Abstract
Purpose
Gastric cancer is the most common malignancy worldwide and is the third leading cause of cancer-related deaths, urgently requiring an early and non-invasive diagnosis. Circulating extracellular vesicles may emerge as promising biomarkers for the rapid diagnosis in a non-invasive manner.
Methods
Using high-throughput small RNA sequencing, we profiled the small RNA population of serum-derived extracellular vesicles from healthy controls and gastric cancer patients. Differentially expressed microRNAs (miRNAs) were randomly selected and validated by reverse transcription-quantitative real-time polymerase chain reaction. Receiver operating characteristic curves were employed to assess the predictive value of miRNAs for gastric cancer.
Results
In this study, 193 differentially expressed miRNAs were identified, of which 152 were upregulated and 41 were significantly downregulated. Among the differently expressed miRNA, the expression levels of miR-21-5p, miR-26a-5p, and miR-27a-3p were significantly elevated in serum-derived extracellular vesicles of gastric cancer patients. The miR-21-5p and miR-27a-3p were closely correlated with the tumor size. Moreover, the expression levels of serum miR-21-5p and miR-26a-5p were significantly decreased in gastric cancer patients after surgery.
Conclusions
The present study discovered the potential of serum miR-21-5p and miR-26a-5p as promising candidates for the diagnostic and prognostic markers of gastric cancer.
Introduction
Gastric cancer (GC) ranks as the world's fifth most common cancer and the third leading cause of cancer-related death. 1 GC poses a significant global health challenge. With a 5-year survival rate ranging only from 20% to 40%, 2 GC poses a significant global health challenge. It is highly endemic in Japan, Korea, and China, and it is estimated that 860,000 people worldwide die every year from GC.3,4 Serological and histomorphological biomarkers offer valuable tools for stratifying the risk of GC.5,6 Due to the disease's prolonged incubation period and the absence of effective early diagnostic methods, it is difficult to achieve early detection. Despite recent advances in the treatment approaches, 5 the overall survival of patients remains poor. The need to explore innovative biomarkers for early GC detection persists.
Extracellular vesicles (EVs) represent a kind of lipid membrane vesicle found in various body fluids, including blood, urine, cerebrospinal fluid, and amniotic fluid. 7 Among these, exosome-like EVs have garnered significant attention in cancer research due to their remarkable and pivotal role in intercellular communication, shaping the tumor microenvironment and fostering tumor progression, as well as their potential in cancer diagnosis. 8 Emerging evidence highlights that EVs isolated from the body fluids of cancer patients, or the culture medium of cancer cells, encapsulate various tumor-specific molecules, such as specific proteins, functional mRNAs, or small RNAs (sRNAs).7,9,10 Profiling cancer-derived EVs in the body fluids of cancer patients may offer an efficient strategy for diagnostic candidate selection.
microRNAs (miRNAs), a class of small, single-stranded, non-coding molecules, exert regulatory effects on gene expression at the transcriptional or post-transcriptional level. 11 Their involvement in the tumor growth, metastasis, and development underscores their significance in cancer biology.12,13 Recent studies have shown that miRNAs are enriched in EVs and affect the biological processes of recipients through cell-to-cell interactions. 14
In this study, we utilized a deep-sequencing approach to investigate EV-derived miRNA profiles in GC patients. Our findings offer insights into the potential use of serum EV-derived miRNAs as diagnostic candidates, thereby expanding the scope of their applications in the evaluation of GC diagnosis and prognosis.
Materials and methods
Ethics statement
Human studies were approved by the Ethics Committee of the Gansu Provincial People's Hospital (2021–144) and the Baiyin First People's Hospital (YL-KY-2021-030). Written informed consent was obtained from all adult research participants.
Human serum samples
The GC patients were confirmed by biopsy, electron gastroscopy, and postoperative pathology. Other systemic tumors and other underlying diseases were excluded, as well as preoperative chemotherapy or radiotherapy. For high-throughput sRNA-sequencing, 10 sera from healthy controls (HCs) (n = 5) and GC patients (n = 5) were collected. For the diagnosis of GC patients, 33 serum samples of GC patients, 33 postoperative patients, and 33 HCs were collected. A total of 10 mL of whole blood were used for serum isolation. After centrifuging at 3000 × g for 15 min at 4°C, the supernatant (serum) was immediately transferred into a clean tube and stored at −80°C for subsequent use.
Enrichment and characterization of EVs
EVs were isolated as previously reported with some modifications. 15 In brief, 6 mL of sera were first centrifuged at 10,000 × g for 30 min at 4°C, filtered through 0.22 µm syringe filters, and then centrifuged at 100,000 × g for 2 h at 4°C. The pellets were then washed with filtered phosphate-buffered saline (PBS), centrifuged at 100,000 × g for 90 min, and finally resuspended in small amount of PBS for further analysis.
Nanoparticle tracking analysis
The size distribution of enriched EVs was assessed using by nanoparticle tracking analysis (NTA). Briefly, 5 uL of EVs were diluted with PBS to a final 1 mL (1:200), and the mixture was thoroughly mixed. The diluent sample was then loaded into a Zetasizer Nano (Malvern Panalytical, Malvern, UK), where the Browne motion of each nanoparticle was observed and used to calculate the size of enriched EVs.
Transmission electron microscopy
The morphology of enriched EVs was characterized by transmission electron microscopy (TEM). Briefly, 10 μL EV drops were placed on the carbon-coated copper grid and incubated for 5 min. The remaining EVs were absorbed with filter paper. After being washed, the grids were stained with 3% phosphotungstic acid and visualized under JEM–2010 TEM.
