Abstract
OBJECTIVE:
This study aimed to explore the correlation of long non-coding RNA taurine-upregulated gene 1 (lncRNA TUG1) expression with clinicopathological features and its predictive value for treatment response and survival profiles in refractory or relapsed acute myeloid leukemia (R/R AML) patients.
METHODS:
Seventy three R/R AML patients who received cladribine combined with cytarabine and granulocyte colony-stimulating factor (G-CSF) (CLAG) or fludarabine combined with cytarabine and G-CSF (FLAG) based chemotherapy and 37 non-malignant controls were recruited. LncRNA TUG1 expression was detected in bone marrow sample obtained before treatment. Complete response (CR), partial response (PR), overall response rate (ORR) and overall survival (OS) were evaluated.
RESULTS:
LncRNA TUG1 expression was upregulated in R/R AML patients compared to controls. It was also elevated in R/R AML patients with age
CONCLUSIONS:
In conclusion, lncRNA TUG1 expression was elevated in R/R AML patients, and it might serve as a potential biomarker for poor prognosis in R/R AML patients treated with CLAG or FLAG based chemotherapy.
Introduction
Acute myeloid leukemia (AML), a common myeloid disease, is characterized by the proliferative, clonal, abnormally differentiated, or occasionally poorly differentiated cells penetrating into the bone marrow, blood and other tissues [1, 2]. The hemopoietic progenitor cells in AML lose the normal differentiation function and the regular respond to proliferation, and such a disorder results in fatal infection, bleeding or organ infiltration [2]. Although the complete response (CR) rate reaches 80% in patients receiving initial induction chemotherapy, there are still some patients classified as refractory or relapsed AML (R/R AML) patients: approximately 20% to 50% AML patients ultimately diagnosed as refractory disease, and 20% to 70% patients relapse after achieving CR with initial induction chemotherapy [3, 4, 5]. For R/R AML patients, the 1-year survival is only 29% and 5-year survival is as low as 11% [6]. Hence, effective chemotherapy for R/R AML patients is needed.
One of the common therapies for R/R AML patients is the chemotherapy regimen consisting of cytarabine, purine analogs and granulocyte colony-stimulating factor (G-CSF) [3, 7]. For purine analogs, fludarabine and cladribine are widely applied in treating R/R AML [8, 9, 10]. The regimen of fludarabine combined with cytarabine and G-CSF (FLAG) achieves CR of 46% to 63% in R/R AML patients [4, 11, 12]. Meanwhile, another frequently-used regimen, cladribine with cytarabine and G-CSF (CLAG), has obtained CR in approximately 78.8% [13, 14]. For Chinese R/R AML patients, CLAG has been proposed as the first-line treatment [13, 14]. These chemotherapies elevate the remission rates in R/R AML patients, whereas only around 20% of patients attain long-term relapse-free survival [15]. Increasing evidence has demonstrated that knowledge of prognostic biomarkers for R/R AML would contribute to extending survival of these patients. Thus, exploration of novel biomarkers for prognosis is important to improve the treatment outcomes of R/R AML patients [16].
Long non-coding RNA (lncRNA) is a class of endogenous RNA molecules with transcript length more than 200 nucleotides [16]. As reported, lncRNA lacks protein-coding potential and plays a crucial role in hematological malignancies [16, 17]. LncRNA taurine-upregulated gene 1 (TUG1), located at chromosome 22q12, is initially identified as a key regulator in the formation of photoreceptors and retinal development [18, 19]. Of late, the fact that lncRNA TUG1 takes part in carcinogenesis in a number of cancers such as breast cancer, bladder cancer and glioma is reported [18, 20]. Whereas, the expression and biologic effect of lncRNA TUG1 in R/R AML patients are rarely investigated. Hence, this study was conducted to evaluate the correlation of lncRNA TUG1 expression with clinicopathological features. Furthermore, the predictive value of lncRNA TUG1 for treatment response and survival profile in R/R AML patients treated by CLAG or FLAG chemotherapy was investigated.
