (ab172730, abcam, Cambridge, UK) overnight. And the complexes were incubated with protein A/G beads (88802, Thermo Scientific, USA) for another 3 h at room temperature. Then the immunoprecipitate was resuspended in 2x SDS loading buffer were boiled for 5 minutes and subjected to western blotting assay using LILRB1 (ab185325, abcam, Cambridge, UK) and SHP1 antibodies according to the above method.
Differences between clinical information and LILRB1 expression was analyzed by GraphPad 6.0 using chi-square and Fisher’s exact tests. Pearson correlation coefficient was used for analyzing the relation between LILRB1 and SHP1 or SHP2 by SPSS 25.0 software. Different degrees of staining of histological staining were analyzed by image pro plus. Quantitative analysis of protein and RNA was conducted by Quantity One and Graph Pad Prism using one-way ANOVA, followed by Student’s
Results
The expression of LILRB1 is depressed in liver cancer cells and liver cancer tissue array
LILRB1, as an immunoregulatory receptor, functions mainly as a key regulator in immune cells and hematologic malignant, but is seldom involved in the progress of solid cancer types [21]. To investigate the role of LILRB1 on HCC, we firstly determined its expression pattern in normal liver LO2 cells, and three liver cancer cell lines including MHCC97L, Huh7 and HepG2. As shown in Fig. 1A, the protein level of LILRB1 was drastically reduced in three liver cancer cell lines compared to that in the LO2 cells (Fig. 1A). Next, qRT-PCR was employed to further evaluate the transcription level of the LILRB1 gene in these cell lines. The results showed that relative expression of LILRB1 mRNA was significantly lower in the three cancer cell lines than that in normal liver cells (Fig. 1B).
