Abstract
PURPOSE:
The purpose of this study was to establish a risk scoring system based on miRNAs to evaluate the prognosis in pancreatic adenocarcinoma.
METHODS:
Using a miRNA microarray dataset (179 pancreatic adenocarcinoma specimens and 4 normal control specimens) from TCGA, differentially expressed miRNAs were identified. Cox proportional hazards regression analysis was used to identify significant prognostic miRNAs, with which a risk scoring system was established and tested on a validation set. Cox regression analysis was performed to identify independent predictors of survival from clinical characteristics. Stratified Cox regression analyses were conducted to unravel the associations of clinical characteristics with survival. Differentially expressed genes (DEGs) were screened followed by functional annotation of the DEGs.
RESULTS:
Eight miRNAs (miR-1301, miR-598, miR-1180, miR-155, miR-496, miR-203, miR-193b, miR-135b) were independent predictors for survival. A risk scoring system was established with the 8 signature miRNAs. Upon Cox multivariate regression analysis, risk score, new tumor and targeted molecular therapy were independent predictors of prognosis. Stratified Cox regression analyses found that targeted molecular therapy and new tumor are associated with survival of patients. Survival-related DEGs were significantly enriched with regulation of transforming growth factor beta receptor, potassium ion transport and MAPK signaling pathway.
CONCLUSIONS:
The study proposes 8-miRNA expression-based risk scoring system to predict prognosis in pancreatic adenocarcinoma. New tumor and targeted molecular therapy were independent predictors of prognosis. Transforming growth factor beta receptor, potassium ion transport and MAPK signaling pathway may be related to prognosis in pancreatic adenocarcinoma.
Introduction
Pancreatic cancer, one of leading cancers, is estimated to cause 41,780 deaths in the United States in 2016 [1]. Its most common type is pancreatic adenocarcinoma. Early diagnosis of pancreatic cancer remains a challenge, and patients have a very poor prognosis with a five-year survival rate of 5–6% [2]. Surgical removal as the only cure is possible in only 20% of new cases [3]. In order to improve outcome of patients, identification of molecular prognostic markers have been an active area to investigate.
MicroRNA (miRNA) is a small non-binding RNA molecule (19–25 nucleotides) binding to 3’-UTR of target messenger RNAs (mRNAs) to regulate human genes [4]. They are involved in gene regulation, development, pathogenesis of various malignances and other biological processes [5]. MiRNAs are very exclusive to different types of tissues or cells, thus raising the prospects that they are of prognostic value for predicting outcome of cancer patients. Increasing studies have recommended panels of miRNA in serum as biomarkers for early diagnosis of pancreatic cancer [6, 7, 8]. A meta-analysis provides evidence that tumoural miR-21 is a promising predictor of prognosis after pancreatic ductal adenocarcinoma resection [9]. However, correlations of miRNAs with prognosis of pancreatic cancer have not been fully elucidated. Although Zhou et al. [10] have suggested a 13-miRNA signature as an independent prognostic marker of PC. However, prognostic value of the 13-miRNA signature was not tested in a validation set, and the underlying mechanisms were not further studied.
Comparing to the study by Zhou et al., this study not only developed a risk score system based on survival-related miRNAs from The Cancer Genome Atlas (TCGA), but also validate the risk score system on a Gene Expression Omnibus (GEO) dataset. Besides, stratified Cox regression analyses were conducted to identify significant clinical variables associated with survival. In addition, survival-related differentially expressed genes (DEGs) were identified. Gene ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) enrichment analysis were performed for function annotation of the DEGs identified. This study would provide novel insights into the associations of miRNAs with survival of patients with pancreatic cancer.
Materials and methods
Datasets
A miRNA-seq dataset (platform: Illumina HiSeq 2000 RNA Sequencing) and a mRNA-seq dataset (platform: Illumina HiSeq 2000 RNA Sequencing) were downloaded from TCGA (
As a validation dataset, a gene expression dataset (GSE62498) [11] was downloaded from GEO database (
Baseline and clinical characteristics of patients in TCGA and GSE26498 datasets
Baseline and clinical characteristics of patients in TCGA and GSE26498 datasets
“–” means information not available; SD, standard deviation; TCGA, The Cancer Genome Atlas.
