Abstract
BACKGROUND
: Colorectal cancer is the fourth cause of cancer related death. Drug resistance and toxicity remain major clinical issues. HOTAIR and MALAT1 are long non-coding RNAS that affect cellular proliferation, apoptosis and drug resistance; their up-regulation has been linked with a poor prognosis.
OBJECTIVE:
Investigation of the association between rs4759314 HOTAIR and rs3200401 MALAT1 polymorphisms and irinotecan-based chemotherapy in terms of drug efficacy and toxicity.
METHODS:
Samples from 98 patients receiving different regimens of irinotecan-based therapy were included. Efficacy and toxicity were evaluated. KRAS mutation, rs3200401 HOTAIR and rs4759314 MALAT1 polymorphisms genotyping in the tumors and peripheral blood respectively were performed with PCR.
RESULTS:
Neither rs3200401 MALAT1 nor rs4759314 HOTAIR polymorphism are associated with response to treatment regimens. Rs4759314 was also not associated with increased toxicity in patients receiving irinotecan-based regimens. CT genotype of rs3200401 was associated with significantly reduced overall survival. An association between KRAS mutation and AG/GG genotypes in the rs4759314 was detected.
CONCLUSIONS:
CT genotype of rs3200401 MALAT1 polymorphism could serve as a toxicity biomarker. Carriers of the G allele of the rs4759314 HOTAIR are more likely to be carriers of KRAS mutations too. However, further studies in larger patient populations are required.
Introduction
Colorectal cancer (CRC) represents a major cause of mortality and morbidity globally; CRC is the third most commonly diagnosed malignancy and the fourth cause of cancer-related death [1, 2]. Despite screening efforts, a significant proportion of CRC patients are diagnosed when metastasis is already present. Furthermore, targeted therapies have been developed and progress has been made in the field of cytotoxic drugs. However, drug resistance and the prediction of drug toxicity remain major clinical issues, especially at advanced disease stages [2, 3]. There are several mechanisms that are responsible for drug resistance: limitation of drug transport, dysregulation of important cellular pathways (including apoptotic pathways and DNA repair mechanisms) and epigenetic factors including DNA methylation, histone modification and changes in the expression of non-coding RNAs [4] Research has shown that long non-coding RNAs (lncRNAs) can alter tumor resistance to chemotherapy in many ways, including DNA damage repair mechanisms and inhibition of apoptosis [5]. HOTAIR and MALAT1 are two lncRNAs that have been extensively studied and that are both connected to drug resistance [6, 7]. Polymorphisms of these lncRNAs have also been found to increase gastrointestinal and hematological toxicity in platinum-based chemotherapy [8].
HOTAIR is a newly discovered lncRNA with multiple roles in important aspects of cancer including cellular proliferation, survival, genomic stability and drug resistance. This lncRNA has been connected to several solid malignancies including lung cancer, hepatocellular carcinoma, gastric and colorectal cancer [9]. Recently, a meta-analysis has found a correlation between HOTAIR polymorphisms and cancer risk [10]. In colon cancer, HOTAIR appears to promote oncogenesis and metastasis acting as an oncogene [11]. However, Tian et al. [12], in a meta-analysis reported that no significant associations between the HOTAIR polymorphisms (rs920778, rs4759314 and rs1899663) and cancer risk were observed. This difference may be explained by the fact that at this meta-analysis only one study on colorectal cancer in Asian population was included and it is known that many differences between different ethnic groups exist [13]. Luo et al. observed that tumors with increased HOTAIR expression tend to be chemotherapy-resistant; upregulation of HOTAIR indicates a poor prognosis [14]. Furthermore, three new studies showed that HOTAIR is involved in cisplatin, crizotinib and imatinib resistance in lung adenocarcinoma, in non-small cell lung cancer and in chronic myeloid leukemia respectively [15, 16, 17].
