Abstract
PURPOSE:
DNA methylation plays an important role in thyroid oncogenesis. The aim of this study was to investigate the connection between global and local DNA methylation status and to establish the levels of important DNA methylation regulators (TET family and DNMT1) in thyroid tumours: follicular adenoma-FA, papillary thyroid carcinoma-PTC (classic papillary thyroid carcinoma-cPTC and papillary thyroid carcinoma follicular variant fvPTC).
METHODS:
Global DNA methylation profile in thyroid tumours tissue (41 paired samples) was assessed by 5-methylcytosine and 5-hydroxymethylcytosine levels evaluation (ELISA), along with TETs and DNMT1 genes expression quantification. Also, it was investigated for the first time TET1 and TET2 promoter’s methylation in thyroid tumours. BRAF V600E mutation and RET/PTC translocation testing were performed on all investigated samples. In vitro studies upon DNA methylation in K1 thyroid cancer cells were performed with demethylating agents (5-AzaC and vitamin C).
RESULTS:
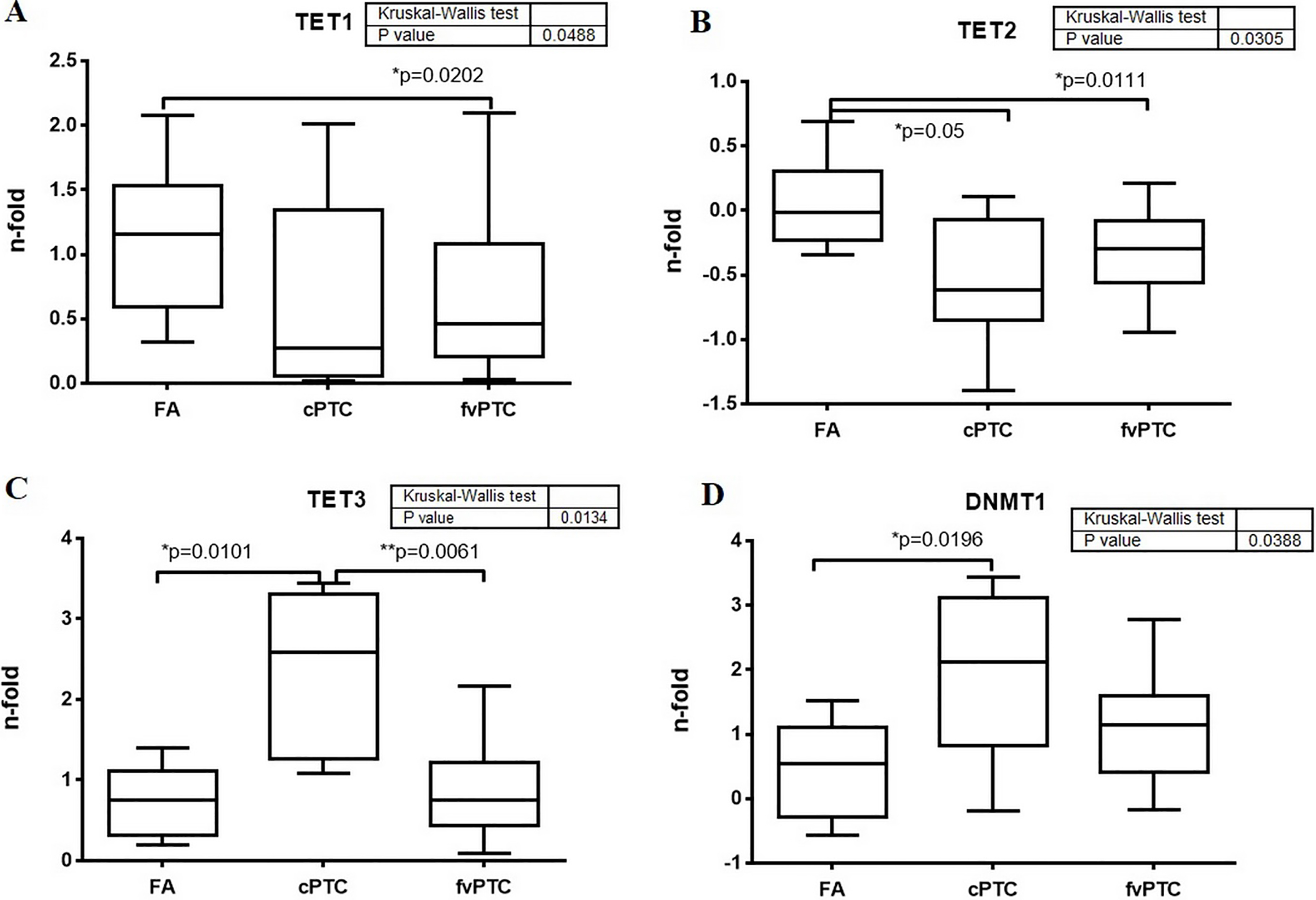
TET1 and TET2 displayed a significantly reduced gene expression level in PTC, while DNMT1 gene presented a high level of expression. PTC samples presented increased levels of 5-methylcytosine and low levels of 5-hydroxymethylcytosine. 5-methylcytosine levels were associated with TET1/TET2 expression levels. TET1 gene expression was significantly lower in patients positive for BRAF mutation and with RET/PTC rearrangement. TET2 gene was found hypermethylated in thyroid carcinoma patients overall, especially in PTC-follicular variant samples (
CONCLUSIONS:
