Abstract
BACKGROUND:
Circular RNAs (circRNA) play key regulatory roles in cancer progression. Human circRNA microarray was performed to screen for abnormally expressed circRNA in gastric cancer tissues. In this study, we found circRNA102958 was up-regulated in gastric cancer tissues and cell lines.
METHODS:
The hsa_circRNA_102958 levels were detected by quantitative reverse transcription-polymerase chain reaction (qRT-PCR) in gastric tissue and cell. Then, the association between the expression level of hsa_circRNA_102958 and the clinicopathological features of patients with gastric cancer was further analyzed. Finally, a network of hsa_circRNA_102958-miRNA-mRNA interactions was predicated.
RESULTS:
In this study, we analyzed 30 patients and found that hsa_circRNA_102958 was significantly overexpressed in gastric cancer tissues compared with paired adjacent normal tissues. Clinicopathological features showed that hsa_circRNA_102958 level in GC tissues was positively associated with TNM stage (
CONCLUSIONS:
hsa_circRNA_102958 may be a potential novel and stable biomarker for the diagnosis of gastric carcinoma.
Introduction
Gastric cancer (GC) is a leading cause of tumor-associated morbidity and mortality worldwide. Although the treatment of gastric cancer has gradually improved, a poor prognosis for GC patients due to tumor metastasis and recurrence [1, 2]. One important reason is lack of effective molecular targets for diagnosis and treatment.
