Abstract
Defensin alpha 6 (DEFA6) is a member of the alpha defensin family of microbicidal and cytotoxic peptides that defend against bacteria and viruses. Here, we provide a novel function of DEFA6 in tumorigenesis of colorectal cancer (CRC) in vitro and in vivo. Specifically, DEFA6 is highly expressed in both CRC cancer cell lines as well as patient-derived samples at the level of RNA and protein. By shRNA-mediated loss of function of DEFA6, we found that proliferation, migration, invasion, colony forming ability of CRC cell lines were impaired in the absence of DEFA6 in vitro. Furthermore, DEFA6-deficient cancer cells exhibited significantly reduced growth rates compared to control cells in vivo. More importantly, by analyzing 352 patient-derived samples, we revealed that DEFA6 is associated with overall survival rate of CRC patients and thus an independent prognostic marker for CRC. These results suggest that DEFA6 plays an essential oncogenic role in CRC and serves a good therapeutic target for the disease.
Keywords
Introduction
Primers used in this study
Primers used in this study
Colorectal cancer (CRC) is one of the most common cancers worldwide [1]. There has been a significant increase in reported incidence rates over the last decade, especially in the Republic of Korea [2]. Despite the identification and development of new anticancer agents and therapeutic regimens, CRC is still a relatively aggressive malignancy with a high level of recurrence and metastasis. In particular, survival rates of patients with CRC are highly dependent on the occurrence of distant metastases [3]. Thus, predicting prognoses is very important to establish efficient therapeutic plans for patients with CRC. Nowadays, there are various CRC screening tests, including non-invasive stool or blood tests and more invasive imaging or colonoscopy [4]. Fecal tests, which includes fecal occult blood tests (FOBTs), fecal immunochemical tests (FIT), DNA-, RNA-, and protein-based biomarker tests, and microbiome-based biomarkers, are varied in sensitivity, but rapidly increasing [4]. The correlation between gut microbiota and CRC is also gaining a strong interest [5]. TNM (tumor/node/metastasis) classification and staging system can aid in forming a prognosis for patients with CRC, but it can be still difficult to predict prognoses precisely [6]. Thus, it has been suggested that the discovery and application of more sensitive and specific biomarkers could improve outcomes for patients with CRC [7]. Despite extensive research, few useful biomarkers used to predict a patient outcome have been identified until yet.
