Abstract
BACKGROUND:
Altered expressions of miR-186, miR-494 and miR-3651 in whole blood of OSCC patients have already been demonstrated. However, nothing is known about their expression in tumor tissues. This study aimed at evaluating differences in miRNA expression in OSCC tissues compared to healthy oral mucosa and at checking if the altered expression is reflected in corresponding blood.
METHODS:
In 53 OSCC and 40 healthy mucosal tissues the expression of miR-186, -494 and -3651 was determined by RT-qPCR and expression levels were compared between the groups. The altered expression in tissues was compared with that determined for corresponding blood. MiR-3651 target genes were identified using TargetScan software.
RESULTS:
Differential expression of miR-186 and miR-494 could not be observed in tumor tissues compared to normal mucosa (
CONCLUSIONS:
Decreased expression of miR-3651 in OSCC tissues may be a diagnostic biomarker. The opposite change in expression level in the blood might indicate different functions of this miRNA in circulation and tissue.
Background
Oral squamous cell carcinoma (OSCC) is one of the most common malignant tumors of the head and neck region. Despite the improvements in treatment, OSCC is associated with poor prognosis [13]. Early diagnosis and best possible treatment drastically improve patients’ outcome. However, there is a lack of clinically proven biomarkers for diagnosis and prognosis of OSCC and this limits therapeutic decisions [5]. Hence, suitable tumor biomarkers are urgently needed [34]. Their altered expression in tissues could help to better predict the course of disease and improve tumor treatment. In this context, tumor-specific changes that can also be detected in body fluids, are particularly valuable, because they could be useful minimally invasive diagnostic tools for an early diagnosis and oral cancer screening [37] as well as for monitoring the therapeutic response to different therapies by serial measurements [35]. However, today most biomarkers lack a high degree of specificity and sensitivity and cannot be used as reliable tools in diagnostic routine [38]. In addition, conventional therapies of OSCC failed especially at late stages of the disease. Therefore, the development of new effective therapies based on molecular tools, is also required.
MicroRNAs (miRNA) are global players controlling the expression of cellular genes and have an enormous impact in fundamental cellular behavior [3]. Their aberrant expression rates are essentially associated with the tumorigenesis of several carcinomas and are thought to be potential biomarkers for different solid tumors including OSCC [4, 23, 45]. Altered miRNA expression patterns can also be detected in body fluids including plasma, serum and whole blood and represent a potential tool for minimally invasive methods that may aid in risk assessment, diagnosis, prognostic assessments, detection of recurrence and in predicting or monitoring treatment response as well as in making therapy decision [2, 21, 30, 31, 35, 44]. Furthermore, due to their role as intracellular mediators that are necessary for various biological processes, they could serve as targets for therapeutic interventions against cancer development and progression. For a possible therapeutic use, however, the possible functions and the involvement of the miRNAs in particular signaling pathways must be clarified.
Recently, aberrant expression levels of miR-186, miR-494 and miR-3651 in whole blood samples of OSCC patients were verified. It was postulated that they might serve as promising novel biomarkers for screening and an early detection of OSCC by a minimally invasive method based on whole blood samples [31, 32]. However, it is not known whether these miRNA are also differentially expressed in tumor tissues making them to biomarkers and therapeutic tools of OSCC in tissue specimens, too.
Alterations in cancer cells are thought to be reflected in the extracellular space. However, although some miRNA display the same expression patterns in blood and tissues, it was also observed that changed expression levels in blood of cancer patients are not necessarily the same ones that are differentially expressed in cancer tissue samples [16, 18, 19, 42, 46, 47]. Due to the different expression patterns of defined miRNA distinct roles and different functions of the same miRNA in circulation and in tissue were claimed [42, 46]. Hence, expression analysis may help to elucidate their different regulatory effects beyond the tumor cell itself, including interactions with the immune system and tumor stromal cells and could help to provide deeper insights into the function of these miRNA in OSCC tumorigenesis.
The present study aimed at evaluating the difference in miRNA expression patterns in tumor tissues compared to oral mucosa in order to evaluate the utility of these miRNAs as molecular biomarkers in OSCC tissues. Additionally, it was checked whether their dysregulation is reflected in whole blood of OSCC patients. Possible target genes of the miR-3651 should be determined in order to get first insights how this miRNA might affect tumorigenesis.
Demographic characteristics and clinico-histopathological features for the OSCC group and the control group
Demographic characteristics and clinico-histopathological features for the OSCC group and the control group
SD: Standard deviation. *MWU test, **Chi-squared test, ***grouped.