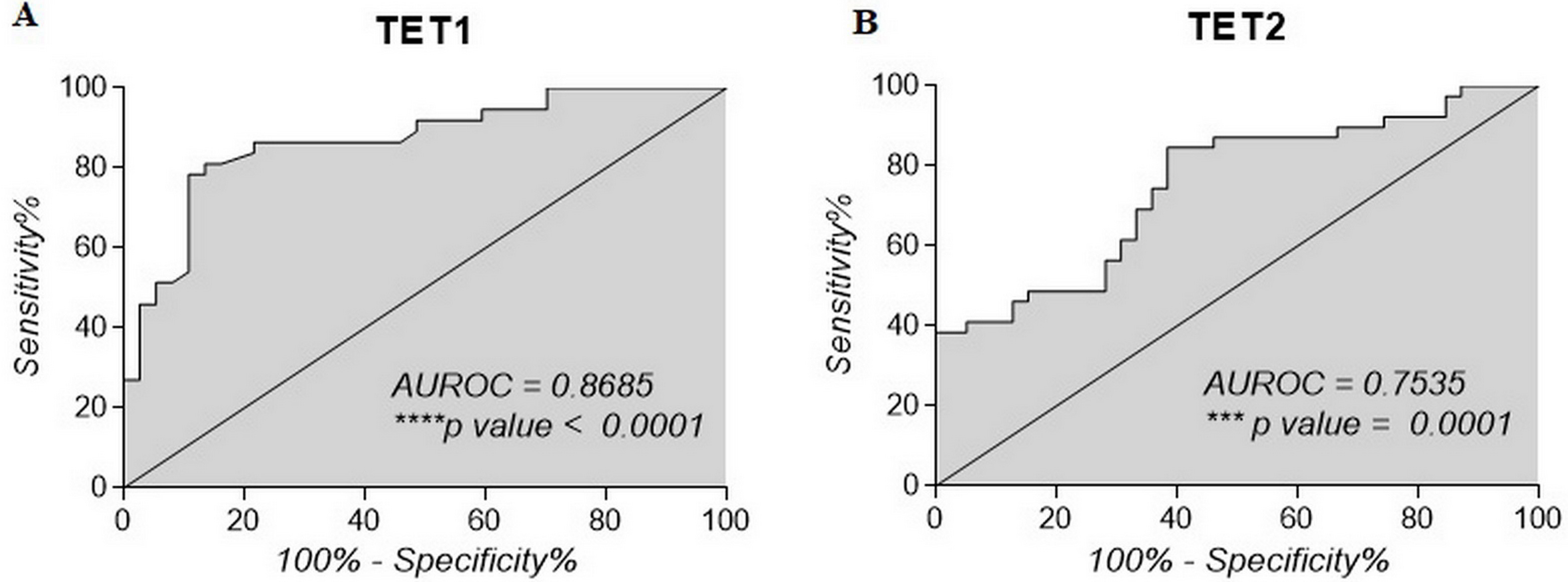
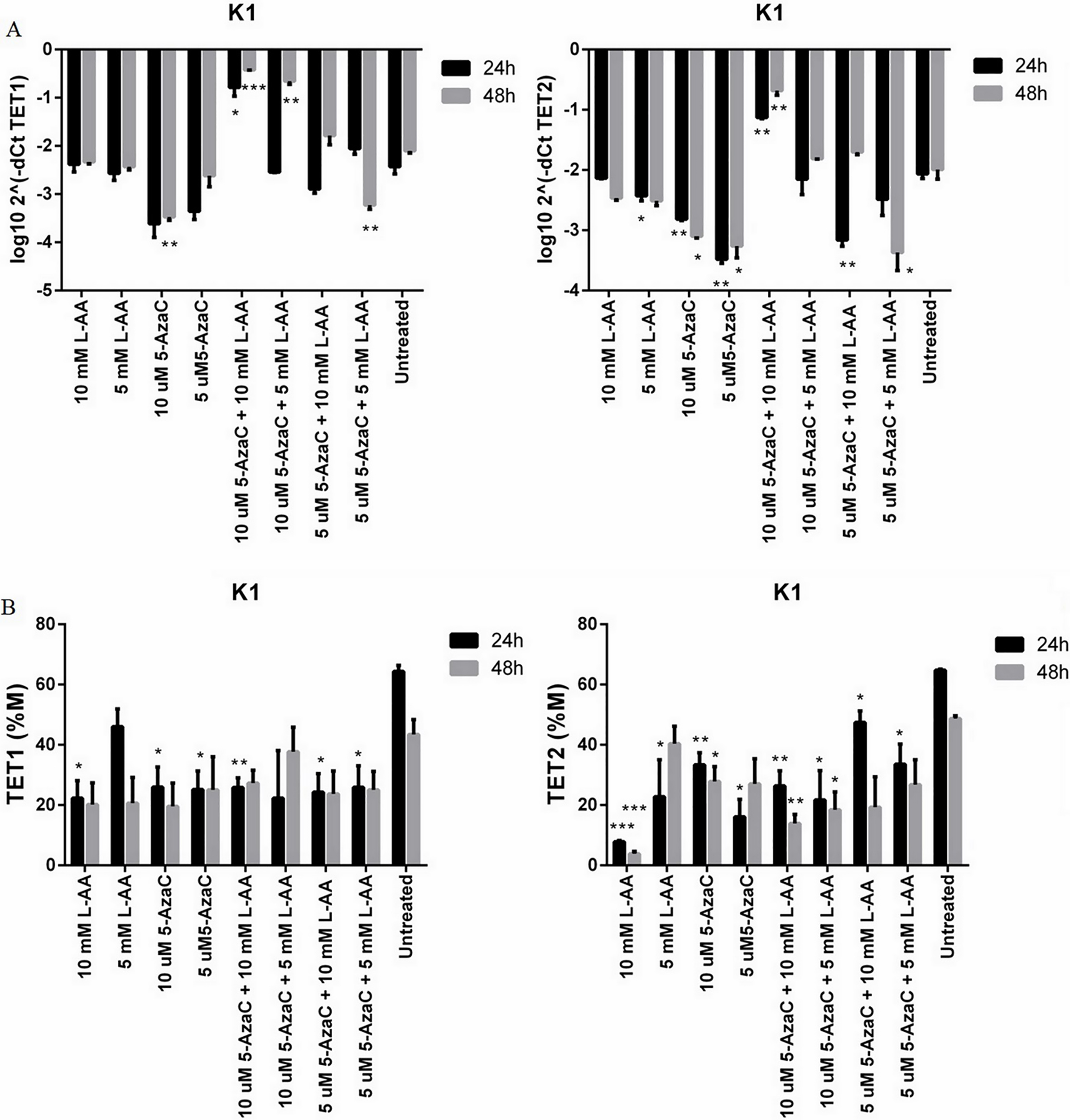
These results suggest that TET1/TET2 gene expression and methylation may serve as potential diagnostic tools for thyroid neoplasia. Our study showed that the methylation of TET1 increases in malignant thyroid tumours. fvPTC patients presented lower methylation levels compared to cPTC and could be a discriminatory factor between two cancer types and benign lesions. TET2 is a poorer discriminator between FA and fvPTC, but it can be useful for cPTC identification. K1-cells treated with demethylating agents showed a demethylation effect, especially upon TET2 gene. The cumulative effect of L-AA and 5-AzaC proved to have a potent combined demethylating effect on genes promoter’s activation and could open new perspectives for thyroid cancer therapy.