Recent years, circular RNA (circRNA), rather than linear RNA molecules, play key regulatory roles in various cellular physiological processes and may contribute to pathological processes such as cancer formation and progression [3]. CircRNAs as a novel type of ncRNAs derived from exons, introns or intergenic regions have a covalently closed continuous loop, display cell or tissue specific expression and are conserved across species due to resistance to RNase R [4, 5], The expression of circRNAs is highly stable in comparison with their linear counterparts, and is predominantly localized in the cytoplasm, indicating important functions for circRNAs in human diseases [5, 6]. Emerging evidence shows that some circRNAs as miRNA sponges modulate gene transcription and interact with RNA binding proteins (RBPs) involved in tumorigenesis [7].
In this study, we focused on a circRNA, hsa_circ_ RNA102958, from tissue microarrays. Based on the microarray findings, a series of clinical validation experiments were performed to explore the role of hsa_circ_RNA102958 in gastric cancer from tissue and cell levels, respectively. In addition, a prediction and annotation of hsa_circ_RNA102958 molecular function with was miRNAs or mRNAs was performed on relevant database.
Materials and methods
Patients and sample collection
A total of 30 GC tissues and paired adjacent nontumorous tissues were collected in this study during 2017–2018 at Jiangsu Cancer Hosptital affiliated with the Nanjing Medical University. None of these patients had received adjuvant chemotherapy or radiotherapy before surgery. All specimens collection and preprocessed were according to previously described protocol and preserved at
Cell culture procedures
Normal human gastric epithelial cell line GES-1 and GC cell lines (SGC-7901, MKN-45, MGC- 803, AGS, BGC-823) were from Shanghai Institutes for Biological Sciences, Chinese Academy of Sciences(Shanghai, China). GES1 and SGC7901 cells were cultured in Dulbecco’s Modified Eagle medium (DMEM) medium supplemented with 10% heat-inactivated fetal bovine serum (FBS), 100 U/ml of penicillin, and 100