Western blot
The EV samples were homogenized in 50 μL of cold RIPA lysis buffer. Followed by centrifugation to determine the total protein concentration using the bicinchoninic acid assay method. Approximately 30 μg of protein was loaded per lane on a Bis-Tris Sure PAGE gel and transferred to a polyvinylidene fluoride membrane. After incubating with 5% skim milk in PBS for 1 h, the membranes were immunoblotted with CD9, CD63 antibodies (Abcam, Cambridge, MA, USA) at 4°C overnight. Subsequently, were washed and incubated with the second antibody and were visualized by using the Clinx-6100 Imaging System (Clinx, Shanghai, China).
RNA extraction and deep sequencing
Total RNAs were extracted from the serum-derived EVs using Trizol, purified using RNeasy columns. Small RNA sequencing was carried out in Ruibo. A total of 1 ug total RNA of each example was prepared for the construction of sRNA sequencing libraries. We performed the following bioinformatical analysis. Briefly, the raw reads were processed by filtering out the adaptor sequences and low-quality and smaller than 15nt reads; only reads perfectly mapped the human genome were used for the following analysis. Human known miRNAs were identified by BLAST against the Mir GeneDB database. The relative miRNAs expression levels were analyzed using the DEGseq method. Differentially expressed miRNAs were identified based on a fold change threshold of ≥2 and a significance level of the P-value (<0.05). TargetScan, miRanda, and RNAhybrid were used to predict target genes of identified miRNAs. KOBAS was used to further the Gene Ontology (GO) and KEGG (Kyoto Encyclopedia of Genes and Genomes) pathways.
Quantitative real-time polymerase chain reaction
Total RNAs extracted from EVs and sera were used for miRNA complementary DNA (cDNA) synthesis using miRNA First-Strand cDNA Synthesis Kit 2.0 (Igenen Biotech, Guangdong, China). A total of 10 uL of synthetic cel-miR-39 was added to each sample as an external reference. RT-PCRs were carried out using the miRNA qPCR Kit (Igenen Biotech, Guangdong, China) in a 7500 Real-Time PCR system. All the primers were synthesized from the Igenen Biotech (Guangdong, China). Relative fold change of miRNAs was calculated as 2−ΔΔCt. Results are expressed as mean ±SD.
Data analyses
Data analyses were performed using GraphPad Prism 8.4.3 software. The significance of the relative expression of miRNA was analyzed using t-test and one-way analysis of variance. A P value < 0.05 was considered significantly different. The expression of each EV miRNA was plotted against the subject's receiver operating characteristic (ROC) curve.
Results
Isolation and characterization of serum-derived EVs from GC patients
The enriched EVs isolated from HC and GC patients exhibited remarkable physical homogeneity, and was approximately 30–150 nm in diameter by TEM (Figure 1(a)). Furthermore, NTA results demonstrated that these EVs were highly homogeneous in size, with a diameter of 30–150 nm and a peak at approximately 100 nm (Figure 1(c)). There were no significant differences in size and morphology between the EVs derived from the two groups. Western blot results revealed that both CD9 and CD63 were more abundant in these EVs (Figure 1(b)).
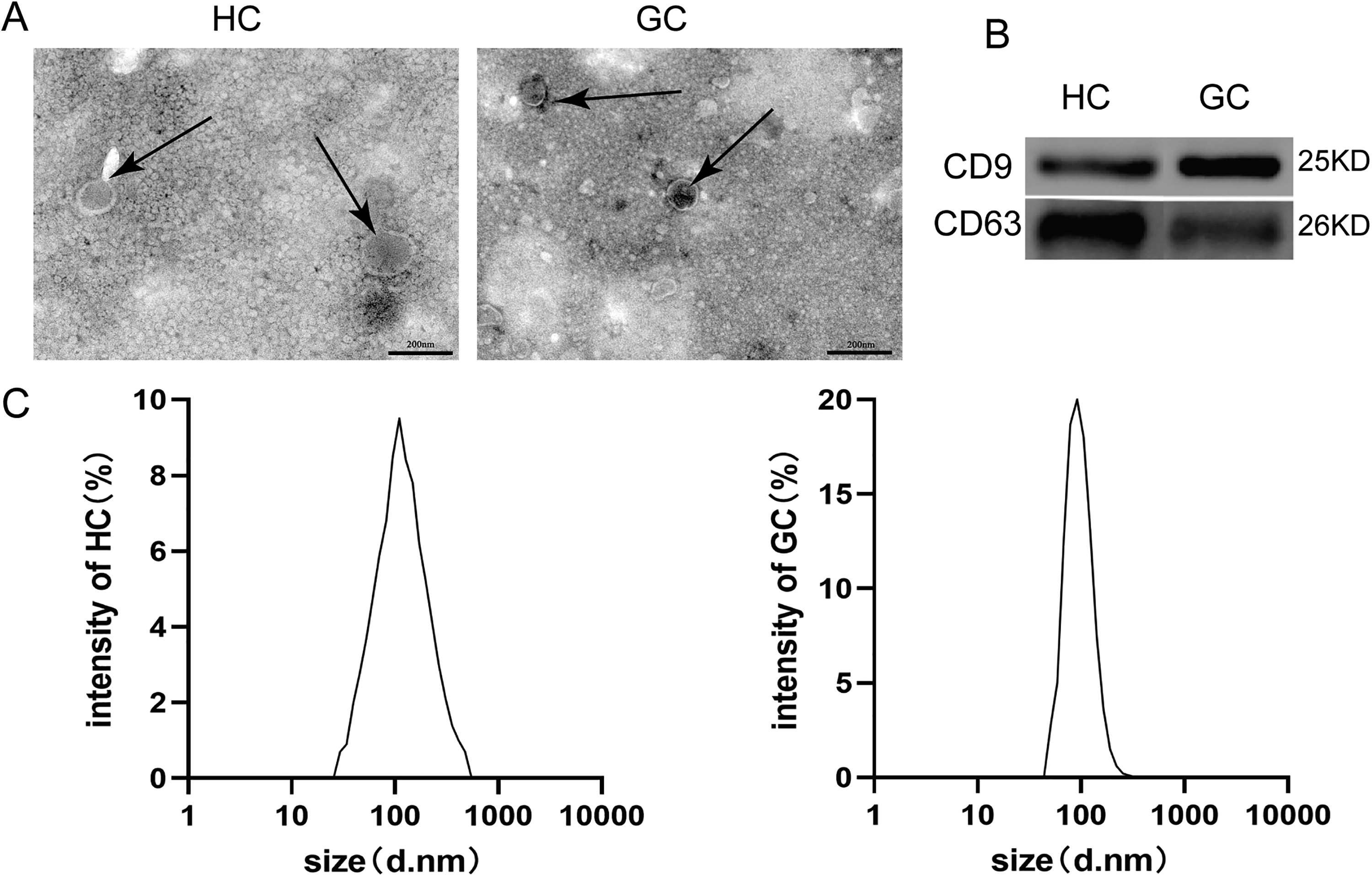
Identification and characteristics of healthy controls (HC) and gastric cancer (GC) serum-derived extracellular vesicles (EVs). (a) Transmission electron microscopy of the enriched EVs isolated from the sera of HC and GC patients. (b) Western blotting analysis of EV-marker CD9 and CD63 in the purified EVs. (c) The diameter distribution of the purified EVs isolated from the sera of HC and GC patients.
