Abstract
OBJECTIVES:
To study the expression pattern of long non-coding RNA FGFR3 antisense transcript 1(FGFR3-AS1) and the cell proliferation inhibition, apoptosis, and motility changes induced by silencing FGFR3-AS1 in bladder cancer.
METHODS:
The differential expression levels of FGFR3-AS1 and FGFR3 in tumor tissues and paired normal tissues were determined using Real-Time qPCR in a total of 36 patients diagnosed with bladder cancer (urothelial carcinoma). Pearson’s coefficient correlation was used for expression correlation assay. Expression differences of FGFR3-AS1 were analyzed according to grading and staging. FGFR3 protein was detected by western blot assay. Human bladder cancer T24 and 5637 cell lines were transiently transfected with FGFR3-AS1-specific siRNA or negative control siRNA. The cell proliferation changes of transfected bladder cancer cells were determined using CCK-8 assay. Apoptosis caused by knockdown of FGFR3-AS1 was evaluated using ELISA assay. Motility changes induced by knockdown of FGFR3-AS1 were measured using wound healing assay and transwell assay.
RESULTS:
Both FGFR3-AS1 and FGFR3 were overexpressed in bladder cancer tissues compared to matched normal tissues. They were also positively expressed in bladder cancer. FGFR3-AS1 expression levels were higher in high grade tumors than those in low grade tumors. FGFR3-AS1 expression levels were higher in invasive tumors than those in non-invasive tumors. Cell proliferation inhibition, increased apoptosis, and decreased motility were observed in FGFR3-AS1 siRNA-transfected T24 and 5637 cell lines.
CONCLUSIONS:
FGFR3-AS1 plays an oncogenic role in human bladder cancer. Knockdown of FGFR3-AS1 may provide a potential new therapeutic approach to this disease.
Keywords
Introduction
As one of the most common malignancies worldwide, urothelial carcinoma of bladder displays a broad histologic spectrum ranging from superficial and well differentiated lesions to invasive and poorly differentiated tumors [1, 2]. High rates of recurrence and progression have prompted investigation into the molecular mechanisms underlying carcinogenesis and progression in bladder cancer. The genetic and epigenetic changes that contribute to development of bladder cancer are continuing to be identified and it is necessary to identify novel biological targets for bladder cancer [3, 4].
Recently, the studies on long non-coding RNAs (lncRNAs) in cancer progression and metastasis have led to new approaches in treating cancers [5]. As non-protein coding RNA transcripts, lncRNAs are larger than 200 nt in length and can serve as scaffolds or regulators [6, 7] to regulate cellular signaling pathways. They also play critical roles in the programming and regulation of mammalian genome [8]. As a widespread phenomenon, some lncRNAs have function as antisense transcripts and could regulate the expression of sense genes through RNA-RNA interaction [9, 10, 11]. For example, antisense Uchl1 increases Uchl1 translation through an embedded SINEB2 repeat [9]. The dysregulation of lncRNAs is related to many types of human cancer, including FGFR3 antisense transcript 1 (FGFR3-AS1) [12]. FGFR3-AS1 (previous name: lncRNA-BX537709) oriented in antisense direction with respect to FGFR3, and formed a ‘tail-to-tail’ pairing pattern with 1053 nucleotides full complementarity. FGFR3-AS1 increases the expression of its target gene FGFR3, which was previously identified as a potential biomarker for detection of bladder cancer [13, 14]. Somatic point mutations of FGFR3 were significantly associated with low grade/stage in bladder cancer [15]. Genomic alterations of FGFR3, such as translocations and activating mutations, have been found in advanced and metastatic urothelial carcinoma [16]. It has also been reported that FGFR3 decreases cell-cell and cell-matrix adhesion, and induces pro-proliferative and anti-apoptotic events in bladder cancer [17]. However, the relationship between FGFR3-AS1 and bladder cancer is still unclear.
In this study, our results showed that FGFR3-AS1 was overexpressed in bladder cancer compared to matched normal bladder tissue. The expression levels of FGFR3-AS1 in high grade tumors were higher than those in low grade tumors. FGFR3-AS1 RNAs were more abundant in invasive tumors than those in non-invasive tumors. Knockdown of FGFR3-AS1 inhibited bladder cancer cell proliferation, induced apoptosis and decreased cell motility.