List of miRNAs and snRNAs (endogenous controls) analyzed by miScriptPrimerAssay (Qiagen, Hilden); #
Patients and sample collection
The study was approved by the Ethics Committee of the University of Erlangen-Nuremberg, Erlangen, Germany (approval number: 3962) and patients’ written informed consent was obtained.
Tissue specimens and peripheral whole blood samples of 53 OSCC patients (OSCC group) and 40 healthy volunteers (control group) were collected. Age and gender of all study participants were recorded. Patients were only included in this study if OSCC occurred for the first time. All samples were taken before any treatment occurred. Healthy volunteers were selected based on the absence of any tumor disease, general disease and acute or chronic inflammations in normal oral mucosa. Tissue samples of healthy volunteers were obtained during minor routinely done surgery avoiding additional incisions at the Department of Oral and Maxillofacial Surgery of the Friedrich-Alexander University Erlangen-Nürnberg (FAU). After harvesting the samples were immediately transferred into RNAlater
All samples were collected at the Department of Oral and Maxillofacial Surgery of the Friedrich-Alexander University Erlangen-Nürnberg (FAU) between 2010 and 2014. In order to confirm the diagnosis, all tumor tissue samples were examined by two independent pathologists. Grading (G1–G3; differentiation status), clinical UICC-stage (I–IV) and TNM classification of OSCC were documented according to the guidelines of the World Health Organization and the International Union Against Cancer [25]. Afterwards, tumors were grouped as early (stages I and II) and late (stages III and IV) clinical stages and as N0 and N
Demographic and clinico-histopathological characteristics of the study participants
None of the healthy volunteers showed remarkable oral mucosal changes, like inflammation, hyperplasia or dysplasia. The median values concerning the age of the participants were equal in both groups. There were no statistically relevant differences (
Sampling of tissue and whole peripheral blood
Specimens of healthy volunteers and tumor tissue samples were taken during intraoral surgery and tumor resection, respectively. After harvesting all tissue samples were immediately transferred into RNAlater
MiRNA isolation and RT-qPCR analysis
Whole RNA was extracted using the miRNeasy mini-Kit (Qiagen, Hilden, Germany) for tissue samples and the PAXgeneBlood miRNA Kit (PreAnalyticX GmbH, Hombrechtikon, Switzerland) for blood samples. RNA concentration und purity was determined using a NanoDrop spectrometer (PeqLab, Erlangen, Germany). The RNA samples were stored at
