Abstract
Background:
The distribution of components in fermented dairy products forms the microstructure which influences final product texture and taste. Confocal Raman microscopy may provide new molecular information on product structure not possible with other advanced microscopy techniques.
Objective:
Dairy products including non-fat and full fat yoghurt, Camembert and Cheddar cheese samples were surveyed and the product microstructure observed using confocal Raman microscopy in order to determine the applicability of the technique to dairy product analysis.
Methods:
Confocal Raman microscopy provided spatially resolved chemical information on the components of fermented dairy products. In conjunction with component analysis and exploratory data analysis, spatially resolved chemical information on the components of fermented dairy products was obtained and compared.
Results:
Yoghurts with differing fat levels displayed different microstructures, consistent with other techniques. The influence of different molecular structures on the Camembert cheese centre and surface was revealed and Raman microscopy also gave new insights on the chemical structures within Cheddar cheese.
Conclusions:
The method provides a new technique for observing the contribution of different components to the product microstructure that may be used to monitor product quality and guide product development.
Introduction
Raman spectroscopy has been applied to study various food systems as a non-destructive method to assess chemical information. As Raman scattering is stronger for non-polar bonds, the technique has traditionally found more applications in lipids and fat-rich products and has been applied to foods such as potato chips as a rapid technique for determining fat content [29], to beef in order to predict sensory quality properties [42] and to determine interactions and migration between food components and packaging [28]. Raman spectroscopy has also been applied to lipid-rich dairy products. Primarily studies have focussed on detecting adulteration with non-dairy ingredients of products such as milk [2,34], dairy powders and infant formula [35,39] or dairy cream [31]. However, some studies have also applied the technique to detect structural changes on heating or freezing in anhydrous milk fat [21] and semi-hard cheese [13]. Chemometric techniques such as multivariate data analysis are often used with Raman spectroscopic data to enable discrimination of datasets.
Fermented dairy products such as cheese and yoghurt consist of complex ingredient interactions that influence the product structure and texture [10]. The microstructure of dairy products containing polysaccharides has been examined by confocal laser scanning microscopy (CLSM) [4], however structural information is limited to those components that can be fluorescently stained and sample preparation can be complicated. Often studies employ CLSM to observe the protein network, which can be easily imaged, and draw conclusions on ingredient interactions based on the protein structural information [8,40]. This method has some limitations, as the unstained ingredients are not directly observed and thus their contribution to the structure is inferred from the stained, usually protein, components.
Spatially resolved information is possible when Raman spectroscopy is combined with a microscope. Raman microscopy can provide label-free data on the molecular structure of components such as lipids, proteins and carbohydrates in a sample with minimal sample preparation. The microscopy technique has found applications to detect differences in ingredient distribution in processed cheese [38] and Cheddar cheese [18] as well as lipid composition in milk fat globules [14]. However, these studies were limited to visual observations of component intensity maps and spectral information as they did not employ chemometrics to interpret the data. When Raman spectroscopic data is combined with chemometric techniques, detection and quantification of components at low concentration in spreadable cheese was possible [36], highlighting the potential of the technique.
The global production of milk and subsequent dairy products is gradually increasing [12]. In particular, the European Union contributes approximately one third of the 2.57 million tonnes of cheese that were exported globally in 2018 [12]. Dairy products are a diverse range of foods that have varying composition, production and final product uses that make their characterisation important for manufacturing and consumer acceptability. Raman microscopy has the potential to provide new direct molecular information on components in fermented dairy products that is not possible through other microscopy techniques without complex sample preparation. The technique may be applied to understand chemical structure and interactions to assist the development of novel structures and textures in dairy products.
In the present study, the distribution of ingredients in fermented dairy products was assessed using Raman microscopy. Greek style full fat yoghurt and non-fat yoghurt were compared as well as Camembert and Cheddar cheese. Variation in components such as lipids, proteins, water and carbohydrates was monitored to obtain spatially resolved structural information. Chemometric techniques were applied to establish differences in molecular information in dairy products and show the potential of the confocal Raman microscopy technique for structural characterisation.