MALAT1 (also known as NEAT2) is a newly discovered lncRNA believed to be a promising marker for the metastasis of lung cancer [4]. Subsequent studies revealed that similarly to HOTAIR, MALAT1 is also involved in proliferation, tumor invasion and apoptosis. Its expression has been found to be upregulated in several malignancies, a fact that also indicates a poor prognosis. MALAT1 emerges as an appealing biomarker for several types of cancer as well as a promising therapeutic target [18]. In a 2012 study, a MALAT1 polymorphism was found to be corelated with CRC [19]. Three more studies showed that MALAT1 is linked to cisplatin, adriamycin and temozolomide resistance in non-small lung cell lung carcinoma, large B cell lymphoma and glioblastoma respectively [20, 21, 22]. MALAT1 upregulation results too in poor response to oxaliplatin (a chemotherapy drug) [23].
In the present study we aimed to investigate the connection between two lncRNA polymorphisms (rs47 59314 HOTAIR and rs3200401 MALAT1) and drug resistance as well as toxicity in two patient groups that were treated with irinotecan-based chemotherapy. Irinotecan is an analogue of camptothecin with anticancer properties. It is a first line antitumor agent used to treat metastatic CRC. However, it is highly toxic and sometimes even life-threatening [24, 25]; thus, approaches for personalized irinotecan doses need to be developed [26, 27]. Currently, therapeutic strategies are based on whether a patient is classified as “fit” or “unfit”, to determine treatment protocol (combination of cytotoxic drugs and/or a biological agent) [28]. Widely accepted and well-established guidelines have recommended mainly oxaliplatin and irinotecan-based regimens: FOLFOX (oxaliplatin, 5-fluorouracil, and leucovorin); FOLFIRI (irinotecan, 5-fluorouracil, and leucovorin); CAPOX (capecitabine and oxaliplatin); CAPIRI (capecitabine and irinotecan); and FOLFOXIRI (infusional5-FU/LV, oxaliplatin and irinotecan) [29]. Continuous infusion of 5-FU added to irinotecan (FOLFIRI) has been shown to be more effective and tolerable than bolus 5-FU. However, this regimen requires either hospitalization or the placement of central venous line. In contrast, the irinotecan-capecitabine combination (XELIRI) appears to be more convenient [30]. Moreover, a recent meta-analysis showed that although triplet chemotherapy (FOLFOXIRI) plus bevacizumab is more effective conversion therapy (converts unresectable metastatic colorectal cancer into resectable) than double chemo- therapy (FOLFOX/FOLFIRI) plus bevacizumab; the latter is associated with a higher risk of neutropenia and diarrhoea [31]. Several targeted therapies that inhibit the angiogenic process (bevacizumab, aflibercept) or the epidermal growth factor receptor (panitumumab, cetuximab) have currently been added to the treatment of mCRC, either as monotherapy or in combination with backbone chemotherapies [28, 29] resulting in further improvements in RR, PFS and OS [32, 33, 34, 35, 36].
Materials and methods
Experimental subjects
CRC adult patients (
Patient characteristics and therapeutic approaches
Patient characteristics and therapeutic approaches
Primer sequences
The new Response Evaluation Criteria in Solid Tumors (RECIST v. 1.1) [38] were used to assess clinical response. This study included patients with measurable disease at baseline. CT and MRI imaging modalities are the best available for measuring response assessment.
The tumor burden was assessed based on target lesions (longest diameter
Toxicity
The evaluation of toxicity was performed in every chemotherapy cycle based on Common Terminology Criteria for Adverse Events (CTCAE) Version 4.0 [39].
Pharmacogenomic analysis
DNA was extracted from frozen blood samples using a DNA extraction kit (Nucleospin Blood, Mache- rey-Nagel, Germany). HOTAIR (rs4759314) genotyping was performed by using allele specific PCR using the KAPA Taq PCR kit, (KAPA Biosystems Pty (Ltd)) with a set of primers for G and A alleles. The same procedure was followed for MALAT1 (rs3200401) for C and T alleles. Electrophoresis was performed on the PCR products using a 2% agarose-98% TAE (1% TAE) gel. All experiments were performed in duplicate and analyzed by two independent observers. Representative PCR products were subjected to sequencing for results verification. The sequences of primers used are shown on Table 2.