Introduction
Thyroid carcinoma is one of the most frequent types of endocrine malignancy. Lately, worldwide thyroid cancer incidence has grown, the malignancy being more common among women and often diagnosed between ages 45–54 [1]. Compared with other types of cancer, it presents a low death rate and in the majority of cases is curable following surgery and treatment. According to WHO 2017 thyroid carcinomas are classified as follows: benign, borderline/uncertain and malignant types. For benign type there is only one subtype: follicular adenoma, borderline/uncertain includes hyalinizing trabecular tumour and other encapsulated follicular patterned tumours. As for malignant type: poorly differentiated thyroid carcinoma, anaplastic thyroid carcinoma, Hurthle (oncocytic) cell tumours, follicular thyroid carcinomas (FTC) and papillary thyroid carcinomas (PTC) subtypes [2].
The majority of thyroid differentiated tumours cases are curable while medullary and anaplastic types are more aggressive, presenting an early metastasis potential and overall a poor prognosis. The most commonly diagnosed type of thyroid cancer is papillary thyroid carcinomas up to approximately 85% of all cases [3].
PTC is characterized by genetic alterations such as mutations of BRAF, RAS genes or RET gene rearrangements (RET/PTC) [4]. For instance, BRAF (V600E) mutation is found in approximatively 45% of PTC cases and it is associated with aggressiveness and drug resistance due to heterogeneity [5, 6].
Besides genetic alterations, a key factor that contributes to cancer development and progression is played by epigenetic changes: DNA methylation, non-coding RNAs expression (specifically microRNAs) and in histone modifications, all of which affect the gene expression pattern regulation [7].
Such an important epigenetic alteration often seen in thyroid neoplasia is represented by DNA methylation. Commonly, in thyroid tumours, it has been shown that many tumour suppressor genes can be epigenetically inactivated by promoter’s methylation [8]. p16, PTEN, RAR
DNA methylation, a reversible process that occurs within the CpG dinucleotides at the position 5 of cytosine ring (C5) is mediated by a family of enzymes, DNA methyltransferases (DNMTs). DNMT1 is mainly involved in maintenance of DNA methylation pattern while DNMT3A and DNMT3B are responsible for the de novo methylation.
On the other hand, DNA demethylation enzymes have been identified, namely Ten-Eleven Translocation (TET) proteins. TET proteins family consists in three members: TET1, TET2 and TET3 and they are accountable to catalysing the conversion of 5-methylcytosine (5mC) to 5-hydroxymethylcytosine (5hmC), to 5-formylcytosine (5fC) and 5-carboxylcytosine (5caC) [15, 16]. DNA methylation, being a dynamic process and subjected to many changes, affects the cytosine forms (5hmC, 5fC, 5caC) distribution throughout the genome. 5mC levels correlate with transcriptional silencing and are generally found at repetitive and heterochromatic regions of the genome while 5hmC is enriched at promoter regions, ORFs (open reading frames) and intergenic regions and it associates with gene transcriptional activity [17, 18].
DNA methylation changes are connected with TETs and DNMTs functions in human cancers, but little is known about this interplay in thyroid cancer. The present study aims to investigate the connection between global and local DNA methylation status and to establish the levels of important DNA methylation re-gulators (TET family and DNMT1) in thyroid tumours: follicular adenoma-FA, papillary thyroid carcinoma-PTC (classic papillary thyroid carcinoma-cPTC and papillary thyroid carcinoma follicular variant fvPTC).
Materials and methods
Clinical samples and characterization of the study group
Forty-one patients (age: 17–74) with thyroid neoplasia were enrolled and 82 fresh tissue specimens (tumoral and matched nontumoral adjacent tissue) were obtained after surgical thyroidectomy. Inclusion criteria included patients presenting thyroid nodule pathology and suspicious for thyroid carcinoma. The pre-sence of any other malignancies in patient’s history or thyroid aggressive malignant types (anaplastic or medullary) represented the exclusion criteria. Based on the histopathological exam, patients were divided into three groups: patients with FA (follicular adenoma;
Clinical and histopathological characteristics of studied groups
Clinical and histopathological characteristics of studied groups
Informed consent was obtained from each subject prior to the study enrolment.