Materials and methods
Dairy samples and sample preparation
Cheese and yoghurt samples were purchased at a local supermarket in Ireland. Two different types of yoghurt were obtained; Greek style full fat and non-fat. A Camembert cheese and a mature Cheddar cheese were also purchased. The composition of the dairy products are summarised in Table 1. Samples were stored at 4°C until immediately prior to analysis.
Composition of dairy samples used for confocal Raman microscopic analysis
Composition of dairy samples used for confocal Raman microscopic analysis
Confocal Raman microscopy measurements were performed using a WITec alpha300R (Ulm, Germany) system equipped with a thermoelectrically cooled CCD detector. A 532 nm laser was used for excitation at 30 mW power. WITec Control 5.0 software was used to operate the system and collect data.
Samples were prepared with a surgical blade to approximately 5 × 5 × 2 mm and transferred to a glass slide immediately prior to imaging. A 50× objective (NA = 0.55) was used to acquire spatially-resolved spectral data of the samples with a calculated spatial resolution of 0.6
The data collection and imaging parameters were slightly different for the various samples. The sampled area of yoghurts and Cheddar cheese was 25 × 25
Data processing and multivariate data analysis
After data collection, cosmic ray removal (filter size 3, dynamic factor 8) and background subtraction (shape size 250, noise factor 1) was conducted using Project Plus software (WITec).
The True Component Analysis package (WITec) was employed to characterise the key regions of interest in each sample and generate component intensity maps as well averaged spectra representative of each component. For yoghurt and Cheddar cheese samples three components were applied to represent the sampled area. For Camembert cheese samples two components were deemed adequate to represent the sampled area.
The averaged component spectra for each sample were extracted and imported into the Unscrambler® 10.1 software (CAMO Software AS., Oslo, Norway). Principal Component Analysis (PCA) using the spectral range 3600–400 cm−1 was used to explore differences between paired samples; the surface and centre of Camembert cheese and the Greek style full fat and non-fat yoghurts as well as to examine differences within Cheddar cheese samples. Further data processing was not deemed necessary in the present case due to the clear observation of spectral features possible in the minimally processed data.
Cryo scanning electron microscopy of yoghurt samples
The Greek style full fat and non-fat yoghurts were analysed by cryo scanning electron microscopy (cryo-SEM) following a previously published method [37]. Yoghurt microstructure was observed using a scanning electron microscope (SEM; Zeiss Supra 40VP field emission, Carl Zeiss AG, Darmstadt, Germany) with a cryo sample preparation system (Gatan Alto 2500, Gatan UK). Fresh samples were rapidly immersed in liquid nitrogen slush (−210°C) and transferred under vacuum to the cryo-preparation chamber maintained at −140°C. The samples were fractured with a scalpel blade and followed by etching at −95°C. Sputter coating was conducted at −140°C and the sample transferred to the SEM cold stage maintained at −140°C. Cryo-SEM micrographs were collected over a range of magnifications with one representative image shown per sample. The yoghurt microstructure observed by cryo-SEM was used as a comparison to that observed using confocal Raman microscopy.
Results and discussion
Confocal Raman microscopy of yoghurt samples
Confocal Raman microscopy images of two yoghurt samples with differing fat contents were collected and compared. Both the Greek style full fat and the non-fat yoghurts contained several characteristic peaks representing the pyranose ring of carbohydrates at 476 cm−1 [38] and associated with proteins at ∼1456 cm−1 and carotenoids at 1520–1530 cm−1 [30] (Fig. 1). The peak at 2432 cm−1 was tentatively assigned to atmospheric CO2 [15].
