Abstract
BACKGROUND:
Cervical lymph node metastases (LNM) in papillary thyroid carcinomas (PTCs) are common and develop in approximately 30–80% of PTCs. The presence of cervical LNM significantly increases the rate of locoregional recurrence in PTCs.
OBJECTIVE:
To search for predictive gene signatures for nodal metastasis in PTCs.
METHODS:
We used unsupervised clustering with unbiased manner to compare molecular profiles between PTCs with nodal metastasis and PTCs without nodal metastasis using mRNA-seq of TCGA data. Using gene ontology (GO) and logistic regression test, we generated 12-predictive genes for nodal metastasis in PTCs.
RESULTS:
Unsupervised clustering of mRNA-seq (training set, N
CONCLUSIONS:
These 12 gene signatures could be used to predict the chance of nodal metastasis in PTCs in preoperative evaluation using fine needle aspiration biopsy (FNAB) so that appropriate plan such as central neck dissection could be made.
Introduction
Papillary thyroid carcinoma (PTC) is the most common type of thyroid cancer. Approximately about 80% of all thyroid malignancies are PTCs [1]. Cervical lymph node metastases (LNM) in PTCs are common and develop in approximately 30–80% of PTCs [2, 3, 4]. The impact of neck node metastases on the prognosis of patients with PTCs is controversial [5, 6, 7]. Although many studies have reported that cervical LNM has no major impact on survival of low-risk patients [8], recently there has been growing recognition that cervical LNM can adversely affect survival, particularly in older patients with larger tumors and extrathyroidal extension (ETE) [9, 10, 11]. In contrast to its impact on cancer prognosis, the impact of cervical LNM on cancer recurrence is undisputed. The presence of LNM significantly increases the rate of locoregional recurrence [12, 13]. Since 60% to 75% of all cancer recurrence in the neck occur in lymph nodes, detection of cervical LNM at initial preoperative evaluation is very important to reduce reoperation rates [14].
Ultrasonography has been reported to be the most sensitive method to detect metastatic lymph nodes in patients with PTC [15, 16, 17]. This technique is capable of detecting metastatic lymph nodes as small as 2 to 3 mm in diameter [17]. Therefore, preoperative ultrasonographic staging of the neck might improve surgical management and reduce recurrence. However, 50% to 90% of subclinical micrometastases could not be detected by preoperative ultrasonographic evaluation [18, 19, 20].
Several studies have reported that clinicopathological variables could predict cervical LNM in PTCs. Koo et al. have reported that PTCs with maximal diameter of greater than 1-cm is associated with a high rate of ipsilateral central neck LNM [21]. Kupferman et al. have reported that multifocal cancer within the thyroid is associated with a high rate of ipsilateral lateral neck LNM [22]. However, whether gene signatures could be used to predict nodal metastasis in PTCs is currently unknown.
Therefore, the objective of this study was to compare gene expression profiles between PTCs with nodal metastasis and PTCs without nodal metastasis using mRNA-seq of TCGA data and search for gene signatures that could predict nodal metastasis in PTCs using multiplatform genomic analysis.
Methods
Genomic and clinical data sets
All genomic data of papillary thyroid carcinoma from TCGA project were obtained from TCGA data
Analysis of gene expression data and unsupervised clustering
BRB-ArrayTools software program (
Selection of specific gene signature in each cluster
To select genes differentially expressed between PTCs with nodal metastasis and PTCs without nodal metastasis, we applied stringent cutoff of
Significant canonical signaling pathways enriched in each subgroup
Pathway analysis was carried by using Ingenuity Pathways Analysis (IPA, Ingenuity, Redwood City, CA, USA). Genes from the dataset that were associated with a canonical pathway in Ingenuity Pathways Knowledge Base were considered for analysis. The significance of the association between differently expressed genes and the canonical pathway was measured using Fischer’s exact test (
Unsupervised clustering heat-map showing two distinct molecular patterns of gene expression between PTCs with nodal metastasis and PTCs without nodal metastasis. A hierarchical clustering of gene expression data from 158 PTCs cases in TCGA data was performed. Genes with expression levels at least 2-fold different in at least 15 cases relative to median value across cases were selected for hierarchical clustering analysis. Data are given in matrix format, in which rows represent individual genes while columns represent each patient. Each cell in the matrix represents the expression level of gene feature in an individual pattern. Color red or green in cell reflects relative high or low expression levels, respectively, as indicated in the scale bar.
Altered canonical pathways in node positive PTCs
PTCs: Papillary thyroid carcinomas.