Materials and methods
Patient samples
Thirty-six patients who diagnosed with urothelial carcinoma of the bladder and received partial or radical cystectomy were included in this study. The bladder cancer tissues and matched histologically normal tissues from patients were snap-frozen in liquid nitrogen immediately after surgical operation. This study was carried out in accordance with the recommendations of the Institutional Review Board of Shenzhen 2
Cell culture
T24, 5637, UM-UC-3, SW-780 and SV-HUC-1 cell lines were obtained from the Institute of Cell Research, Chinese Academy of Sciences, Shanghai, China. They were grown in minimal essential medium (DMEM) (Invitrogen, Carlsbad, CA, USA) supplemented with 10% fetal bovine serum (Invitrogen, Carlsbad, CA, USA) at 37
Real-time quantitative PCR
Total RNA was extracted from the tissue samples or cell lines by using the TRIzol reagent (Invitrogen, Carlsbad, CA, USA) according to the instructions. cDNA was synthesized from total RNA with the RevertAid
FGFR3-AS1 was overexpressed in bladder cancer. A. The genomic organization of FGFR3-AS1 and FGFR3. Arrows show transcription direction and blocks indicate exons. B. T represents tumor, N represents normal. The heights of the columns in the chart represent the log2-transformed fold changes (tumor/normal) in FGFR3-AS1 expression in 36 patients. C. FGFR3-AS1 expression levels were higher in high grade cancers than those in low grade cancers (**
Western blotting was performed as described elsewhere [7]. Antibodies specific to FGFR3 and TBP were obtained from Santa Cruz, Dallas, Texas, USA.
siRNA transfection
FGFR3-AS1 siRNAs were synthesized by Geneph- arma Co., Ltd., Suzhou, China. The siRNA sequen- ces [12]: the siRNA-1 sequence was 5
Cell proliferation assay
Cell proliferation was measured using Cell Counting Kit-8, CCK-8 (Beyotime Institute of Biotechnology, shanghai, China) according to protocols. Cells were cultured in a 96-well plate for 24 h and grown in normal medium. Then cells were transfected with FGFR3-AS1 siRNA or negative control siRNA. At 0, 24, 48 and 72 h after transfection, 10
Cell apoptosis assay
Cell apoptosis was measured by using Caspase 3 ELISA assay kit (R&D, Minneapolis, MN, USA). Bladder cancer cell lines were cultured and transfected with either FGFR3-AS1 siRNA or negative control siRNA. 48 hours after transfection, apoptosis caused by knockdown of FGFR3-AS1 was measured by analyzing the activity of caspase 3 according to the manufacturer’s protocol. A microplate reader (Bio-Rad, Hercules, CA, USA) was used to calculate OD values. Data were shown as the ratios between the OD values of FGFR3-AS1 siRNA transfected cells and those of negative control siRNA transfected cells.
Wound healing assay
Bladder cancer cell lines were cultured and transfected with either FGFR3-AS1 siRNA or negative control siRNA. Cells were scratched in the monolayer and cultured in normal condition. The migrated distances were measured at 0, 24 h after scratching for T24 and 5637 cells respectively.
Transwell assay
About 1
Statistical analysis
The FGFR3-AS1 RNA expression differences between bladder cancer tissues and matched normal tissues were analyzed using paired samples t-test. The expression correlation between FGFR3-AS1 and FGFR3 was analyzed using Pearson’s correlation coefficient. The FGFR3-AS1 RNA expression differences between cancer subgroups were analyzed using independent samples t-test. The differences between FGFR3-AS1 siRNA and negative control siRNA transfected cells in the CCK-8 assay were analyzed using independent samples t-test at each time point. Cell apoptosis assays and wound healing assays were analyzed using independent samples t-test. All these statistical analyses were performed using SPSS (Version 19.0 SPSS Inc.). A P value of less than 0.05 was considered to be statistically significant.
