Abstract
Faced with the increasing colorectal cancer (CRC) cases, the interrogation of pivotal molecules in CRC appears to be vitally important. Long non-coding RNAs (lncRNAs) are well-known regulators of gene expression at transcriptional, post-transcriptional or epigenetic level, among which the competing endogenous RNA (ceRNA) network is a common way that lncRNAs exert their properties. The current study aimed to provide a new insight into improving the outcomes of CRC patients. Our study detected that ELFN1-AS1 expression was elevated in CRC tissues and cells, and ELFN1-AS1 upregulation was correlated with poor prognosis of CRC sufferers. Besides, it was viewed that ELFN1-AS1 knockdown impeded the proliferation and migration abilities as well as activated the apoptosis ability of CRC cells. In subsequence, mechanism assays also displayed that ELFN1-AS1 targeted miR-4644 to augment TRIM44 level. Finally, rescue experiments confirmed that TRIM44 took part in the ELFN1-AS1-medatied promotional influences on CRC cells proliferation and migration. In conclusion, ELFN1-AS1 exerted pro-proliferation, anti-apoptosis and pro-migration functions on CRC cells by acting as a sponge of miR-4644 to increase TRIM44 expression at mRNA and protein level, providing an additional molecule responsible for the carcinogenesis and progression for CRC.
Introduction
In developing countries, the number of new cases and even mortalities of cancers is steadily increasing owing to rapid economic development and aging population. Colorectal cancer (CRC) arouses the attention of researchers for its second highest incidence and third highest mortality among cancers [1, 2]. There are several treatment patterns for CRC, such as neoadjuvant chemoradiotherapy with radical surgery, surgical resection and anal retention [3]. In Asia, the overall cure rate of CRC is still far from satisfaction in past decades and the 5-year survival remains at nearly 60%, which are largely due to cancer recurrence and metastasis [4, 5].
In the human genome, only 1 to 2% genes encodes proteins and the other genes are non-coding RNAs (ncRNAs) involving small ncRNAs (miRNAs, piRNAs and siRNAs) and long ncRNAs (lncRNAs) with at least 200 nucleotides in length [6, 7, 8, 9]. Increasing evidence has reported that aberrantly expressed lncRNAs are associated with the progression of CRC, either play oncogenic roles or develop tumor suppressor functions in the biological processes [10, 11, 12]. For examples, lncRNA ABHD11-AS1 boosts cell proliferation and invasion in CRC through miR-1254-WNT11 pathway [13]; lncRNA SNHG6 regulates EZH2 expression by sponging miR-26ab and miR-214 in CRC [14]; lncRNA BLACAT1 influences cell proliferation by epigenetic silencing of
LncRNA ELFN1 antisense RNA 1 (ELFN1-AS1) is an upregulated lncRNA in CRC in the cancer genome atlas (TCGA) database. Before its official name was assigned, ELFN1-AS1 was studied as UniGene cluster Hs.104073 and UniGene cluster Hs.633957 in several human tumors [16, 17, 18]. Recently, ELFN1-AS1 was identified as a novel primate gene with possible microRNA function expressed mainly in human cancers and predicted to be key lncRNAs in early colon adenocarcinoma (COAD) [19, 20]. However, the detailed role and regulatory mechanism of ELFN1-AS1 in CRC are not known yet.
The purpose of our research is to interrogate the possible function and molecular mechanism underlying ELFN1-AS1 in CRC. In this paper, we firstly tested that ELFN1-AS1 was heightened in CRC and its knockdown hindered cell proliferation and migration as well as activated cell apoptosis of CRC cells. Since that ELFN1-AS1 located in the cytoplasm could mediate TRIM44 expression but not affect the activity of TRIM44 promoter, we further discovered the competing endogenous RNA (ceRNA) network constituted by ELFN1-AS1, miR-4644 and TRIM44, where ELFN1-AS1 targeted miR-4644 to upregulate TRIM44 expression. Rescue experiments confirmed that ELFN1-AS1 promoted CRC cells proliferation and migration via miR-4644/TRIM44 axis.