Total RNA extraction
Total tissue RNA were extracted from cell lines and paired tissues using TRIzol reagent (Invitrogen, Karlsruhe, Germany), according to the manufacturer’s instructions. The purity and concentration of RNA samples were determined with the NanoDrop ND-1000 (Thermo Fisher Scientific, Wilmington, DE). RNA was prepared and stored at
circRNA microarray analysis
Expression profile of circRNAs in GC tissue, as determined by microarray analysis. (A) Unsupervised hierarchical cluster analysis, based on the expression levels of circRNAs. CircRNA microarray expression profiles from three sets of matched GC tissue samples (C3-C5) and adjacent normal tissues samples (N3-N5). CircRNAs in red indicate overexpression; those in green indicate reduced expression. (B) Volcano plot analysis based on circRNA expression levels. The vertical green lines correspond to 1.5-fold increased or decreased expression, and the horizontal green line represents 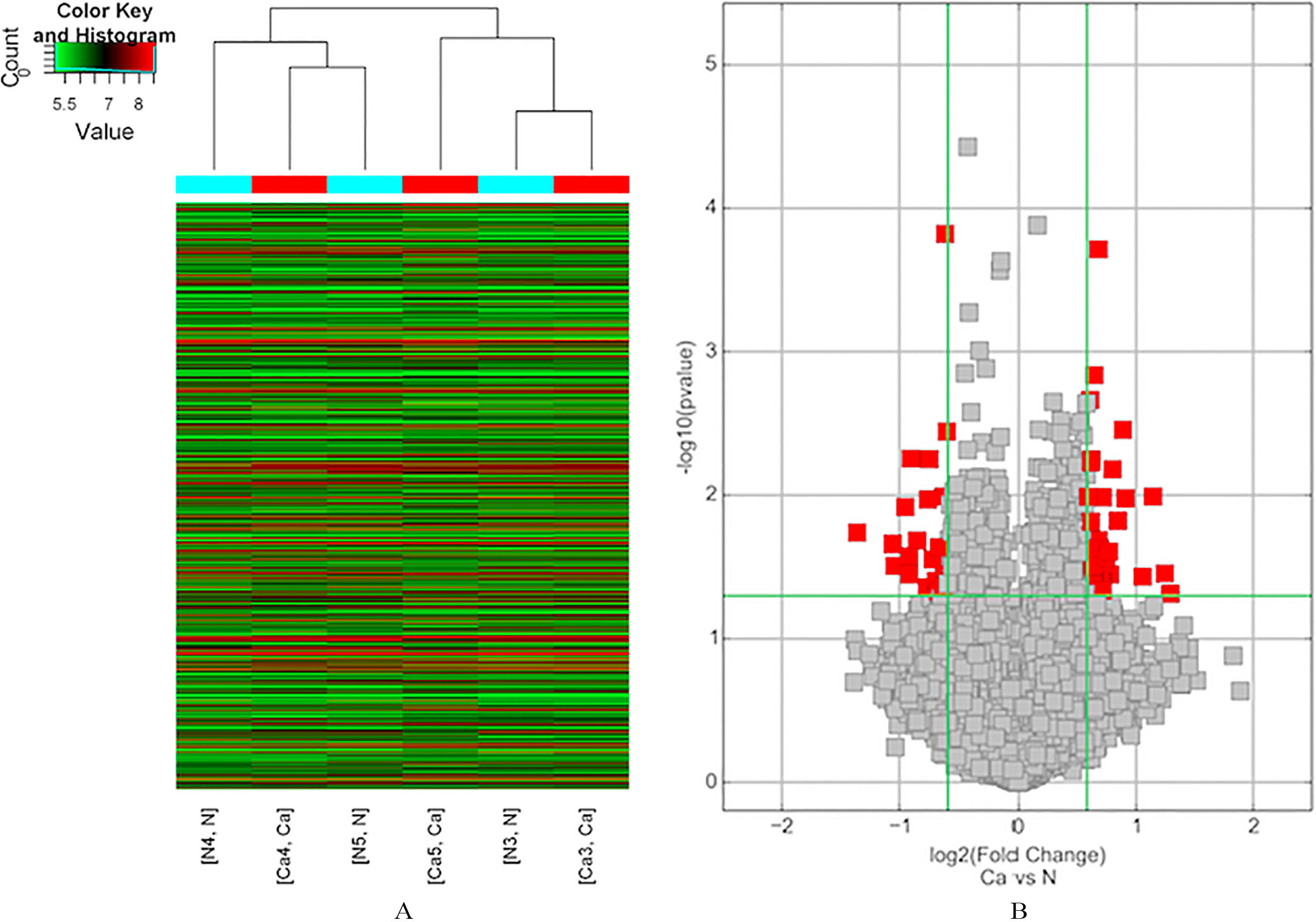
Total RNA from three GC and adjacent normal tissues was quantified using the NanoDrop ND-1000. The sample preparation and microarray hybridization were performed based on the Arraystar’s standard protocols. Briefly, total RNAs were digested with RNase R (Epicentre, Inc.) to remove linear RNAs and enrich circular RNAs. Then, the enriched circular RNAs were amplified and transcribed into fluorescent cRNA utilizing a random priming method (Arraystar Super RNA Labeling Kit; Arraystar). The labeled cRNAs were hybridized onto the Arraystar Human circRNA Array V2 (8