Methods
Participants
This study was a part of the cohort study investigating the efficacy of CLAG compared to FLAG treatment in R/R AML patients. Seventy three R/R AML patients who received purine analogue based chemotherapy regimens (CLAG or FLAG) from Jul 2013 to Jun 2016 at Department of Hematology in Affiliated Hospital of North Sichuan Medical College, were consecutively enrolled in this prospective cohort study. The inclusion criteria were as follows: (1) Diagnosed as AML according to classification of morphology, immunology, cytogenetics, molecular biology (MICM); (2) Refractory or relapsed disease; (3) Bone marrow (BM) blasts above 10%; (4) About to receive purine analogue based chemotherapy regimens (CLAG or FLAG); (5) Able to be followed up regularly. The exclusion criteria were: (1) Previous purine analogue based chemotherapy (which indicated if multiple-lines of purine analogue based chemotherapy were performed in one patient, only the first time would be included); (2) Severe heart dysfunction, severe arrhythmia, severe lung dysfunction, severe hepatic or renal dysfunction; (3) Other solid tumor history; (4) Known allergic or resistant to drugs included in CLAG or FLAG regimens; (5) Patients in pregnancy or lactation, or planned for pregnancy.
Besides, 37 age and gender-matched non-malignant patients who received bone marrow biopsy (including 24 thrombocytopenic purpuras, 8 infectious diseases and 5 health donors) were consecutively enrolled as controls to elevate the expression of lncRNA TUG1 in R/R AML and controls.
The Institutional Review Board of Affiliated Hospital of North Sichuan Medical College approved this study, and written informed consents were obtained from all patients. This study was conducted according to Declaration of Helsinki.
Sample collection
Bone marrow samples were collected before treatment in R/R AML patients, and were also collected in non-malignant controls. Total RNA was extracted with TRIzol Reagent (Invitrogen, USA), according to the manufacturer’s instructions.
Quantitative polymerase chain reaction (qPCR) assay for lncRNA TUG1 expression
The RNA was quantified by OD 260 (Takara, Japan). Then 1
Treatments
The treatments of R/R AML patients were not intervened in this cohort study, and patients received purine analogue based chemotherapy regimens (CLAG or FLAG) “on demand” according to disease condition and patients’ wills. Patients were divided into two groups according to the actual treatment regimens: (1) CLAG group (
Assessments