Materials and methods
Clinical samples
Sixty-two pairs of tumor samples and normal CRC tissues were collected from patients who underwent surgical resection in the Affiliated Hospital of Southwest Medical University. The fresh tissues were immediately frozen in liquid nitrogen and maintained at
Cell culture
Normal colorectal cell (FHC) and CRC cells (HCT116, SW620, HT29, LoVo) were all obtained from the Shanghai Cell Bank of the Chinese Academy of Sciences (China). Cells were all cultivated in RPMI-1640 medium (HyClone, Logan, UT, USA) with 1% penicillin-streptomycin antibiotics (Hyclone) and 10% heat-inactivated fetal bovine serum (FBS; Thermo Fisher Scientific, Waltham, MA, USA). Cells were cultured at 37
Cell transfection
As for transfection, confluent LoVo and HT29 cells at 50–80% were put into 6-well plates. The specific shRNAs of ELFN1-AS1 (shELFN1-AS1#1/2/3) and shNC from Genechem (Shanghai, China) were separately transfected into HT29 and LoVo cells, of which shELFN1-AS1#2 induced the highest interference efficiency and decreased ELFN1-AS1 relative expression from 99.93% to 21.39% on average. shELFN1-AS1#2 displayed the second highest knockdown efficiency, reducing ELFN1-AS1 relative expression from 100% to 20% on average. The mimics and inhibitors of miR-4644 and corresponding NCs were constructed by Genepharma (Shanghai, China). The TRIM44-overexpression vectors (pcDNA3.1/TRIM44, also written as TRIM44) and empty pcDNA3.1 vectors were obtained from Genepharma. Cell transfection which lasted for 48 h was routinely conducted in triplicate via Lipotransfectamine 3000 (Thermo Fisher Scientific).
Quantitative real-time PCR (qRT-PCR)
qRT-PCR was performed to explore the expression of RNA. Total RNA from HT29 or LoVo cells was obtained via TRIzol reagent (Invitrogen, Carlsbad, CA, USA). The reverse transcription of RNA into cDNA was accomplished using PrimeScript RT reagent Kit (Takara, Tokyo, Japan). Subsequently, SYBR Green PCR Master Mix (Roche, Mannheim, Germany) on an ABI7300 real-time PCR machine (Applied Biosystems, Foster City, CA, USA) was carried out. Gene expression was calculated in triplicate by 2
RNA extraction and microarray analysis
Trizol reagent (Invitrogen, Carlsbad, CA, USA) was used to extract total RNA as instructed by the manufacturers. The RNA quality affirmed through formaldehyde agarose gel electrophoresis was quantified by NanoDrop ND-1000. The samples (CRC tissues and non-cancerous tissues) were utilized to synthesize double-stranded cDNA, which was labeled and then hybridized to the LncRNA Expression Microarray (Arraystar 8x60Kv3.0, Rockville, USA). Post hybridization, the arrays were rinsed and the slides were scanned with an Agilent Microarray Scanner (Agilent p/n G2565BA). These data were exported as pair files with the assistance of the Agilent Feature Extraction. The differentially expressed genes were identified by the random variance model and the
Cell counting kit-8 (CCK-8) assay
Briefly, transfected LoVo and HT29 cells at a cell concentration of 2000 per well were seeded into 96-well plates. After 24, 48, 72, 96 h’s incubation, each well was supplemented with CCK-8 solution (Dojindo, Tokyo, Japan) and cultured for extra 4 h. A multi-mode plate reader (Bio-Tek Instruments, Hopkinton, MA, USA) was utilized to measure the absorbance values at 450 nm.
5-ethynyl-2’-deoxyuridine (EdU) staining
The Click-iT Alexa Fluor 488 Imaging Kit (Invitrogen) was utilized in triplicate as the supplier required. Transfected LoVo and HT29 cells were washed twice with 3% bovine serum albumin (BSA; Sigma-Aldrich, Burlington, MA, USA). Then, cells were maintained in EdU-Click reaction-mix and rinsed with 3% BSA. Cell proliferative ability was observed after cell nuclei were stained with DAPI in the dark.
Caspase-3 activity assay
Transfected HT29 or LoVo cells were cultivated with 1 mL PBS (Thermo Fisher Scientific). Cells were centrifuged for 10 min at 4
Transwell assay
After transfection, CRC cells were planted into 24-well plates at a density of 1
Bioinformatics analysis
According to TCGA database (
Luciferase reporter assay
TRIM44 promoter was sub-cloned into downstream of the firefly luciferase gene in the pGL3-basic (Gene-chem). CRC cells were co-transfected with the pGL3-TRIM44 promoter and shELFN1-AS1#2 or shNC to affirm the post-transcriptional regulation of ELFN1-AS1 on TRIM44. The wild-type plasmids of ELFN1-AS1 and TRIM44 (ELFN1-AS1-WT, TRIM44-WT; Genepharma), along with their respective mutant plasmids (ELFN1-AS1-Mut, TRIM44-Mut; Genepharma) were co-transfected with miR-4644 mimics or NC mimics by Lipotransfectamine 3000. After 48 h of incubation, the firefly and Renilla luciferase activities were quantified with Dual Luciferase Report Assay System (Promega, Madison, WI, USA). Experimental results were normalized to Renilla activity and repeated three times.