To quantify the amount of mRNA and circRNA, cDNA was synthesized with the PrimeScript RT Master Mix (Takara, Dalian, China) from 500 ng of RNA. The real-time PCR analyses were performed using SYBR Premix Ex Taq II (Takara). In particular, the divergent primers annealing at the distal ends of circRNA were used to determine the abundance of circRNA. To determine the absolute quantity of RNA, the purified PCR product amplified from cDNA corresponding to the hsa_circ_RNA102958 sequence was serially diluted to generate a standard curve. The sequences of hsa_circRNA_102958 divergent primers were as follows: Forward: 5’-GGATGCCCGCCAGAG TCCAGAT-3’and Reverse: 5’-ACGGGCTCCTTGGT GGGGTTA-3’. The primers’ sequences of glyceralde hyde-3-phosphate dehydrogenase (GAPDH), as a control, were 5’-GGGAGCCAAAAGGGTCAT-3’ and 5’-GAGTCCTTCCACGATACCAA-3’. we performed qRT-PCR three times to measure differences between experimental batches. Then, the correlation of their 2
Statistical analysis
All statistical data were analyzed using GraphPad Prism 6.0 (GraphPad Software, La Jolla, CA). The differences in levels of hsa_circRNA_102958 between gastric cancer tissues and paired adjacent non-tumorous tissues were assessed using the
Results
CircRNA expression profile in gastric cancer tissues
A total of 3 gastric cancer tissues and their matched non-gastric cancer (non-GC) tissues were collected and screened for dysregulated circRNA using human circRNA microarray. The results of hierarchical clustering show distinct circRNA expression profiling among the samples (Fig. 1). A Volcano plot was constructed to visualize the differential expression between tumor and non-tumor tissues (Fig. 2). A fold change