To select gene signatures that could predict nodal metastasis in PTCs, we used 200 differently expressed genes (DEG) between PTCs with nodal metastasis and PTCs without nodal metastasis (Supplementary Data 3). We tried to pick up representative genes for each biologic pathway. For this analysis, we used Gene Ontology Consortium online tools (
Heat-map showing each subtype-specific gene expression pattern. Data given in the matrix represent the expression of a gene feature in an individual patient. We selected nodal metastasis-specific 200 genes. Heat-map is ordered by ratio. Color red or green in cell reflects relative high or low expression levels, respectively.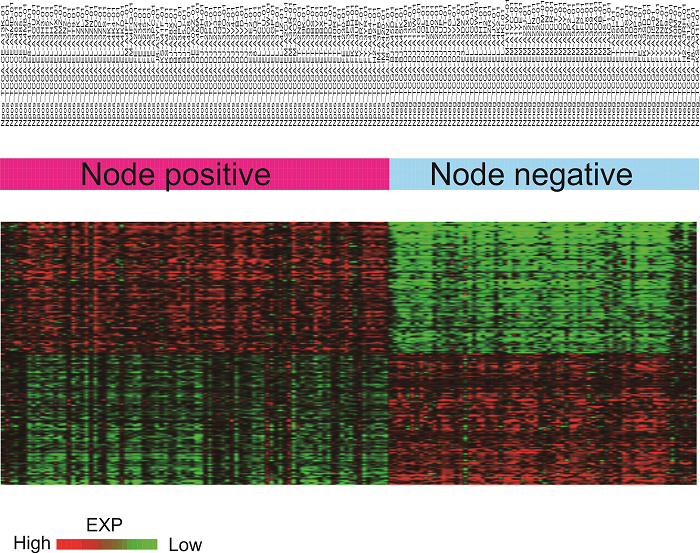
A total of 12 predictive gene signatures were used to evaluate their robustness in predicting nodal metastasis in a validation cohort (N
Statistical analysis
The association of each group with each clinical variable was evaluated using Chi-square test and Fisher’s exact test.
(A) Heat-map showing definitely distinct gene expression pattern between PTCs with node positive (left, red-bar) and PTCs with node negative (right, blue-bar) in training cohort. (B) ROC curve showing the predictive value of 12 predictive genes in training cohort (N 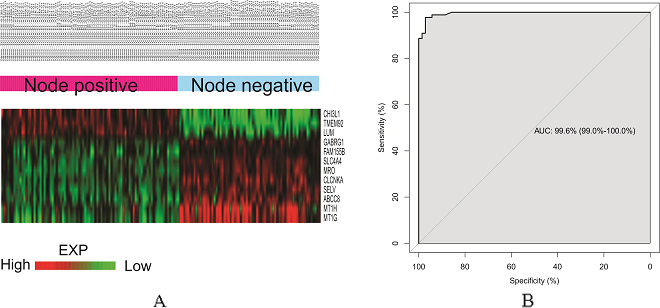
(A) Heat-map showing distinct gene expression pattern between PTCs with node positive (left, red-bar) and PTCs with node negative (right, blue-bar) in validation cohort. (B) ROC curve showing the predictive value of 12 predictive genes in validation cohort (N 
Unsupervised clustering of training cohort (N
158)
To compare gene expression patterns between PTCs with nodal metastasis and PTCs without nodal metastasis (N
Selection of differently expressed genes in each group (PTCs with nodal metastasis vs. PTCs without nodal metastasis)
Next, we tried to select genes that were differentially expressed between PTCs with nodal metastasis and PTCs without nodal metastasis. We applied stringent cutoff of
Significant canonical signaling pathways enriched in PTCs with nodal metastasis
Pathway analysis revealed that PTCs with nodal metastasis group showed different canonical signaling pathways (Table 1 and Supplementary Data 4). PTCs with nodal metastasis showed enriched canonical pathways such as GABA receptor signaling pathway, LXR/ RXR activation pathway, and Wnt/
Selection of specific gene signatures to predict nodal metastasis in PTCs
We used Gene Ontology Consortium online tools and logistic regression test to select representative genes for each biologic pathway. Finally, a total of 12 genes (BCC8, CHI3L1, CLCNKA, FAM155B, GABRG1, LUM, MRO, MT1G, MT1H, SELV, SLC4A4, and TMEM92) were selected (Supplementary Data 5).
Prediction with BCCP for validation cohort
These 12 predictive gene signatures (Supplementary Data 5) were used to predict nodal metastasis in PTCs using a validation cohort (N
Univariate analysis of clinical variables in validation cohort (N
80)
Univariate analysis of clinical variables in validation cohort (N
*means statistically significant
Plot for odds ratios of each variable in validation cohort. Logistic regression with univariate analysis revealed that ETE, multifocality, and 12-gene signatures were significantly correlated with nodal metastasis. Multivariate analysis revealed that these 12 predictive gene signatures had significantly higher odds ratio than other variables.
Results of logistic regression with univariate analysis revealed that ETE, multifocality, and these 12 gene signatures had statistically significant correlations with nodal metastasis. Multivariate analysis revealed that these 12 predictive gene signatures showed more significant odds ratio compared to other variables (Fig. 5 and Table 2).
