Abstract
miR-99a is down-regulated in esophageal squamous cell carcinoma (ESCC), however the role and underlying mechanism are still unknown. We aim to explore the role and mechanism of miR-99a down-regulation in ESCC. The expression of miR-99a in ESCC tissues and cell lines was detected by Human miRNA Microarrays and Real-time PCR. The effects of miR-99a on cell proliferation, migration and invasion were determined by Cell Counting Kit-8 (CCK-8) assay, transwell migration and invasion assay. Target gene of miR-99a were analyzed by target prediction software and validated by Real-time PCR and Western blotting assay. Our microarray results and four Gene Expression Omnibus (GEO) datasets showed lower expression level of miR-99a in ESCC tissues. Overexpression of miR-99a using mimics significantly suppressed cell proliferation, and decreased expressions of CCND1, CCNA2 and CCNE1. We also found that enhanced miR-99a significantly inhibited migration, invasion and epithelial-mesenchymal transition (EMT) of ESCC cells, and down-regulated EMT associated transcription factor Slug, and MMPs including MMP2, MMP7 and MMP13. TargetScan predicted insulin-like growth factor 1 receptor (IGF1R) as the cadidate target gene of miR-99a, and western blotting confirmed the negative correlation between miR-99a and IGF1R. Importantly, we further found that knockdown of IGF1R also significantly inhibited the proliferation, migration, invasion and slug-induced EMT of ESCC cells, and reduced the cell cycle regulatory proteins and MMPs. In conclusion, our findings suggested that loss of miR-99a in ESCC promoted the tumor cell proliferation, migration, invasion and slug-induced EMT through activating IGF1R signaling pathway.
Keywords
Introduction
Esophageal cancer is one of the most common cancers and is the sixth leading cause of cancer death worldwide [1, 2, 3]. Approximately 70% of global esophageal cancer cases occur in China, with squamous cell carcinoma (SCC) being the histopathological form in the vast majority of cases (
miRNAs are short (19–25 nucleotides), single-stranded, non-coding RNAs. They bind to the 3
miR-99a was down-regulated in oral cancer, ovarian cancer, breast cancer, cervical cancer, non-small cell lung cancer and anaplastic thyroid cancer, and was a potential tumor suppressor [11, 12, 13]. miR-99a could suppress the proliferation, migration and invasion of breast cancer cells [14, 15]. And miR-99a could also inhibit the growth and colony formation of osteosarcoma cells through inducing apoptosis [16]. Overexpression of miR-99a could decrease the ability of cholangiocarcinoma cells for cancer stem-cell-like mammosphere generation by regulating CD133 and CD44 [11]. Up to now, AKT1, mTOR, HOXA1 and TNFAIP8 were identified as the direct targets of miR-99a in cancer progression [15, 16, 17, 18]. In ESCC, miR-99a was down-regulated, and the decreased expression of miR-99a was associated with worse overall survival in ESCC patients. Overexpression of miR-99a could inhibit cell proliferation by inducing apoptosis via directly targeting mTOR [19]. However, the role of miR-99a in esophageal cancer progression such as metastasis and the underlying mechanism have never been deciphered.