Hsa_circRNA_102958 was identified to be significantly up-regulated in gastric cancer tissues
Among screened up-regulated circRNA, circRNA_ 102958 ranked second and not found to be reported. Therefore, circRNA_102958 was selected as a target for investigating the relationship with gastric cancer. To further test the expression level of the dysregulated circRNA_102958, quantitative RT-PCR was used to measure its expression in 30 paired primary cancerous and adjacent noncancerous tissues from GC patients. The results indicated that circRNA_102958 expression was significantly up-regulated than those in corresponding noncancerous tissues (Fig. 2).
The expression levels of hsa_circRNA_102958 expression in tumor and adjacent normal tissues in 30 GC patients.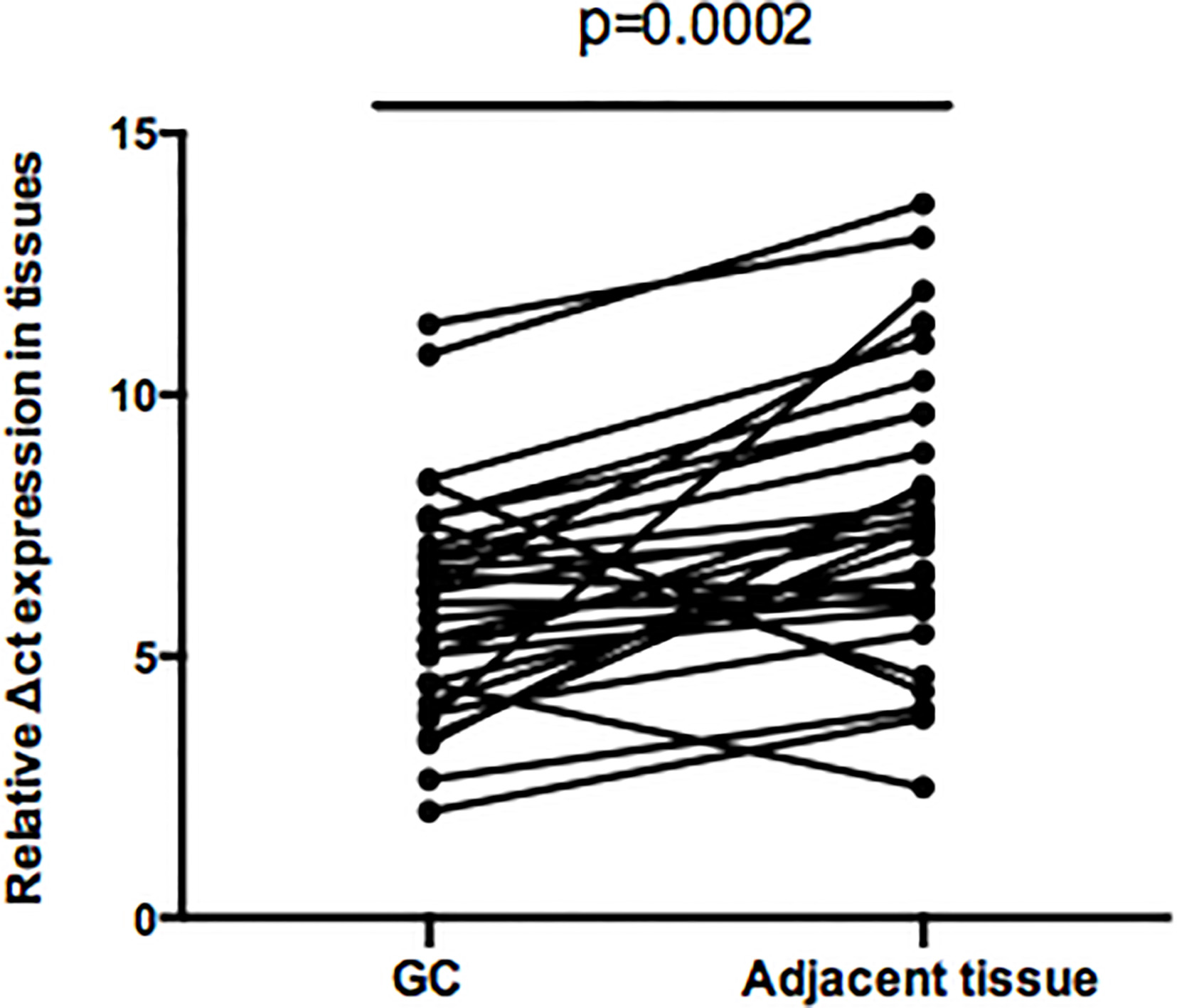
circRNA_102958 expression was also examined by qRT-PCR in five human gastric cancer cell lines and a nontumorigenic gastric epithelial cell line GES-1. This experiment showed that the levels of hsa_circRNA_ 102958 in five human gastric cancer cell lines, MKN- 45, SGC-7901, BGC-823, MGC-803 and AGS, were higher than the levels in the GES-1 human normal gastric epithelial cell line (Fig. 3).
Hsa_circRNA_102958 expression in gastric cancer cells. * means 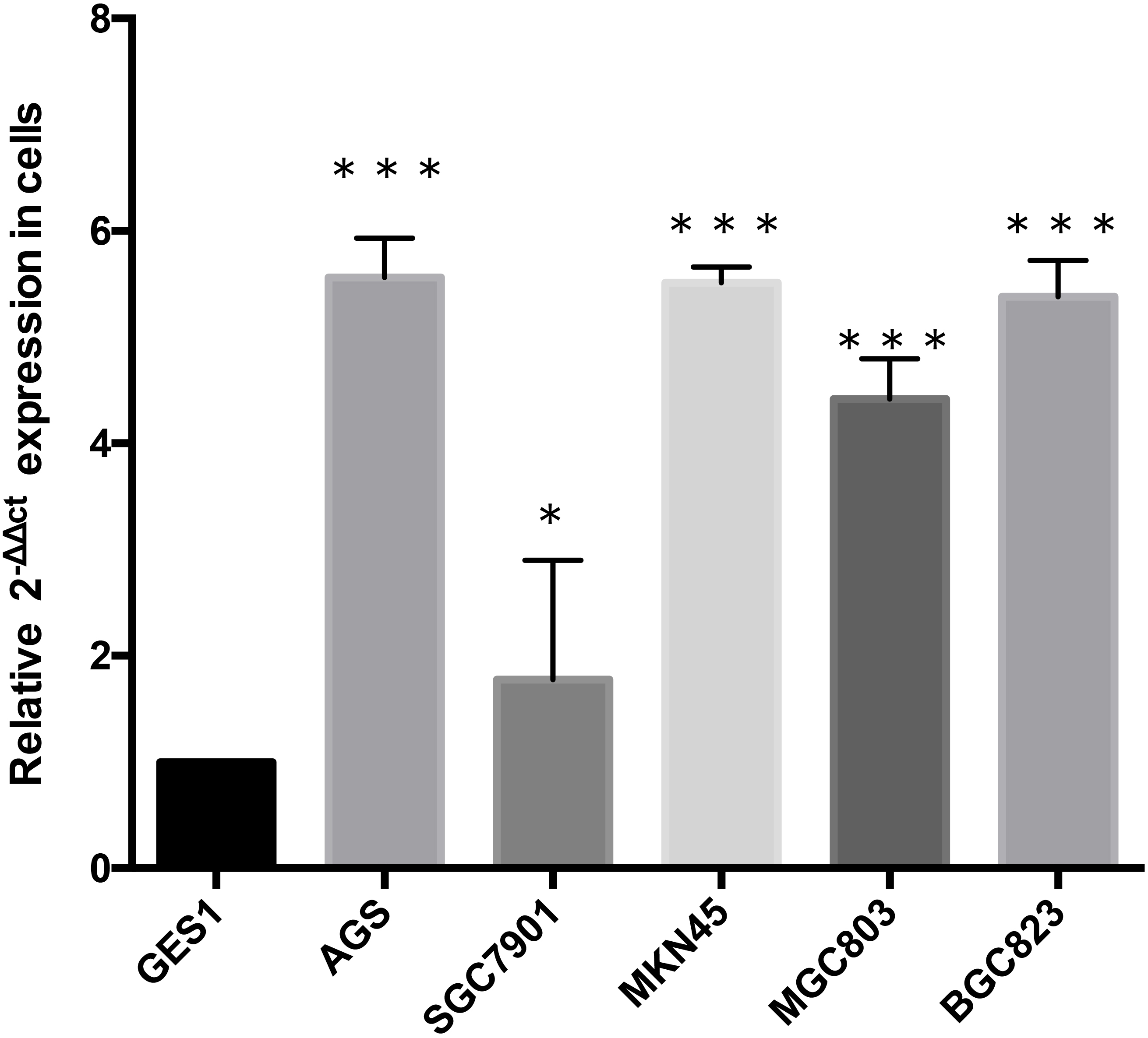
Furthermore, the area under the ROC curve was 0.74 (Fig. 4). The sensitivity and specificity were 0.61 and 0.86, respectively. The ROC is a comprehensive index used to reflect the sensitivity and specificity of continuous variables. When the area under the curve is larger, the diagnostic value of the variable is higher. Due to the limited number of samples of gastric cancer tissues, we only analyzed hsa_circRNA_102958 expression levels in 30 paired gastric cancer tissues samples. More samples should be analyzed in the future.