To further ascertain the expression pattern of LILRB1 in hepatocellular carcinoma, a tissue microarray containing 75 pairs of HCC samples and paired adjacent liver tissue were employed to evaluate the level of LILRB1 in human specimens by immumohistochemical staining. The IHC results showed that the expression of LILRB1 was notably downregulated in most of the HCC samples compared to the paired adjacent liver tissue (Fig. 1C). Through IHC staining, we discovered that a low expression of LILRB1 (
IHC staining of LILRB1 in human HCC and adjacent liver tissues
IHC staining of LILRB1 in human HCC and adjacent liver tissues
–, +, ++, +++ represent different degrees of staining.
Next, we conducted the association between LILRB1 expression and clinicopathological characteristics of HCC patients in the tissue microarray. Among these 75 patients, we observed a significant decrease of LILRB1 in male subjects (
Expression pattern of LILRB1 in human HCC specimens. (A) The analysis of LILRB1 transcription activity in normal, liver cancer precursor and HCC based on the online Oncomine database. (B) The transcript of LILRB1 in normal and liver tumor as analyzed by GEPIA 2. *, 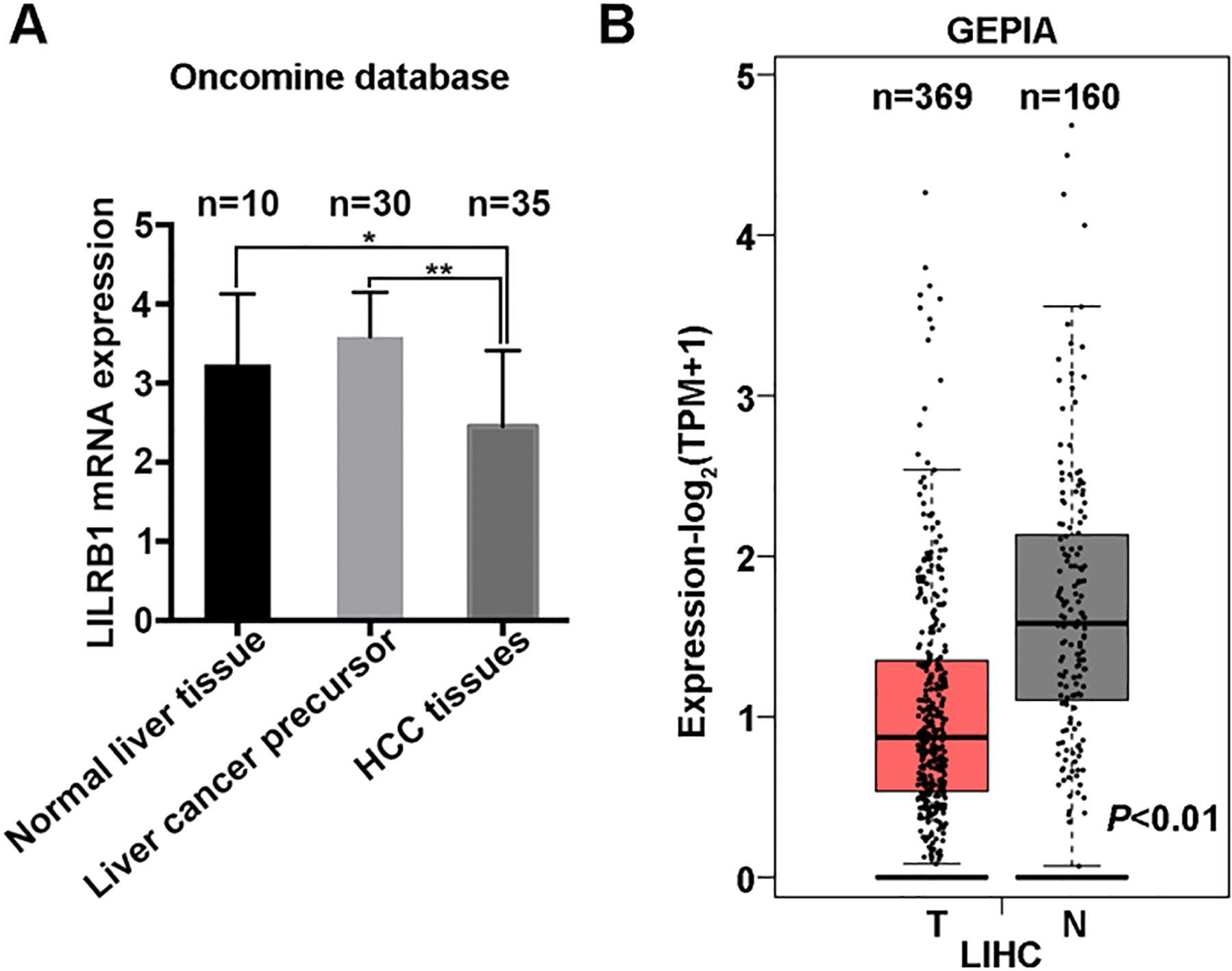
Correlation between LILRB1 expression and clinicopathological characteristics of HCC patients
Survival rates analysis in low and high LILRB1 group. (A) Overall survival and (B) Disease Free survival in HCC patients with low LILRB1 (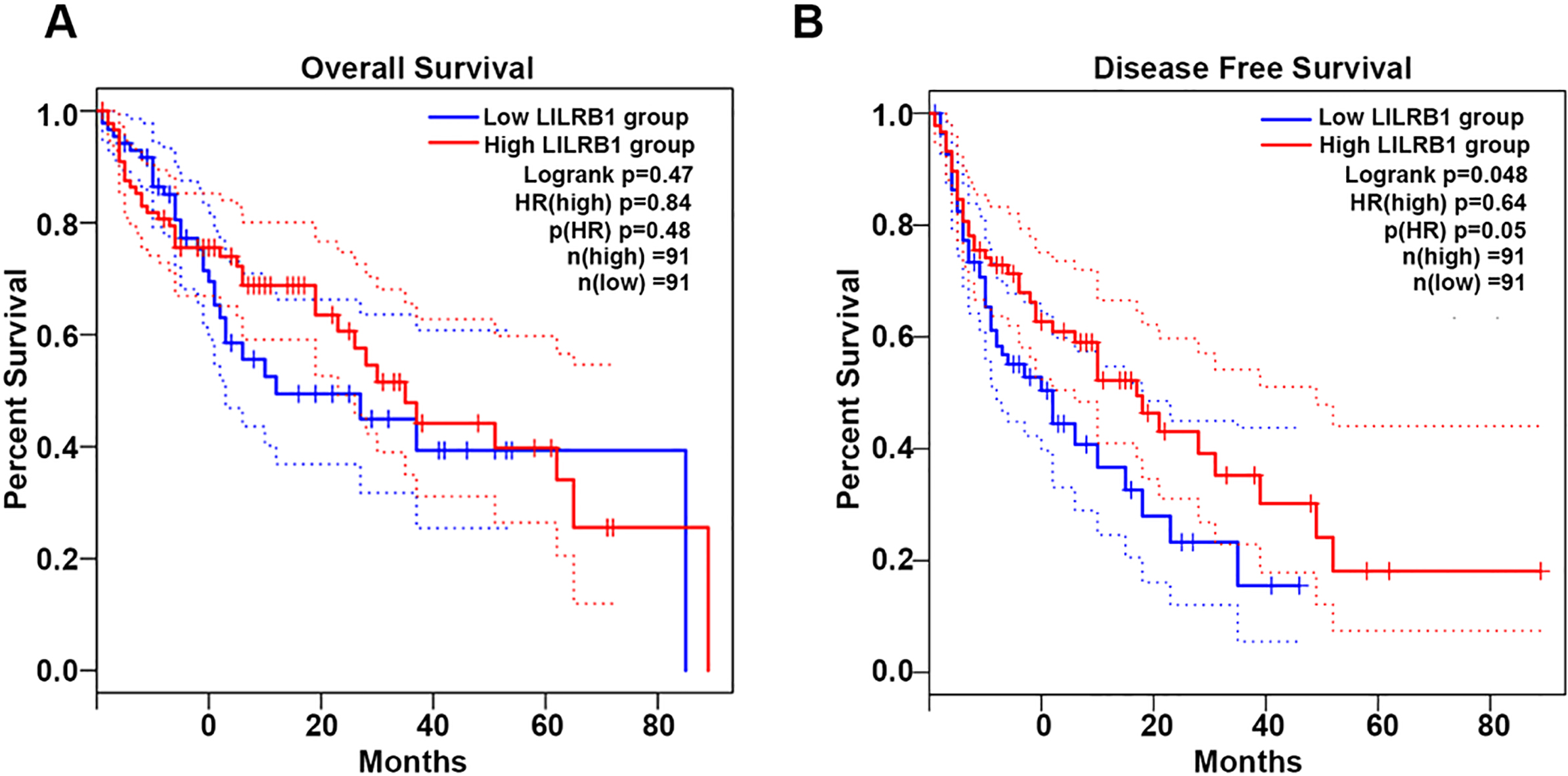
