Abstract
BACKGROUND:
Sal-like protein 4 (SALL4) is recognized as a potential biomarker for assessing cancer prognosis. Many experiments have been done to explore the aberrant expression of SALL4 in cancer patients.
OBJECTIVE:
To explore the association between SALL4 expression and cancer prognosis.
METHODS:
A systematically comprehensive search was performed through PubMed, Web of Science, and CNKI. The prognostic value of SALL4 expression in cancer patients was evaluated by pooled hazard ratios (HRs) and 95% confidence intervals (CIs).
RESULTS:
The overexpression of SALL4 was identified to be significantly associated with a poor prognosis in cancer patients (pooled HR: 2.02, 95% CI: 1.67–2.46). This association was also detected in digestive system neoplasms subgroup (pooled HR: 1.91, 95% CI: 1.52–2.39) and reproductive system neoplasms subgroup (pooled HR: 1.95, 95% CI: 1.17–3.26). In the geography subgroup analysis, a statistical association was confirmed in the Chinese subgroup (pooled HR: 1.982, 95% CI: 1.526–2.576). In the respect of disease progression, there was a certain degree of relationship between higher expression of SALL4 and recurrence-free survival (RFS) (pooled HR: 1.617, 95% CI: 1.190–2.196).
CONCLUSION:
SALL4 is a promising prognostic biomarker for cancer, and is appropriate for the assessment of cancer prognosis in the Chinese people.
Introduction
SALL4, as a zinc-finger transcription factor, was originally considered as a homolog of the Drosophila homeotic gene spalt [1]. Afterwards, it deemed to play an essential role in embryonic development and self-renewal of embryonic stem cells through interactions with OCT3/4, SOX2, and NANOG [2, 3, 4, 5]. The functional loss of SALL4 may result in dysplasia of multiple organs and abortion or embryonic death in mice [2, 6]. In humans, SALL4 gene mutations can lead to an autosomal dominant disease called Okihiro syndrome (Duane radial ray syndrome), which causes multiple organ dysfunction [7, 8, 9, 10]. As tissues and organs mature, SALL4 expression gradually reduced [11]. However, accumulating evidence indicates that SALL4 expression is reactivated in numerous human malignancies. It was the first time that SALL4 was confirmed as an oncogene in leukemia [12]. Consecutively, the overexpression of SALL4 has been detected in a variety of solid cancers and indicated a poor clinical prognostic value in a few of them, such as germ cell tumor [13, 14], hepatocellular carcinoma [15], colorectal cancer [16], gastric cancer [17], lung cancer [18] and breast cancer [19]. However, some researcher drew the inconclusive conclusion [20]. Thus, whether higher expression of SALL4 predicts poor cancer prognosis remains confused. In view of the relatively small sample sizes of studies and the different results, we conducted this meta-analysis of all available studies in order to acquire more reliable conclusion about the relationship between SALL4 expression and the prognosis influence in cancer patients.
Materials and methods
Literature search
A systematic literature search was carried out by means of the databases PubMed, Web of Science and CNKI for studies in humans of the association between SALL4 expression and cancer prognosis. The search strategy used the terms: ([sal-like protein 4 or SALL4] AND [carcinoma or tumor or cancer] AND [prognosis or survival or outcome]) covering all relevant studies published up to March 2016. Meanwhile, the population, sample size, time period or types of clinical trials on the retrieved studies were not limited, and only studies in English or Chinese were included. Reference lists from relevant articles were inspected manually to further find additional eligible studies. Two investigators performed the search independently. When some overlapping studies were checked, we chose a better quality study or one with more detailed data.
Inclusion criteria
To be eligible for inclusion, all the studies have to meet the following criteria: (1) definite diagnosis of cancer in humans, (2) trials published as a full paper in English or Chinese literature, (3) evaluating the association between SALL4 and cancer prognosis, (4) SALL4 quantity (protein or mRNA) analysis of the primary tumor (not in metastatic tumor or in tumor adjacent tissues) or in the serum, (5) sufficient patient data for eligible HRs with 95% CIs or other available data for HR with a 95% CI (including data extracted from Kaplan-Meier curves). The major reasons for exclusion of studies were: (1) overlapping data; (2) abstract, comment, editorials letters, and review; (3) studies without detailed data. The study selection flow diagram was shown in Fig. 1.