Of the 179 pancreatic cancer specimens, 15 did not have survival results and 19 were alive with survival record shorter than 6 months, both of which thus were deleted from the study. The remaining 146 specimens were selected as training set and divided into two groups: bad prognosis group in which patients died in 6 months after diagnosis, and good prognosis group in which patients lived longer than 24 months. Differentially expressed miRNAs were screened between the bad prognosis group, and the good prognosis group by using Limma package [12] in R3.1.0 language, with false discovery rate (FDR)
Prognosis-related miRNA screening
In order to identify significant miRNAs associated with overall survival of pancreatic adenocarcinoma patients, univariate Cox proportional hazards regression analysis [13] was performed on the differentially expressed miRNAs screened using Survival package in R3.1.0 language. The log-rank test
Risk scoring system
A risk score for a patient corresponded to a patient’s risk of death and was calculated as a linear combination of miRNAs expression levels weighted by regression coefficient (
Analysis of correlations of patients’ clinical characteristics with survival
Cox univariate and multivariate regression analysis was performed to assess the associations of survival of patients in two sets with their clinical variables including age, gender, pathologic_M, alcohol, tobacco, chronic pancreatitis history, diabetes history, pathologic_N, pathologic_T, pathologic_stage, radiation therapy, new tumor, targeted molecular therapy, and risk score.
Stratified analyses
Correlations of each clinical variable with expression of each signature miRNA included in the risk scoring formula was evaluated by using Cox univariate regression analysis with survival package in R language [15]. Stratified Cox regression analyses were performed in high and low risk patients according to each clinical variable as well as comparison between patients stratifed by each clinical variable in high and low risk groups. Survival curves were made by the Kaplan-Meier and log-rank method.
GO and KEGG enrichment analyses
Following identification of mRNAs corresponding to the miRNAs in the training set based on individual specimen number, DEGs between the high risk group and the low risk group were identified using Limma package (2) in R3.1.0 language using FDR
Results
Identification of differentially expressed miRNAs
After cutting one miRNA with averaged expression less than 1 reads per million (RPM), there were total 318 miRNAs left in the training set. Of remaining 146 specimens in the training set, 18 were included in the bad prognosis group and 19 were included in the good prognosis group. Consequentially, a total of 36 differentially expressed miRNAs were selected between the good prognosis group and the bad prognosis group.
Survival of patients in different groups stratified by low and high risk in TCGA (A) and GSE62498 (B).
Comparison of 8 prognostic miRNAs expression between high and low risk patients in TCGA (A) and GSE62498 dataset (B). *0.01 
Univariate Cox regression analysis found that 16 out of the 36 differentially expressed miRNAs were significantly related to prognosis of patients. With the 16 significant miRNAs, multivariate Cox regression analysis was employed and found 8 miRNAs were independent predictors of survival, including miR-1301, miR-598, miR-1180, miR-155, miR-496, miR-203, miR-193b, miR-135b (Table 2). With the 8-miRNA prognostic signature, a risk scoring system for prediction of prognosis was formulated as follow:
Eight signature miRNAs in the risk scoring system
Eight signature miRNAs in the risk scoring system
HR, hazard ratio.
Kaplan-Meier survival curves for high and low risk patients in different ages. A, Kaplan-Meier curves for patients with age 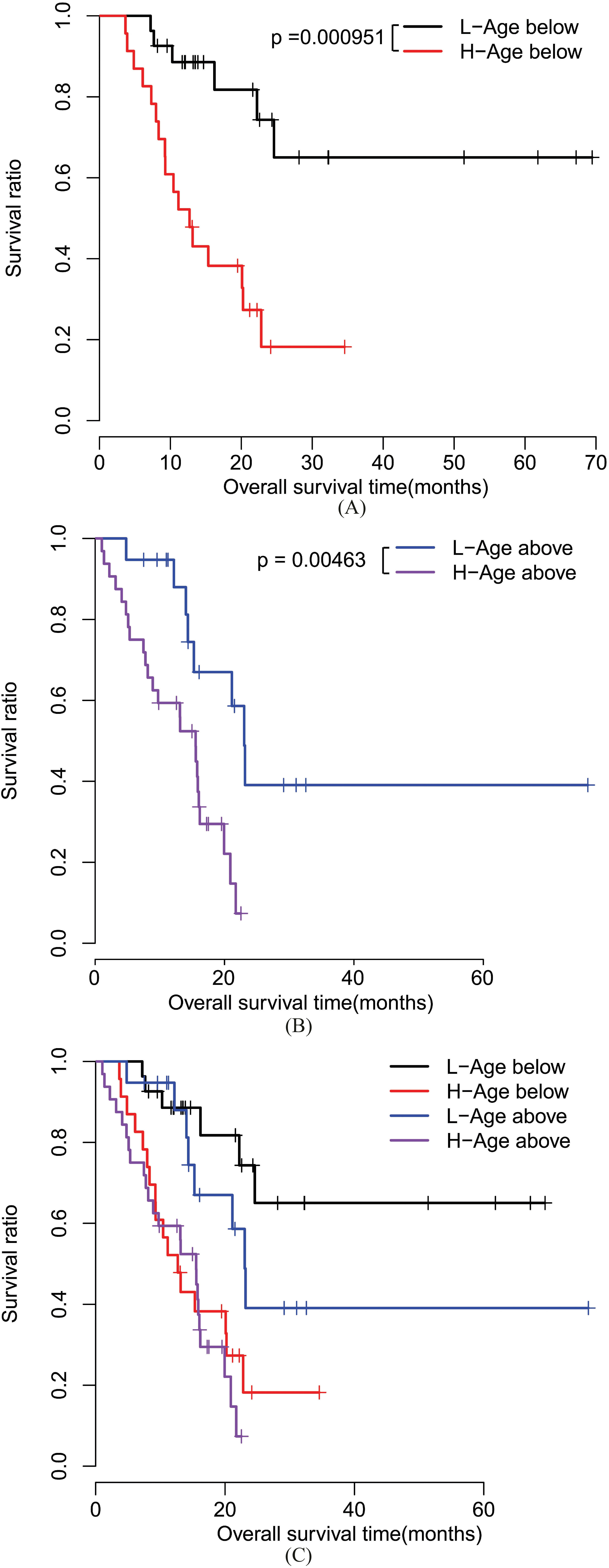
Risk score calculated for each individual patient in the training set (TCGA dataset) and the median risk score was 0.944. All patients in the training set were thus grouped into high risk patients (risk score
As shown in Fig. 3, in both TCGA and GSE62498 dataset, expression levels of the 8 prognostic miRNAs were obviously different between the high risk group and the low risk group.
Results of Cox regression analyses on clinical characteristics and risk score
Univariate Cox regression analysis for each signature miRNA expression in patients stratified by clinical features
Stratified Cox regression analyses for survival ratio between high and low risk patients among different subgroups
Upon univariate and multivariate Cox regression analyses, the study found that risk score, new tumor and targeted molecular therapy were independent predictors of prognosis of pancreatic cancer patients (Table 3).
Univariate Cox regression analysis was performed on correlations of each signature miRNA expression with each clinical characteristic. Table 4 showed that miR-203 expression was significantly different between patients
As shown in Table 5, patients in both training and validation sets were stratified according to each clinical characteristic, separately. Stratified cox regression analyses found that survival ratio was significantly different between the high risk patients and the low risk patients regardless of age, pathologic_N, targeted molecular therapy, and new tumor (
Kaplan-Meier survival curves for patients stratified by chronic pancreatitis history. A, Kaplan-Meier curves for the patients without chronic pancreatitis history. B, Kaplan-Meier curves in the patients with chronic pancreatitis history. C, a combined image of A and B. L, low risk; H, high risk.
Kaplan-Meier survival curves for patients stratified by diabetes history. A, Kaplan-Meier curves for the patients without diabetes history. B, Kaplan-Meier curves for the patients with diabetes history. C, a combined image of A and B. L, low risk; H, high risk.
Kaplan-Meier survival curves for patients stratified by alcohol. A, Kaplan-Meier curves for the patients not drinking alcohol. B, Kaplan-Meier curves in the patients drinking alcohol. C, a combined image of A and B. L, low risk; H, high risk.
Kaplan-Meier survival curves for patients stratified by pathologic_N. A, Kaplan-Meier curves for pathologic N0 patients. B, Kaplan-Meier curves for pathologic N1 patients. C, a combined image of A and B. L, low risk; H, high risk.
Kaplan-Meier survival curves for patients stratified by targeted molecular therapy. A, Kaplan-Meier curves for patients not receiving targeted molecular therapy. B, Kaplan-Meier curves for patients receiving targeted molecular therapy. C, a combined image of A and B. L, low risk; H, high risk.