Real-time quantitative reverse transcription-PCR (RT-qPCR) analysis
MiRNA was reverse transcribed using the miScript II RT kit according to the manufacturer’s recommendations (Qiagen, Hilden, Germany). The amplification was performed using the miScript SYBR-Green PCR Kit and miRNA-specific quantitative RT-PCR primer sets on an ABI 7300 Sequence Detection System (ThermoFisher Scientific Inc., Waltham, MA, USA) as formerly described [31]. The features of the miRNAs are summarized in Table 2.
The values of RT-qPCR analyses were normalized by the
Mean values and standard deviations of the CT values of the normalizers RNU6-2 and SNOD44 for the populations (OSCC and healthy controls) and source of the samples (tissue and blood)
Mean values and standard deviations of the CT values of the normalizers RNU6-2 and SNOD44 for the populations (OSCC and healthy controls) and source of the samples (tissue and blood)
The averaged value of both controls was applied as the normalization value. Relative quantification of differences in expression (FC
TargetScan version 7.1 (June 2016) was used to predict regulatory and biological targets of the miR-3651 by searching for the presence of conserved 8 mer, 7 mer, and 6 mer sites that match the seed region by using cumulative weighted context score (CS) of the sites [1]. The importance of the targets was checked by alignment using different algorithmic programs (miRWALK, mRanda, MiR22, TargetScan) that specify the selected target genes, too. Additionally, miRTarBase (
Statistical analysis of RT-qPCR
For statistical evaluation of the RT-qPCR analyses, the program IBM
The correlation between the expression rates of the miRNAs in tissue and blood is illustrated in a scatter plot (Fig. 3). If the pattern of dots slopes from lower left to upper right, it points out a positive (rising) correlation between the variables. If the pattern of dots slopes from upper left to lower right, it refers a negative (falling) correlation. An identity line (trend line) is drawn as a reference. The statistical significance of the correlation was ascertained by the Spearman’s rho test. For all statistical tests,
Box-Whisker Plots of median expression rates of the examined miRNAs. No aberrant expression between the two groups could be detected in tissues for miR-186 (A) and miR-494 (B). A statistically significant twofold downregulation of miR-3651 was observed in OSCC tissues compared to healthy oral mucosa (C). The by ROC curve for miR-3651 calculated AUC value indicates that the two groups could be distinguished by expression analysis of the marker in tissues. The highest Youden (Y) index (circle) could be determined (D).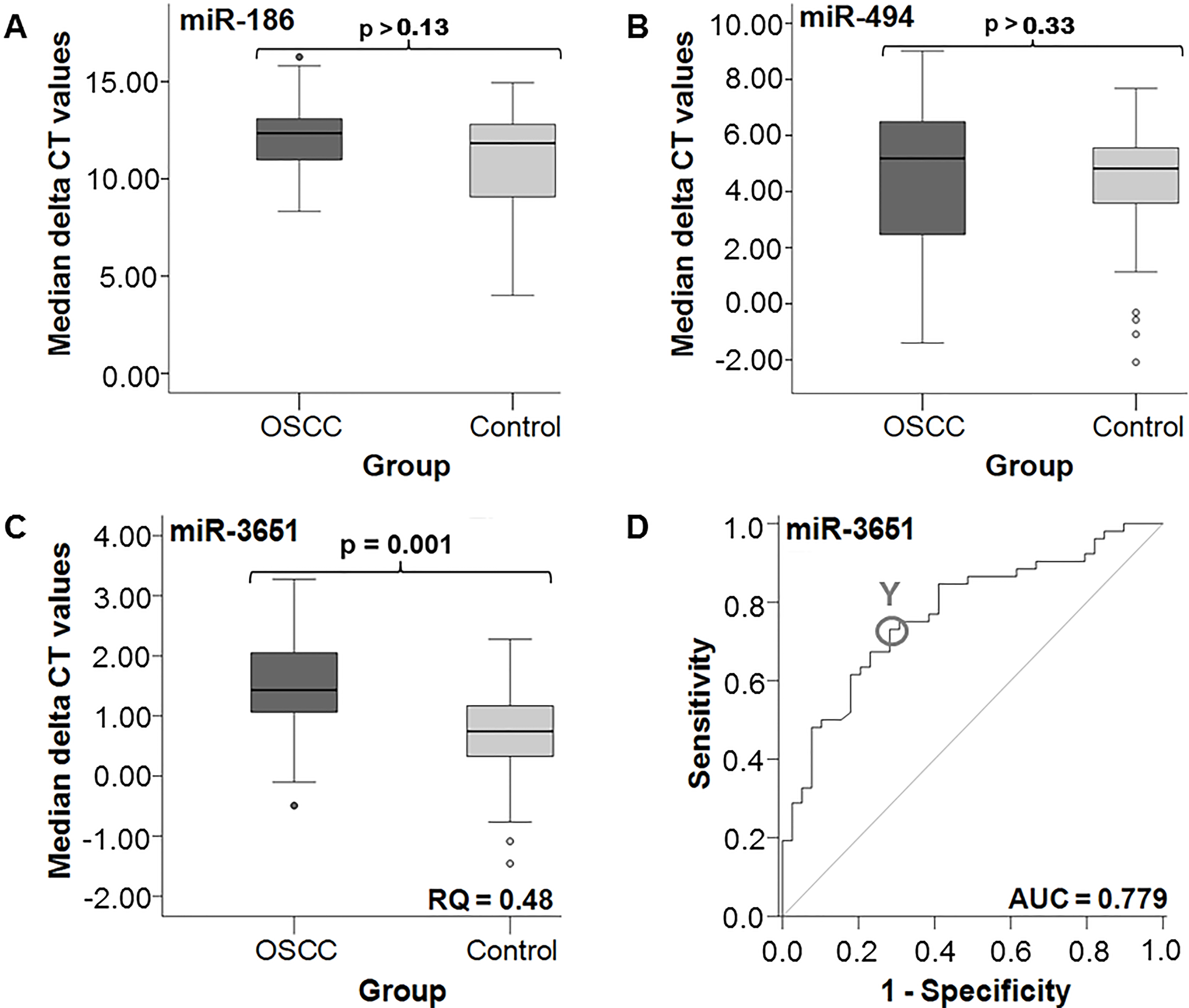