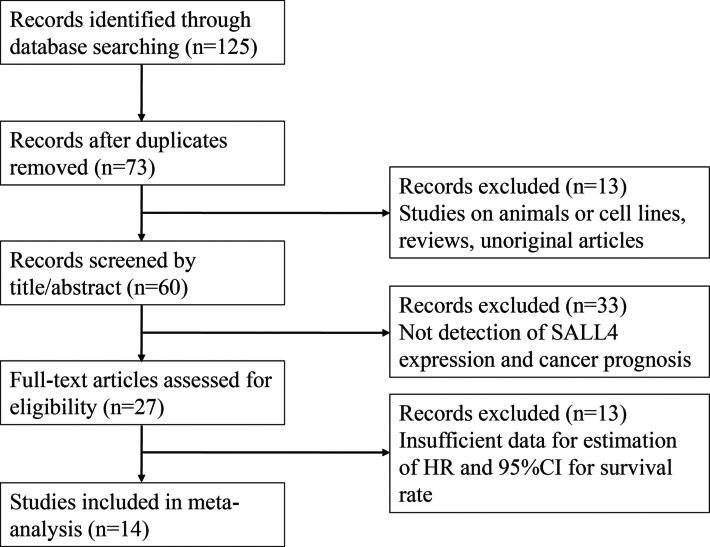
Flow chart of the selection of publications included in the meta-analysis.
Main characteristics of studies included in the meta-analysis
OS, overall survival; DFS, disease-free survival; RFS, recurrence-free survival; PFS, progression-free survival; NA, not available; SC, survival curve; HR, hazard ratio; NOS score, Newcastle-Ottawa Scale score.
Two investigators extracted the data independently according to the standard approach [21]. In case of discordant studies, the third investigator was invited to further review the original data until an agreement was reached. We collected the following data from each eligible publication: first author’s name, publication year, case nationality, sample size, mean age, cancer type, follow-up months, detecting method (immunohistochemistry (IHC) or tissue microarray (TMA) or enzyme-linked immuno sorbent assay (ELISA)), specimen, and cut-off value, respectively. Most importantly, the HR with the corresponding 95% CI for overall survival (OS), disease-free survival (DFS), recurrence-free survival (RFS) or progression-free survival (PFS) were extracted from every included study. Since the study only provided RR (risk ratio) with 95% CI, the RR is approximately considered as HR [22]. When the HRs with 95% CIs were described in text or tables, we acquired them directly [22, 23, 24, 25, 26, 27]. Otherwise, we obtained the necessary data from the only provided K-M survival curves using Engauge Digitizer 4.1 [28] by two independent researchers [29, 30, 31, 32, 33, 34, 35, 36]. For studies providing more than one HR, the HRs from multivariate models with the most completed adjustment for confounding factors were extracted for further analysis.
Study quality assessment
The quality assessment of the studies included in present meta-analysis was independently evaluated by two investigators according to the Newcastle-Ottawa quality assessment scale (NOS) [37]. Each study was assessed on the base of three broad perspectives: selection, comparability and exposure with a score ranging from 0 to 9. Studies with a score of 7 or greater were considered high quality studies. Discrepancies were adjudicated by discussion or through consultation with a third investigator.
Statistical analysis
HR with a 95% CI was assessed for the relationship between SALL4 expression and cancer prognosis (OS and DFS/RFS/PFS, respectively). An observed HR
HRs and 95% CIs for patient survival in association with SALL4 expression in enrolled studies
HRs and 95% CIs for patient survival in association with SALL4 expression in enrolled studies
The source of HR (hazard ratio) and 95% CI (confidence interval) is represented as derived from univariate analysis (U) or multivariate analysis (M). *HR and 95% CI calculated from K-M survival curves. OS, overall survival; DFS, disease-free survival; RFS, recurrence-free survival; PFS, progression-free survival; HR, HR (higher vs lower); NA, not available; IHC, immunohistochemistry; TMA, tissue microarray; ELISA, enzyme-linked immuno sorbent assay.