Insulin-like growth factor 1 receptor (IGF1R) is a transmembrane tyrosine kinase receptor with a heterodimer of
In the present study, we focus on the roles of down-regulation of miR-99a in ESCC, and further revealed the underlying mechanisms.
Materials and methods
Bioinformatics analysis
The expression level of miR-99a was lower in 8 ESCC tissues than their adjacent non-malignant tissues using Agilent Human miRNA Microarrays. The expression level of miR-99a was analyzed using GEO datasets (GSE66274, GSE43732, GSE59973 and GSE61047) (
Cell culture
Esophageal squamous cell carcinoma (ESCC) cell lines such as KYSE30, KYSE150, KYSE180, KYSE 410 and KYSE510 were cultured in RPMI-1640 (Invitrogen corporation, USA) with 10% fetal bovine serum (FBS) (Hyclone, USA). All of these cells were maintained at 37
Cell transfection
Hsa-miR-99a mimics were designed and provided by GenePharma Biotech (Shanghai, China). Specific siRNAs targeting IGF1R were custom-designed and provided by GenePharma Biotech (Shanghai, China). Transfection of Hsa-miR-99a mimics or IGF1R siRNAs or control siRNAs was performed using Lipofectamine 2000 (invitrogen) according to the manufacturer’s instructions. Transfected cells were incubated at 37
The sequences of miR-99a mimics were:
Sense: 5
The sequences of IGF1R siRNAs were:
Si-IGF1R-1: Sense: 5
The sequences of negative control siRNAs were:
Sense: 5
Down-regulation of miR-99a in ESCC tissues. (A) miR-99a was down-regulated in ESCC tissues compared to normal tissues using microarray analysis. **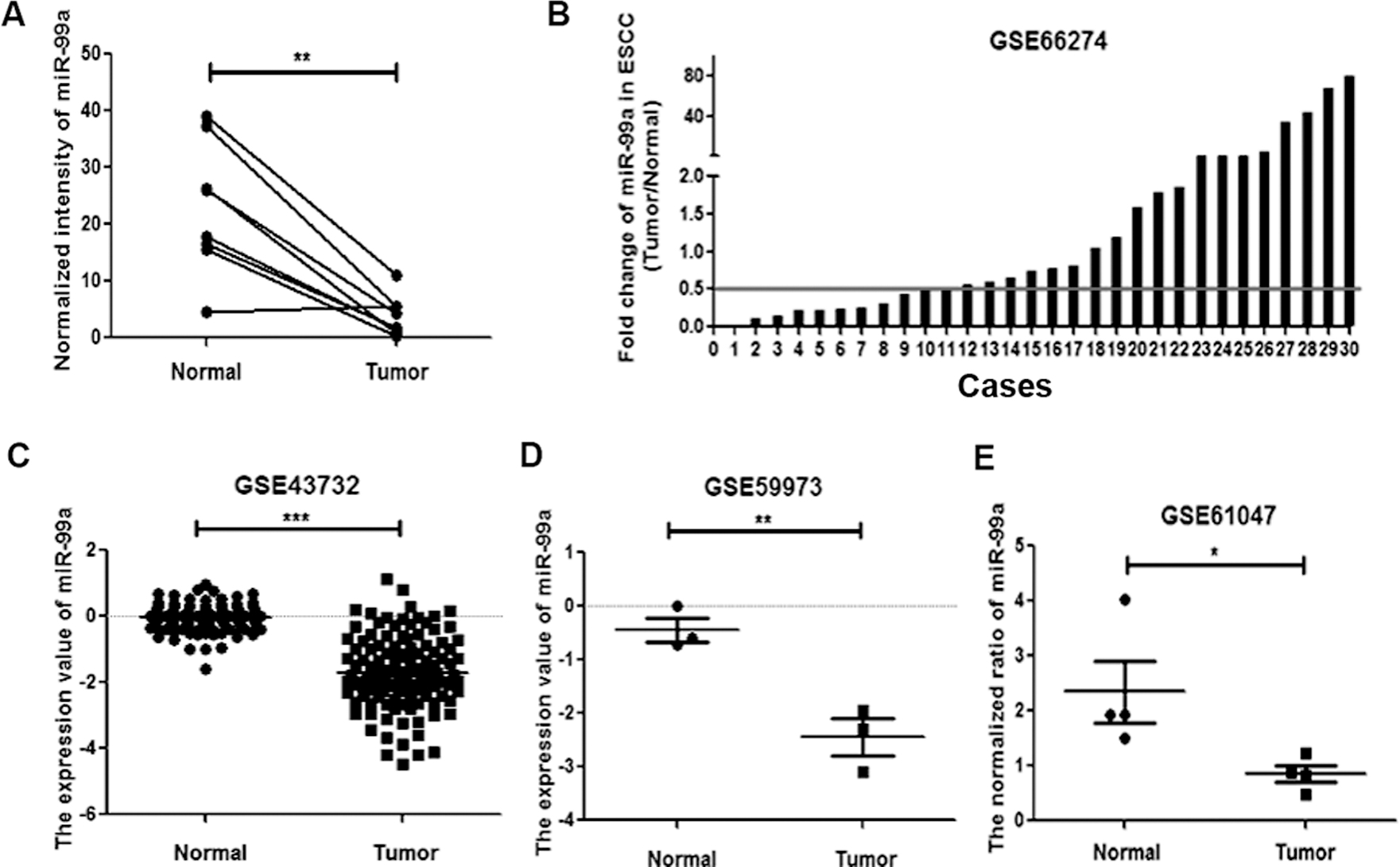
Overexpression of miR-99a suppressed cell proliferation and the expressions of cell cycle proteins in KYSE150 and KYSE510 cells. (A) Level of miR-99a in KYSE150 and KYSE510 cells transfected with miR-99a mimics or their negative control was detected by using Real-time PCR analysis. miR-99a expression was normalized against an endogenous control U6. (B) Cell proliferation was analyzed in KYSE150 and KYSE510 cells with ectopic miR-99a expression and negative control. (C) CCND1, CCNA2 and CCNE1 mRNA levels in KYSE150 and KYSE510 cells were detected by real-time PCR. Bars represent mean mRNA levels normalized to GAPDH mRNA. Data represent means 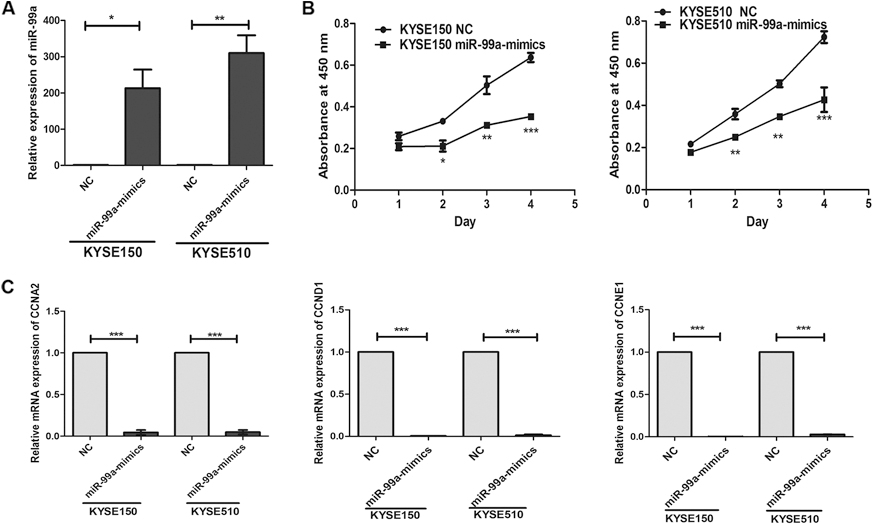
Cell viability was assessed by the Cell Counting Kit-8 assay (CCK-8; Dojindo, Kumamoto, Japan). After being transfected with miR-99a mimics or negative control siRNAs (NC), KYSE150 and KYSE510 cells were cultured in 96-well plated at a density of 5,000 cells/well. The viability of the cells was determined at the indicated time (0, 24, 48 and 72 h) according to the manufacturer’s instructions, and the absorbance value at a wavelength of 450 nm was detected by an enzyme-linked immunosorbent assay reader.