Kaplan-Meier survival curves for patients stratified by new tumor. A, Kaplan-Meier curves for patients without new tumor. B, Kaplan-Meier curves for patients with new tumor. C, a combined image of A and B. L, low risk; H, high risk.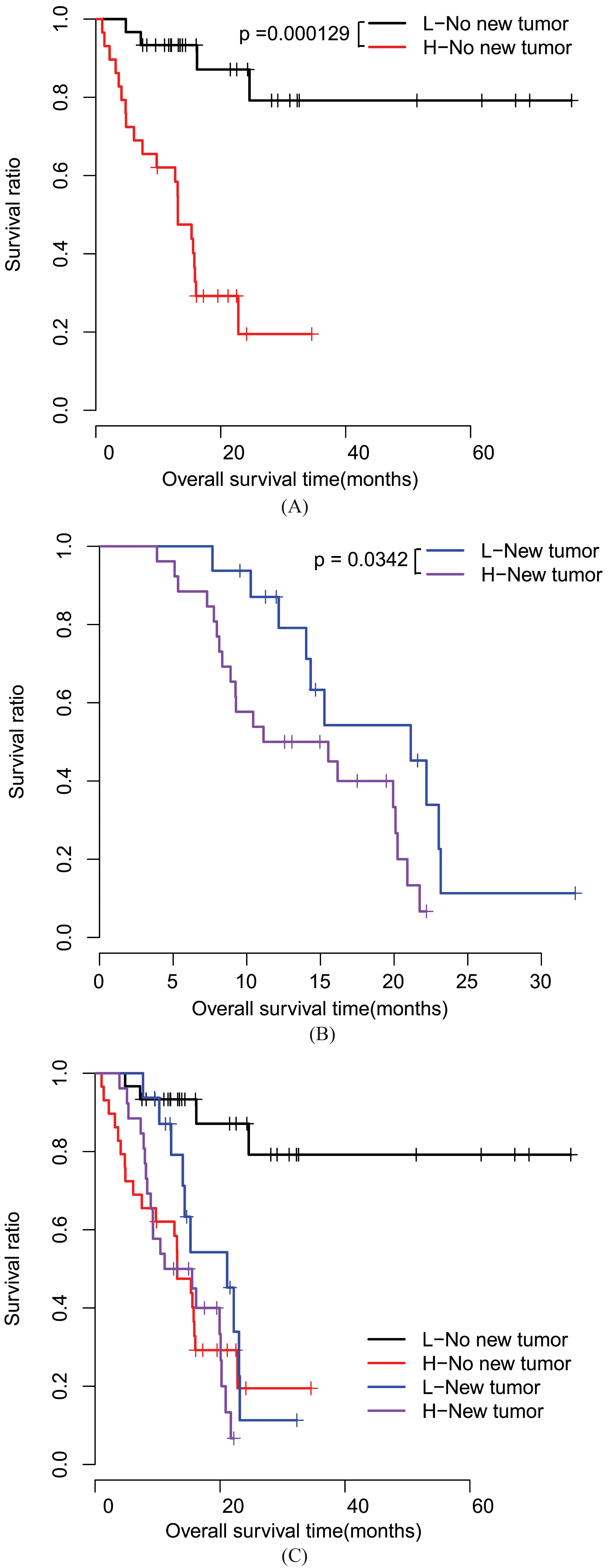
Moreover, stratified analyses found that survival ratio was significantly different (Table 6) between low risk patients with or without new tumor (
Besides a heatmap displaying the expression of the 6 prognostic miRNAs, distribution of risk score and survival time of each patient in the training set and the validate set was shown in Fig. 6.
Stratified cox regression analyses for survival ratio in high and low risk patients
The study identified a total of 143 DEGs between high risk and low risk patients in the training set (FDR
Discussion
Pancreatic cancer ranks fourth in the list cancer-related deaths in the western world with a poor prognosis [19]. The current study defined a panel of 8 signature miRNAs using data from TCGA, and established a risk scoring system based on the 8-miRNA signature, which could strongly predict the survival of patients with pancreatic adenocarcinoma. Prognostic value of the risk scoring system was validated on a GEO dataset. Moreover, multivariate Cox regression analysis identfied risk score as an independent risk predictor of prognosis.
Kaplan-Meier survival curves for high or low risk patients stratified by targeted molecular therapy. A, Kaplan-Meier curves for low risk patients stratified by targeted new therapy. B, Kaplan-Meier curves for high risk patients stratified by targeted new therapy. C, a combined image of A and B. L, low risk; H, high risk.
Kaplan-Meier survival curves for high or low risk patients stratified by new tumor. A, Kaplan-Meier curves for low risk patients stratified by new tumor. B, Kaplan-Meier curves for high risk patients stratified by new tumor. C, a combined image of A and B. L, low risk; H, high risk.
Kaplan-Meier survival curves for high or low risk patients stratified by radiation therapy. A, Kaplan-Meier curves for low risk patients stratified by radiation therapy. B, Kaplan-Meier curves for high risk patients stratified by radiation therapy. C, a combined image of A and B. L, low risk; H, high risk.
Significant GO terms and KEGG pathways
GO, gene ontology; KEGG, Kyoto Encyclopedia of Genes and Genomes.