Analysis of altered miRNA expression rates in tissues
Statistical evaluation was done using the determined
Due to these results of altered expression in tissues, in further statistical investigations only expression analyses of miR-3651 were taken into account and are summarized in Table 4. The determined AUC value of 0.779 confirmed the statistical relevance of miR-3651 expression analyses for the discrimination between OSCC and healthy tissues (Fig. 1D). At the same time the parameter reached a high sensitivity (76.1%) and specificity (70.3%). The highest Youden index calculated for miR-3651 implicated the optimal threshold value (COP) to
Statistical results of the comparison of the miRNA expression between the OSCC and the control group in tissue and the blood samples of the same collective of the study participants
Statistical results of the comparison of the miRNA expression between the OSCC and the control group in tissue and the blood samples of the same collective of the study participants
Statistical association was determined by the Mann-Whitney U-test. The AUC and RQ values for the comparison of miRNA expression of miR-186 and miR-494 in OSCC tissues compared to the control group was not determined (n.d.). FC
Associations of aberrant miR-3651 expression in tissue and whole blood with diagnosis and clinico-histopathological data of OSCC patients based on the determined COPs
The determined
A significantly different expression of all tested miRNAs of the used patients collective was already ascertained between the OSCC and the control group in whole blood samples (
Splitting of the OSCC and control group into positive and negative subgroups based on established COPs (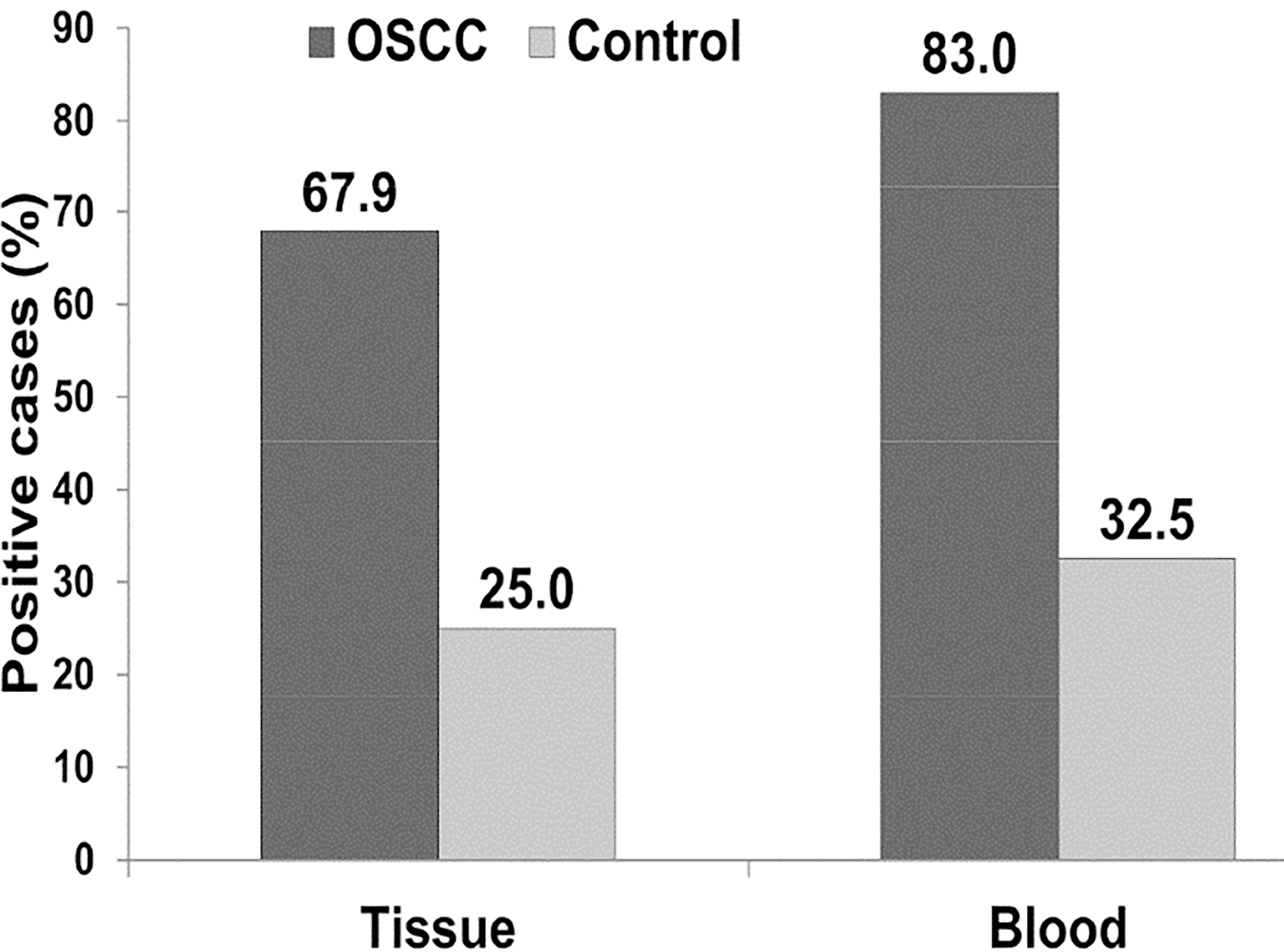
Scatter plot for the correlation between the expression rates of miR-3651 in blood and corresponding tissue. The pattern of dots slopes from upper left to lower right and the identity line exhibits a negative value. That points to a negative correlation. Additionally, the “Spearman’s rank correlation coefficient” (Spearman’s rho) indicates a weak negative correlation (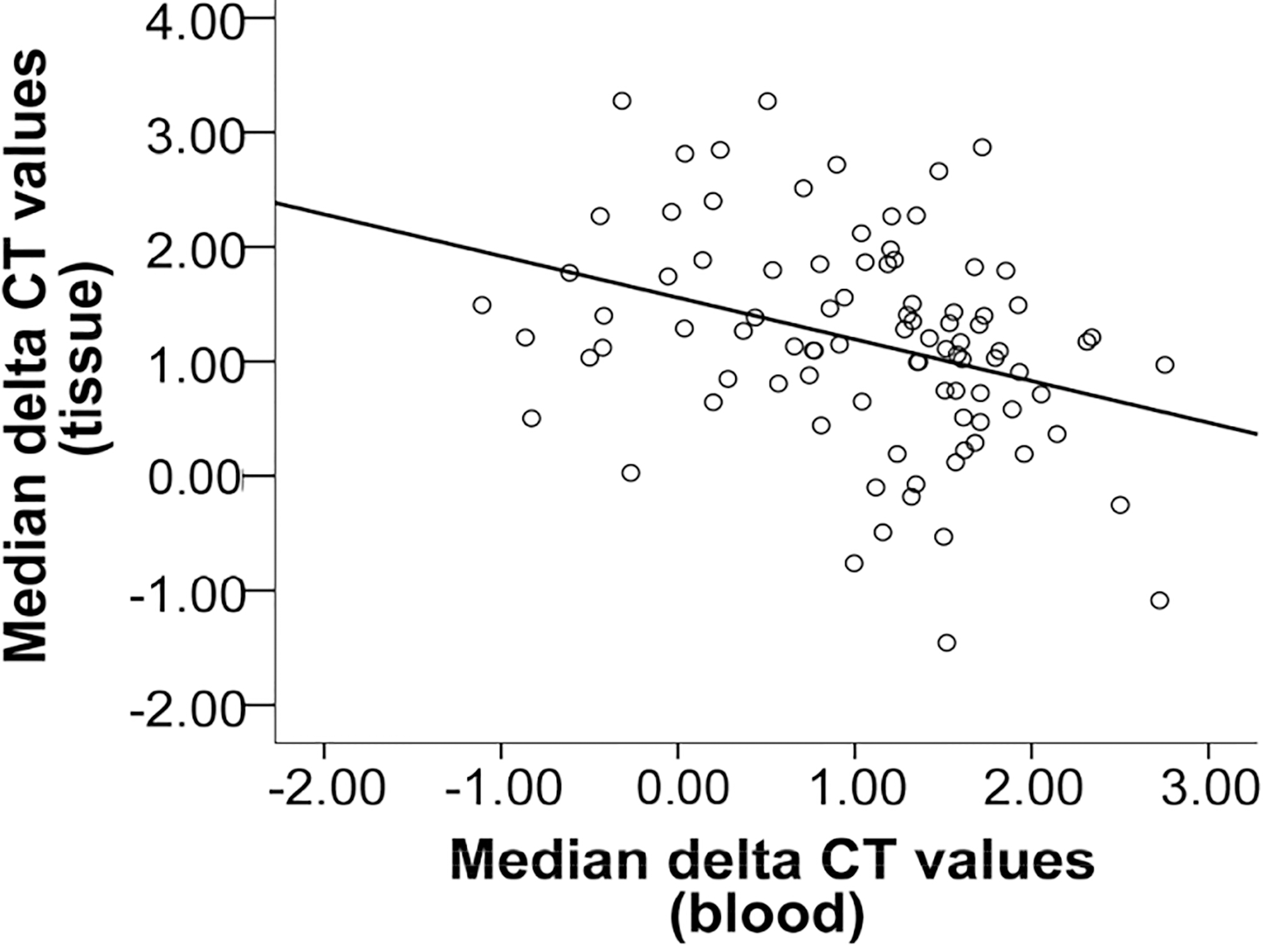
Due to the fact that only the changed expression of the miR-3651 was important for tumor diagnosis in tissue specimens, only the results of further statistical analyses of this miRNA are shown (Table 5, Fig. 2). Based on the determined COP (