Cell migration and invasion assay
A typical transwell assay (Costar, 6.5 mm diameter, 8
Real-time PCR assay
The Real-time PCR was performed using the Premix Ex Taq kit (Takara) and a 7300 Real-time PCR system (life Technologies) according to the manufacturer’s instructions. The mRNA expression levels of Cyclin A2 (CCNA2), Cyclin E1 (CCNE1), Cyclin D1 (CCND1), Matrix metalloproteinase 2 (MMP2), Matrix metalloproteinase 7 (MMP7) and Matrix metalloproteinase 13 (MMP13) were normalized to the endogenous expression of GAPDH. Primers were provided by TSINGKE. Hairpin-itTM miR-99a qRT-PCR Primer Set (GenePharma) was used for the measurement of the relative quantity of hsa-miR-99a. The mRNA expression of miR-99a was normalized to the endogenous expression of U6. The 2
Western blotting assay
Cells from each group were detached with trypsin, centrifuged and washed 2 times with pre-chilled PBS. Cell lysis buffer was subsequently added and incubated on ice for protein extraction. Protein concentration was determined using the BCA Protein Assay Kit (Beyotime Biotechnology, China). Equal amounts of proteins were separated via 12% SDS-PAGE and then transferred to a PVDF membrane (Millipore Corporation, Billerica, MA, USA). The membrane was soaked in 10% skimmed milk (in PBS, PH 7.2, containing 