The study suggest that the 8 prognostic miRNAs, including miR-1301, miR-598, miR-1180, miR-155, miR-496, miR-203, miR-193b and miR-135b were significantly associated with prognosis of pancreatic adenocarcinoma. Unsurprisingly, previous researchers have found that some of the 8 miRNAs are involved in pancreatic cancer.
Risk score (upper), survival time (middle) and expression of 8 signature miRNAs (lower) in the training set (A) and the validation set (B). Red points stand for dead specimens; black points stand for alive specimens.
Functional annotation of DEGs. A, a heat map of top 20 DEGs. Vertical axis represents genes, and horizontal axis represents samples with low to high risk. B, GO terms enriched with negative (upper pie chart) or positive DEGs (lower pie chart). C, significant signaling pathways. In a pie chart, each sector corresponds to the number of genes enriched with each GO term or pathway.
For instance, it has long been reported that miR-155 is overexpressed in pancreatic cancer [20]. Besides, miR-155 plays a part in regulating pancreatic cancer invasion and migration [21], and promotes cellular oxidative stress [22]. Moreover, Liu et al. provide evidence that miR-155 is related to prognosis of pancreatic cancer [23], which is in accordance with the finding of the present study. Similarly, Greither et al. have reported that up-regulated expression of miR-155 and miR-203 is correlated with poor survival of patients with pancreatic tumor [24]. Furthermore, increasing studies have found that miR-203 is involved in modulating tumor cells proliferation, apoptosis, cell cycle progression, migration and invasion in pancreatic cancers [25, 26]. Previous studies have established that miR-193b plays a role in tumor growth and metastasis in pancreatic cancer [27, 28]. Munding et al. conducted global microRNA expression profiling of tumor tissues and suggest that miR-135b is a promising biomarker for pancreatic ductal adenocarcinoma [29]. Nonetheless, few previous studies have reported the associations of miR-1301, miR-598, miR-1180 and miR-496 with prognosis of pancreatic adenocarcinoma.
In the present study, new tumor and targeted molecular therapy were defined as independent predictors of prognosis. Similarly, results of stratified cox regression analyses showed that targeted molecular therapy and new tumor are associated with survival of patients. In line with these findings, recent studies have established that individualized molecular therapy is particular pertinent for pancreatic cancer [30, 31]. Furthermore, the study found that presence of new tumor was significantly associated with miR-1301, miR-203 and miR-193b expression, while targeted molecular therapy was significantly related to miR-1180 expression. These observations indicate that new tumor and targeted molecular therapy have a bearing on survival of patients with pancreatic adenocarcinoma partly through regulating miR-1301, miR-203 and miR-193b expression.
In the present study, DEGs between high and low risk patients were significantly enriched for regulation of transforming growth factor beta receptor, potassium ion transport and MAPK signaling pathway, suggesting that transforming growth factor beta receptor, potassium ion transport and MAPK signaling pathway may play a role in prognosis. Previous studies have reported that expression of transforming growth factor beta receptor correlates with survival of patients with pancreatic cancer [32] and participate in regulating pancreatic cancer cell metastasis [33]. Recently, Sauter et al. reports that two-pore potassium channels are key drivers of tumor progression and thus recommended as novel therapeutic target in pancreatic cancer [34]. A rich body of evidence have proved that MAPK pathway is implicated in pancreatic cancer growth, invasiveness and progression [34, 35]. Noticeably, in a study by Ikeda et al. [36]. MAPK signaling pathway activation induced deregulated expression of 183 miRNAs, of which miR193b was most remarkably inhibited. It indicates that MAPK activation may be partly responsible for the aberrant expression of miRNAs in pancreatic cancer. MAPK pathway might affect prognosis by regulating expression of miRNAs.
There are some limitations in the study. Firstly, its sample size is limited. Further studies with a larger number of patients are warranted to verify the prognostic value of the risk scoring system. Secondly, this study does not include experiments that are needed to validate the results of the study extracted from analysis of miRNA expression profiles from TCGA. Further validation experiment would provide exact information about pathogenesis of pancreatic adenocarcinoma.
This study suggests a risk scoring system based on 8-miRNA signature for assessment of prognosis of patients with pancreatic adenocarcinoma. New tumor and targeted molecular therapy were defined as independent predictors of prognosis. Transforming growth factor beta receptor, potassium ion transport and MAPK signaling pathway may be associated with prognosis in pancreatic adenocarcinoma. This study proposes a novel method to predict the survival of patients, which would pave a way for the determination of a suitable individualized therapy for patients. It is necessary to establish the clinical application of this risk scoring system for pancreatic adenocarcinoma.
Footnotes
Conflict of interest
None.