Selected putative target transcripts of miR-3651 identified by TARGETSCAN 7.0 and verified by alignment using different algorithmic programs and their putative function in oral carcinogenesis
CSC
Based on the COP for miR-3651 in blood and tissues the positivity of the OSCC group for the parameter was assessed and a significant correlation to diagnosis was ascertained (
The statistical evaluation revealed that there was no association of miR-3651 underexpression in tissue and tumor size (
Correlation between the expression rates of miR-3651 in blood and tissues
An opposite change in miR-3651 expression rates in blood samples compared to its expression rates in corresponding tissue samples could be ascertained. The correlation between the two variables is illustrated in a scatter plot (Fig. 3). The identity line shows a negative slope. The “Spearman’s rank correlation coefficient” (Spearman’s rho) indicates a significant negative correlation (
Putative target transcripts of miR-3651
Totally, 1710 putative targets transcripts of miR-3651 were identified by TargetScan. Most of them have functions controlling pathways which allow the cell to reach the so-called “hallmarks of Cancer” [10]. Through these signaling pathways the miR-3651 could be involved in the regulation of fundamental cellular behavior like proliferation, differentiation, apoptosis, migration, invasion/EMT, angiogenesis and immortalization as well as in cellular energetics. Further targets are also genes which give the cell stem cell properties. In addition, there are also indications for interactions between this miRNA with genes that control the immune system and the immune modulation. Selected gene candidates with their assumed function that have been identified on their relevance to oral carcinogenesis through a search of current scientific literature are summarized in Table 6.
Other different mRNA target prediction algorithms were applied in order to identify possible targets of miR-3651. Using the software programs miRWALK, miRANDA and RNA22 not all targets recognized by TargetScan could be confirmed. Hence, from the artists acting in the immunomodulation pathways only 6 genes, CRISP3, AKT2, TNFSF9, TLR8, CD84 and CD226, are recognized by 4 and 3 algorithmic tests, but not the other target genes identified by TargetScan. This also applies to other genes involved in almost all signaling pathways. The number of different software programs that specify the selected target genes is supplemented in the Table 6.
The first analysis using the miRTarBase (
Discussion
Changes in miRNA expression in tumor tissues and whole blood