KRAS mutation analysis
The analysis was performed on formalin-fixed paraffin embedded tumor tissue removed from each patient. The percentage of tumor cells, in each sample was 30% at minimum. DNA extraction was performed using a commercial kit (NucleospinTissue, Macherey-Nagel, Germany). An enriched polymerase chain reaction followed by restriction fragment length polymorphism (PCR-RFLP) was performed as previously described [40]. Two mutations were detected, the first one at codon 12 and the other at codon 13 of KRAS.
Statistical analysis
Genotype frequencies were compared using the
The rs4759314 and rs3200410 genotypes distributions and the patients’ response status. No significant associations were observed
The rs4759314 and rs3200410 genotypes distributions and the patients’ response status. No significant associations were observed
The rs4759314 and rs3200410 haplotype analysis and the patients’ response status. No significant associations were observed
The rs4759314, and rs3200401 polymorphisms in association with KRAS status
The relationship between the rs4759314, and rs3200401 polymorphisms, and the toxicity status
A total of 98 CRC patients, were included in the analysis and successfully genotyped for HOTAIR (rs4759314) and MALAT1 (rs3200401) polymorphisms. The allelic frequencies observed for both polymorphisms were in accordance with probability limits of Hardy-Weinberg equilibrium. No statistical significance existed between the clinicopathological parameters and the examined polymorphisms. Until August 2018, 4 (4.08%) patients were characterized as CR, 30 (30.6%) as PR, 32 (32.6%) as SD, and 32 (32.6%) as PD. Regarding KRAS mutation status, 40 of all patients tested were carriers of codon 12 mutated allele, and 5 were carriers of the codon 13 mutated allele. We did not observe statistically significant differences between carriers of KRAS mutated alleles between CR, PR, SD, and PD patients’ group. As illustrated in Tables 3 and 4, no significant difference was found between response rates and both rs4759314 and rs3200401 genotypes and haplotypes. Additionally, as indicated in Table 5 KRAS status is statistically correlated with the carriers of rs4759314 AG and GG genotypes, whereas is not correlated with the host of rs3200401 genotypes.
The most frequent major toxicities observed were nausea (29 patients) and neutropenia (20 patients). Our findings did not indicate significant differences between the toxicity profile of the treatment combinations used. No statistically significant difference was observed between toxicity and rs4759314 genotypes. However, as indicated in Table 6 a significant association (
The relationship between the rs4759314, and rs3200401 haplotypes, and the toxicity status
The relationship between the rs4759314, and rs3200401 haplotypes, and the toxicity status