ELFN1-AS1 silencing inhibited the proliferation and migration and activated the apoptosis in CRC. (A) TCGA result of the markedly elevated expression of ELFN1-AS1 in colon adenocarcinoma (COAD) tissues. (B) The up-regulation of ELFN1-AS1 in CRC cells was detected by qRT-PCR. (C) The inference efficiency of sh-ELFN1-AS1#1/2/3 in LoVo and HT29 cells was determined by qRT-PCR. (D and E) The influences of ELFN1-AS1 deficiency on cell proliferation were examined by CCK-8 and EdU assays. (F) Caspase-3 activity assay was employed for test of cell apoptosis in sh#2 (sh-ELFN1-AS1#2)-transfected cells. (G) Transwell migration assay was adopted for examination of cell migration. 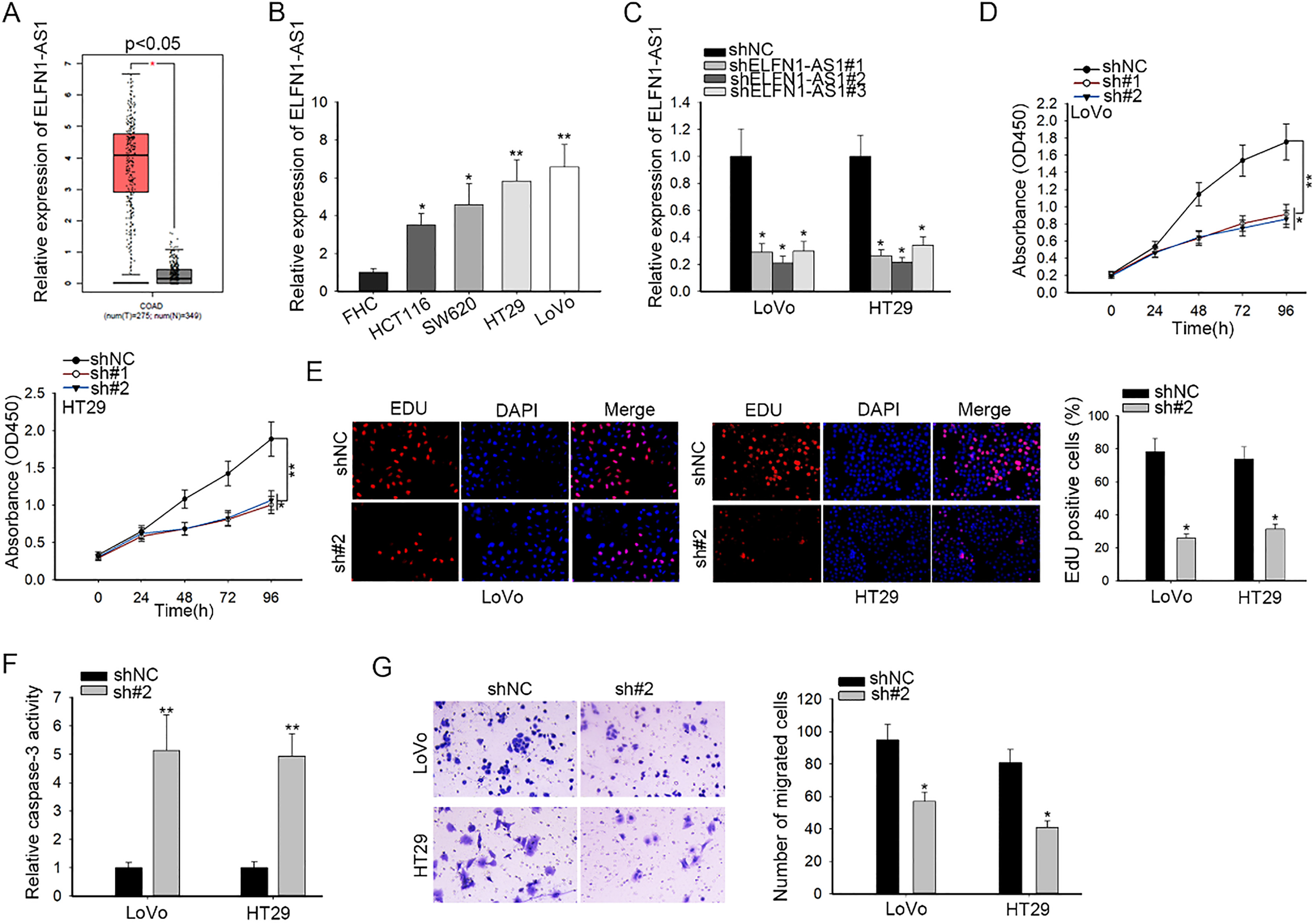
Ago2-RIP assay was carried out via Magna RIP™RNA Binding Protein Immunoprecipitation Kit (Millipore, Bedford, MA, USA) in line with the standard method. Cells at a confluence of 80–90% were cultivated in RIP lysis buffer (Solarbio, Beijing, China) containing magnetic bead conjugated with anti-Ago2 or anti-IgG antibody (Millipore) overnight at 4
RNA fluorescent in-situ hybridization (FISH) Assay
In this experiment, RNAscope Multiplex Fluorescent Reagent Kit v2 (Advanced Cell Diagnostics, Newark, CA, USA) was utilized. Cells were fixed in 4% paraformaldehyde for 30 min, followed by hybridization with the ELFN1-AS1 probe at 40
ELFN1-AS1 regulated TRIM44 expression post-transcriptionally. (A) Five mRNAs containing TRIM44, DPEP1, CDH3, CDK6 and RUNX2 were significantly overexpressed in CRC tissues, as shown in the heatmap. (B) In LoVo and HT29 cells, the impacts of ELFN1-AS1 inhibition on the expression of mRNAs above were affirmed through qRT-PCR. (C) TRIM44 was also