0.1% Tween-20) for 2 h and incubated with an appropriate amount of primary antibody (working dilutions of antibodies: E-cadherin 1:1000, N-cadherin 1:1000, Vimentin 1:1000, Slug 1:1000, IGF1R 1:2000,
Statistical analyses
All results were confirmed in at least three independent experiments, and all quantitative data are presented as mean
Overexpression of miR-99a inhibited the migration and invasion of ESCC cells. (A) Migration and invasion of cells were analyzed at 48 h post-transfection. Photographs represented the migrated and invaded cells. (B) The protein expression levels of Slug, E-cadherin, N-cadherin and Vimentin were detected by western blotting assay at 48 h post-transfection and normalized to that of 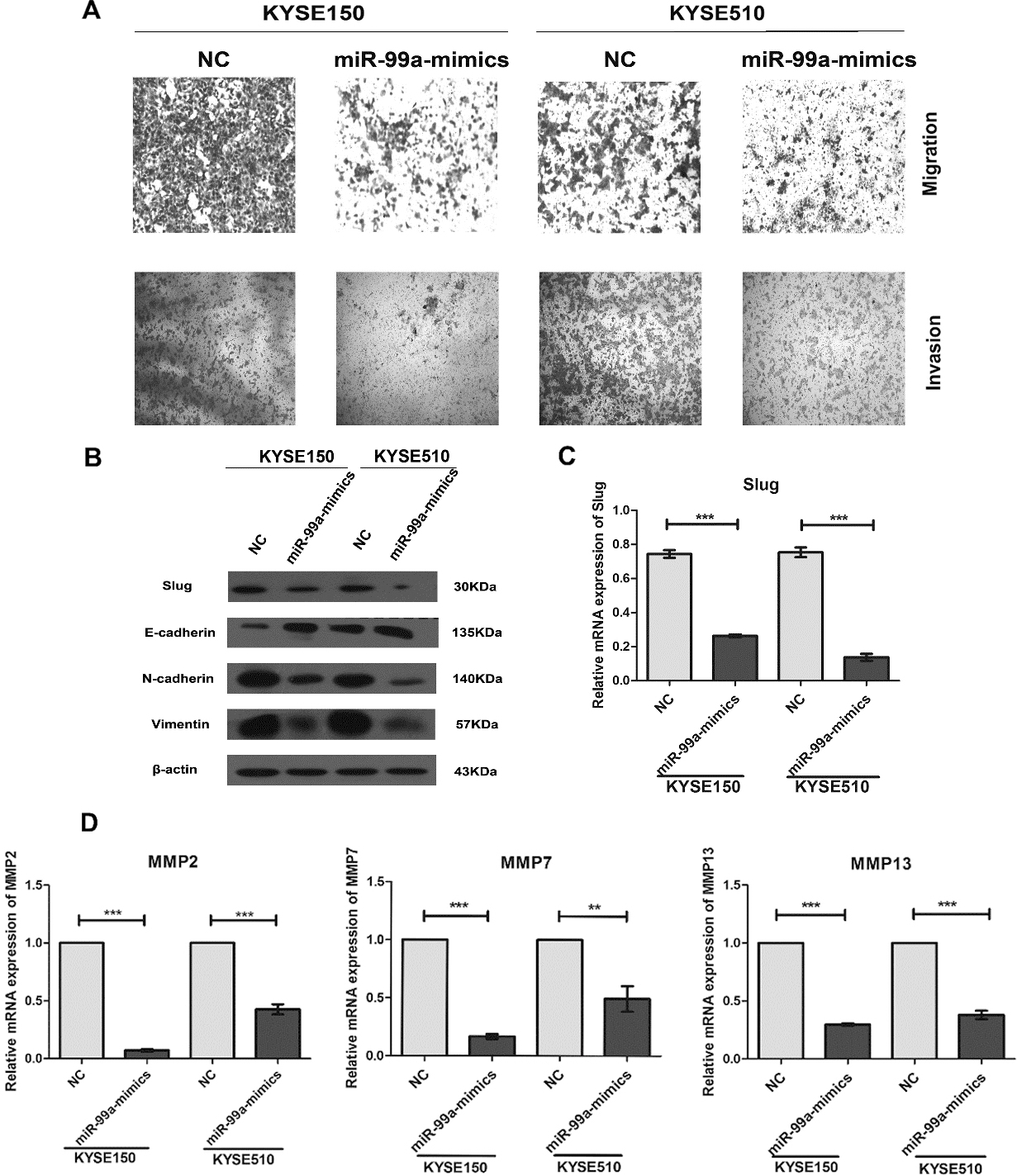
IGF1R is the direct target of miR-99a, and knockdown of IGF1R suppressed the cell proliferation and the mRNA expressions of cell cycle proteins in KYSE150 and KYSE510 cells. (A) Predicted targeting sequence of miR-99a at nucleotides 9750-9769 of the IGF1R 3’UTR. (B) Western blotting assay revealed that overexpression of miR-99a significantly inhibited the expression of IGF1R in KYSE150 and KYSE510 cells. (C) Efficacy of RNA interference was verified by western blotting. (D) Down-regulation of IGF1R by si-IGF1R-1 and si-IGF1R-2 