The expression of a large number of miRNAs is altered in tissues, plasma, serum, saliva, and whole blood of OSCC patients [23, 30, 31, 45]. Hence, expression analysis of these markers could aid in the diagnosis, prognostic prediction and planning of patient-specific treatments, thus improving the clinical outcome and survival of patients. Additionally, altered expression patterns of circulating miRNAs may serve as minimally invasive screening tests for the detection, prognostication, determination of the host immune response to cancer cells, and monitoring of treatment response in OSCC [30, 35, 44].
In former studies we were able to show that the three miRNAs, miR-186, miR-494 and miR-3651, are differentially expressed in whole blood of OSCC patients when compared to that of healthy volunteers. The changed expression rates were significantly associated to diagnosis of OSCC. Hence, these parameters could be potential blood based minimally invasive biomarkers for OSCC [31, 32]. However, whether their expression rates are altered in tissues and whether expression analyses of these miRNAs are useful diagnostic and prognostic tools in tissue has not yet been investigated.
The expression analyses of miR-186 and miR-494 in tissue samples showed no significant alteration in their expression rates between healthy mucosa and tumor specimens. A significant downregulation of miR-3651 in OSCC tissues compared to healthy tissues could be ascertained. The occurrence of this downregulation was significantly associated to OSCC diagnosis. Thus, altered expression of this miRNA may be useful for diagnostic applications using tissue samples.
It was hypothesized that alterations of miRNAs in cancer cells are reflected in the extracellular space and body fluids because malignant cells deliver upregulated miRNAs or fail to release apparently downregulated species. However, it was already shown that this is not the fact for all changes and also contradicting expression levels in tissue and blood could be assigned [16, 18, 19, 42, 46, 47]. This study indicates that the altered expression rates of the investigated miRNAs in blood were not reflected in tumor tissues of the patients. The altered expression of miR-494 and miR-186 could not be proved in tissue. Moreover, unlike to the already demonstrated, significantly increased expression in the blood of OSCC patients, the miR-3651 was significantly decreased in OSCC tissue. Additionally, the formerly determined prognostic significance of its altered expression in blood could not be ascertained in tissues. These observations suggest that the miR-3651 may have different regulatory features in blood and in tissue.