Characteristics of the studies
The flow chart describing the process of study selection was presented in Fig. 1. From a total of 125 potentially relevant articles through searching from the databases, we excluded 58 duplicates and 13 irrelevant articles. After a rough review of the titles and abstracts, 33 studies were excluded because of no relationship between SALL4 expression and cancer prognosis. In the remaining 27 records, with a systematical review of the full texts, another 13 articles were excluded for lacking of insufficient data. Eventually, 14 eligible articles (16 studies) containing 2,060 patients were included in this meta-analysis [22, 23, 24, 25, 26, 27, 29, 30, 31, 32, 33, 34, 35, 36]. Yong and Kilic’s studies included two different survival analyses due to two different case nationality and two different pathological types, separately [24, 36].
The main characteristics of all the eligible studies are summarized in Tables 1 and 2. Of the 16 studies, 14 studies were extracted for OS [22, 23, 24, 25, 26, 27, 29, 31, 32, 33, 34, 35, 36] while 8 studies were obtained for DFS/RFS/PFS [23, 24, 25, 27, 30, 35, 36]. Except Oikawa’s study [29] based on both Chinese and White, 13 studies (1683 patients: 81.7%) were conducted in Asian area, and only two studies (267 patients: 13.0%) were performed in European area. The malignant tumors assessed in these studies included ovarian malignant