Identification of differentially expressed known miRNAs in serum-derived EVs of GC patients
Serum-derived EVs from five HCs and five GCs were subjected to high-throughput sequencing. The size distribution of sRNA reads was similar in 8 sRNA libraries, and the majority of these sRNA were 21–22 nt in length (Figure 2(a)). These results indicated that the miRNA species of sRNA libraries were highly enriched. A total of 888 and 1152 known miRNAs were identified in the HC and GC groups, respectively, and a total of 819 unique miRNAs were identified from both groups (Figure 2(b)). Among these, 193 differentially expressed miRNAs were identified (fold change ≥2 and P-value < 0.05), with 152 miRNAs upregulated and 41 miRNAs downregulated (Figure 2(c) and (d); Table 1). Analysis of target genes of differentially expressed miRNAs revealed enrichment in cellular components such as focal adhesion and cell-substrate junctions, molecular functions including cell adhesion molecule binding and DNA-binding, and biological processes such as gland development and regulation of cell morphogenesis (Figure 3(a)). KEGG pathway analysis highlighted the enrichment of top 30 pathways, including the PI3K-Akt signaling pathway, insulin signaling pathway, and proteoglycans in cancer (Figure 3(b)). A miRNA-messenger RNA (mRNA) prediction network illustrated the complex regulatory interactions, where individual miRNAs could target multiple mRNA targets, and conversely, a single mRNA could be targeted by multiple miRNAs (Figure 3(c)).
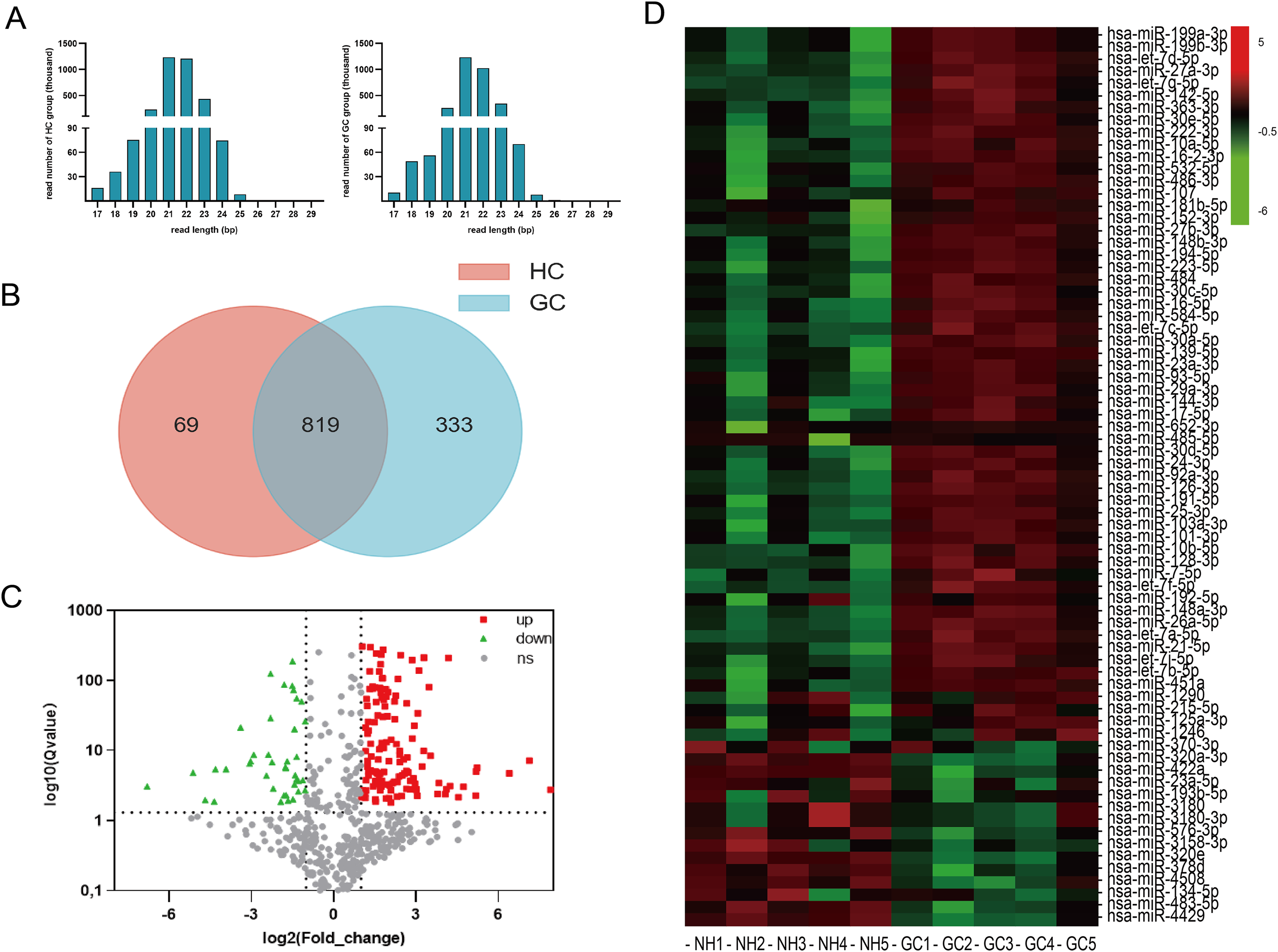
Identification of differentially expressed miRNAs in the HC and GC serum-derived EVs. (a) miRNA particle size. (b) The distribution of 888 and 1152 unique miRNAs identified in the serum-derived EVs of both HC (left, red circle) and GC sera (right, blue circle). The overlapping region indicates co-expressed unique miRNAs. (c) Volcano map of differentially expressed miRNAs between the HC and GC serum-derived EVs. (d) Heatmap of differentially expressed miRNAs between the HC and GC serum-derived EVs.