ROC curve of hsa_circRNA_102958.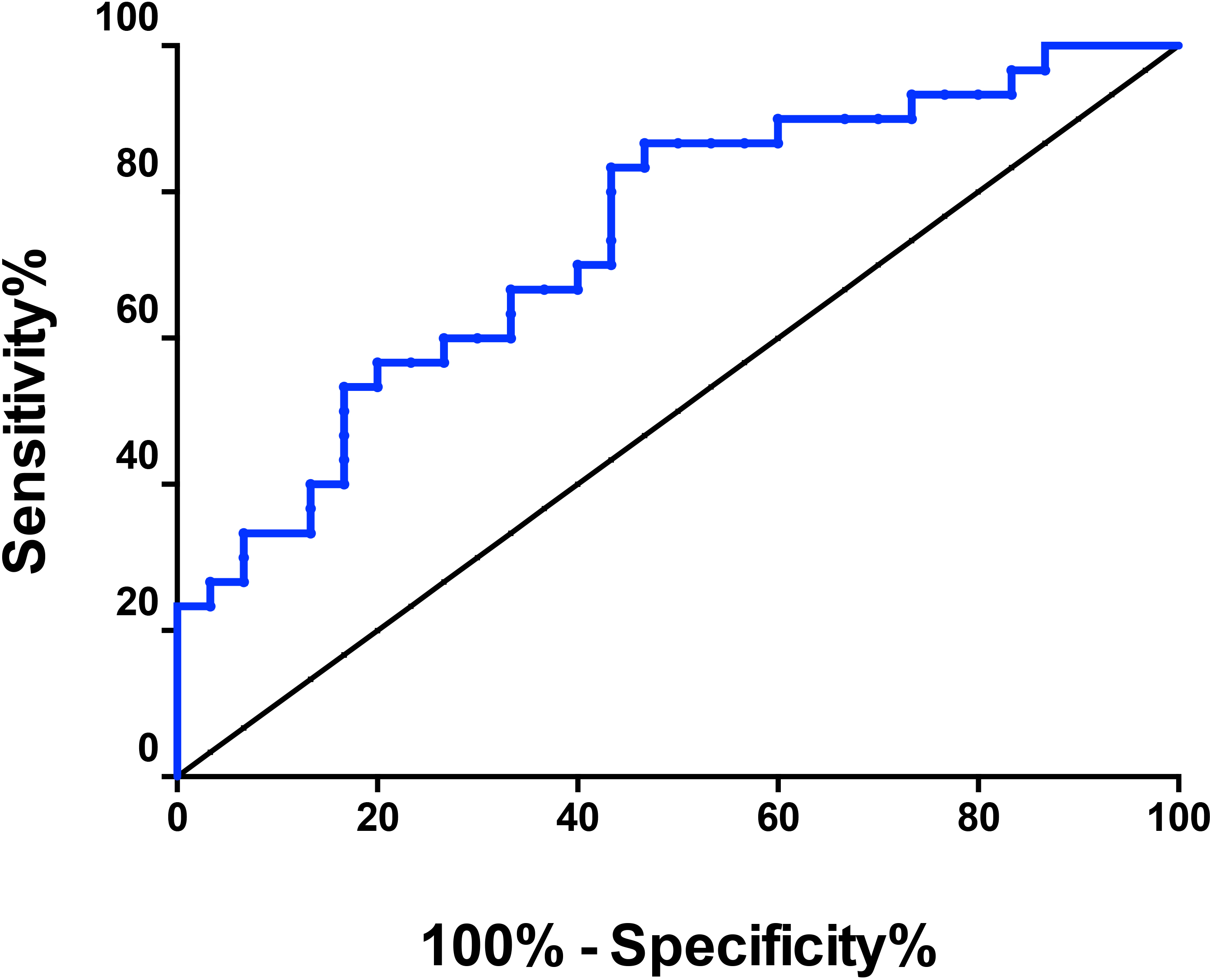
The predicted hsa_circRNA_102958 targeted miRNA and mRNA network.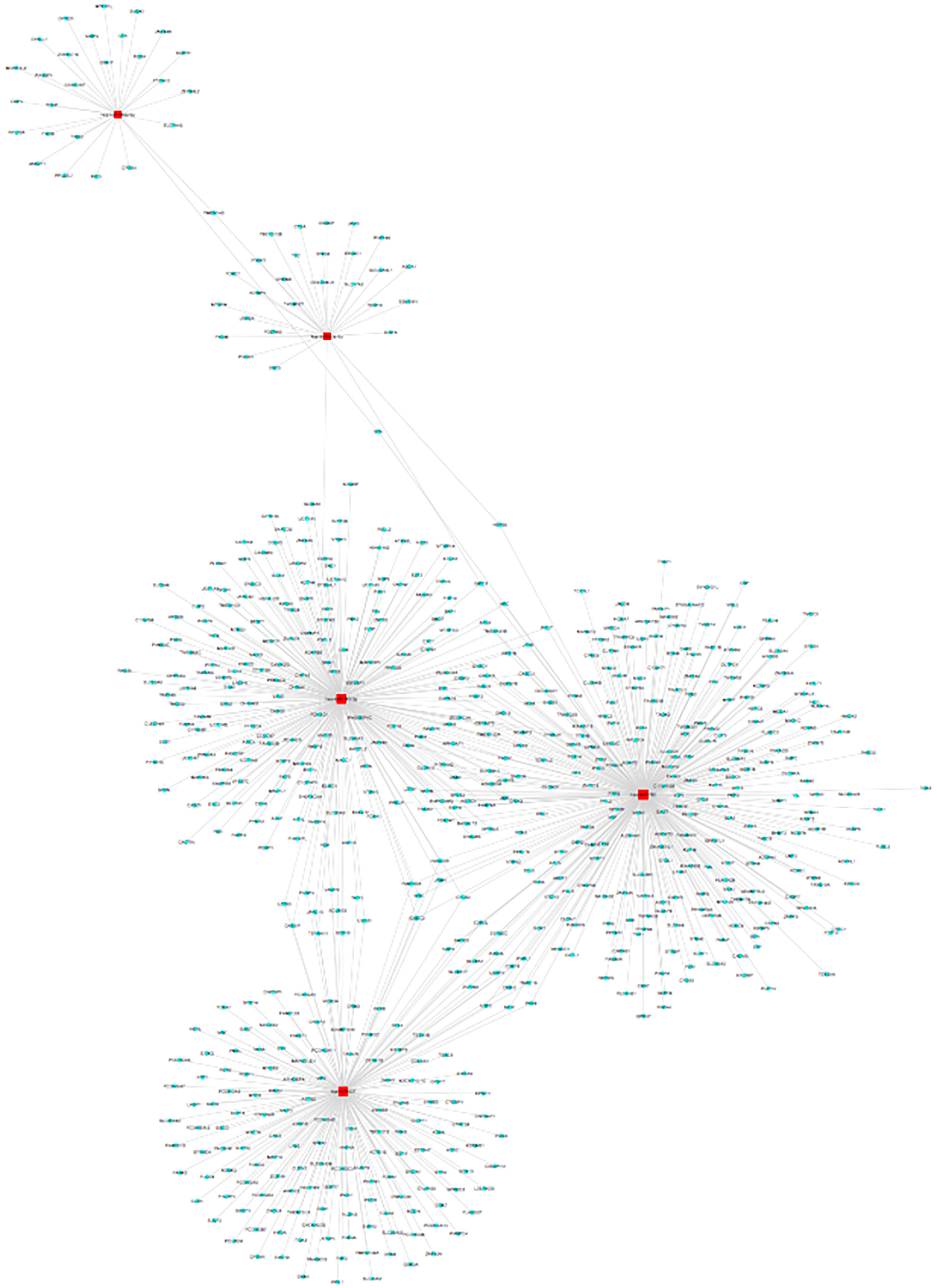
The results showed that hsa_circRNA_102958 expression was associated with several clinicopathological features of patients with gastric cancer. As shown in Table 1, hsa_circRNA_102958 levels was related to TNM stage (
A pathological feature of 30 promary GC and nontumor samples was analyzed. As shown in Table 1.
A pathological feature of 30 promary GC and nontumor samples was analyzed
A pathological feature of 30 promary GC and nontumor samples was analyzed
Next, the circRNA-miRNA-mRNA axis in cancer-related pathways was predicted. hsa-miR-146b-3p, hsa-miR-23a-5p, hsa-miR-185-3p, hsa-miR-762, hsa-miR-637 and their corresponding target mRNAs were predicted to have an interaction with hsa_circRNA_ 102958 in this study.
Discussion
Circular RNAs are newly found endogenous noncoding RNAs, which had been considered as the results of transcriptional noise [8]. With the development of bioinformatics’ methods and sequencing technology, lots of circular RNAs were predicted. CircRNAs have highly conserved sequences and a high degree of stability in mammalian cells [9, 10]. Due to their closed-loop structure, circRNAs are not affected by RNA exonuclease and are more stable and less degradable [11]. Many study shown that circRNA may act as endogenous sponge RNAs to interact with miRNAs and influence the expression of miRNA target genes [8, 12, 13, 14]. Compared with other noncoding RNA, these characteristic provide