germ cell tumor [23], hepatocellular carcinoma (HCC) [24, 25, 26, 29, 30, 32], combined hepatocellular carcinoma and cholangiocarcinoma, [35] small cell lung cancer, [31] breast cancer, [33] endometrial cancer [34], glioma [22], esophagus cancer [36] and serous ovarian carcinoma [27]. IHC was used to detect the expression of SALL in 13 studies, except that Yong1 and Oikawa’s methods were performed by TMA [24, 29] and Han’s were detected via ELISA [32].
Evidence synthesis
SALL4 expression and OS
Fourteen studies with a total of 1,801 patients which provided survival results in the form of OS were enrolled in this meta-analysis. Since the heterogeneity was not statistically significant (
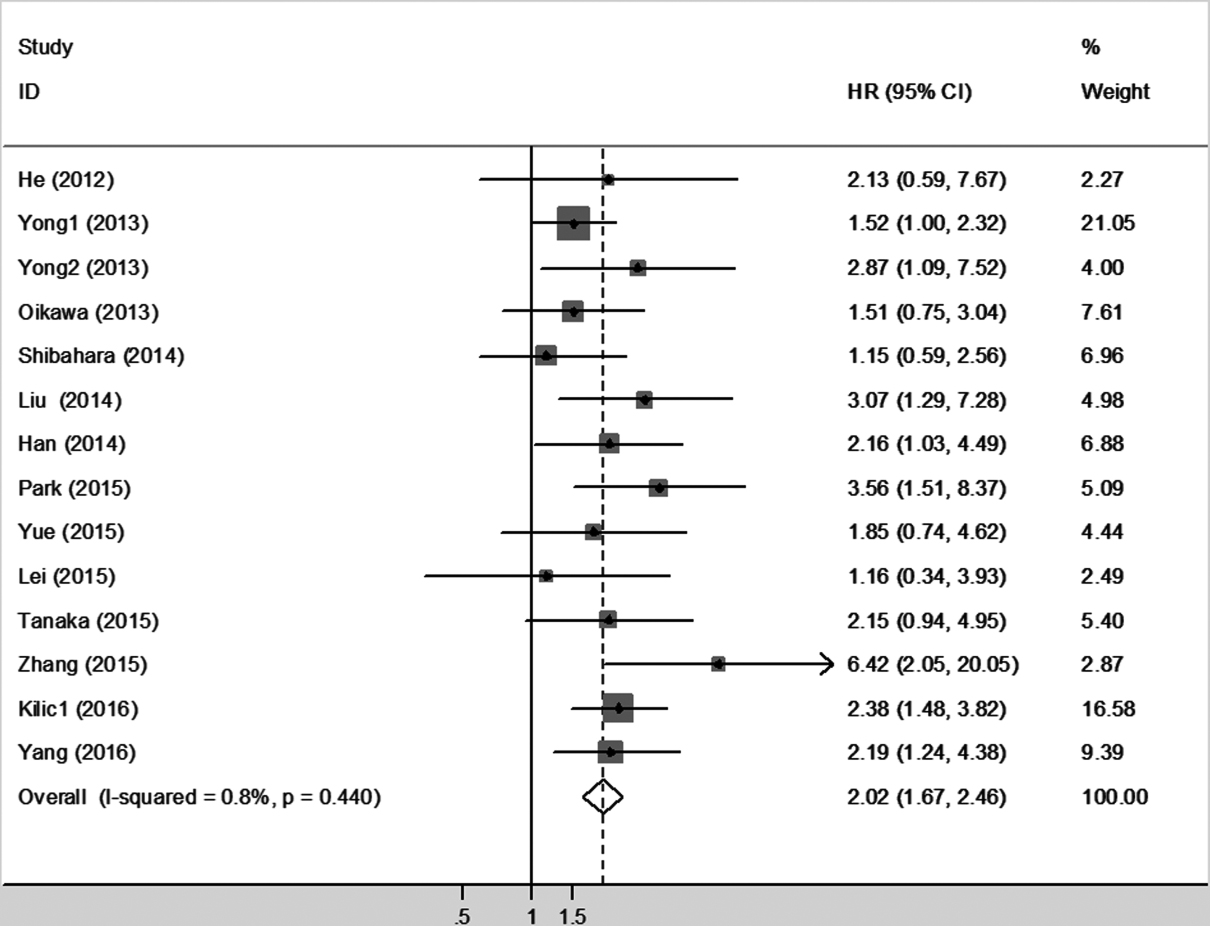
Forest plot of HR for the association between higher SALL4 expression and OS in patients with malignancies.
To explore the prognostic role of SALL4 in different carcinomas, we divided 14 studies providing results of OS into subgroups by human body system. The results suggested that higher SALL4 expression was an infaust prognostic marker in patient with digestive system neoplasms (HR: 1.91, 95% CI: 1.52–2.39,
Main results of meta-analysis
PFS, progression-free survival; SALL4
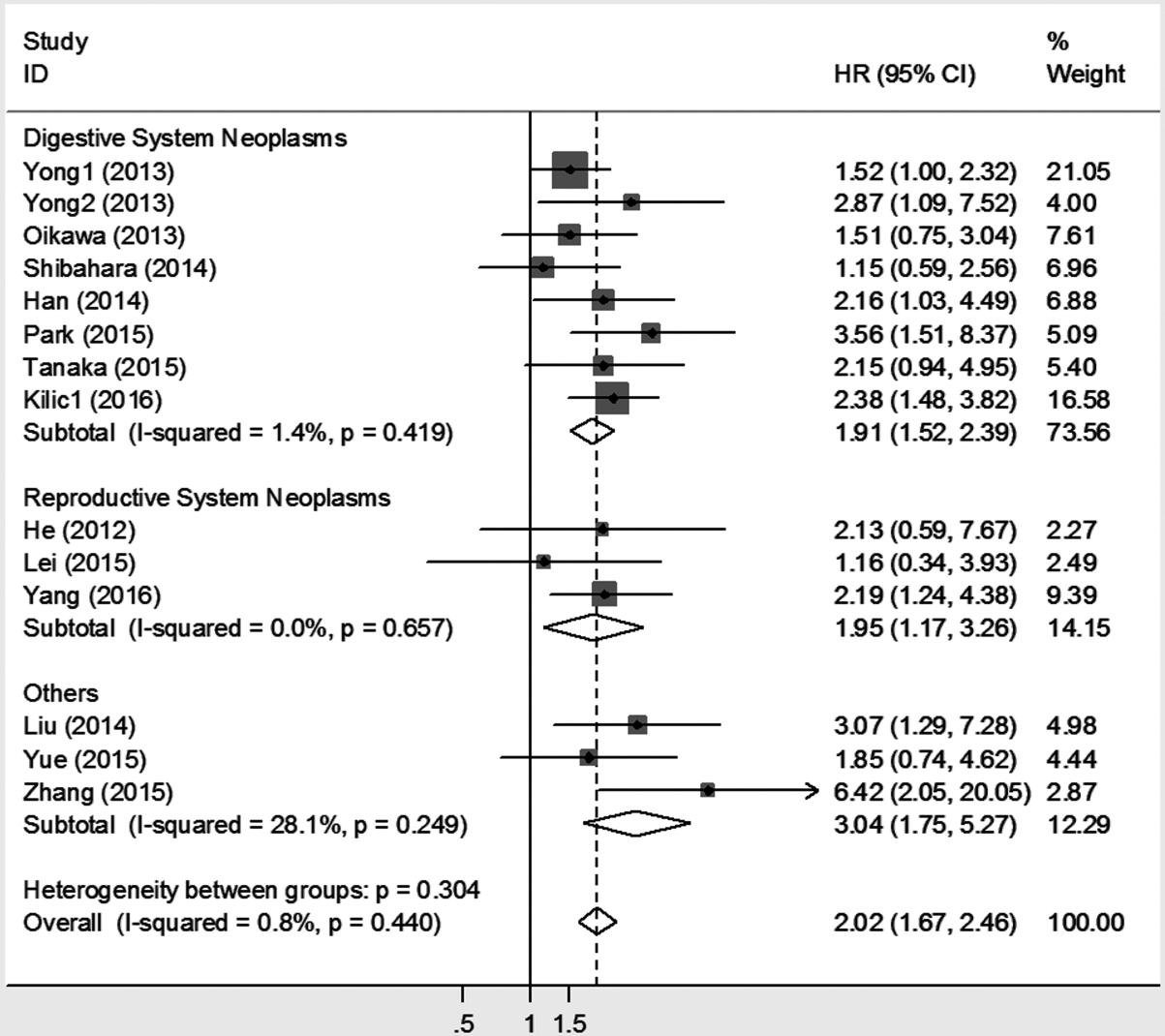
Meta-analysis (Forest plot) of the eligible studies assessing SALL4 expression and OS in cancer patients stratified by human body system.