Defensin alpha 6 (DEFA6) is expressed in the lysozyme-rich granules of Paneth cells found in the small intestine [8]. It is a member of the alpha defensin family of microbicidal and cytotoxic peptides that defend against bacteria and viruses. Although DEFA6 has a poor antibacterial potency, it does offer some protection against invasion by enteric pathogens through the self-assembly of fibrils and nanonets that surround and entangle bacteria [9]. As the primary role of DEFA6 is defending the intestine, high DEFA6 expression is typically found in chronic inflammatory conditions of the colon. However, unexpectedly, there have been some reports showing the association of defensin with tumors. For example, DEFA6 is shown to be involved in colonic adenomas [10], and several defensins were detected in epithelial tumors [11, 12, 13]. Further, it has been shown that human neutrophil peptides (HNP1-3), known as
Cell culture
Human colorectal cancer cell lines (HCT116, SW480, and SW620) were obtained from the Korean Cell Line Bank (Seoul, Korea). Cells were cultured in RPMI1640 medium supplemented with 10% FBS and 1% of penicillin-streptomycin at 37
shRNA transfection and analysis of mRNA and protein expression
CRC cells were grown to 60–70% confluence in RPMI1640 medium (contained 10% FBS) before being transduced with DEFA6-specific short hairpin RNA (shRNA) lentiviral particles (Origene, MD, USA). shRNA transduction was performed according to the manufacturer’s instructions. After transduction, transduced cells were selected with 2
Total RNA was purified from control and transfected cell samples using a Hybrid-R
Clinicopathological features in patients with DEFA6 expression
Clinicopathological features in patients with DEFA6 expression
shRNA-mediated DEFA6 depletion in colorectal cancer cell lines. A and B. Colorectal cancer cell lines including HCT116, SW480 and SW620 were transduced with either control or DEFA6 shRNA. RNA and proteins were isolated to determine the expression of DEFA6. A. RT-PCR was performed to determine DEFA6 mRNA expression in cancer cell lines. B. Immunoblot with anti-DEFA6 antibody was conducted to analyze DEFA6 protein level. GAPDH was used as the loading control for RT-PCR and immunoblot (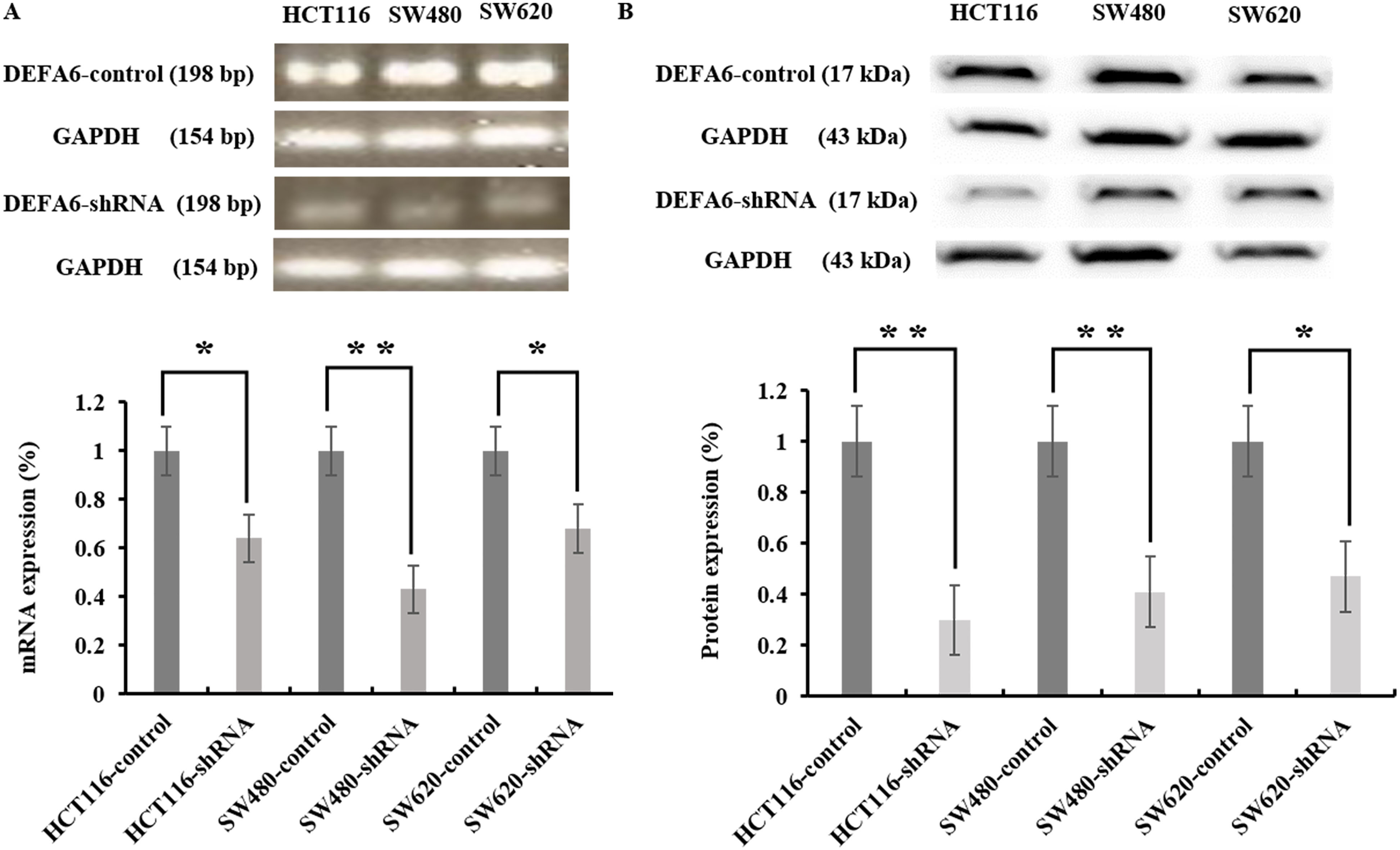
Protein expression levels were assessed by immunoblot analysis of cells lysates prepared using Pro-Prep
Untransfected control and DEFA6 shRNA transfected cells were seeded onto 96-well plates. After 24, 48, and 72 h of culture, 10
Migration and invasion assays