circRNAs with the potential to become ideal biomarkers in the diagnosis of cancers. Pan et al. demonstrated that ciRS-7 may act as a prospective prognostic biological marker and a promising therapeutic target for GC [15]. Chen et al. resulted that hsa_circ_0000190 may be a novel non-invasive biomarker for the diagnosis of gastric cancer [16]. Zhang et al. found that circRNA_100269 and miR-630 comprise a novel pathway that regulates proliferation of GC cells [17]. Huang et al. concluded that hsa_circ_0000745 plays an important role in GC and its expression level in plasma in combination with CEA level is a promising diagnostic marker [18]. We have shown in this study that the expression level of hsa_circRNA_102958 is upregulated in GC tissues when compared to the matched adjacent gastric tissues, and had potential diagnostic value. In our study, we found that high expression levels of hsa_circRNA_102958 in gastric cancer were significantly associated with TNM stage. Compared with the diagnostic value of hsa_circRNA_102958, whose AUC, sensitivity, and specificity were 0.74, 0.61, 0.86, respectively. However, due to the limited number of samples of gastric cancer tissues, we only analyzed hsa_circRNA_102958 expression levels in 30 paired gastric cancer tissues. More samples should be analyzed in the future. More importantly, by further studying the functions of circRNAs and circRNA-microRNA-mRNA axis, we could improve our understanding of the mechanisms of disease associated with circRNAs and improve the diagnosis and prevention of circRNA-associated diseases.
In conclusion, our data indicate that hsa_circRNA_ 102958 may represent a potential novel biomarker for the diagnosis of gastric cancer.