Knockdown of FGFR3-AS1 expression. T24 and 5637 cells were cultured and transfected with either FGFR3-AS1 siRNAs or negative control siRNA and cultured in normal medium. Forty-eight hours after transfection, the cells were collected. Then the FGFR3-AS1 RNA expression levels in siRNA transfected T24 (A) and 5637 (B) cells or the LETM1 mRNA expression levels in T24 cells (C) were analyzed. C. Data are indicated as mean 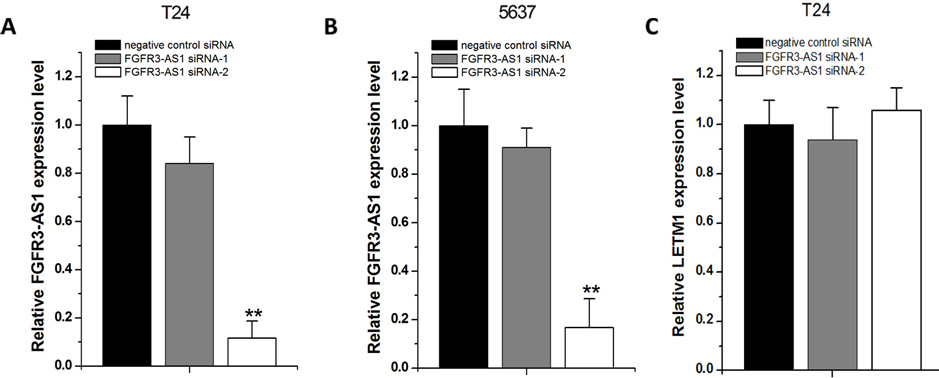
Knockdown of FGFR3-AS1 inhibited cell proliferation. Cell proliferation was measured by CCK-8 assay. After transfection of FGFR3-AS1 siRNA or negative control siRNA, OD values were measured and converted to cell numbers. Independent t test was used for comparing the differences between FGFR3-AS1 siRNA and negative control siRNA at each time point. A. Cell proliferation inhibition was observed in bladder cancer T24 cells (**
Summary of clinicopathological features of tissues of bladder cancer
Pt No.: patient number; M: male; F: female; Grade: the World Health Organization 2004 classification; H: high; L: low; Stage: the American Joint Committee on Cancer TNM classification.
Knockdown of FGFR3-AS1 induced apoptosis. Forty-eight hours after transfection of FGFR3-AS1 siRNA or negative control siRNA, the cell apoptosis changes were determined by ELISA. Cell apoptosis induction was observed in FGFR3-AS1 siRNA-transfected bladder cancer T24 (**
FGFR3-AS1 was overexpressed in bladder cancer tissues
The genomic organization of FGFR3-AS1 andFGFR3 was shown in Fig. 1A. The relative expression of FGFR3-AS1 was detected using Real-Time qPCR in a total of 36 patients with bladder cancer. FGFR3-AS1 was overexpressed in bladder cancer compared to matched normal tissue. The FGFR3-AS1 expression fold change (bladder cancer tissue/matched histologically normal tissue) in each patient is indicated in Fig. 1B. The clinical features of this cohort of patients (from No. 1 to No. 36) are provided in Table 1.
We analyzed expression differences according to grading and staging. FGFR3-AS1 expression levels were higher in high grade tumors than those in low grade tumors. The expression fold change (high/low grade tumor) was 3.18 (Fig. 1C). FGFR3-AS1 expression levels were higher in invasive tumors than those in non-invasive tumors. The expression fold change (invasive/non-invasive tumor) was 2.63 (Fig. 1D).