heightened in CRC cells, as measured by qRT-PCR. (D) qRT-PCR analysis of TRIM44 expression under repressed ELFN1-AS1 expression. (E) Luciferase reporter validation of the impact of ELFN1-AS1 knockdown on the activity of TRIM44 promoter. (F and G) FISH and nuclear-cytoplasmic fractionation determined the position of ELFN1-AS1 in CRC cells. 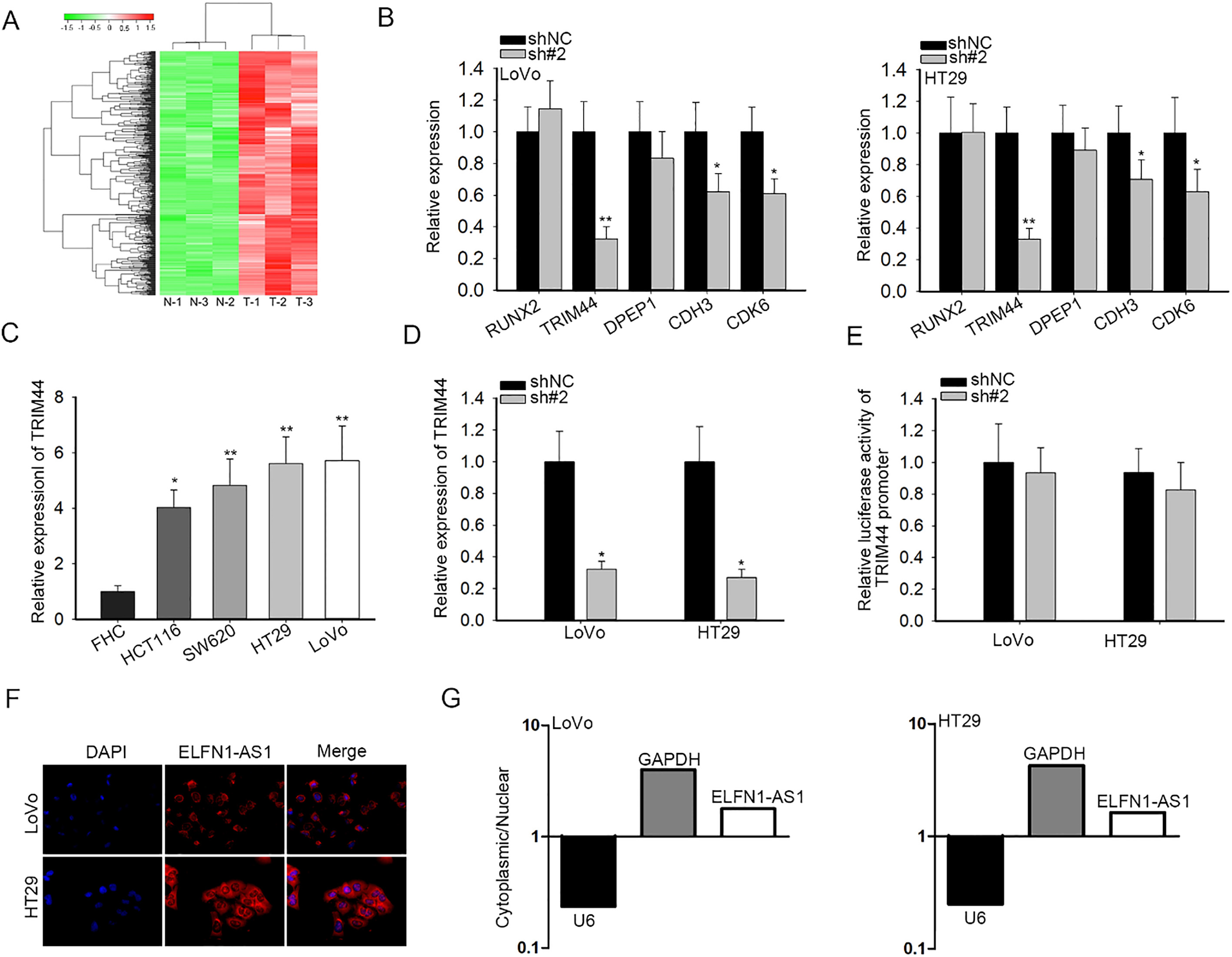
Cells were grown in 10 cm dishes and isolated into two tubes for subcellular fractions and whole-cell extraction. After 5 min of centrifugation at 1000
Western blotting
Cells were lysed on ice for 30 min in RIPA buffer (Sigma-Aldrich). Pierce BCA Protein Assay kit (Ther-mo Fisher Scientific) was utilized to measure the protein concentration based on the specification. Protein samples were separated by SDS-PAGE on a 15% gel and then transferred to PVDF membranes (Millipore). Following blocking with 5% BSA, membranes were treated with anti-TRIM44 antibody (1/2000, ab23 6422, Abcam) and anti-GAPDH antibody (1/1000, ab8245, Abcam) at 4
Statistical analysis
Data from at least three independent experiments were expressed as mean
Results
The silencing of ELFN1-AS1 inhibited the proliferation and migration and activated the apoptosis in CRC
The significantly elevated expression of ELFN1-AS1 in colon adenocarcinoma (COAD) tissues (
ELFN1-AS1 modulated TRIM44 expression at post-transcription level
We employed microarray analysis to analyze the top 500 upregulated genes in CRC tissues, among which TRIM44, DPEP1, CDH3, CDK6 and RUNX2 were the top five mRNAs (Fig. 2A). To validate whether ELFN1-AS1 could regulate the expression of these genes, we performed qRT-PCR assay. And the results discovered that when