For migration and invasion assays, the capacity for either untransfected control cells or DEFA6 downregulated cells to migrate through a 24-well Transwell chamber (Corning, NY, USA) was assessed. For these assays, 2.5
Immunohistochemistry for DEFA6 expression in colorectal cancer tissue. A–D. Three hundred and fifty-two colorectal cancer samples were stained with DEFA6 antibody and graded based on both staining intensity and staining frequency. Representative images of tissue microarray and immunohistochemistry are shown (A, Negative DEFA6 expression; B, Weak; C, moderate; D, Strong). Original magnification 
A soft agar colony-forming assay using 6-well plates was used to assess anchorage-independent cell growth. Untransfected control cells or DEFA6 knockdown cells (2.5
Tumorigenicity in BALB/c nude mice
Untransfected control CRC cells (2
Tissue specimens and immunohistochemistry (IHC)
Colorectal cancer tissues were obtained from 352 patients who were diagnosed and underwent subsequent surgery at the Soonchunhyng University Cheonan Hospital, Korea. This study was conducted in accordance with the ethical standard of Ethics Committee of Soonchunhyang University Cheonan Hospital. An overview of patient information is shown in Table 2. Collected tissues were embedded in paraffin and cut into 4
DEFA6 depletion impairs cell proliferation in colorectal cancer cell lines. A–C. Control or DEFA6-depleted (shRNA) colorectal cancer cell lines such as HCT116, SW480, and SW620 were analyzed every 24 hours for 3 days to determine the proliferation rate by MTT assay. A. HCT116 cells (48 h; 
DEFA6 depletion impairs migration and invasion of colorectal cancer cell lines in vitro. A–D. Control or DEFA6-depleted (shRNA) colorectal cancer cell lines were seeded on a matrigel-uncoated and coated transwell, followed by incubation for 24 and 48 hrs for assessing migration and invasion, respectively. Cells that had migrated to the lower surface of the transwell were stained and quantified. A and B. Imaging was done using an inverted microscope and representative images are shown. DEFA6-depleted cells (shRNA) displayed significantly reduced ability of migration (C. HCT116; 
The results from each experiment were analyzed using SPSS version 18.0 (SPSS Inc., Chicago, IL, USA). The relationships between DEFA6 expression and various clinicopathological factors were evaluated with
Results
DEFA6 is highly expressed in both colorectal cancer cell lines and tissues, and associated with clinicopathological features