Contradictory findings of miRNA profiles in the peripheral blood and tumor tissue have been shown throughout previous studies [6, 7, 9]. In order to explain this discrepancy in miRNA expression patterns several hypotheses were formulated. It is postulated, that the concentration of miRNAs in tissue cells is directly regulated by increased or decreased transcription of the miRNAs. However, a decreased amount could also be caused by the active and selective export of specific molecules. It has already been shown, that beside the energy-free passive outflow of miRNAs from broken cells under condition of tissue injury or cell apoptosis there exists an active, energy-dependent and selective segregation by cells as a response to altered gene expression or other various stimuli [28]. On this path miRNAs could be selectively released as non-membrane bound miRNAs associated with protein complexes (argonaute proteins) or via cell-derivated microparticles and exosomes into blood and tissue microenvironment what leads to decreased miRNA concentrations in cells and to increased concentrations in blood [28, 46]. Hence, the tumor cells could have developed a special mechanism to reduce miRNA. The raised concentration of miR-3651 in blood, by its concurrent decrease in tumor cells, could be an indication for such a mechanism. Another explanation is that potential biomarker candidates in fact originate from other sources than the primary tumor itself, such as blood cells, especially immune cells [27, 29, 36]. Our investigation of miRNAs in whole blood samples includes miRNAs from blood cells, released tumor cells, exosomes as well as all circulating, cell-free miRNAs released from the tumor [40]. Since the proportion of circulating miRNAs of the total miRNAs in blood samples is small, it was purposed that the greater amount of cellular miRNAs contained in blood cells may mask or dilute the tumor-specific cell-free miRNAs associated to exosomes or to protein complexes [8]. Additionally, whole blood miRNA profiles were reflect by miRNA expression patterns of peripheral blood mononuclear cells (PBMC) [22]. Hence, the variations in miRNA concentrations are mainly caused by the blood cells and may be the result of systemic reactions linked to the tumor disease. Thus, the tumor characteristic miRNA pattern may more likely results from a specific response of the innate immune system to the tumor rather than the miRNA expression profile of the tumor itself [7, 11, 12, 17]. Therefore, it is not surprising that when analyzing tissue miRNA, we did not find an upregulation of miRNAs which have been previously been described to be differentially expressed in whole blood of OSCC patients.
Nevertheless, one may argue that the primary carcinoma could not be diagnosed by this minimal-invasive method. However, it was already postulated that the contribution of neoplastic cells to total circulating extracellular miRNAs in plasma and serum is relatively small and often undetectable. Consequently, any confirmed changes in circulation must be traced instead to responses to the presence of cancer [44]. Due to our experimental settings for expression analyses in blood different cell types of the immune system are enclosed that cause, in addition to the secretion by tumor cells, an early and measurable change in miRNA expression pattern in whole blood. Therefore, the method might be more sensitive for the detection of a malignancy [12]. Hence, information contained in the cellular fraction could compensate limitations in peripheral detection of the tumor that could be linked to an undetectable level of tumor miRNAs.
The role of miR-3651 in OSCC
To our best knowledge, our work is the first that has found underexpression of miR-3651 in OSCC tissues. Although there is little information about the role of miR-3651 in human cancer, it seems to be involved in tumorigenesis and progression. So, the biomarker was already recommended to be an independent prognostic and diagnostic tool for esophageal, lung and colorectal SCC [26, 41]. Additionally, a correlation between the overexpression of miR-3651 in whole blood of OSCC patients and prognosis could be shown [30, 31].
MiRNAs have classically been categorized as either oncogenic or tumor suppressive. However, today there are many conflicting reports as to whether specific miRNAs are exclusively oncogenic or tumor suppressive [39]. This can be explained if one considers the fact that a single miRNA molecule is able to target hundreds of different mRNAs which encode tumor suppressor as well as oncogenic proteins. Therefore, they influence many different and partly opposite signaling pathways as suppressive or oncogenic miRNAs. Additionally, miRNAs may have different functions in different body compartments and the detection of varying alterations in expression levels and prognostic impact of miRNAs in tissues and blood possibly indicate different effects [42, 46]. Thus, expression analysis of miRNAs in both tissues and blood is necessary to investigate their clinical role in various body components and to elucidate their importance for various regulatory effects beyond the tumor cell itself including interactions with the immune system [46].