Combined with the online Oncomine database, we further explored the expression profile of LILRB1 mRNA in HCC. The results showed that the expression of LILRB1 mRNA was much lower in HCC tissues (
HCC patients with low LILRB1 have a short period of Disease Free Survival
To further prove the crucial determinant of LILRB1 in the development of HCC, we analyzed the impact of LILRB1 on survival rates. As shown in Fig. 3, among the 182 HCC patients including low LILRB1 group (
Association analysis of LILRB1, SHP1 and SHP2 in HCC. (A) Protein expression of LILRB1, SHP1 and SHP2 in different cells as determined by western blotting assay. (B) Grayscale quantitation of LILRB1, SHP1 and SHP2 protein expression in different cells. (C) Correlation analysis between LILRB1 and SHP1 as evaluated by GEPIA 2 in liver, normal, tumor and the three combination. (D) Correlation analysis between LILRB1 and SHP2 as evaluated by GEPIA 2 in liver, normal, tumor and the three combination. R represent Pearson’s correlation coefficient. (E) Evaluation of the interaction between LILRB1 and SHP1 in liver LO2 cells using co-immunoprecipitation assay. The antibody against SHP1 was used to immunoprecipitate specific complexes. Then the immunoprecipitate was subjected to SDS-polyacrylamide gel electrophoresis for protein detection. *, 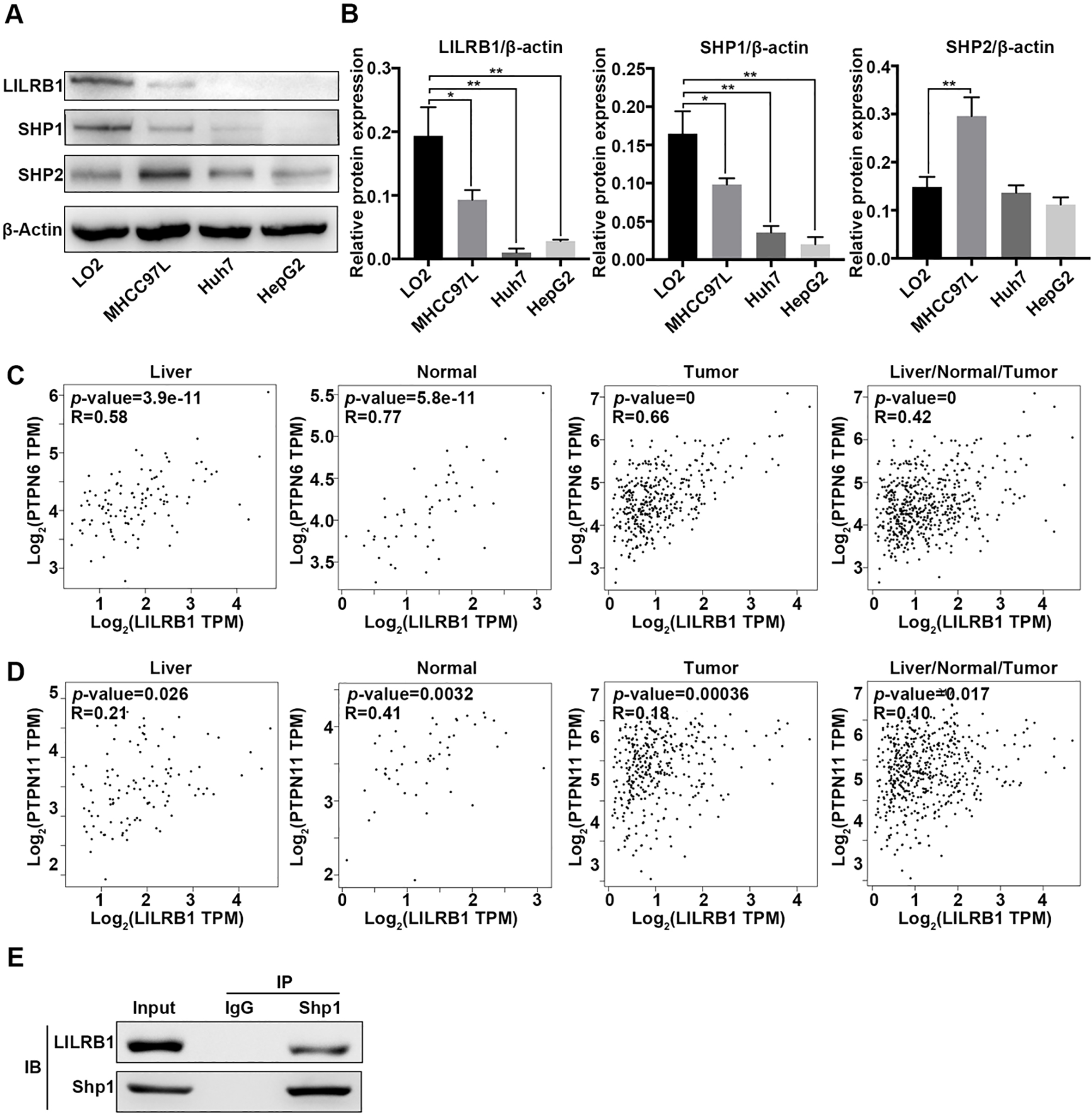
The SH2 domain of SHP1/2 phosphatases constitutively bind to the cytoplasmic ITIMs of LILRB1, which play a pivotal role in cellular homeostasis and tumor development [26]. We therefore extensively searched the expression association of LILRB1, SHP1 and SHP2 in different liver subtypes. First, we determined