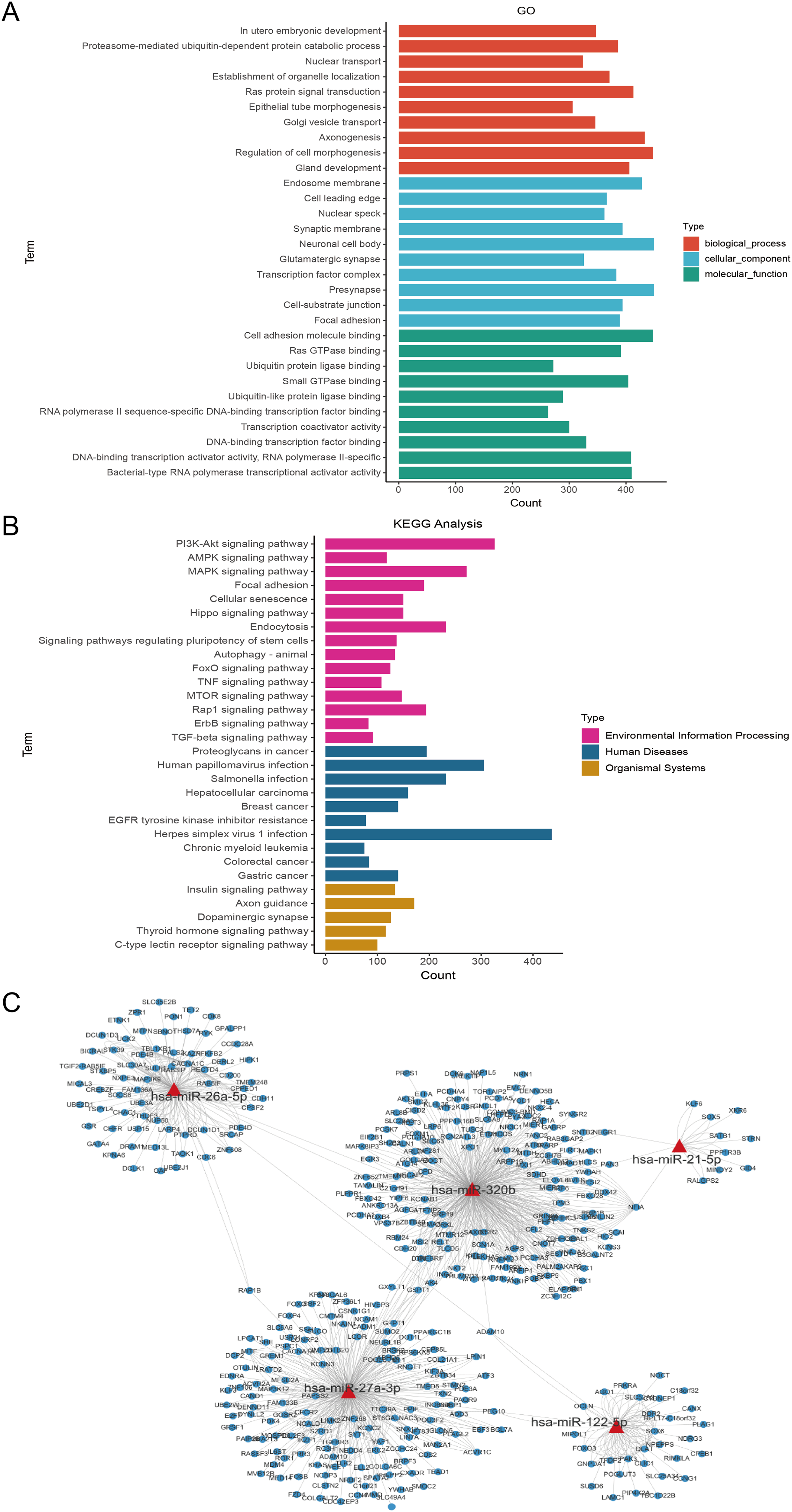
KEGG and GO enrichment analysis of the target genes of differentially expressed miRNAs. (a) GO molecular function annotations of the target genes of differentially expressed miRNAs. According to the P value, 30 top GO terms of biological process, molecular function and cellular component were shown, respectively. (b) KEGG pathway enrichment analyses of the target genes of differentially expressed miRNAs. (c) The prediction network of five differentially expressed miRNAs, including miR-21-5p, miR-26a-5p, and miR-27a-3p (marked in red) and those targeted genes (marked in blue).
Upregulation of the top 20 differential miRNAs.