To determine the expression pattern of DEFA6 by RT-PCR and western blotting, we used RNA and protein obtained from CRC cell lines. We found that both the RNA and protein level of DEFA6 are highly expressed in CRC cell lines (Fig. 1A and B). This was confirmed by immunohistochemistry (IHC) analysis of patients-derived colorectal cancer tissues with DEFA6 protein being expressed in the cytoplasm of cancer cells. We classified patients into two groups based on their DEFA6 expression score, either a low expression group (Fig. 2A and B) or a high expression group (Fig. 2C and D). One-hundred eighty-one of the 352 patients with CRC (51.4%) showed a high level of DEFA6 protein expression, while 171 patients (48.6%) displayed a low level of DEFA6. We next analyzed the relationship between DEFA6 expression and various clinicopathological parameters in patients with CRC (Table 2). This demonstrated that DEFA6 expression was significantly associated with pN stage (
DEFA6 is essential for the progression of CRC in vitro and in vivo
As we found that DEFA6 is highly expressed in both CRC cell lines and patient-derived tissues, we reasoned that DEFA6 may be involved in the progression of CRC. To understand a progressive role of DEFA6 in patients with CRC, we assessed the functional importance of DEFA6 in various CRC cell lines. After shRNA-mediated knocking down of DEFA6, DEFA6 mRNA and DEFA6 protein levels were verified by RT-PCR and immunoblotting, respectively. As shown in Fig. 1, DEFA6 was significantly knocked down in DEFA6 shRNA-transduced CRC cells compared to untreated control cells (each cell type comparison,
The results presented in Fig. 3 demonstrate that the proliferation of DEFA6 shRNA-transduced cell line, SW480, was significantly decreased 24 h after DEFA6 knockdown (
To investigate the function of DEFA6 in cancer progression, we performed Transwell migration and invasion assays with a selection of CRC control cells and transfectants (HCT116, SW480, and SW620). These results indicated that the migration of DEFA6 knockdown cells was significantly lower than that of control cells (HCT116,
To determine if DEFA6 has a role in tumorigenicity in vitro, soft agar colony formation assays were performed (Fig. 5). Both control cells and shRNA-mediated DEFA6 knockdown cells were plated in soft agar, and colony formation was scored after 15 days of incubation. Cells with shRNA-mediated knockdown of DEFA6 formed significantly fewer colonies compared to control cells (HCT116, 324
Finally, to establish if DEFA6 is involved in CRC tumor growth in vivo, control cells and shRNA-mediated DEFA6 knockdown cells were subcutaneously inoculated into athymic nude mice. Tumors induced by shRNA-mediated DEFA6 knockdown cells exhibited reduced growth rates (5 weeks, 65.6%,
Univariate and multivariate Cox regression analysis of the relative risk of death according to DEFA6