the protein expression of SHP1, SHP2 and LILRB1 in different liver cell lines. We observed a moderate expression of SHP1, SHP2 and LILRB1 in normal liver cells. However, both the protein levels of SHP1 and LILRB1 were robustly declined in liver cancer cell lines such as MHCC97L, Huh7 and HepG2 cells (Fig. 4A and B). Though a moderate increase in the expression of SHP2 was observed in MHCC97L cells compared to LO2, no alteration appeared in Huh7 and HepG2 cells compared with LO2 cells (Fig. 4A and B). Additionally, based on search results from the online Oncomine database, we observed that transcription levels of LILRB1 had a significantly positive correlation with SHP1 in normal liver tissues (
Discussion
Compared with traditional surgery, radiation therapy, and chemotherapy, immunotherapy focuses on the activation of immunogenicity of tumor cells, which improves the recognition of tumor cells by immune cells and enhances anti-tumor immune response [27]. The well-known targets of tumor immunotherapy include cytotoxic T-lymphocyte-associated antigen 4 (CTLA-4), PD-1 and its ligand PD-L1, and lymphocyte activation gene 3 (LAG-3) [28]. However, owing to the complex mechanism of tumor progression, the anti-tumor effects against these targets is not sufficient to satisfy the clinical demand. In the present study, we found that immunosuppressive receptor LILRB1 was depressed in hepatocellular carcinoma cells and had a positive correlation with SHP1 that is a key switch in the progression of hepatocellular carcinoma, implying that LILRB1 may act as a potential therapeutic target for hepatocellular carcinoma immunotherapy.