inhibited cell proliferation of KYSE150 and KYSE510 cells by CCK-8 assay. (E) CCNA2, CCND1 and CCNE1 expression levels in KYSE150 and KYSE510 were analyzed by real-time PCR. Data represent means 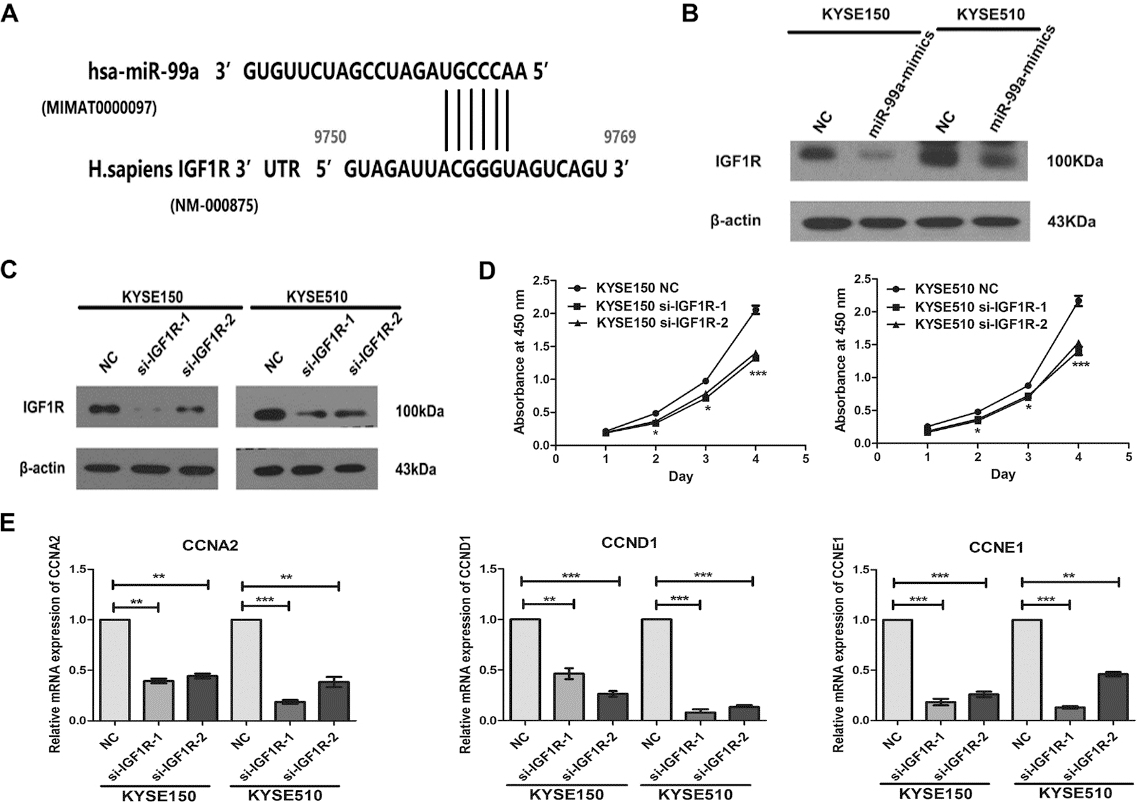
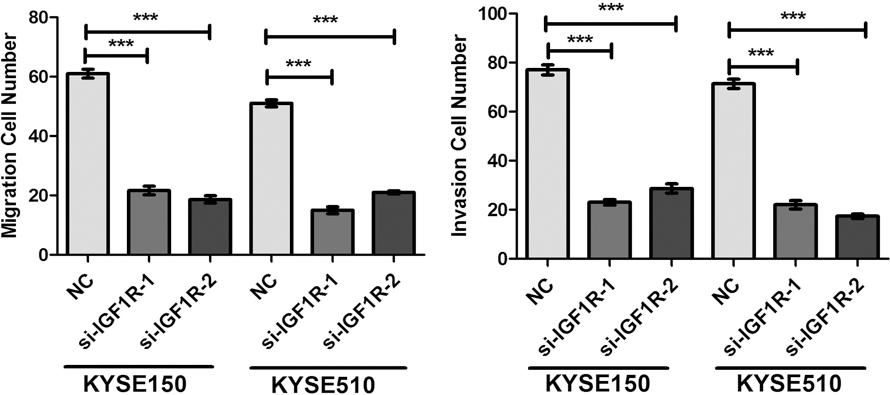
Knockdown of IGF1R inhibited the migration and invasion of ESCC cell. (A) Down-regulation of IGF1R by si-IGF1R inhibited migration, and invasion of KYSE150 and KYSE510 cells. (B) The expression levels of Slug, E-cadherin and N-cadherin proteins were detected by western blotting assay at 48 h post-transfection and normalized to that of 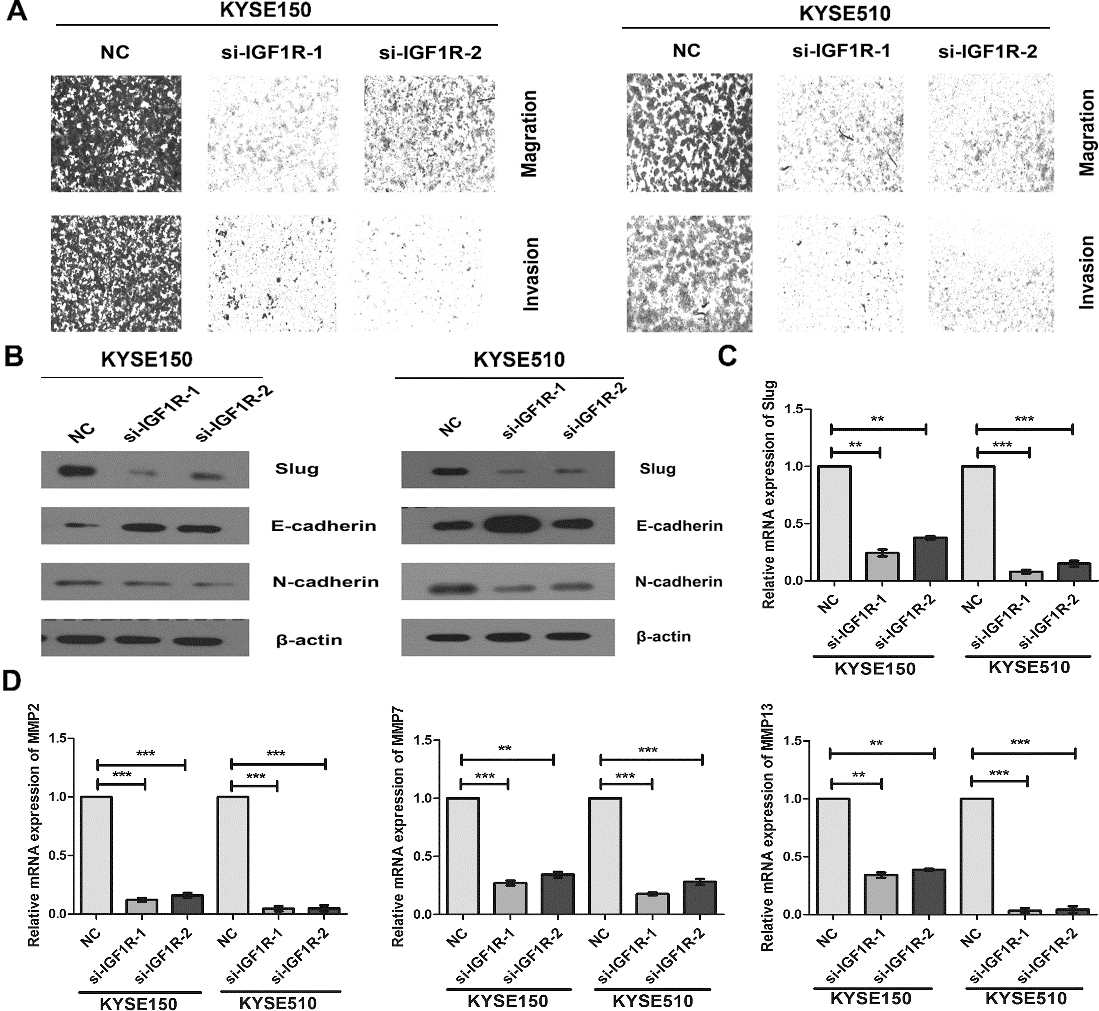
Low expression of miR-99a in ESCC tissues
In this study, we found that the expression level of miR-99a was lower in ESCC tissues than their adjacent non-malignant tissues using Agilent Human miRNA Microarrays(Fig. 1A). And we further analyzed the expression of miR-99a in four datasets (GSE66274, GSE43732, GSE59973 and GSE61047) of ESCC (Fig. 1B–E). miR-99a was significantly downregulated in 9/30 ESCCs with fold change below 0.5 (Tumor/Normal) in the dataset of GSE66274 (Fig. 1B). Importantly, another three datasets (GSE 43732, GSE59973 and GSE61047) were also explored to verify the decrease of miR-99a expression in tumor samples (Fig. 1C–E).