ELFN1-AS1 was silenced, TRIM44 was the most remarkably downregulated gene (Fig. 2B). Since TRIM44 is an oncogene in multiple cancers, including CRC [21], we further explored whether ELFN1-AS1 influenced CRC progression by TRIM44. We then tested the expression level of TRIM44 by qRT-PCR and the results showed that TRIM44 was distinctly upregulated in CRC tissues (
ELFN1-AS1 sponged miR-4644 to relieve TRIM44
Considering that lncRNAs can regulate some mRNAs expression through targeting their shared miRNAs in ceRNA network [22, 23], we hypothesized that ELFN1-AS1 might target some miRNA to modulate the expression of TRIM44. The Venn diagram was produced by searching the miRNAs binding with ELFN1-AS1 on DIANA tools and the miRNAs targeting TRIM44 on starBase v3.0, which exhibited the only common miRNA of ELFN1-AS1 and TRIM44: hsa-miR-4644 (Fig. 3A). MiR-4644 was selected for further investigation owing to its tumor-suppressor roles in human carcinomas [24]. Subsequently, qRT-PCR results elucidated that miR-4644 was expressed at low levels in CRC tissues (
ELFN1-AS1 sponged miR-4644 to relieve TRIM44. (A) The miRNAs targeting TRIM44 from starBase v.3.0 and the targets of ELFN1-AS1 from DIANA tools were gathered to generate the Venn diagram. (B) qRT-PCR results of the expression of miR-4644 in CRC cells and normal human colorectal FHC cells. (C–E) qRT-PCR was used to evaluate the influences of ELFN1-AS1 silencing on miR-4644 or miR-4644 overexpression on TRIM44. (F) The assumed binging sites between miR-4644 and ELFN1-AS1 or TRIM44 and the corresponding mutant binding sites were displayed. (G) Luciferase reporter assays were performed to examine the effect of miR-4644 