CR, partial response (PR) and overall response rate (ORR) were evaluated in 62 patients (available for assessment). Percentage of patients achieving allo-HSCT was also assessed in all patients. Overall survival (OS) was calculated from the date of treatment to the time of death from any cause. The median follow-up duration was 10.0 months (1/4–3/4 quartiles: 4.0–15.5 months), and the last follow up month was Feb 2017.
Definitions
Refractory AML was defined as (1) patients did not achieve CR after two courses of induction chemotherapy by standard protocol; (2) patients relapsed within 6 months after the first CR; (3) patients relapsed at 6 months or above after first CR and fail by the subsequent induction chemotherapy; (4) patients relapsed more than 2 times; (5) extra-medullary infiltration of leukemia. Relapsed AML was defined as leukemic cells reappeared in peripheral blood or the percentage of bone marrow blasts over 10%. Risk stratification was assessed by cytogenetics and molecular abnormalities according to NCCN guidelines. CR was defined as BM with at least 20% cellularity and BM blasts below 5% at steady state after treatment, without cytological evidence of leukemia, no transfusion requirement, leucocyte count above 1
Baseline characteristics of R/R AML patients
Baseline characteristics of R/R AML patients
Data were presented as a mean value
Statistical analysis was carried out by SPSS 21.0 software (IBM, USA) and Graphpad Prism 6.01 software (GraphPad Software Inc, USA). Data was mainly presented as mean value
Results
Baseline characteristics
Comparison of lncRNA TUG1 relative expression between R/R AML patients and controls. LncRNA TUG1 relative expression was remarkably higher in R/R AML patients compared with the controls. Comparison between groups was performed by Wilcoxon rank sum test. 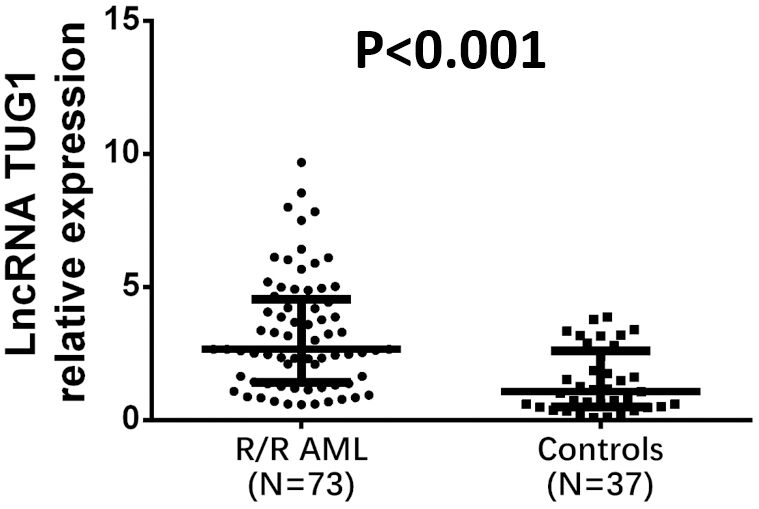
Totally 73 R/R AML patients with mean age of 48.73
Comparison of lncRNA TUG1 relative expression was carried out between R/R AML patients (
Analysis of lncRNA TUG1 expression in subgroups. Expression of lncRNA TUG1 was elevated in patients with age 
Comparison of CR and ORR between patients with lncRNA TUG1 high expression and lncRNA TUG1 low expression. Patients with lncRNA TUG1 high expression achieved numerically lower CR than patients with lncRNA TUG1 low expression without significant difference (A). Patients with lncRNA TUG1 high expression obtained decreased ORR compared with patients with lncRNA TUG1 low expression(B). Comparison between groups was performed by Chi-square test. 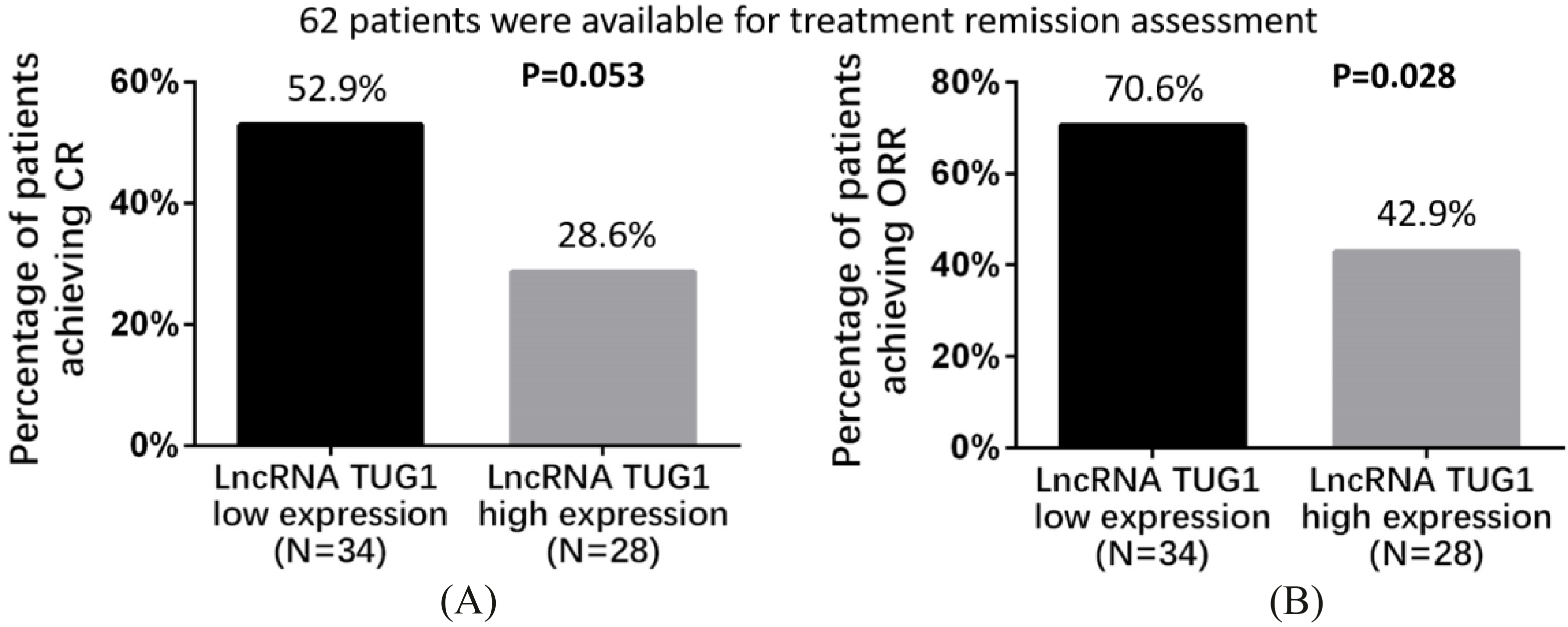
Factors affecting CR achievement by logistic regression analysis
Data were presented as
Factors affecting ORR achievement by logistic regression analysis