Human thyroid carcinoma cell line K1 purchased from European Collection of Authenticated Cell Cultures (ECACC 92030501) is a papillary thyroid carcinoma cell line derived from GLAG-66 cell line [19, 20] and shown to exhibit BRAF V600E mutation [21]. Cells were grown in accordance with manufacturer’s indications: in DMEM Ham’s F12: MCBD (2:1:1) medium supplemented with 2 mM Glutamine and 10% Foetal Bovine Serum (FBS) at 37
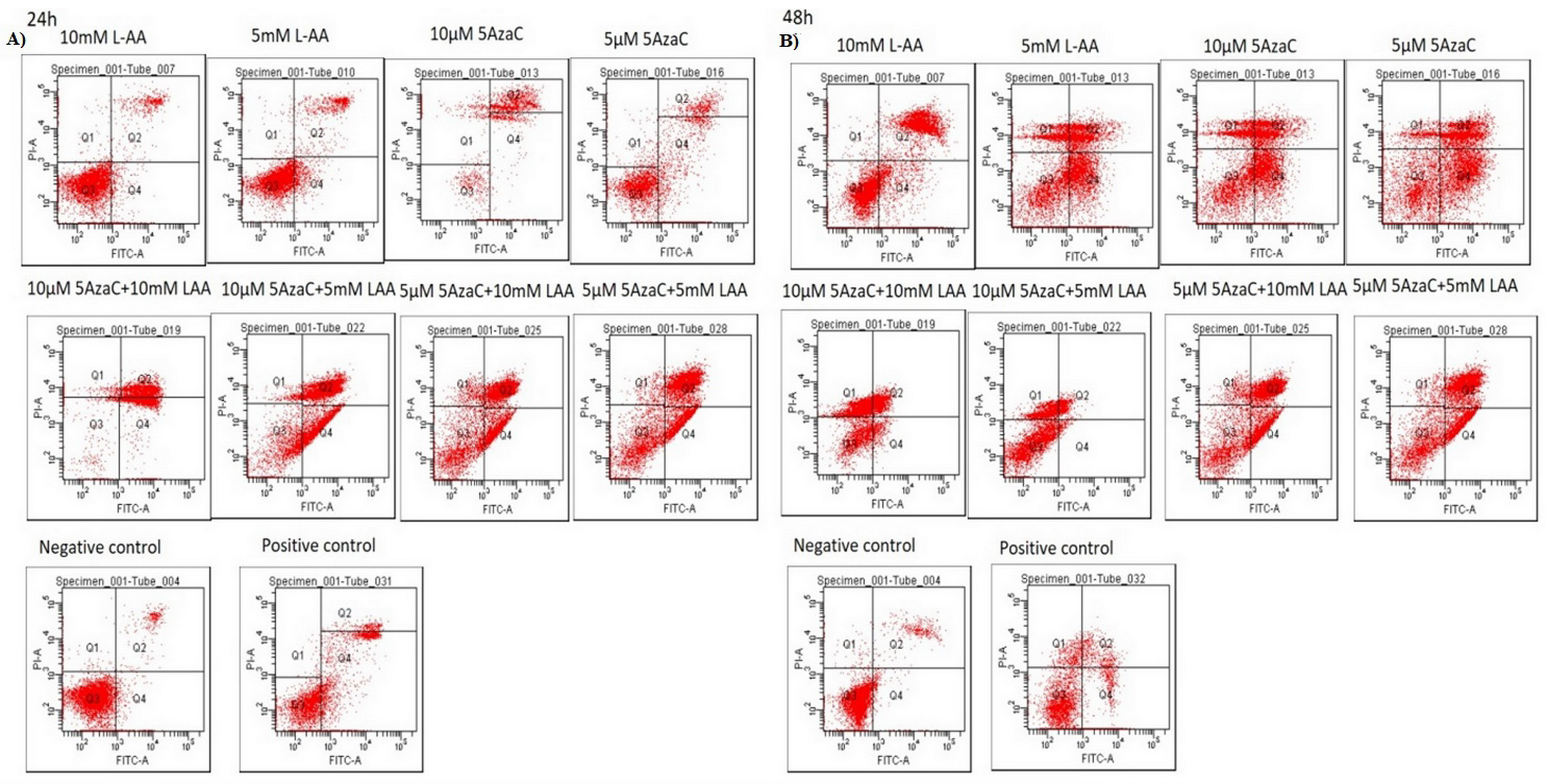
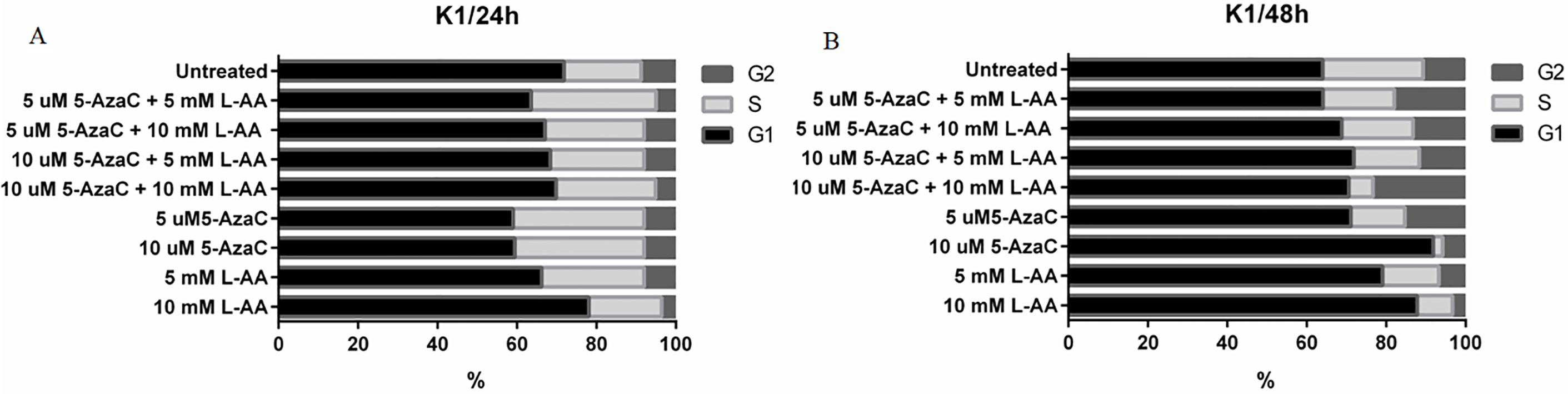
Drug cytotoxicity assays and analysis of cell viability, apoptosis and cell cycle
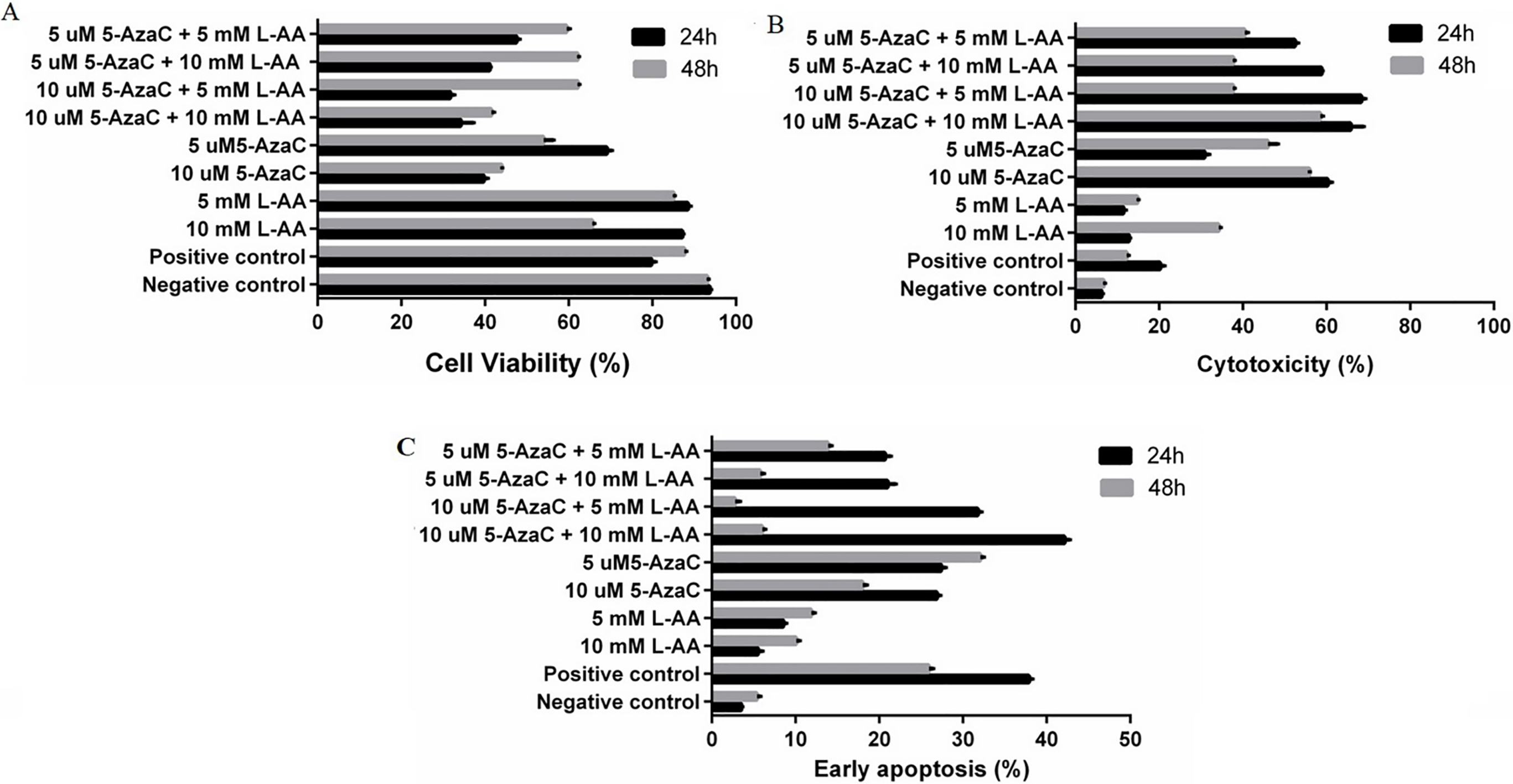
This analysis was performed using MTT assay (Sigma-Aldrich; Germany) according to manufacturer’s indications and as previously described [22]. This experiment was done in triplicate and following this, the optimal concentrations and conditions were selected for cells treatment: 5 and 10