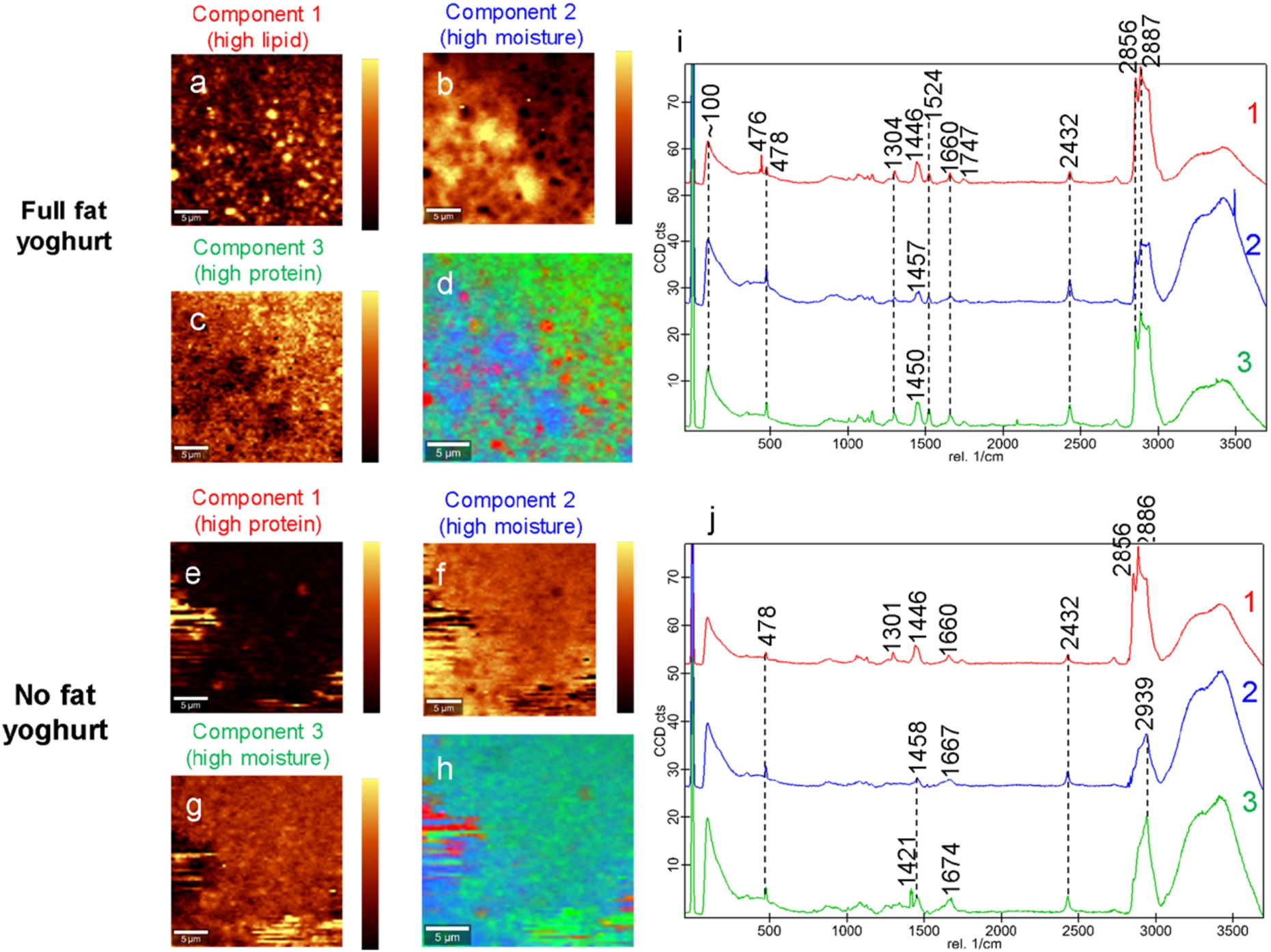
Confocal Raman images showing the distribution of three components for Greek style full fat (a–c) and non-fat (e–g) yoghurt. Light areas indicate high component intensity and dark areas indicate low component intensity. The combined overlay images are also shown for full fat (d) and non-fat (h) yoghurt. The average spectra for each of the identified components are shown for full fat (i) and non-fat (j) yoghurt samples. The scale bars are 5
In addition, the Greek style full fat yoghurt samples had characteristic peaks associated with lipids that were not present in the non-fat yoghurt sample, namely at 1660 cm−1 assigned to lipids as well as amide I proteins [30], at ∼1750 cm−1 and 2856 cm−1 associated with lipids and at ∼2887 cm−1 associated with proteins and lipids [30].