promotion on the luciferase activity of ELFN1-AS1-Wt or TRIM44-Wt reporter in LoVo and HT29 cells. (H) The relationship among ELFN1-AS1, miR-4644 and TRIM44 was determined via RIP assay. (I–K) The co-effects of ELFN1-AS1 and miR-4644 on TRIM44 expression in mRNA and protein levels were analyzed through qRT-PCR and western blot. 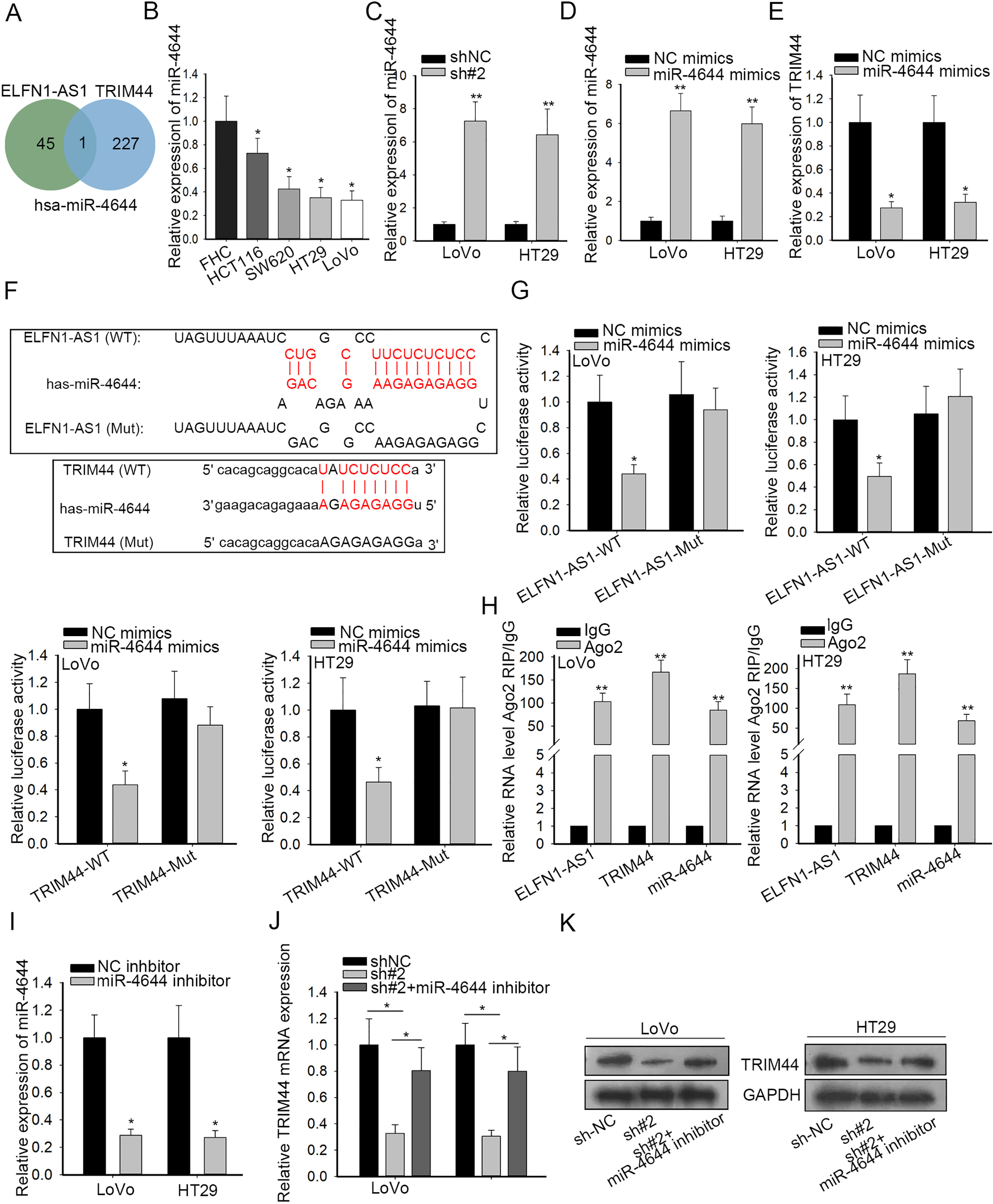
ELFN1-AS1 affected the biological functions of CRC via miR-4644/TRIM44 axis. (A) The co-transfection efficiency in LoVo cells were dissected by qRT-PCR. (B and C) CCK-8 and EdU experiments were performed to examine cell proliferation in different transfected cells. (D and E) Caspase-3 activity and transwell assays were adopted to estimate cell apoptosis and migration, respectively. 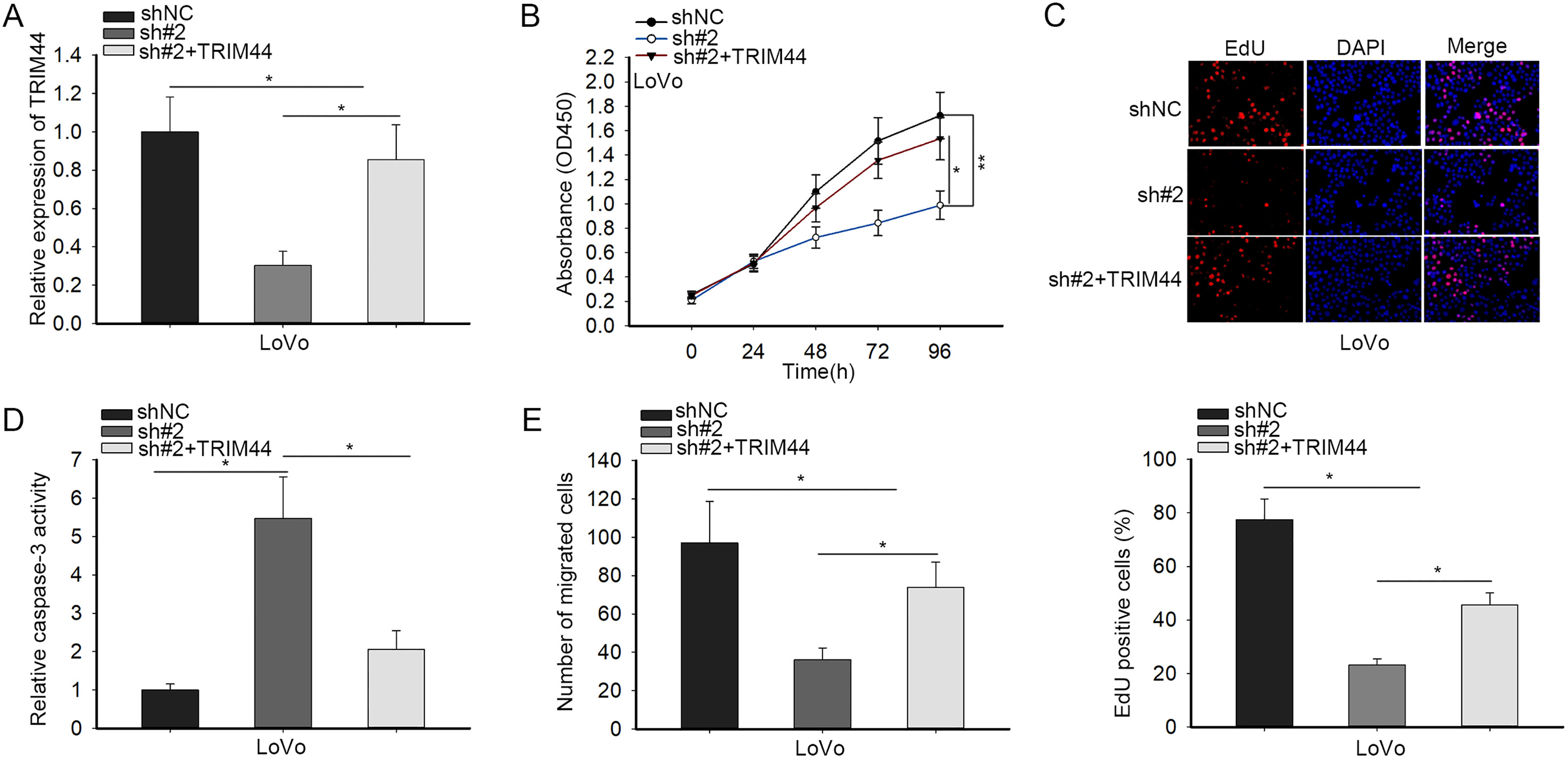
Rescue experiments were carried out to affirm the whole modulation mechanism underlying ELFN1-AS1. LoVo cells were co-transfected with shNC, sh#2 or sh#2
Discussion
The high expression of ELFN1-AS1 in CRC was once revealed but the function role of ELFN1-AS1 is not clearly explained. In this study, we found that ELFN1-AS1 was overexpressed in CRC and ELFN1-AS1 depletion was an obstruction for physiological processes by repressing the proliferation and migration, and boosting the apoptosis of CRC cells. Furthermore, a ceRNA network among ELFN1-AS1, miR-4644 and TRIM44 was demonstrated. ELFN1-AS1 functioning as a ceRNA targeted miR-4644 for the activation of TRIM44. We indicated the existence of ELFN1-AS1/miR-4644/TRIM44 axis in CRC.