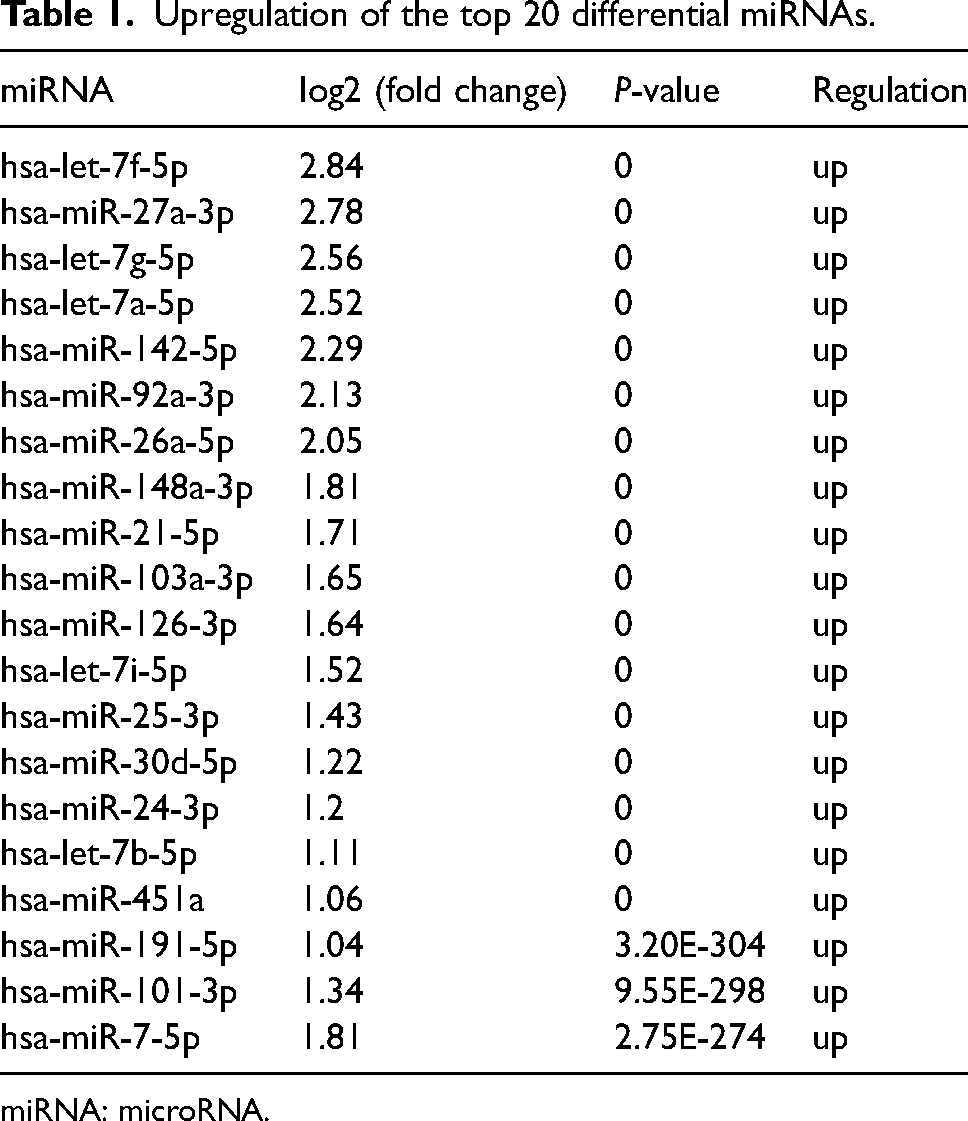
miRNA: microRNA.
Values of differentially expressed miRNAs in the diagnosis of GC patients
To evaluate their diagnostic values, three differentially expressed miRNAs (including miR-21-5p, miR-26a-5p, and miR-27a-3p) with high abundance and low P value were selected for assessing GC diagnostic performance by qPCR. Consistent with the sequencing data, the expression levels of miR-21-5p, miR-26a-5p, and miR-27a-3p were significantly elevated in serum-derived EVs from GC patients compared to those from HC patients (P < 0.05) (Figure 4(a)). Notably, miR-21-5p and miR-26a-5p exhibited superior diagnostic performance for GC patients, with area under the curve (AUC) values of approximately 0.7648 (95% confidence interval (CI) 0.6197, 0.9099), and 0.7382 (95% CI 0.5727, 0.9036), respectively. Furthermore, miR-21-5p (sensitivity: 53.58%; 95% CI 35.46,71.24%; specificity: 100%; 95% CI 87.13, 100%), significantly outperforming miR-26a-5p demonstrated a sensitivity of approximately 69.23% (95% CI 50.01, 83.50%) and specificity of 100% (95% CI 87.13, 100%), (Figure 4(b)). Considering the diagnostic potential of miR-21-5p, miR-26a-5p, and miR-27a-3p in serum from GC patients, we further evaluated their diagnostic utility in post-surgery sera. In serum samples, the expression of the three miRNAs were higher in GC patients compared to the HC group (P < 0.05), with miR-21-5p and miR-26a-5p significantly downregulated post-surgery (P < 0.05). However, there was no significant difference in the expressions of miR-27a-3p post-surgery (P > 0.05) (Figure 5(a) and (b)).