Using the True Component software, the two different yoghurt samples were both described by three components each (Fig. 1). In the full fat yoghurt, the first component tended to describe lipids in the sample, as supported by the lipid peaks at ∼1750 cm−1 and 2856 cm−1 with peaks at 1660 cm−1 and ∼2887 cm−1 covering both lipids and protein absorbance (Fig. 1a). The second component tended to describe moisture or water, as evidenced by the broad OH band at 3600-3300 cm−1 (Fig. 1b). The third component described mostly protein in the sample, with some peaks also associated with lipids here (Fig. 1c). All components in the full fat yoghurt also had peaks associated with carbohydrates (478 cm−1) and atmospheric CO2 (2432 cm−1).
For the non-fat yoghurt, the first component tended to describe proteins and residual lipids in the sample (Fig. 1e). The second and third components both contained a large amount of moisture as well as peaks associated with proteins and lipids such as at 1421 cm−1, 1458 cm−1 and ∼1670 cm−1 (Fig. 1f–g). All components in the non-fat yoghurt had peaks associated with carbohydrates (478 cm−1) and atmospheric CO2 (2432 cm−1). As noted in the non-fat yoghurt composition (Table 1), residual lipids of less than 0.5% were likely present in the sample and may account for the presence of lipid related peaks in the Raman spectra.
In order to confirm the microstructure of yoghurt samples observed by confocal Raman microscopy, cryo-scanning electron microscopy (cryo-SEM) was also employed. The microstructure of Greek style full fat yoghurt contained a dense protein network that appeared well interconnected (Fig. 2a). The compact protein network in the Greek style full fat yoghurt contained small serum pores and was interspersed with fat globules. In contrast, the non-fat yoghurt structure appeared relatively open and porous (Fig. 2b). The non-fat yoghurt contained comparatively larger protein strands with correspondingly larger serum pores and likely fewer connections or cross-linking of protein strands. These microstructures are consistent with the literature [32,33].