We also conducted subgroup analysis by geography and detecting methods. The results indicated that there were significant correlations between SALL4 overexpression and poor OS in studies with Chinese (HR: 1.982, 95% CI: 1.526–2.576,
Three studies for DFS, four studies for RFS and only one study for PFS were performed separately in our meta-analysis due to different survival type owning different definition. With a random-effects model due to significant heterogeneity (
Sensitivity analysis
To assess the robustness of SALL4 expression and patient overall survival, a sensitivity analysis was adopted through excluding the enrolled studies one by one and analyzing the influence and homogeneity of the remaining studies. Both random effects model and fixed effects model were used and the two models did not cause any significant effect on the results. The sensitivity analysis results demonstrated that pooled HRs was not materially altered, indicating that no significant changes were existed in the HRs when ruling out any of the studies.
Publication bias
Begg’s funnel plot and Egger’s test were used to evaluate the publication bias of all included studies providing OS. The graphical funnel plots showed no evidence of asymmetry. Then, Egger’s test was applied to provide statistical evidence of funnel plot symmetry. The
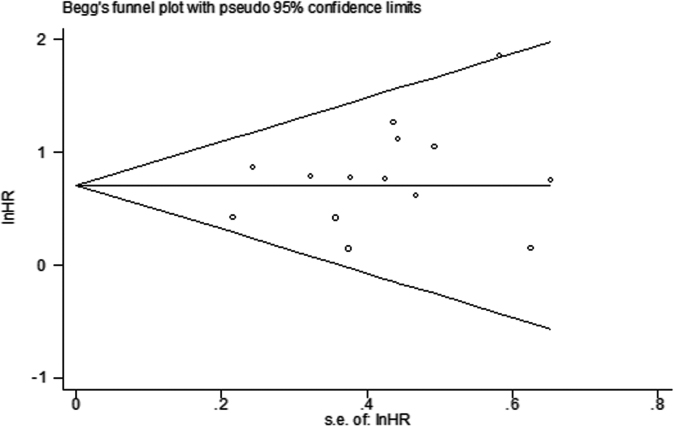
Begg’s funnel plot of SALL4 expression and OS in cancer patients.