Rs4759314, and rs3200401 genotypes in association with overall survival
Table 8 shows the association of rs4759314 and rs3200401 genotypes with overall survival. During the study period, there were 60 deaths among subjects who were included in the study. The survival was calculated from the date of the first treatment with irinotecan-based combinations until death from any cause. Regarding the rs4759314 polymorphism no significant association with survival was found, whereas the rs3200401 CT
Association of overall survival with genotypes of HOTAIR rs4759314 (A) and MALAT1 rs3200401 (B) polymorphisms. Kaplan-Mayer curve represents the percentage of survival in months of patient’s different genotypes of the rs1561927 and rs3200401 polymorphisms.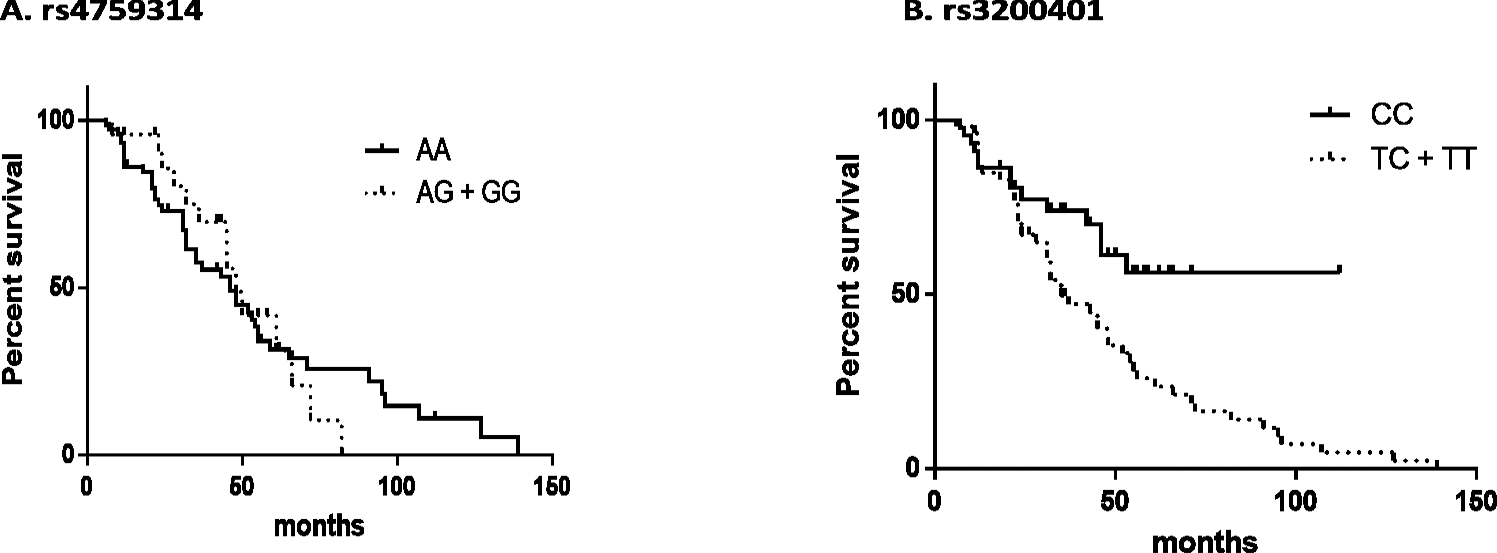
Irinotecan is currently one of most frequently prescribed anticancer drugs used in the mCRC treatment as monotherapy or in chemotherapy schemes (including FOLFIRI and XELIRI), which have significantly improved RR, PFS and OS [41, 42, 43, 44, 45]. However, irinotecan exhibits dose dependant and wide interindividual toxicity which is associated with the patient’s genetic profile [3, 24, 25, 26, 27]; genetic factors are investigated towards their role in the therapeutic outcome and in drug toxicity of irinotecan-based regimens [46, 47, 48, 49, 50].
The role of lncRNAs is also investigated. Li et al., demonstrated that HOTAIR contributes to 5-FU resistance and that high HOTAIR expression was corelated with shorter OS and RFS in CRC patients receiving 5FU-based therapy [51]. Until now, no previous study has investigated the correlation between HOTAIR expression or the polymorphism under study and irinotecan resistance.
Polymorphisms of MALAT1, which is also linked to cancer progression [52, 53], are investigated for the development of several clinical manifestations. A recent study showed that the rs3200401 CT/TT genotypes exhibited significantly lower levels of total cholesterol [54] and rs3200401 CT and CT/TT genotypes in MALAT1 had better prognosis than rs3200401 CC genotype, among lung cancer patients [55]. On the contrary, it has been suggested that carriers of the heterozygous rs3200401 CT genotype are more susceptible to Pb exposure than subjects with homozygous CC genotype, indicating that MALAT1 SNPs may play a critical role in Pb derived toxicity [56]. Given that cellular levels of MALAT1 were found to be increased during G1/S and M phases of the cell cycle suppressing apoptosis and/or increased DNA repair efficiency [57] and that irinotecan acts via inhibiting DNA topoisomerase I this could explain tumour resistance to irinotecan. Direct associations between MALAT1 SNPs and irinotecan resistance have not been investigated in previous studies.