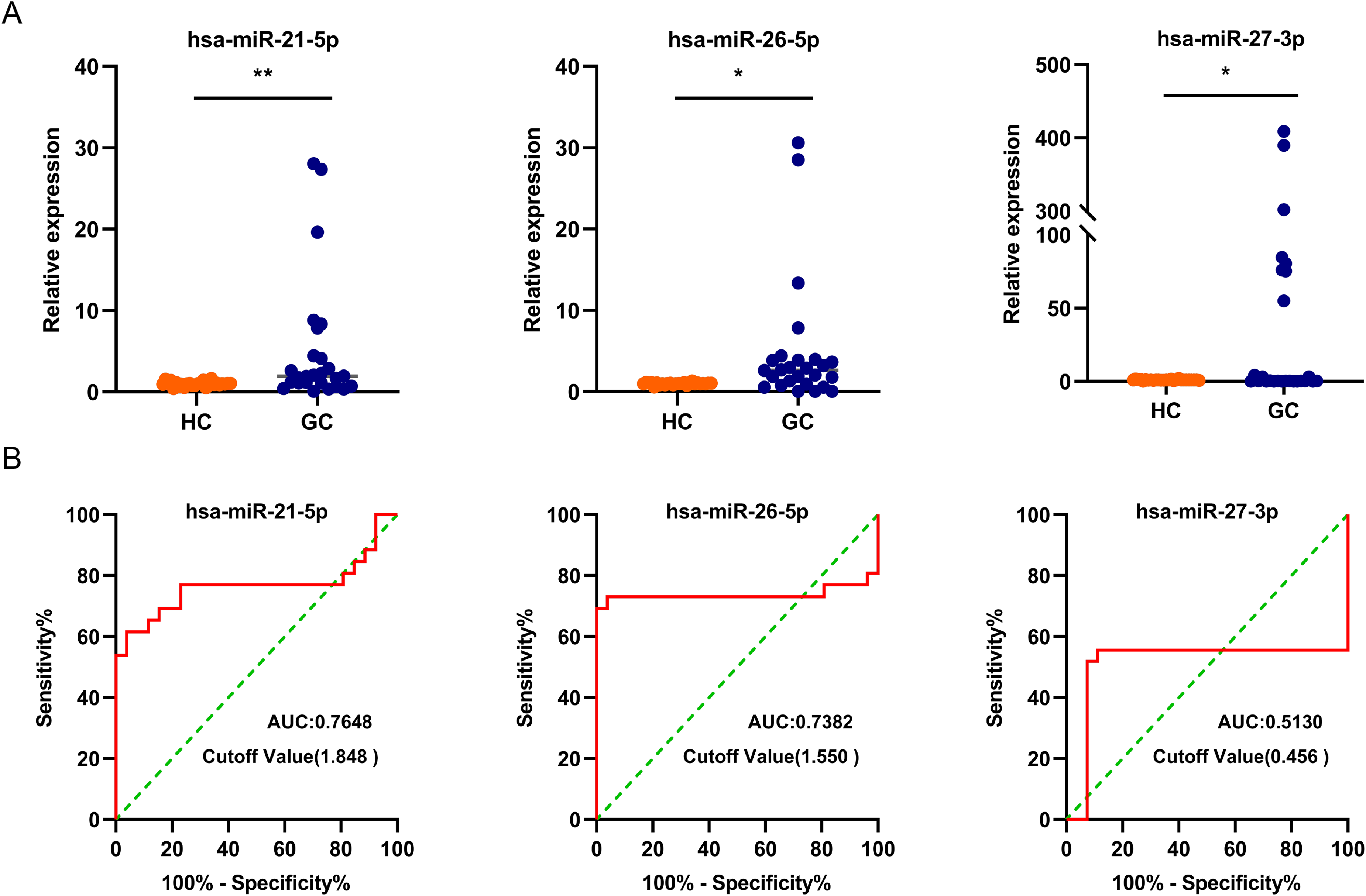
The diagnostic values of miR21-5p, miR-26a-5p, and miR-27a-3p in GC patients. (a) Relative expression of miR-21-5p, miR-26a-5p, and miR-27a-3p in the serum-derived EVs from GC patients. (b) Receiver operating characteristic (ROC) curve for miR-21-5p, miR-26a-5p, and miR-27a-3p in the serum-derived EVs from GC patients.
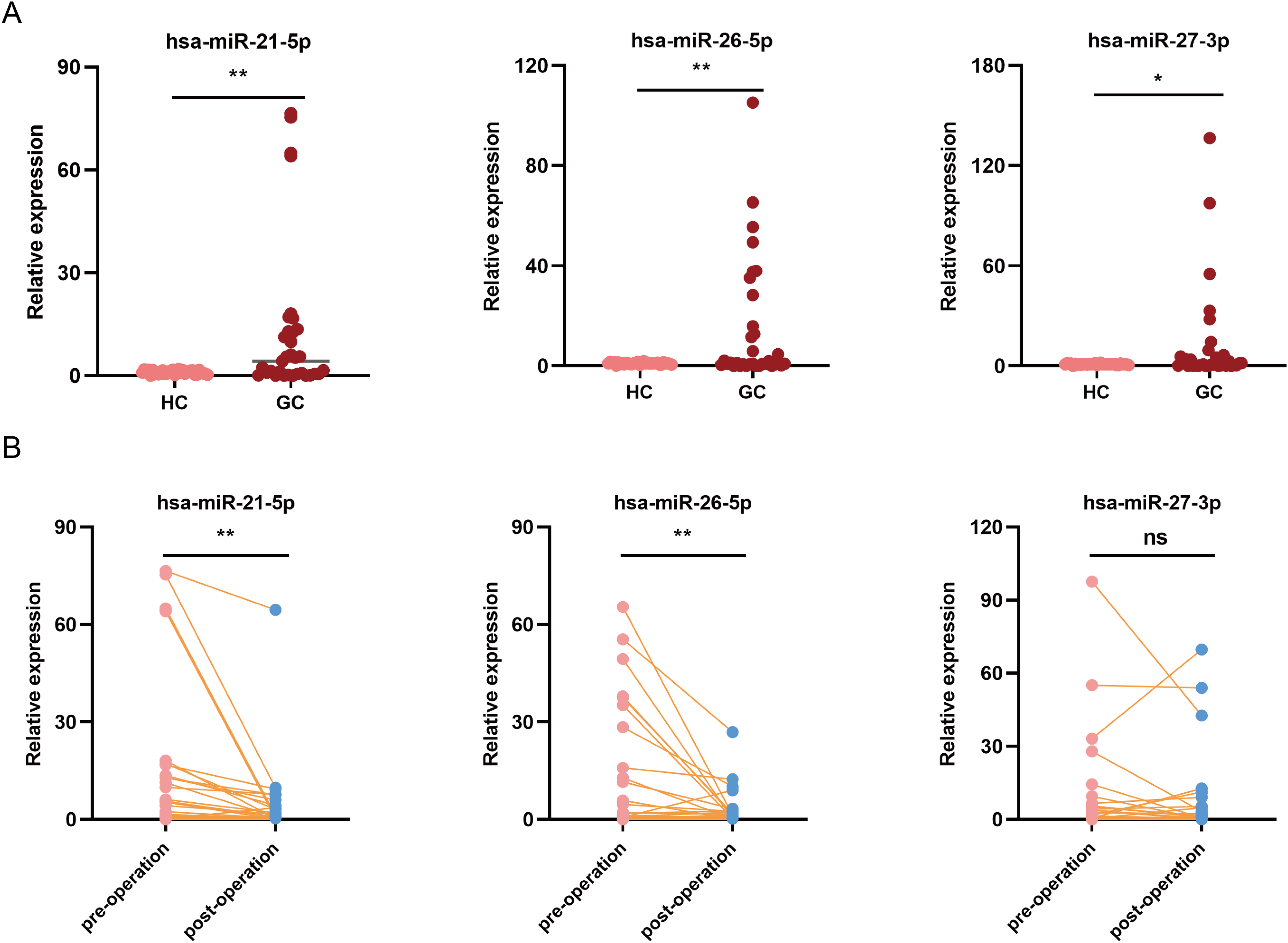
The pre- and postoperative values of serum miR-21-5p, miR-26a-5p, and miR-27a-3p in the GC patients. (a) Relative expression of those miRNAs in the sera of GC patients. (b) Relative expression of those miRNA in the in sera of GC patients after surgery.