Discussion
Cervical LNM is common (30% to 80%) in patients with PTC [1, 2]. Although the effect of nodal metastasis on survival is marginal, the presence of LNM significantly increases the rate of locoregional recurrence [2, 3, 4, 5, 6]. For this reason, preoperative detection of metastatic node is important for selecting treatment options.
Several reports have described predictive factors for central nodal metastasis in patients with different thyroid cancers. Extracapsular spread (ECS), lateral LNM, and tumor size exceeding 5 mm are significantly associated with central compartment nodal metastasis in papillary thyroid carcinoma [1]. A maximal diameter exceeding 1 cm has been associated with a high rate of ipsilateral central LNM [11]. However, there has been no report about gene signatures for predicting cervical nodal metastasis in PTCs.
In this study, we generated 12-gene signatures and found that they could predict nodal metastasis of PTC. Their predictive value was evaluated in another cohort and odd ratios of these gene signatures were compared to those of other clinical variables. Odd ratios of these gene signatures for predicting nodal metastasis of PTC was found to be much higher than those of other clinical variables such as ETE or multifocality. With these gene signatures, we can predict the chance of nodal metastasis in PTCs in preoperative evaluation using FNAB to make appropriate plan including central neck dissection.
Recently, large-scale multiplatform analysis of genomic data including gene expression, copy number alteration, somatic mutation, and methylation data has been performed, making it possible to compare two or more tumors with different clinical behavior at molecular level. These genomic studies have revealed that same-tissue origin cancers could be divided into multiple molecular subtypes [27]. These sub-classifications of cancer type make it possible to not only predict clinical behavior, but also suggest appropriate targeted therapy considering specific pathway involved in cancer development and progress [27]. Using gene expression data of PTCs in TCGA data, we suggested 12 gene-signatures in this study to predict nodal metastasis in PTCs. When we used heat-map and ROC curve to evaluate the predictive value of these 12 gene signatures, heat-map in training cohort (N
Cost for microarray analysis of the whole genome has sharply decreased as the price of genome analysis was $100 million in 2001 but now (2017) is less than $500. Development of multiplatform genetic techniques made it possible to reduce the price of genomic analysis. In the near future, the price of genomic analysis might be cheaper than ultrasound or computerized tomography (CT). This is why we believe the study of genomic analysis is absolutely needed even with little higher cost.
Prophylactic central neck dissection of PTCs may decrease locoregional recurrence cause by occult nodal metastasis [28]. Opponents of prophylactic neck dissection contend that microscopic nodal metastasis can be treated with radioactive iodine ablation therapy after total thyroidectomy without neck dissection [29]. In addition, operation-related morbidity such as recurrent nerve injury and parathyroid injury might be caused by prophylactic neck dissection [29]. Thus, it is most likely that prophylactic neck dissection is beneficial at the time of initial surgery for selected high-risk patients. Determining which patients are at high-risk before surgery remains difficult and important. In this study, we found that 12 gene-signatures could predict nodal metastasis in PTCs. Therefore, these gene signatures could be used in preoperative FNAB procedure. Using biopsy sample, we can get array data and exact information about nodal metastasis. Such prediction of nodal metastasis will be really helpful in planning surgical extent. Sometimes, preoperative ultrasound or other radiologic evaluation fail to detect small metastatic lymph nodes of PTC. In this case, if these 12 predictive gene signatures showed a high risk pattern, more comprehensive preoperative ultrasound or CT evaluation should be needed to detect occult metastatic nodes. Also, as for the surgical extent of thyroid cancer surgery, we could perform elective central neck dissection for those high-risk patients.
Zhao et al. have suggested that methylation of DACT2 promotes PTCs metastasis by activating Wnt/
This is the first study that uses genomic data to suggest gene signatures for predicting nodal metastasis of PTCs and compares these gene signatures to other clinical variables that might have prognostic effect on nodal metastasis. Our results could be used to predict nodal metastasis in PTCs and plan surgical extent such as elective central neck dissection even if there is no evidence of nodal metastasis in preoperative radiologic evaluation. The limitation of this study is its retrospective nature. A prospective and functional study is needed in the future to obtain precise information about the role of these 12 gene signatures in predicting nodal metastasis in PTCs.
In conclusion, using gene expression data and bioinformatic analysis, we generated 12 gene signatures that could predict nodal metastasis of PTC. With these gene signatures, we can predict the chance of nodal metastasis in PTCs in preoperative evaluation staging using FNAB. This can benefit appropriate planning, including central neck dissection.
Footnotes
Acknowledgments
We appreciated patients and their families who generously donated their tissues to TCGA, as well as the members of TCGA who collected and disclosed valuable data. This research was supported by Hallym University Research Fund 2017 (HURF-2017-82).
Conflict of interest
The authors declare no competing financial interests and no other conflicts of interest.
Supplementary data
The supplementary files are available to download from