miR-99a suppressed proliferation of ESCC cells by regulating cell cycle regulatory proteins
To understand the role of miR-99a in esophageal carcinogenesis, we evaluated the expression of miR-99a in 5 ESCC cell lines (KYSE30, KYSE180, KYSE 450, KYSE150 and KYSE510) by Real-time PCR (Supplement Fig. 1). The result indicated that KYSE 150 and KYSE510 exhibited lower levels than other cell lines. After transfection of miR-99a mimics or negative control, we evaluated the cell viability using CCK-8 kit. Overexpression of miR-99a significantly inhibited the proliferation of KYSE150 and KYSE510 cells (Fig. 2B). Interestingly, mRNA levels of cell cycle regulatory proteins including CCNA2, CCND1 and CCNE1 were significantly decreased in the miR-99a overexpressed ESCC cells (Fig. 2C). The above observation indicated that miR-99a inhibited the proliferation of ESCC cells via down-regulating CCND1, CCNA2 and CCNE1.
miR-99a inhibited the migration and invasion by suppressing Slug-induced EMT and MMPs in ESCC cells
Using transwell assay, we found that overexpression of miR-99a led to a significant reduction of migrated and invaded cell numbers in KYSE150 and KYSE510 cells (Fig. 3A and Supplement Fig. 2). A well-recognized mechanism for initiating tumor cell invasive and metastatic behavior is epithelial-mesenchymal transition (EMT), in which polarized epithelial cancer cells acquire a motile mesenchymal phenotype [23]. The hallmarks of EMT include the decreased expression of the epithelial marker E-cadherin and increased expressions of mesenchymal markers, such as Vimentin and N-cadherin [24]. In our study, overexpression of miR-99a using mimics significantly increased the E-cadherin expression, and reduced the expressions of N-cadherin and Vimentin in KYSE150 and KYSE510 cells (Fig. 3B). Slug is reported as one of the critical regulators for EMT and as the transcription repressor of E-cadherin [25]. Our results also showed that overexpression of miR-99a could down-regulated the mRNA and protein levels of Slug (Fig. 3B and C).
Matrix metalloproteinases (MMPs), a family of zinc-
binding proteins including MMP2, MMP7, MMP9 and so on, have been shown to play a central role in tumor cell invasion and metastasis due to their ability to degrade the extracellular matrix [26]. We evaluated the expressions of five MMPs (MMP2, MMP7, MMP9, MMP13 and MMP14) by Real-time PCR. miR-99a significantly decreased the mRNA expressions of MMP2, MMP7 and MMP13 (Fig. 3D), but with no significant effect on the expressions of MMP9 and MMP14(Supplement Fig. 3). Taken together, overexpression of miR-99a suppresses the migration and invasion of ESCC cells via inhibiting Slug-induced EMT and decreasing the MMPs.
miR-99a negatively regulated IGF1R expression, and knockdown of IGF1R phenocopied the effects on cell proliferation, migration and invasion like overexpression of miR-99a
Next, we performed an in silico study to search for potential targets of miR-99a by TargetScan. IGF1R is the candidate target of miR-99a (Fig. 4A). Our result showed that overexpression of miR-99a significantly decreased the IGF1R protein level by western blotting assay (Fig. 4B).
To confirm whether miR-99a induced inhibition of cell proliferation, migration and invasion was mediated by IGF1R, we detected the effects of silencing IGF1R on ESCC cells. Western blotting analysis confirmed that transfection with si-IGF1R could effectively reduce the IGF1R expression both in KYSE150 and KYSE510 cells (Fig. 4C). Knockdown of IGF1R significantly inhibited the proliferation of KYSE150 and KYSE510 cells (Fig. 4D), and also reduced the expressions of CCND1, CCNE1 and CCNA2 by Real-time PCR assay (Fig. 4E).