Clinical significance of serum miRNAs in GC patients
Considering the diagnostic values of miR-21-5p, miR-26a-5p, and miR-27a-3p in the sera of GC patients, we further investigated the relationship between serum miRNA expression levels and clinical pathological features of GC patients. No statistical significance in age was observed among the study groups (Table 2). The expression levels of miR-21-5p, miR-26a-5p, and miR-27a-3p were significantly elevated in the sera of GC patients compared to the HCs (P < 0.001). Additionally, with the exception of miR-26a-5p, the expression levels of miR-21-5p and miR-27a-3p were significantly correlated with tumor size (Table 2).
A correlation between serum miRNAs and clinicopathological features in GC patients.
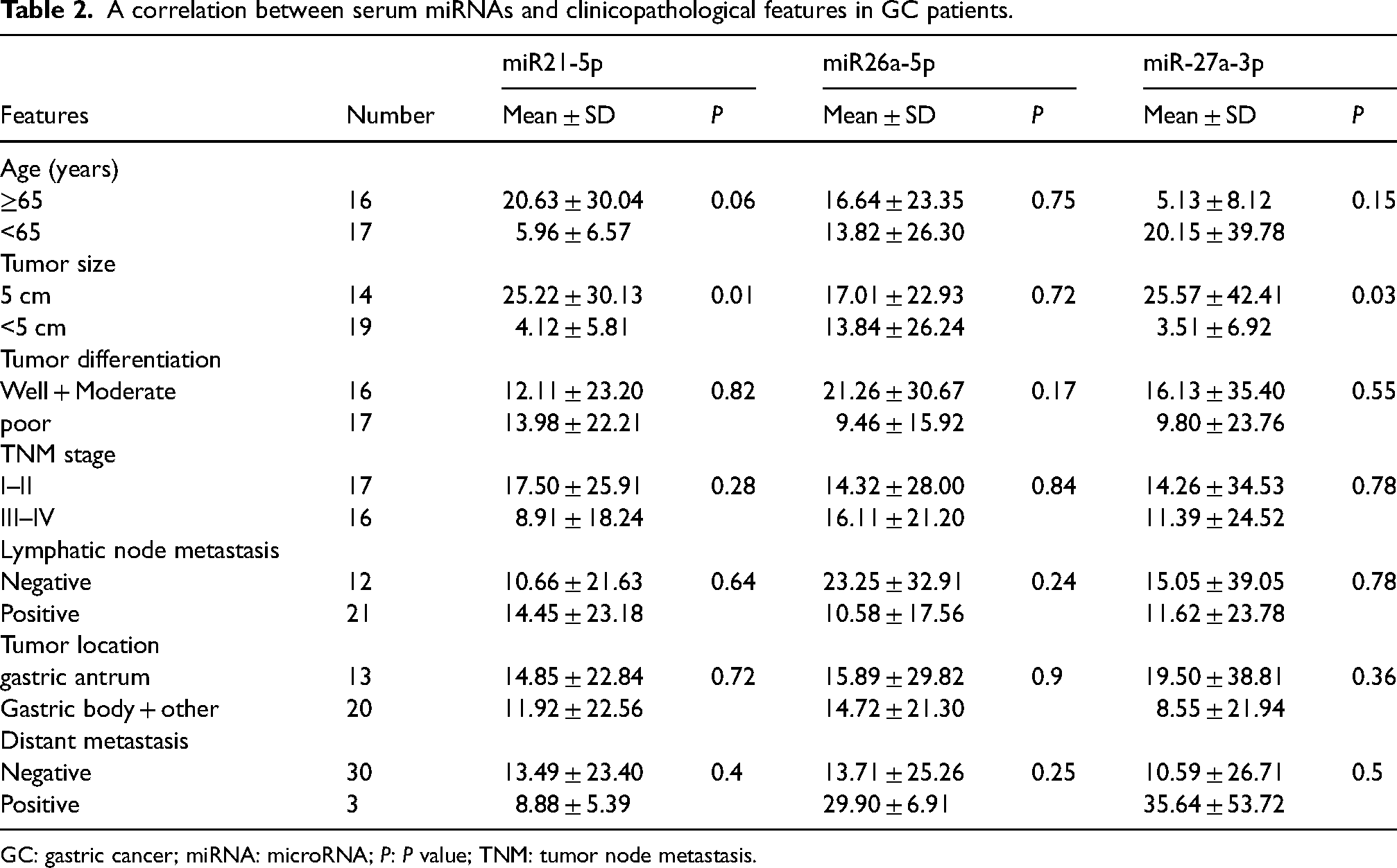
GC: gastric cancer; miRNA: microRNA; P: P value; TNM: tumor node metastasis.