Data were presented as
Comparison of lncRNA TUG1 expression was performed between subgroups divided by clinicopathological features (Fig. 2). LncRNA TUG1 expression was increased in patients with age
Treatment response in patients with high lncRNA TUG1 expression and patients with low lncRNA TUG1 expression
Totally 62 patients were included in treatment remission assessment and were divided into lncRNA TUG1 high expression group and low expression group according to the median value of lncRNA TUG1. Patients with lncRNA TUG1 high expression achieved CR of 28.6%, which was numerically lower than that in patients with lncRNA TUG1 low expression (52.9%), while no significant difference was observed (
Analysis of factors affecting CR
Factors affecting CR were assessed by univariate logistic regression and exhibited in Table 2, which revealed that lncRNA TUG1 high expression was numerically correlated with a lower possibility of CR (
Factors affecting OS by Cox’s proportional hazards regression analysis
Factors affecting OS by Cox’s proportional hazards regression analysis
Data were presented as
OS in patients with lncRNA TUG1 high expression and patients with lncRNA TUG1 low expression. OS of patients with lncRNA TUG1 high expression was greatly lower than that in patients with lncRNA TUG1 low expression. K-M curves were used to display OS and log-rank test was conducted to compare OS between two groups. 
As shown in Table 3, lncRNA TUG1 high expression predicted unfavorable ORR (
Comparison of OS between lncRNA TUG1 high expression and low expression groups
As shown in Fig. 4, the median value of OS in patients with lncRNA TUG1 high expression was 6.0 (95% CI 3.6–8.4) months, which was remarkably reduced compared with patients with lncRNA TUG1 low expression (14.0 (95% CI 8.9–19.1) months) (
Analysis of factors affecting OS
Univariate Cox’s proportional hazards regression was used to assess the factors influencing OS, which disclosed that lncRNA TUG1 (
Discussion
In the present study, we compared the treatment outcomes between R/R AML patients with high lncRNA expression and R/R AML patients with low lncRNA expression receiving CLAG or FLAG chemotherapy. The major results we obtained were as follows: (1) R/R AML patients possessed elevated lncRNA TUG1 expression compared with controls; (2) Patients with lncRNA TUG1 high expression achieved lower ORR and numerically decreased CR, as well as shorter OS compared to patients with lncRNA TUG1 low expression. When further evaluated by multivariate analysis, lncRNA TUG1 high expression was an independent predictive factor for the absence of ORR and reduced OS.