We further found that knockdown of IGF1R significantly inhibited the migration and invasion of ESCC cells, and suppressed the EMT with up-regulation of E-cadherin and down-regulation of N-cadherin. Interestingly, silencing IGF1R decreased the slug protein expression (Fig. 5A–C and Supplement Fig. 4). The mRNA expressions of MMP2, MMP7 and MMP13 showed a markedly reduction in si-IGF1R treated KYSE150 and KYSE510 cells (Fig. 5D). Taken together, our results suggested that down-regulation of miR-99a promoted the proliferation, migration, invasion and slug-induced EMT of ESCC cells via down-regulating IGF1R.
Discussion
Previous studies characterized the miRNA expression profiles associated with the stage and prognosis of ESCC [19, 27, 28], and dysregulation of miRNAs played the important role in esophageal carcinogenesis. miR-675-5p was overexpressed in ESCC, and its overexpression significantly promoted the proliferation, colony formation, migration and invasion via targeting REPS2 [29]. Liu et al. reported that miR-373 could promote migration and invasion of ESCC cells by inhibiting TIMP3 expression [30]. Sun et al. found that miR-99a was lower expression in ESCC, and overexpression of miR-99a inhibited cell proliferation by inducing apoptosis via directly targeting mTOR [19]. However, the role of miR-99a in esophageal cancer progression such as metastasis and the underlying signaling pathways have never been illuminated.
miR-99a, which belongs to the miR-99 family (including other two members miR-99b and miR-100), is located in the intron 13 of the C21orf34 gene at 21q21.1 [31]. In our study, we confirmed down-regulation of miR-99a in ESCC tissues by microarray and GEO datasets. Functional experiments further revealed that overexpression of miR-99a significantly inhibited the proliferation of ESCC cells, and reduced the cell cycle regulatory proteins including CCNA2, CCND1 and CCNE1. In multiple myeloma, miR-99a
Metastasis is the main reason leading to death of ESCC patients. We first found that enhanced miR-99a expression inhibited the migration and invasion of ESCC cells in vitro. In epithelial cancers, EMT is regarded as one of the major mechanisms that promote migration, invasion and metastasis [33, 34]. Loss of E-cadherin is a fundamental step of EMT by which polarized epithelial cells lose the tight cell–cell junction and enhance the migratory capacity. In our study, miR-99a significantly suppressed the EMT with up-regulation of E-cadherin and down-regulation of N-cadherin as well as Vimentin. Several EMT-inducing transcription factors, such as Snail, Slug and ZEB1, are reported to inhibit E-cadherin to promote invasion and metastasis [35]. Slug could bind to the E-box hexa-nucleotide DNA motif in E-cadherin promoter by a resembling structure with a highly conserved carboxy-terminal domain [36]. Interestingly, we found that miR-99a reduced the expression of Slug. Matrix metalloproteinases (MMPs) are proteolytic enzymes that play a pivotal role in the transformation and progression of tumors at all stages, especially during the invasion and metastasis [37]. And in ESCC, miR-99a significantly inhibited the MMP2, MMP7 and MMP13. All these results suggest that decreasing expression of miR-99a in ESCC promotes the migration and invasion of tumor cells via inhibiting EMT and MMPs.
IGF1R is the candidate target of miR-99a. In our study, miR-99a negatively regulates the expression of IGF1R by western blotting analysis. IGF1R is overexpressed in several malignancies and plays a crucial role in promoting cell proliferation and metastasis [38]. Recent study reported IGF1R as a marker for prognosis and a therapeutic target in human ESCC [39, 40]. Blockade of IGF1R was involved in the suppression of gastrointestinal cancer cell invasion through downregulation of matrilysin [41]. In KYSE150 and KYSE510 cells, knockdown of IGF1R significantly inhibited cell proliferation, migration and invasion, and also reduced cell cycle proteins, EMT markers and MMPs. These results indicated that miR-99a suppressed the proliferation, migration and invasion of ESCC cells via negatively regulating IGF1R.
In conclusion, we demonstrated that miR-99a significantly inhibit ESCC cell proliferation, migration and invasion through suppressing IGF1R signaling pathway. The newly identified miR-99a/IGF1R axis provides novel insight into the pathogenesis of ESCC, and represents a potential therapeutic target for the treatment of ESCC.
Conflict of interest
The authors declare no conflicts of interest.
Footnotes
Acknowledgments
This study was supported by the National Natural Science Foundation of China (No. 81460425) and the Yunnan Provincial Research Foundation for Basic Research, China (No. 2013FD012) and Foundation for the Talents of Kunming University of Science and Technology (No. KKSY201226099).