DNA isolation and sodium bisulfite treatment
Genomic DNA from all patient’s fresh tissue spe-cimens (nontumoral adjacent/tumoral) as well as from cell cultures treatments was extracted using High Pure PCR Template Preparation kit (Roche, Mannheim, Germany). Nanodrop ND-1000 Spectrophotometer (Thermo Fisher Scientific, Waltham, MA, USA) was used for DNA concentration quantification for each sample. Afterwards, the samples presenting A260/280 value of approximately 1.8–2 and a quantity of 1
Global DNA methylation assay
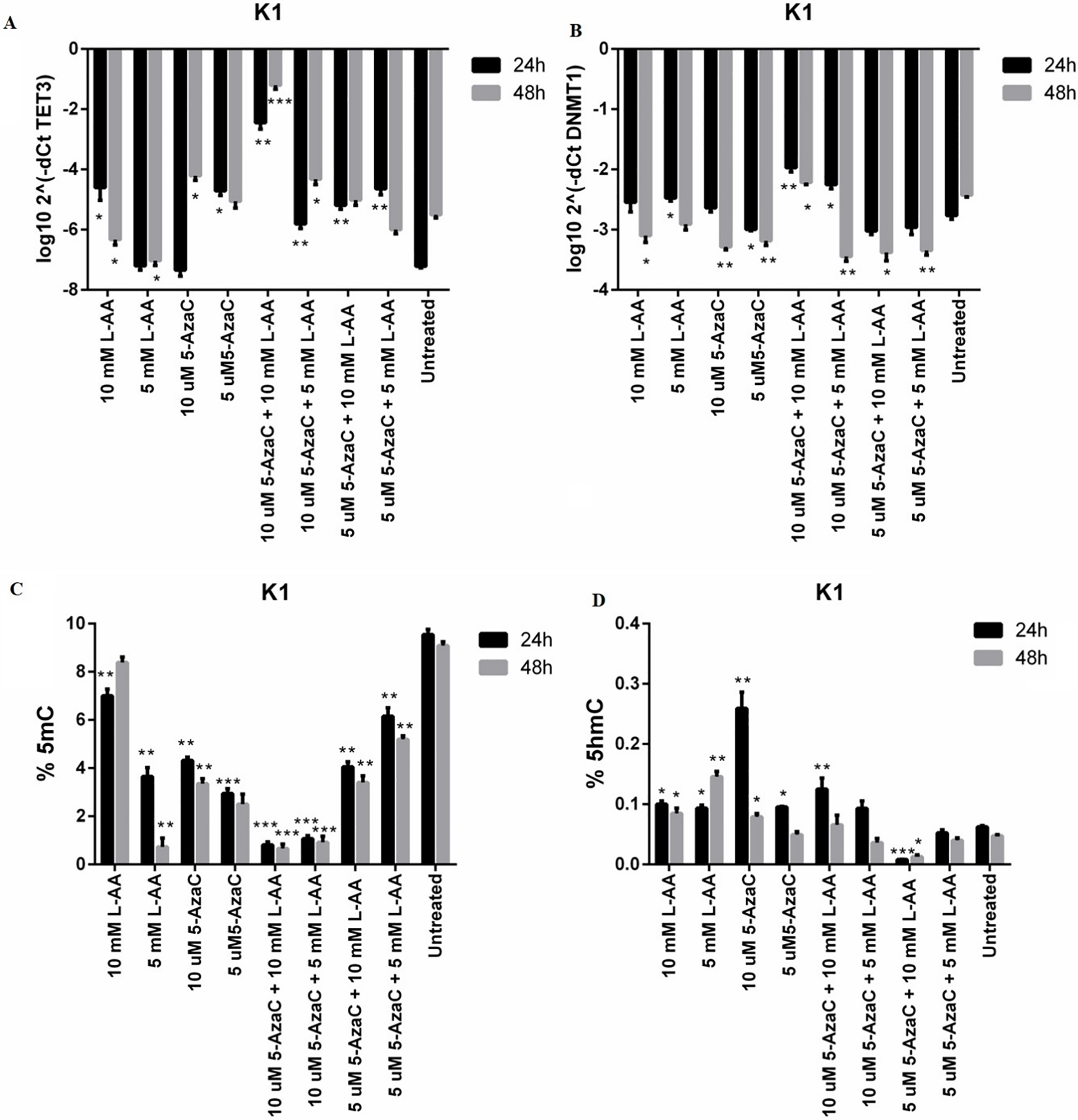
Global DNA methylation and hydroxymethylation evaluation was performed on DNA isolated from thyroid tissues through quantification of 5mC and 5hmC genomic levels using ELISA method 5-mC DNA ELISA kit and Quest 5-hmC DNA ELISA kit (Zymo Research, CA, USA). 