Univariate and multivariate Cox regression analysis of the relative risk of death according to DEFA6
DEFA6 depletion impairs anchorage-independent colony forming ability of colorectal cancer cell lines in vitro. A and B. Control or DEFA6-depleted (shRNA) colorectal cancer cell lines (HCT116, SW480, SW620) were seeded on semi-solid agarose, followed by incubation for 15 days to assess colony forming ability. A. Representative images of colonies were shown. B. Number of colonies from each cell lines were quantified (HCT116; 
DEFA6 depletion impairs the propagation of colorectal cancer cells in vivo. A–D. Control or shRNA-mediated DEFA6 knockdowned HCT116 cells were subcutaneously inoculated into athymic nude mice and tumor size was measured every 7 days. A. Representative images of tumor size are shown. B. Tumor size was determined up to 7 weeks (5 weeks, 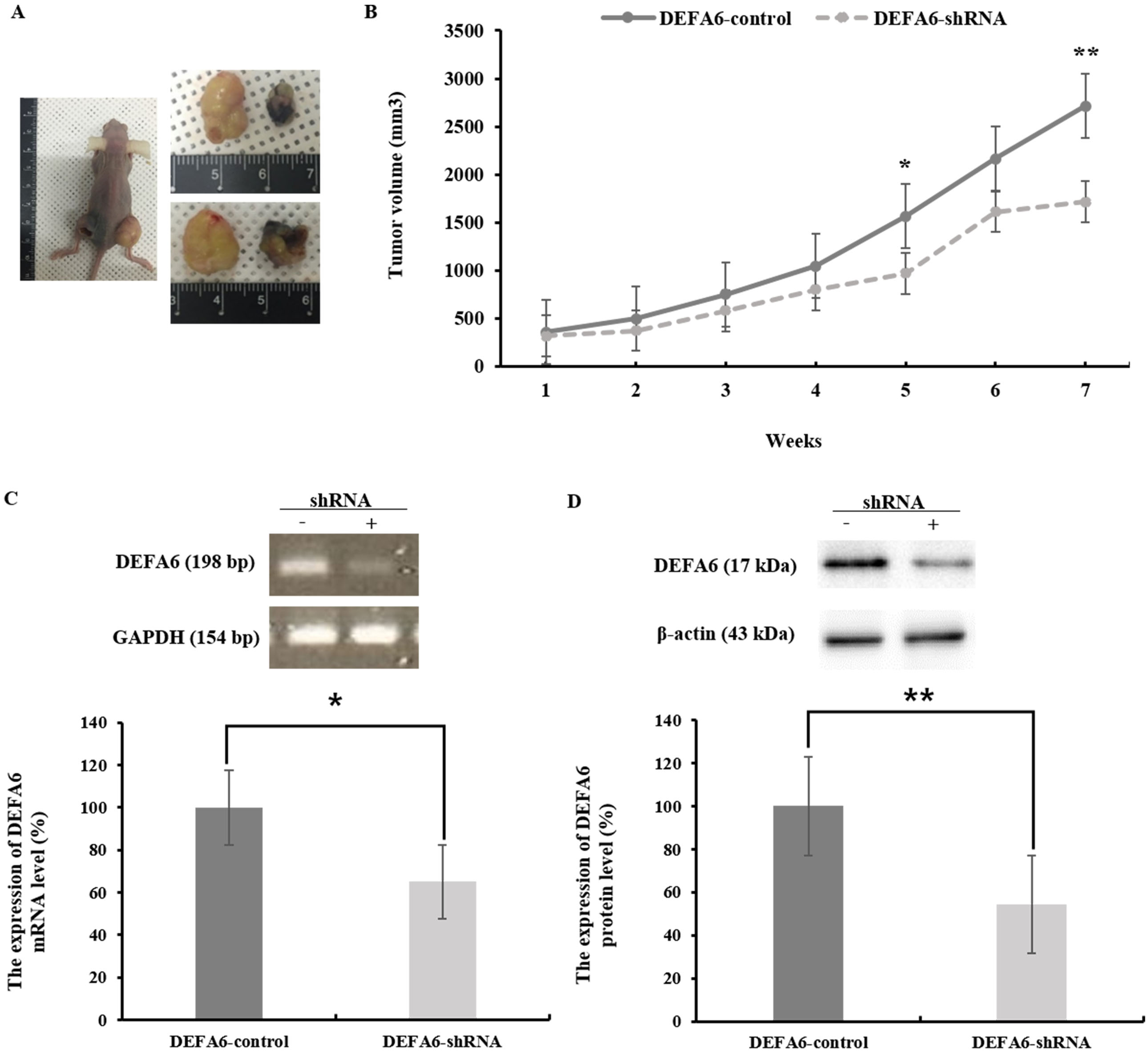
After in vitro and in vivo confirmation of the aggressiveness and tumor-growth effects of DEFA6 using CRC cell lines and a mouse model, we next investigated the correlation between DEFA6 expression and patient outcome to check the relevance of DEFA6 in patient-derived samples. To assess the prognostic significance of DEFA6 expression, we used Kaplan-Meier survival curves and found a significant positive correlation between overall survival and DEFA6 expression levels (log-rank test,
DEFA6 expression is associated with a poor prognosis of colorectal cancer patients. Three hundred and fifty-two colorectal cancer samples were stained with DEFA6 antibody and graded based on both staining intensity and staining frequency. Survival rate was determined based on the expression of DEFA6 expression using Kaplan-Meier method (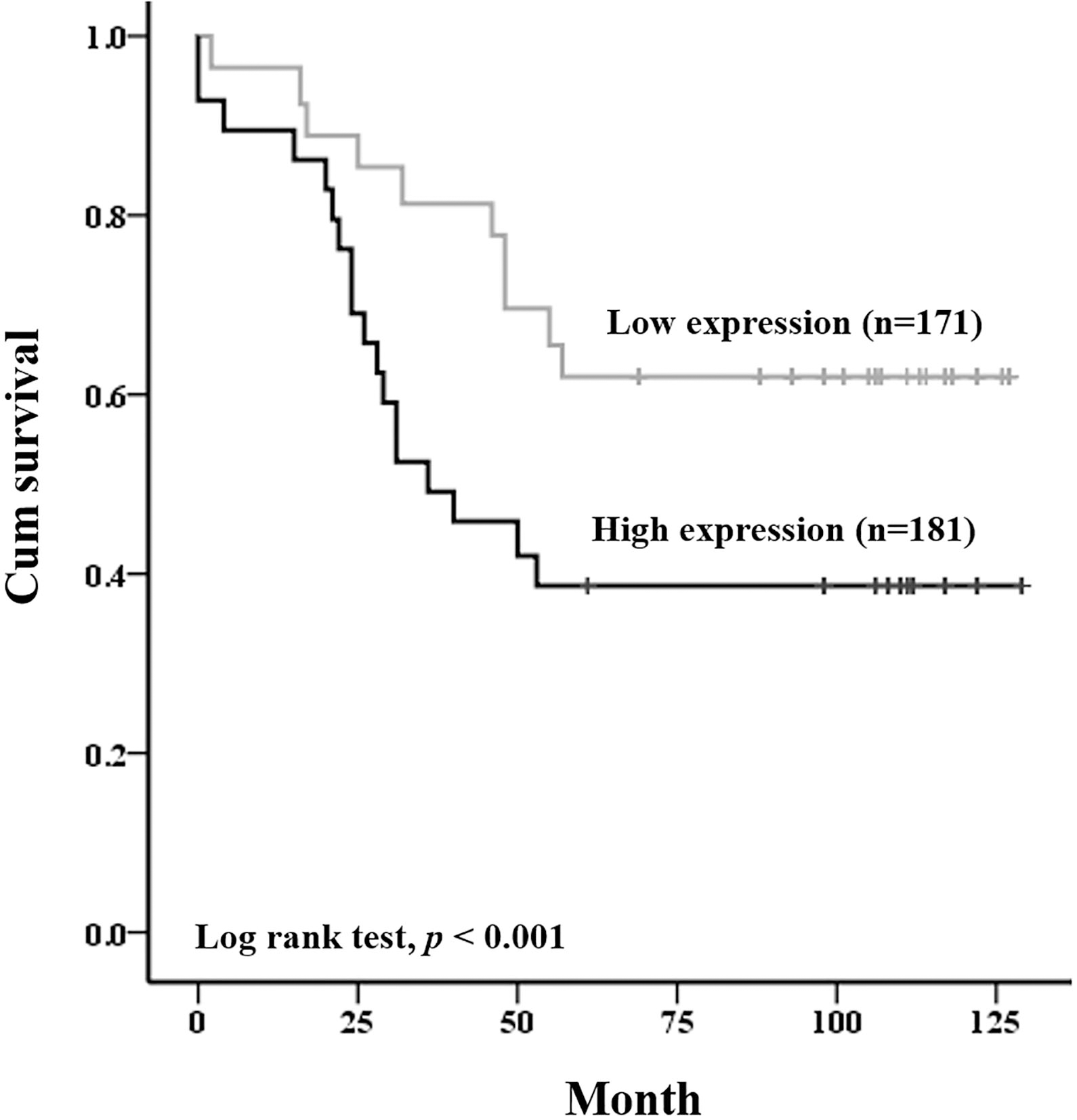
Univariate Cox-regression analysis (Table 3) revealed that vascular invasion (Hazard Ratio [HR]
Previously, we showed the correlation between DEFA6 and patient prognoses using a small number of patient-derived samples [17]. Here, using CRC cancer cell lines as well as IHC in a CRC-TMA dataset containing 352 CRC specimens with complete follow-up data, we confirmed and expanded the previous data to show that DEFA6 is essential and a novel prognosis marker in CRC. In human CRC cell lines such as HCT116, SW480 and SW620, we showed that DEFA6 is highly expressed. shRNA-mediated DEFA6 knockdown significantly inhibited the proliferation, invasion, migration, and even anchorage-independent colony formation of these CRC cell lines compared to controls. In addition, knockdown of DEFA6 in HCT116 inhibited tumorigenesis in vivo. High DEFA6 expression was identified in 51.4% (181/352) of primary CRC tissue samples and was shown to significantly correlate with pT stage (