LncRNA is the common RNA molecule comprising 200 nucleotides to 100 kilobases in length [21, 22, 23]. Although lncRNA is not able to encode functional proteins, it plays a key role in maintaining physiological processes such as proliferation, differentiation and apoptosis of cells through various mechanisms, including: (1) reconstruction of chromatins; (2) regulation of recruiting transcription factors or co-activators; (3) negative modulation of the RNA polymerase II activity; (4) alternative mosaicking of pre-mRNAs; (5) disruption of the mRNA stability; (6) isolation of microRNAs [24, 25, 26]. Accumulating evidence has revealed that lncRNA is considered as a biomarker in predicting treatment outcomes in various cancers, such as B-cell malignancies, bladder cancer, hepatocellular carcinoma and osteosarcoma and AML [20, 27, 28]. As to AML, a previous study explores the influence of lncRNA growth arrest-specific 5 (lncRNA GAS5) expression on OS in 313 AML patients, which demonstrates that relatively higher expression of lncRNA GAS5 predicts reduced OS in AML patients compared with the relatively lower expression of lncRNA GAS5 [29]. Another study investigating the role of lncRNA HOTAIRM1 in 215 immediate-risk AML patients reveals that increased HOTAIRM1 expression (vs. lower HOTAIRM1 expression) is independently correlated with worse OS and LFS as well as increased incidence of relapse [30]. These previous studies indicate dysregulated lncRNA contributes to the disease progress as well as prognosis in AML patients.
Among cancer-related lncRNAs, lncRNA TUG1 is a rising star, which is recently identified as oncogenic lncRNA in some cancers such as B-cell malignancies, oesophageal squamous cell carcinoma, bladder cancer, hepatocellular carcinoma and osteosarcoma [20, 27, 28]. A study carried out by Tan et al. shows that lncRNA TUG1 is upregulated in bladder cancer cell line compared to normal human bladder cell line, and its downregulation suppresses the metastasis of cancer cells [31]. Also, raised lncRNA TUG1 level in osteosarcoma patients is associated with worse OS compared with lower lncRNA TUG1 level [32]. Aforementioned studies demonstrate lncRNA TUG1 is an oncogenic gene in various cancers and associates with poor prognosis. However, limited studies evaluate the predictive value of lncRNA TUG1 in AML patients. A study conducted by Isin et al. discovers that, compared with healthy controls, lncRNA TUG1 presents with remarkably higher expression in patients with multiple myeloma, a hematological malignancy, while only 62 patients with multiple myeloma are enrolled in this previous study, and whether lncRNA TUG1 high expression predicts treatment outcomes is not clarified [33]. Regarding R/R AML, no study has explored the effect of lncRNA TUG1. Hence, our study investigated the expression and prognostic effect of lncRNA TUG1 in R/R AML patients, and we discovered that lncRNA TUG1 expression was elevated in R/R AML patients, moreover, its high expression was correlated with lower ORR and numerically decreased CR as well as worse OS. The possible reasons were as follows: (1) lncRNA TUG1 bound to polycomb repressive complex 2 (PRC2), which was able to hinder the transcription of cell cycle regulatory genes, thereby decreasing malignant cellular proliferation in AML [34]; (2) overexpression of lncRNA TUG1 might contribute to chemoresistance through regulating multiple cell-survival pathways, and finally it led to worse treatment outcomes [35].
Our study also existed some limitations: (1) sample size was relatively small in this study, which just enrolled 73 R/R AML patients, so the statistical power was diminished to some extent; (2) the median follow-up duration in this study was only 10.0 months (1/4–3/4 quartiles: 4.0–15.5 months), which was relatively short, thus the effect of lncRNA TUG1 on long-term treatment outcomes was not evaluated, however, R/R AML presented with short OS, so long-term follow-up duration was hard to be executed as well; (3) detailed mechanisms of lncRNA TUG1 in AML was not investigated.
In conclusion, lncRNA TUG1 expression was elevated in R/R AML patients, and it might serve as a potential biomarker for poor prognosis in R/R AML patients treated with CLAG or FLAG based chemotherapy.
Footnotes
Acknowledgments
This study was supported by General Project Foundation for Natural Science of Sichuan Provincial Education Department (No. 17ZB0167).
Conflict of interest
None.