Previous studies have shown that LILRB1 is not only expressed on immune cells, but also in tumor cells, in which LILRB1 is responsible for the regulation of tumor growth, drug resistance and cancer stem cell activities [29]. It has been proved that LILRB1 is expressed on acute myeloid leukaemia (AML) cells [30], neoplastic B cells (including B-cell leukemia, B-cell lymphoma, and multiple myeloma cells) [22], T cell leukemia and lymphoma cells [31]. However, LILRB1 plays a dual role in the occurrence and development of hematological tumors. LILRB1 can inhibit the cytotoxicity effects of immune cells and promote tumorigenesis. On the other hand, LILRB1 is able to ameliorate the progression hematological carcinoma by inhibiting the proliferation of lymphoma cells [32]. It is demonstrated that the expression of LILRB1 in NK cells was negatively correlated with the function of NK cells, and LILRB1 depletion could revive the immune function of NK cells in triple-negative breast cancer patients [33, 34]. In our study, LILRB1 was downregulated in liver cancer cells and HCC patients with different ages, especially in male HCC patients. But whether LILRB1 expression in liver cancer was associated with patients’ gender needed to further explore based on large scale clinical trials. Moreover, the low expression of LILRB1 was more significant when occurring in patients with high pathological grades and clinical stages. These results indicate that the declined LILRB1 may be involved in the occurrence of liver cancer. However, the decrease of LILRB1 in HCC tissue is not limited to hepatic parenchymal cells. It may be due to the downregulation of LILRB1 in many other cell types such as stellate cells, macrophages and other inflammatory cells, and this still needs further investigation.
Paired immunoglobulin-like receptor B (PirB), as the ortholog of LILRB2, participates in the process of immune response by regulating SHP1 or SHP2 [35, 36]. Removal of PirB leads to a decrease of phosphorylation of SHP-1 and SHP-2 in acute myeloid leukemia [37]. LILRB2 inhibition in non-small cell lung cancer (NSCLC) cells blunts the proliferation and colony formation viability of A549 cells by targeting the SHP2/CAMKI/CREB signal pathway [38]. Besides, Leukocyte-associated Ig-like receptor (LAIR1), also containing the immunoreceptor tyrosine-based inhibition motif (ITIMs), is crucial for the development of AML through activating SHP-1/CAMKI/CREB signal transduction [39]. LILRB1 recruiting SHP2 but not SHP1, as an anti-tumor regulator, significantly arrests the cell-cycle of human-activated T cells [40]. In our data, we detected that LILRB1 was positively correlated with SHP1 but not SHP2 in both HCC tissues and normal liver tissues. There had strong interaction between LILRB1 and SHP1 in liver cells. Therefore, the interaction between LILRB1 and SHP1 in liver cells may be involved in the pathogenesis of liver cancer. Besides, we confirmed that high LILRB1 had a good prognosis through affecting the period of DFS of HCC patients. Given that SHP1 had been proved to affect the prognosis of several cancer types [41, 42], we thought that the LILRB1-mediated improvement of prognosis was involved with SHP1. However, we can’t deny that SHP2 is also very important for HCC progress. SHP2 facilitates HCC cell differentiation and cancer stem cell expansion by promoting
In summary, this study demonstrated that LILRB1 was expressed on liver cells and little expression in HCC patients and liver tumor cells. Low levels of LILRB1 had a significant positive association with SHP1 level and with part of clinicopathological data of HCC patients. LILRB1 level also affected the period of DFS. These data implied that LILRB1 might participate in tumorigenesis and therapy of HCC. Possibly, LILRB1 acts as an anti-cancer factor by interacting with SHP1 in the pathological process of HCC.
Footnotes
Acknowledgments
We thank the support of The Youth Innovation Team of Shaanxi Universities and Gene Expression Profiling Interactive Analysis (GEPIA). This work was supported by the Research Foundation of Xi’an Medical University (Grant No 2017DOC18 and 2018GJFY04), Natural Science Basic Research Plan in Shaanxi Province of China (Grant No. 2018JQ 8043), Young Talent Fund of University Association for Science and Technology in Shannxi (Grant No. 20190309) and Natural Science Foundation of China (81802332).
Conflict of interest
The authors declare that they have no competing interests.