By searching TCGA database, we discovered the upregulation of ELFN1-AS1 in colon adenocarcinoma (COAD) tissues and this phenomenon was also validated in CRC tumors and cells, which were in consistent with the data from past researches that ELFN1-AS1 was overexpressed in COAD and possibly participate the pathological activities of CRC cells [19, 20]. To further explore the role of ELFN1-AS1 in CRC, we carried out loss-of-function assay, which, for the first time, revealed that ELFN1-AS1 downregulation dramatically obstructed the cellular processes of CRC cells.
Extensive evidence has proved that long non-coding RNAs (lncRNAs) contribute to physiological activities of multiple carcinomas via various mechanisms, such as ceRNA network, RNA-binding protein (RBP) interactions and epigenetic regulation [25, 26, 27, 28]. In the ceRNA network, lncRNAs sponge miRNAs as decoys to relieve target mRNAs at post-transcriptional level [29, 30]. In this study, we unexpectedly found the post-transcriptional regulation of ELFN1-AS1 on differentially overexpressed tripartite motif containing 44 (TRIM44), a famous oncogene in multiple carcinomas, CRC contained. For instances, elevated TRIM44 boosts the progression of intrahepatic cholangiocarcinoma through inducing EMT by MAPK signaling [31]; LINC00265 accelerates glycolysis and lactate production of CRC by miR-216b-5p/TRIM44 axis [21]; miR-101-3p restrains EMT to weaken proliferation and metastasis in glioblastoma via decreasing TRIM44 [32]. Given that ELFN1-AS1 was detected to be mainly localized in the cytoplasm, we further confirmed the potential ceRNA role of ELFN1-AS1 in CRC. Has-miR-4644, the only one miRNA shared by ELFN1-AS1 and TRIM44, was gained from the Venn diagram produced by venn website. Mechanism experiments validated that ELFN1-AS1 sponged miR-4644 to modulate TRIM44 expression.