DEFA6 is a member of the alpha defensin protein family that encompasses six known members, human neutrophil peptide 1-4 (HNP1-4), human alpha defensin 5 (DEFA-5), and human alpha defensin 6 (DEFA6) [18]. Both DEFA5 and DEFA6 are expressed by Paneth cells in the small intestine and are known to primarily act via the innate immune system [18]. Among the alpha defensins, HNP1-3 are reported to associate with various tumors [19, 20, 21, 14, 22, 23] and their expression is increased in CRC tumors and the plasma of patients. They are also described as biochemical markers for metastatic CRC [24]. Compared to the other alpha defensins, DEFA6 possesses a slightly different structure and function in that it has less basic residues and lacks antimicrobial properties [25]. There have been a limited number of studies linking DEFA5 and DEFA6 to human tumors. For example, Nomura et al. [22] reported that DEFA5 decreased E-cadherin (CDH1) expression in gastroesophageal junctions, accelerating development of Barrett’s esophagus (BE) and promoting adenocarcinoma. Radeva et al. [10] reported that high DEFA6 expression occurs in adenoma (approximately 60-fold compared to normal tissues) and can be used to distinguish between benign adenomas or carcinomas in the colon. Andreu et al. also reported that DEFA6 mRNA expression was elevated in tumors in which there was an accumulation of Myc (MYC) and Cyclin D (CCND1) [26]. Myc and Cyclin D are both well-known cellular proliferation markers and downstream targets of the Wnt signaling pathway, an important CRC carcinogenic pathway. Based on these reports, it is therefore not surprising to predict that DEFA6 may play a role in CRC tumorigenesis. However, the functional importance of DEFA6 has not been released yet except our study. The exact pathogenetic molecular mechanisms that underlie DEFA6-mediated tumorigenesis remain to be elucidated though.
In summary, our study is the first report to demonstrate that DEFA6 plays an oncogenic role in tumorigenesis of CRC in vivo and in vitro. Furthermore, DEFA6 expression levels are an independent prognostic factor for CRC and may also serve as a future therapeutic target.
Footnotes
Acknowledgments
The present study was supported by the grant of the Korea Health Technology R&D Project through the Korea Health Industry Development Institute (KHID), funded by the Ministry of Health and Welfare, Republic of Korea (grant number: HI17C0031).
Conflict of interest
The authors declare no conflict of interest.