Cancer remains the most public health burden, attributed to multiaspect reasons like stress, lifestyle, environment, psychological/personality, and many health -related factors [42]. It is of great importance to look for reliable and favorable prognostic biomarkers for cancer patients and further provide available information for clinical decision making. SALL4 is considered to be a valuable and specific cancer biomarker. SALL4 expression is reported in numerous malignancies and correlates with overall survival and disease progression [16, 17]. In an early clinical stage of breast cancer, SALL4 is expressed at a high level, indicating that SALL4 may be conducive to breast cancer screening [19]. In human HCC, SALL4 expression is restored in tumor tissues as well as serum levels. According to both tumor recurrence and overall survival rate, HCC patients with higher SALL4 expression are significantly associated with poor prognosis, suggesting that SALL4 is a novel and independent prognosis biomarker for HCC patients [15, 29, 32, 43]. Moreover, the level of SALL4 expression is positively correlated with worse patient survival and disease characteristics such as invasion and metastasis in endometrial cancer [43], colorectal carcinoma [16, 44], esophageal carcinoma [45], and gastric cancer [17, 44]. Thus, to address the prognostic value of SALL4 expression, we conducted this meta-analysis.
To our knowledge, this is the first time that a meta-analysis has been performed to focus on the association between elevated expression of SALL4 and the prognosis of patients with cancers. An aggregate amount of 16 eligible studies, including 2,060 cases, were selected and analyzed in the present meta-analysis. The results revealed that higher SALL4 expression was significantly associated with a poor prognosis (HR 2.02, 95% CI: 1.67–2.46,
In order to seek for more detailed conclusions, we stratified the enrolled studies for further analysis. Eight studies were enrolled in the digestive system neoplasms subgroup (pooled HR 1.91, 95% CI 1.52–2.39,
Concerning DFS in this meta-analysis, however, we failed to explore any significant association between SALL4 expression and DFS (HR 1.436, 95% CI 0.599–3.443,
Although our study is the first meta-analysis about the association between SALL4 expression and patient survival, some limitations should be taken into account. Firstly, our meta-analysis only contained a total of 16 studies, thus the results might be reached by accident because sample error of eligible studies could result in insufficient statistical power. Secondly, our meta-analysis did not encompass all human tumor types. Although SALL4 is a promising biomarker, the correlation still need further research and large-scale studies are required for further credible results. Thirdly, since most of the enrolled studies used IHC to detect SALL4 levels, a technique bias in different studies may exist for various dyeing operation, antibody concentration and cut-off value of different tissues. Fourthly, because the survival data were not presented directly in some included studies, we had to extract HRs and 95% CIs from survival curves and these data might be less reliable than those given directly. Finally, in spite of no significant difference detected in the results of sensitivity analysis and publication bias assay, publication bias cannot be utterly eliminated.
In conclusion, there was a positive correlation between SALL4 overexpression and poor prognosis of patients with cancers from the present meta-analysis. SALL4, therefore, may serve as a promising prognostic marker and an emerging potential therapeutic target for cancer patients. In order to intensify our conclusion, it is strongly suggested that more standardized randomized controlled trials (RCTs) and prospective studies with high quality should be carried out to dig into the molecular mechanisms of SALL4 and the relationship between higher SALL4 expression and prognosis for cancer patients.
Abbreviations
SALL4: sal-like protein 4,
CNKI: China National Knowledge Infrastructure,
HR: hazard ratio,
CI: confidence interval,
RFS: recurrence-free survival,
DFS: disease-free survival, OS: overall survival,
PFS: progression-free survival,
RR: risk ratio,
IHC: immunohistochemistry,
TMA: tissue microarray,
ELISA: enzyme-linked immuno sorbent assay,
NOS: Newcastle-Ottawa quality assessment scale,
HCC: hepatocellular carcinoma,
RCTs: randomized controlled trials.
Footnotes
Acknowledgments
This study was funded by the National Natural Science Foundation of China (81272470).
Conflict of interest
The authors declare that they have no conflict of interests.