In the end, rescue assays were conducted to affirm the whole mechanism underlying ELFN1-AS1 in CRC, which demonstrated that ELFN1-AS1 enhanced cell proliferation, migration and suppressed apoptosis in CRC via miR-4644/TRIM44 axis. These findings suggested that ELFN1-AS1/miR-4644/TRIM44 axis provides a novel insight into the treatment of CRC patients.
Footnotes
Acknowledgments
We thank the contributions from all participators in this research.
Conflict of interest
There are no conflicts of interest.
Supplementary data
PCR primer sequences
Gene name
PCR primers
ELFN1-AS1
F: AGTGAATTCGGGGTGCAGAG
R: TTCCTGGCTGAAGAACCTGC
TRIM44
F: CTGATGAGGAGCAGAAGGCC
R: GCTTGGGCCATCTGAGTCAT
DPEP1
F: TGTGGTGAAGGAGCTGAACC
R: CCGAGGAGTGGCTGAAGATG
CDH3
F: TAGTGGTCCACGTGGAGGAT
R: GTAGACACACACAGGCTCCCA
CDK6
F: ACTATAGATGCGGGCAAGGC
R: TCACACCGAGTAGTGCATCG
RUNX2
F: CTCCAACCCACGAATGCA
R: TAGGTGTGGTAGTGAGTGGTGG
miR-4644
F: TGGAGAGAGAAAAGAGACAGAAGTG
R: CTCTACAGCTATATTGCCAGCCAC
GAPDH
F: CCACCTGGTGCTCAGTGTAG
R: CGTTCAGCTCAGGGATGAC
U6
F: CTCGCTTCGGCAGCACA
R: AACGCTTCACGAATTTGCGT
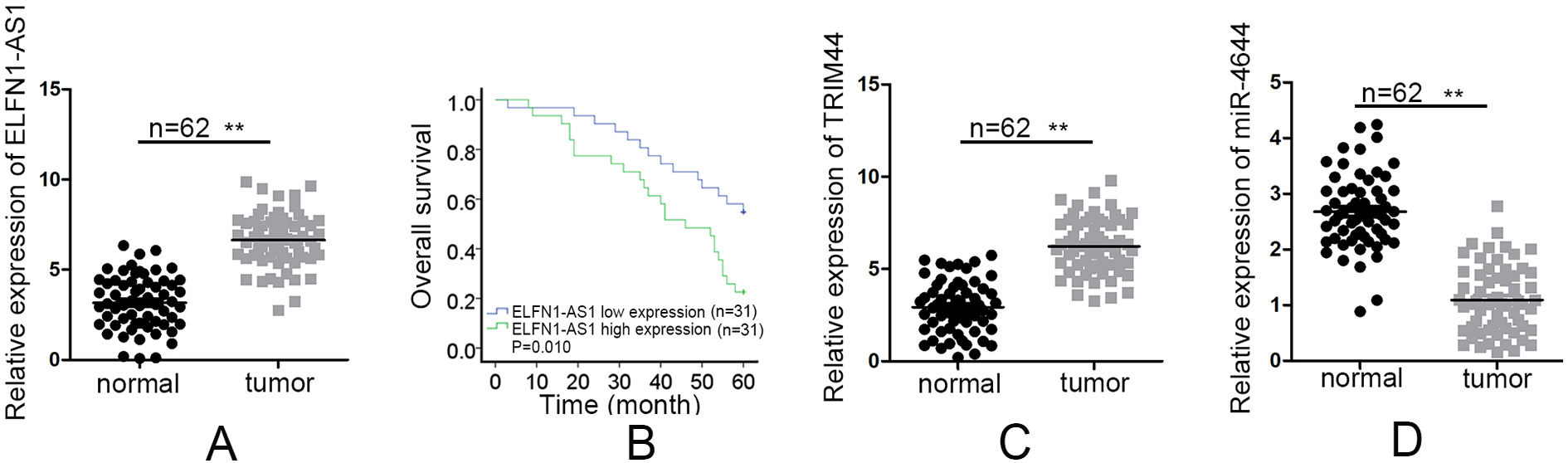
(A) qRT-PCR quantified ELFN1-AS1 expression in CRC tissues (