In our study the results of expression analyses in tissue and blood are contrary suggesting that the miR-3651 may have different regulatory features in the various body compartments [42]. Moreover, the different alterations of the miR-3651 expression suggests, that it may function as tumor suppressor as well as oncogene, depending on the kind of tissue type as it was shown for other tumor entities [41]. MRNAs that are targeted by miRNAs are reduced by degradation. This means that upregulation of a certain miRNA leads to decrease of the target protein products. Hence, in tissues the miR-3651 may work as a tumor suppressor as its concentration is decreased in malignant cells. Consequently, protein products of the targeted mRNAs acting as oncogenes are more frequently translated and drive on carcinogenesis [39, 44]. In contrast miR-3651 seems to exhibit an oncogenic function in blood.
Putative target genes of miR-3651 and possibly affected signaling pathways
Of great importance, but mostly still unknown, is the physiological role of almost all miRNAs [15]. Due to its ability to regulate the expression of a variety of genes, one miRNA can hypothetically interfere with almost all essential processes of tumorigenesis like cell cycle, proliferation, apoptosis, achievement of stem cell properties, invasion, metastasis, angiogenesis, immortalization [3] and energetics including the metabolism of mitochondria could be assigned causing the Warburg effect [14]. Furthermore, many miRNAs play pathological roles in the immune system [24] and might aid to avoid or to suppress the immune response of the host against cancer cells. This event is one of the most important steps in tumorigenesis. Hence, all miRNAs contribute in the achievement of the “hallmarks of cancer” which are involved potentially in oral cancer [20].
In order to get first clues of how the miR-3651 might intervene into signaling pathways of cancer, potential target genes have been identified. There are various computational approaches to predict targets of miRNAs and one of these, the TargetScan, was used [1, 15]. Indeed, the identified targets can be attributed to all the different biological abilities of cancer hallmarks. Of particular note is that many target genes of miR-3651 are involved in inflammation processes or modulate immune response. Indeed, some elevated circulating miRNAs have been found to be related to cancer-specific molecular pathways in immune cells or to influence the individual’s immune response to the tumor. Particularly noteworthy in this context are three genes that regulate the transcription of the immune checkpoint protein PD-L1 whose overexpression on antigen presenting cells is responsible for tumor induced immunosuppression [44]. Thus, two activators of PD-L1 transcription, AKT 2 and mTOR, are targets of miR-3651. It could be supposed that the decreased level of the miRNA in tissues leads to increased levels of these proteins and consequently to increased PD-L1 expression in tumor cells and therefore to immune suppressive effects [33]. On the other hand overexpression of the target gene PTEN inhibits PD-L1 transcription. Hence, the overexpression of miR-3651 in blood cells could lead to degradation of the inhibitor and subsequently to evaluated levels of PD-L1 and to systemic tumor immune tolerance [33]. This hypothesis is supported by the recent observation that PD-L1 is upregulated in whole blood of patients suffering from advanced OSCC. Moreover, an association between more aggressive behavior of the tumor and a systemic state of immune tolerance was postulated [43].
However, these postulations have to be revisited. Other different mRNA target prediction algorithms were applied in order to identify possible targets of miR-3651 and to check the importance of the targets. Using the software programs miRWALK, miRANDA and RNA22 not all targets recognized by TargetScan could be confirmed. Additionally, other additional targets were found. Hence, although the genes CRISP3, AKT2, TNFSF9, TLR8, CD84 and CD226 recognized by 4 and 3 algorithmic tests, appear to play a role in immunomodulation, but not the other target genes recognized in TargetScan. This also applies to other genes involved in almost all signaling pathways. Therefore, the results show that the TargetScan alone is not sufficient to uniquely determine a target gene. Additionally, the function and target genes of the miR-3651 are still poorly understood. Therefore, the first analysis using the miRTarBase (
Conclusion
The detection of a significant downregulation of the miR-3651 in tumor tissues might be a valuable biomarker for cancer detection in tissue specimens. The altered expression levels of all examined miRNAs in tumor tissues were not reflected in the corresponding whole blood samples. This opponent alteration might be a hint that the miR-3651 plays distinct roles in tissues and circulation.
Footnotes
Acknowledgments
Erlangen University Hospital; Friedrich-Alexander University (FAU). Authors thank Mrs. E. Diebel, Mrs. S. Schönherr and especially Mrs. A. Krautheim-Zenk for their valuable technical support.
Conflict of interest
The authors report no declarations or conflict of interest.