Taken these into consideration, 98 samples were genotyped from patients with incurable, mCRC and the connection between germline polymorphisms in HOTAIR and MALAT1 and response to irinotecan-based regimens was investigated. More specifically, the impact of rs4759314 HOTAIR and rs3200401 MALAT1 polymorphisms were evaluated regarding clinical response and toxicity on irinotecan-based schemes. We found that both rs4759314 HOTAIR and rs3200401 MALAT1 polymorphisms were not associated with response to treatment regimens. Interestingly, although MALAT1 expression is associated with poor re- sponse to oxaliplatin-based regimens and chemore-sistance [23], our findings indicated that the SNP rs3200401 does not affect the response to irinotecan-based schemes. The second polymorphism of our study, rs4759314 HOTAIR, also showed no correlation with irinotecan induced toxicity, a finding which agrees with Gonk et al. who also observed no association between rs4759314 HOTAIR and platinum-based toxicity in Chinese patients with lung cancer [8].
RAS mutations have been implicated in CRC metastasis, and it has been reported that they are associated with increased risk of lung and ovarian metastases in patients treated with irinotecan-based regimens [53]. In our study, no significant association between KRAS mutation status and response to treatment was detected. This finding is in accordance with data supporting that KRAS mutations are not considered as independent predictors of survival and only a sub-group of patients benefit from drugs targeting EGRF, due to the complex aetiology of colorectal cancer and the resistance to these therapeutic agents [58, 59]. However, a statistically significant correlation between patients with AG/GG genotypes in the rs4759314 HOTAIR and KRAS mutation was detected, suggesting that carriers of the mutated rs4759314 HOTAIR G allele is more likely to be carriers of KRAS mutations too. This is the first study to have found this correlation.
Based on our data, there was a statistical significance between the CT/TT genotype in the rs3200401 MALAT1 and increased toxicity to irinotecan-based regimens; this finding was not associated with a better response to treatment, making it applicable as an independent toxicity biomarker for patients receiving irinotecan-based regimens. Our results are following Qian et al. observation that rs3200401 CT genotype is more susceptible to toxicity development [56]. Qian et al. suggested that the heterozygous CT genotype of MALAT1 rs3200401 polymorphism may influence lead-induced neurotoxicity via the MAPK or the P53 signalling pathway. However, it should be noted that irinotecan does not exhibit significant neurotoxic effects. In fact, only very few cases have been reported [60] and the exact cause of irinotecan-induced CNS toxicity remains unknown. Therefore, further studies are needed to clarify the role of this specific SNP on irinotecan toxicity mechanisms. To the best of our knowledge, this is the first study to investigate the association of this MALAT1 polymorphism and resistance or toxicity in irinotecan-based treatment of CRC.
Moreover, our results demonstrate that the rs3200401 CT
However, our results need to be validated in future studies so as for larger populations of metastatic colorectal cancer to be included. Additionally, it should be noted that the precise biological function and mechanisms of the lncRNA SNPs in toxicity development and response remain unknown and need further investigation. We should also reckon that irinotecan-based chemotherapy toxicities may result from multiple genetic factors. Therefore, apart from the identification of novel biomarkers, multilevel studies need to be conducted, in order to better understand both the response and toxicity development in patients treated with irinotecan-based regimens.
In conclusion, our study provides the first evidence of the association between lncRNA MALAT1 rs3200401 polymorphism, toxicity development and effect in OS in mCRC patients treated with irinotecan-based regimens. Additionally, gene expression profiling analysis revealed that carriers of the mutated G allele of rs4759314 HOTAIR is more likely to be also carriers of KRAS mutations.
Footnotes
Acknowledgments
This study was supported by a non-profit organization of Greek Society of Cancer Biomarkers and Targeted Therapy.