Because FGFR3-AS1 was reported to increase the expression of its sense gene FGFR3 [12], we then determined the relative expression of FGFR3 mRNA using Real-Time qPCR in the same bladder cancer samples (
Knockdown of FGFR3-AS1 decreased cell migration. After transfection of FGFR3-AS1 siRNA or negative control siRNA, wound healing assay was used to detect cell migration in bladder cancer cells. A. Representative images of wound healing assay in T24 cells. B. Decreased cell migration was observed in T24 cells (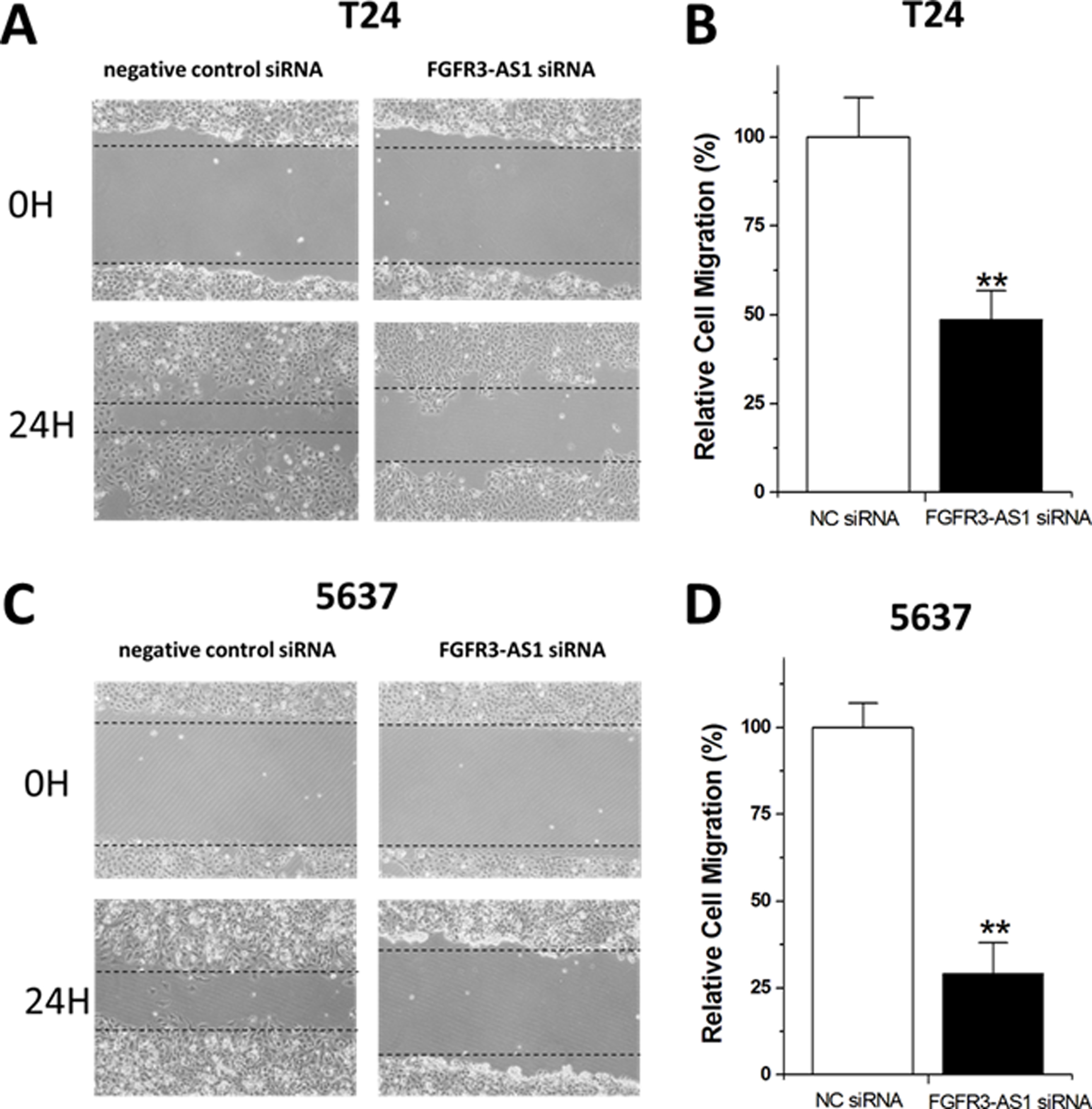
Knockdown of FGFR3-AS1 decreased cell invasion. After transfection of FGFR3-AS1 siRNA or negative control siRNA, transwell assay was used to detect cell invasion in bladder cancer cells. A. Representative images of transwell assay in T24 cells. B. Decreased cell invasion was observed in T24 cells (
We also determined the expression pattern of FGFR3-
AS1 in several well-known bladder cancer cell lines (T24, 5637, UM-UC-3 and SW-780) and one normal urothelial cell line (SV-HUC-1) using Real-Time qPCR. The data showed that FGFR3-AS1 expression levels were much higher in bladder cancer T24 and 5637 cell lines than in other 3 cell lines (Fig. 1H). Therefore, we performed functional experiments in T24 and 5637.
According to the previous reports [12], we have used two different siRNAs (siRNA-1 and siRNA-2) targeting FGFR3-AS1 and tested their efficiencies in bladder cancer T24 and 5637 cells transfected with FGFR3-AS1 siRNAs or negative control siRNA. 48 hours after transfection, the FGFR3-AS1 RNA expression levels were analyzed. Although FGFR3-AS1 siRNA-1 only induced minimal knockdown effects, FGFR3-AS1 siRNA-2 largely reduced FGFR3-AS1 RNA levels in both T24 (Fig. 2A) and 5637 (Fig. 2B) cells. To exclude the possibility that the siRNAs we designed also reduce the expression of LETM1, a gene downstream of FGFR3, we also measured the relative level of LETM1 using qPCR after transfection of these siRNAs. The results confirmed that siRNAs have no influence on LETM1 expression (Fig. 2C). Therefore, we performed functional experiments using FGFR3-AS1 siRNA-2.