Discussion
GC is a prevalent malignancy characterized by high incidence, high malignancy, and unfavorable prognosis for patients. 16 Early diagnosis and treatment of GC remain challenging due to the lack of specific clinical symptoms in the early stages of the disease. 17 Hence, there is an urgent need for novel non-invasive diagnostic methods. Accumulating evidence in recent years suggests that serum-derived EVs hold promise as biomarkers for the rapid and non-invasive diagnosis. In this study, we conducted comprehensive miRNA profiling of serum-derived EVs from GC patients. Our findings highlight serum miR-21-5p and miR-26a-5p as promising candidates for diagnostic and prognostic markers of GC. Consistent with the previous reports, 18 the enriched EVs exhibited identically spherical in morphology, with diameters ranging from 30 to 150 nm. EV-specific markers, including CD9 and CD63 were remarkably detected in these EVs. Deep sequencing analysis revealed that 193 miRNAs were significantly altered in GC patients. Among these differently expressed miRNA, miR-21-5p, miR-26a-5p, and miR-27a-3p were significantly elevated in serum-derived EVs from GC patients. Both miR-21-5p and miR-26a-5p—recognized as important oncogenic miRNAs—have been reported to be dysregulated in various tumors. Specifically, miR-21-5p has been implicated in promoting the growth and metastasis, while miR-26a-5p and miR-26b-5p have been shown to inhibit proliferation by targeting PDCD10.19–22 The upregulation of miR-21-5p and miR-26a-5p in the sera of GC patients further suggests their potential diagnostic value in GC patients. Additionally, studies by Wang et al.14 and Tang et al. 23 have identified potential biomarkers for GC including miR-10401-3p, miR-1255-5p, and miR-6736-5p, as well as serum EV-derived miRNAs, such as miR-92b-3p, miR-146b-5p, miR-9-5p, and let-7g-5p, which may serve as promising candidates for early GC diagnosis. Remarkably, the expression levels of serum miR-21-5p and miR-26a-5p showed a significant decrease in GC patients following surgery. This observation suggests that serum miR-21-5p and miR-26a-5p hold promise as prognostic indicators in the diagnosis of GC patients. Thus, the identification of suitable EV-associated miRNAs as biomarkers holds crucial significance for early tumor detection. The KEGG pathway analyses revealed that the target genes of differentially expressed miRNAs were enriched in the PI3K-Akt signaling pathway, the insulin signaling pathway, and the proteoglycans in cancer. Notably, the insulin signaling pathway plays a crucial role in energy metabolism signal transduction within tumor cells, while the PI3K-Akt signaling pathway significantly influences tumor proliferation, invasion, and metastasis. Our findings indicated a correlation between circulating miR-21-5p and miR-27a-3p expression and tumor size in GC patients, suggesting its involvement in tumor development and growth. Previous studies have demonstrated that miR-21-5p can enhance growth and metastasis by targeting several tumor suppressors, 19 while miR-21-5p/PDCD4 axis suppressed cell apoptosis via the PI3K/AKT/FOXO1 signaling pathway. 24 Additionally, miRNA-26a-5p has been shown to promote cell migration and invasion by targeting FAF1 in non-small cell lung cancer, although its association with pathological clinical features has not been extensively reported. 5
In summary, our study elucidates the alterations in miRNA profiling in serum EVs isolated from patients with GC and identifies significant upregulation of miR21-5p and miR-26a-5p expression in serum EVs, which notably decrease in GC patients post-surgery. We validate miR-21-5p and miR-26a-5p as promising diagnostic and prognostic markers for GC.
Supplemental Material
sj-docx-1-jbm-10.1177_03936155241261390 - Supplemental material for Serum extracellular vesicle-derived miR-21-5p and miR-26a-5p as non-invasive diagnostic potential biomarkers for gastric cancer: A preliminary study
Supplemental material, sj-docx-1-jbm-10.1177_03936155241261390 for Serum extracellular vesicle-derived miR-21-5p and miR-26a-5p as non-invasive diagnostic potential biomarkers for gastric cancer: A preliminary study by Jun-Hong Wang, Zhao-zhao Bai, Xing-dong Niu, Cheng-lou Zhu, Tong Liang, Yong-li Hu, Zhen-Hua Gao and Ming-Xu Da in The International Journal of Biological Markers
Footnotes
Acknowledgments
The authors are grateful to all the laboratory members of Lanzhou Veterinary Research Institute, Chinese Academy of Agricultural Sciences for their assistance. Thanks also to Guangdong Ruibo Biological Company for its support in sequencing this experiment.
Author contributions
Junhong Wang, Zhao-Zhao Bai, and Xing-Dong Niu performed the experiments. Junhong Wang, and Yong-Li Hu designed the study and made a major contribution to writing the article. Cheng-Lou Zhu, and Tong Liang performed statistical analyses. Mingxu Da and Zhen-Hua Gao designed the study and made a major contribution to writing the article. All authors read and approved the final version.
Consent for publication
All authors have read and agreed to all the contents for publication.
Data availability
The datasets and materials used in the current study are available from the corresponding authors upon request.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This study was supported by grants from the Science and Technology Planning Project of Gansu Province (Grant No. 21CX6FD163), the Second Group of Longyuan Young Talents in Gansu Province (Grant No. 2023-11) and the Science and Technology Planning Project of Baiyin City (Grant No.2021-1-18Y).
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