100 ng of genomic DNA/well from each sample was analysed in accordance with manufacturer’s protocol and absorbance at 420 nm was measured. In order to estimate the amount of 5mC and 5hmC for tested samples two standard curves using kit’s controls were generated. The results are presented as percentage of 5mC and 5hmC in total DNA. The experiments were performed in duplicate.
RNA isolation
Total RNA was extracted from investigated samples and K1 cell line using TRIzol Reagent (Thermo Fisher Scientific, Waltham, MA, USA) and later purified with RNeasy kit (Qiagen, Hilden, Germany). Analysis of isolated RNA (quantity and quality) was performed by using Experion RNA analysis kit (Bio-Rad, CA, USA), samples presenting RQI (RNA Quality Indicator) values
Quantitative real-time PCR and quantitative methylation specific real-time PCR (qMSPCR)
mRNA expression levels of TET family members and DNMT1 were quantified by qRT-PCR on ABI 7300 Real Time PCR System (Applied Biosystems, Foster City, CA, USA), using FastStart SYBR Green Master mix (Roche, Mannheim, Germany) following ma-nufacturer’s instructions. 50 ng of cDNA from each biological and cell cultures sample were used and each sample was run in duplicate. For relative normalization GAPDH gene was used as reference. Double normalization was performed to estimate target genes fold change using 2
For TET1 and TET2 promoter methylation specific primers design and promoter’s CpG islands prediction, MethPrimer software was used [26]. To estimate TET1 and TET2 genes methylation levels for investigated samples, quantitative methylation specific PCR (qMSPCR) was done on ABI 7300 Real Time PCR System (Applied Biosystems, Foster City, CA, USA) using 50 ng/
%5mC and %5hmC levels in Nontumoral adjacent tissue and tumour tissue
%5mC and %5hmC levels in Nontumoral adjacent tissue and tumour tissue