Knockdown of FGFR3-AS1 inhibited cell proliferation
Bladder cancer T24 and 5637 cells were transfected with FGFR3-AS1 siRNA or negative control siRNA and the cell proliferation changes of bladder cancer cells were determined by CCK-8 assay. Cell proliferation arrest was observed in both T24 cells (Fig. 3A) and 5637 cells after knockdown of FGFR3-AS1 (Fig. 3B).
Knockdown of FGFR3-AS1 induced apoptosis
Bladder cancer T24 and 5637 cell lines were transfected with FGFR3-AS1 siRNA or negative control siRNA. Forty-eight hours after transfection, the cell apoptosis changes of bladder cancer T24 and 5637 cells were determined using ELISA analysis (Fig. 4). Induced cell apoptosis was observed in both bladder cancer cell lines after knockdown of FGFR3-AS1.
Motility changes induced by knockdown of FGFR3-AS1
We used wound healing assay to detect cell migration caused by knockdown of FGFR3-AS1 in bladder cancer cells. Decreased cell migration was observed in both T24 cells (Fig. 5A and B) and 5637 cells (Fig. 5C and D).
We also used transwell assay to detect cell invasion. Compared with the negative control group, the cell invasion ability was extremely suppressed in T24 (Fig. 6A and B) cells and 5637 cells (Fig. 6C and D) which were transfected with FGFR3-AS1 siRNA.
Discussion
LncRNAs take participation in nearly all the biological and pathological processes and have been involved in the tumorigenesis, metastasis, or recurrence of bladder cancer. They may serve as biomarkers for diagnosis and prognosis [18]. Although the reason why lncRNAs were activated in bladder cancer is unclear, it has been believed that lncRNAs are powerful endogenous regulators for specific targets [19, 20]. As a recently discovered lncRNA, several previous works were conducted to characterize the oncogenic properties of FGFR3-AS1 [12, 21]. Inducible knockdown of FGFR3 in human bladder cancer cells arrested cell-cycle progression and markedly prevented tumor progression in xenografted mice, suggesting that FGFR3 is an oncogene [22]. FGFR3 also plays an important role in progression of other cancers and was associated with aggressive tumor behavior [23]. In this study, we found that FGFR3-AS1 was overexpressed in bladder cancer compared to matched normal tissue. High level expression of FGFR3-AS1 was associated with high grade and stage bladder cancers. The differential FGFR3-AS1 expression patterns between bladder cancer and control and the association of FGFR3-AS1 with clinicopathological features suggest that long noncoding RNA FGFR3-AS1 emerges as a novel player in the development and progression of the bladder cancer. Interestingly, the expression of FGFR3-AS1 was positively correlated with FGFR3 in bladder cancer. These results were consistent with the well-known conclusion which indicated that FGFR3-AS1 increases FGFR3 mRNA stability and expression through antisense pairing with 3’UTR of FGFR3 mRNA.
To understand the possible impacts of FGFR3-AS1 on bladder cancer, we determined the cell proliferation inhibition, apoptosis, and motility changes induced by knockdown of FGFR3-AS1 in bladder cancer. Cell proliferation inhibition, increased apoptosis, and decreased motility were observed in FGFR3-AS1 siRNA-transfected bladder cancer T24 and 5637 cell lines. These findings suggest that FGFR3-AS1 may play roles in carcinogenesis via upregulating FGFR3 in bladder cancer. Targeting FGFR3-AS1 or FGFR3 may be a promising approach to the treatment of bladder cancer. More works will be needed to determine the potential molecular mechanism of FGFR3-AS1 in bladder cancer. Therapies designed to target FGFR3-AS1 or FGFR3 are also should under intensive investigation.
Footnotes
Acknowledgments
The authors thank all the donors whose names were not included in the author list, but who participated in this program. This research was supported by Scientific Research Project of Shenzhen Health and Family Planning System (201606019).
Conflict of interest
None.