Abbreviations
Supplementary data
Supplement Table S1
Differential expression of LILRB1 in human HCC and adjacent liver tissues
Higher expression in adjacent liver tissues
No variance
Higher expression in HCC tissues
Number of samples
49 (49/75, 65%)
15 (15/75, 20%)
11 (11/75, 15%)
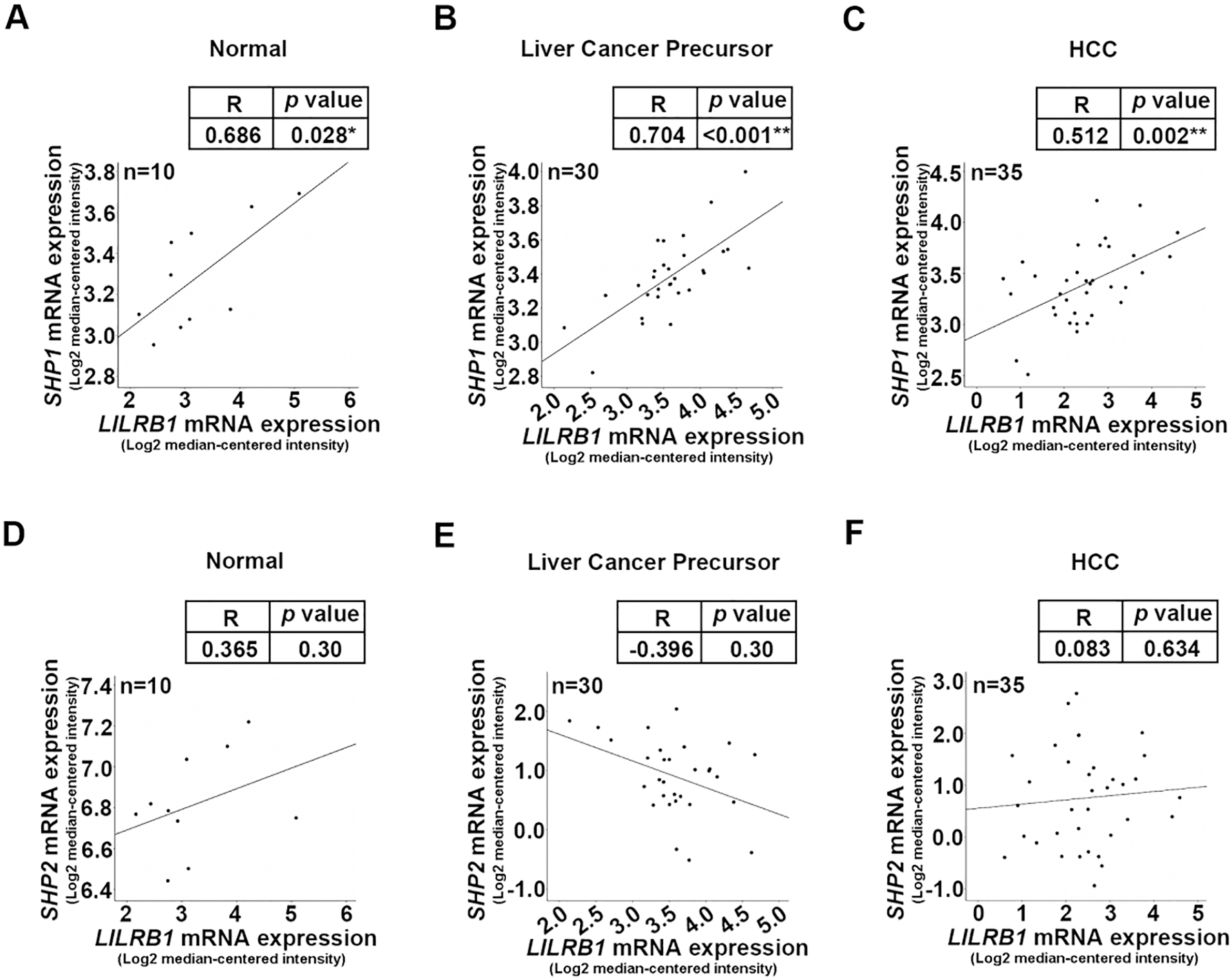
Association analysis ofLILRB1, SHP1 and SHP2 mRNA in human HCC samples. (A) Correlation analysis between LILRB1 mRNA and SHP1 as evaluated by SPSS in normal liver tissues, (B) liver cancer precursor tissues (C) and HCC tissues. (D) Correlation analysis between LILRB1 mRNA and SHP2 as determined by SPSS software in normal liver tissues, (E) liver cancer precursor tissues (F) and HCC tissues. Data represent the means