Supplement material
The mRNA expression levels of miR-99a was detected by Real-time PCR in 5 ESCC cell lines.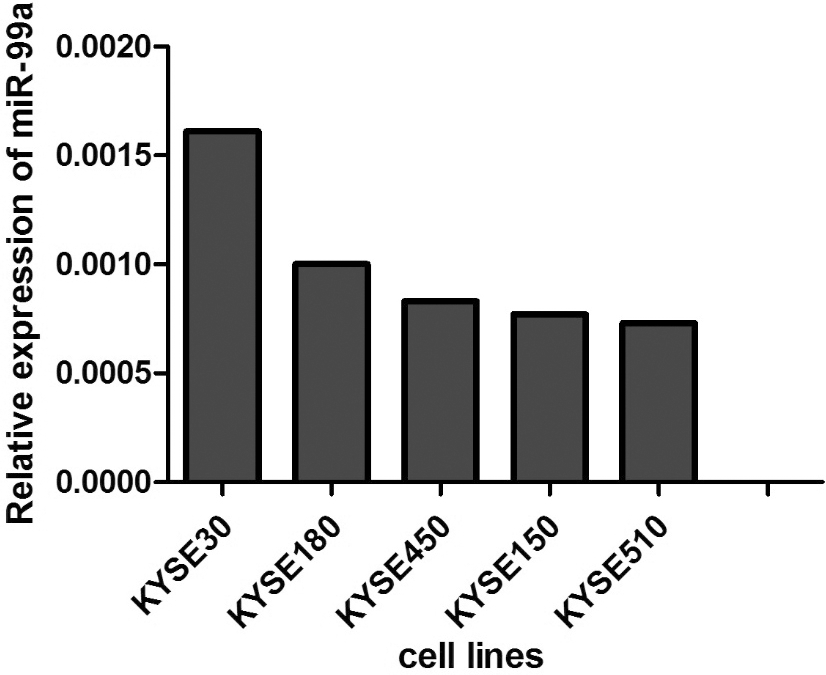
Overexpression of miR-99a led to a significant reduction of migrated and invaded cell numbers in KYSE150 and KYSE510 cells.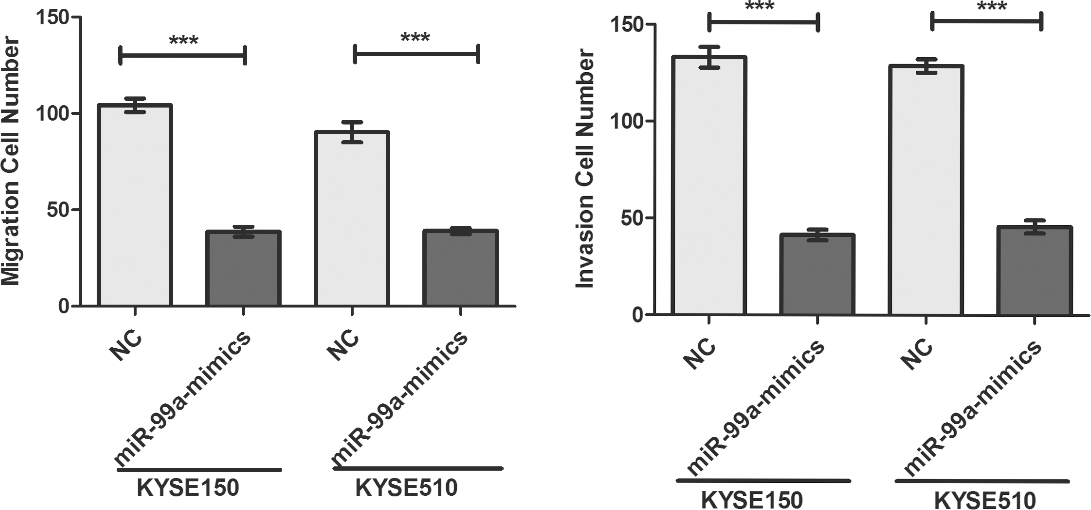
The mRNA expression levels of MMP9 and MMP14 was detected by Real-time PCR, and it exhibited no difference between cells transfected with miR-99a mimics and the cells transfected with negative control.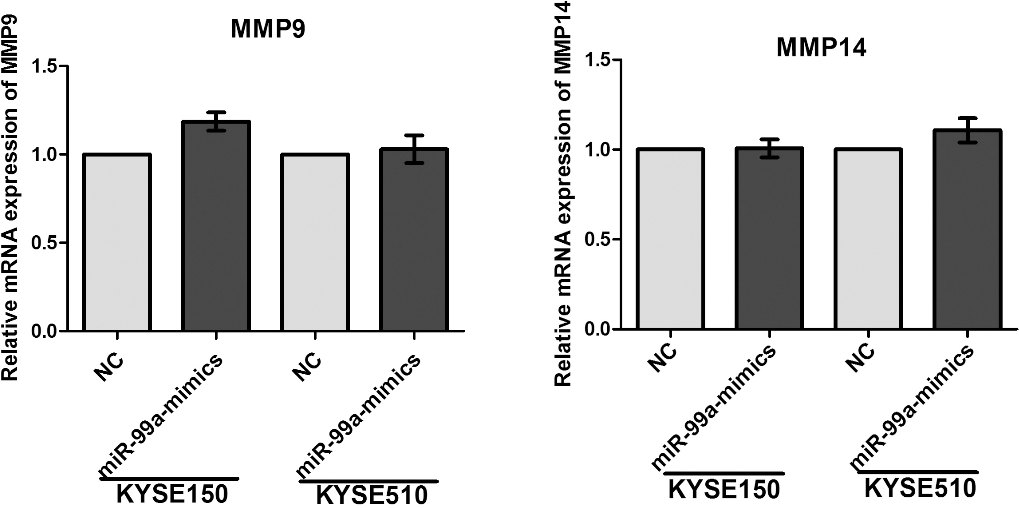
Knockdown of IGF1R led to a significant reduction of migrated and invaded cell numbers in KYSE150 and KYSE510 cells.