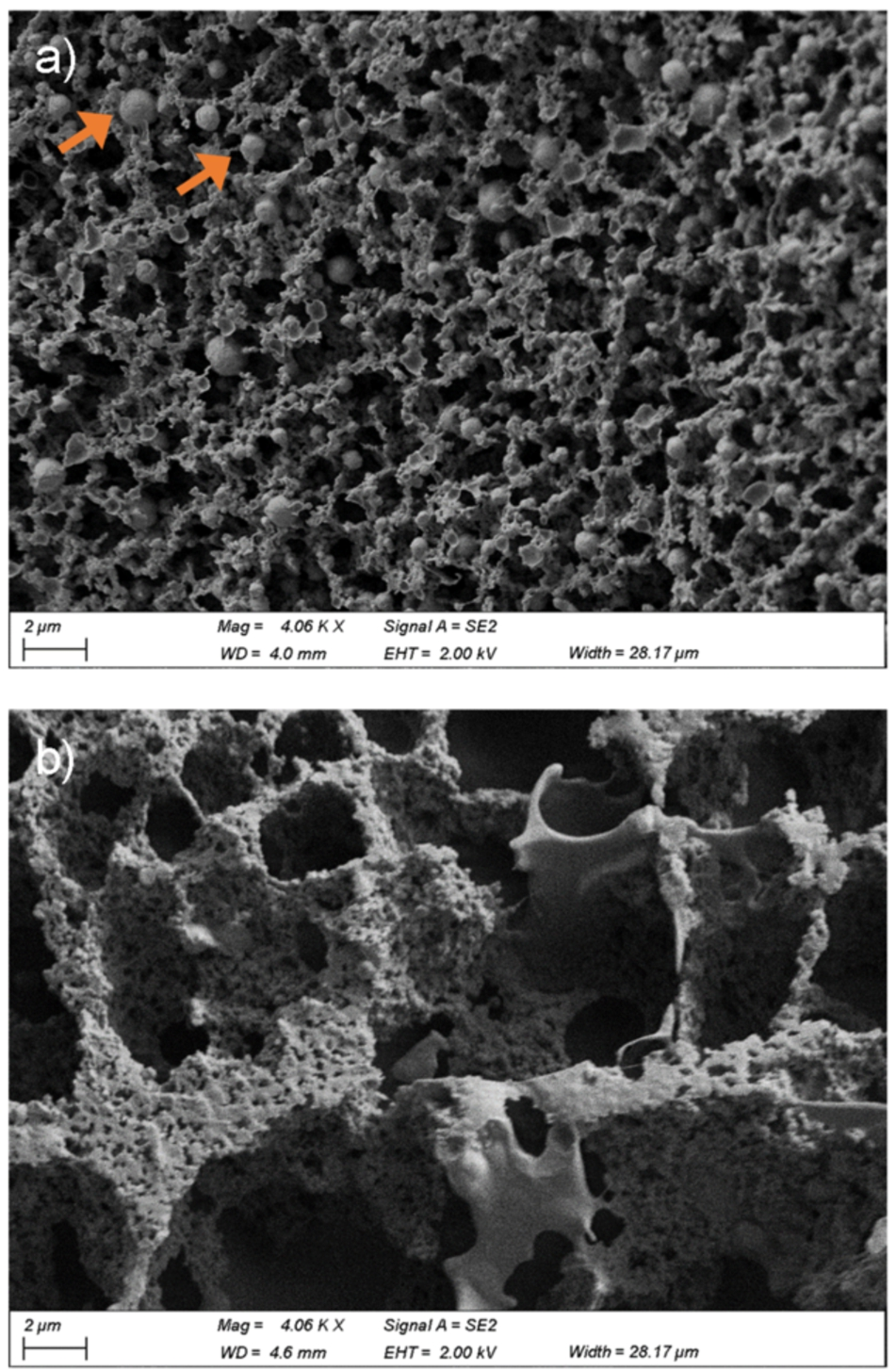

Cryo-SEM micrographs of Greek style full fat (a) and non-fat (b) yoghurts. Orange arrows indicate individual fat globules. Images were taken at approximately 4000× magnification with scale bars 2
The microstructure observed by cryo-SEM is consistent with that observed by Raman microscopy for the Greek style yoghurt, including the presence of individual fat globules. In the case of the non-fat yoghurt, the structural similarity was not as pronounced and it appears that the optical tweezers effect has occurred whilst imaging the sample [25]. The optical tweezers effect occurs where a laser beam provides, in this case, an attractive force to particles that can cause them to be physically moved at a microscopic level [41].
PCA was applied to the spectral region 3600–400 cm−1 to explore differences between samples based on their averaged component spectra. The Greek style full fat yoghurt was clearly distinct from the non-fat yoghurt (Fig. 3a). The two types of yoghurt were separated in PC1 due to lipid related peaks, which described 73% of variation between points and accounted mostly for the variation within the spectra for each yoghurt type (Fig. 3b). The lipid peaks identified in PC1 were 2856 cm−1 and ∼1446 cm−1 as well as the protein dominant peak at ∼2886 cm−1 with lesser contributions from the peak associated with carotenoids at 1530 cm−1 [30,38]. The variation within each yoghurt type was also accounted for in PC2, which described 25% of the variation between data points and the spread between each sample type. The differences in PC2 were mainly attributed to the large protein dominant peak at ∼2886 cm−1.
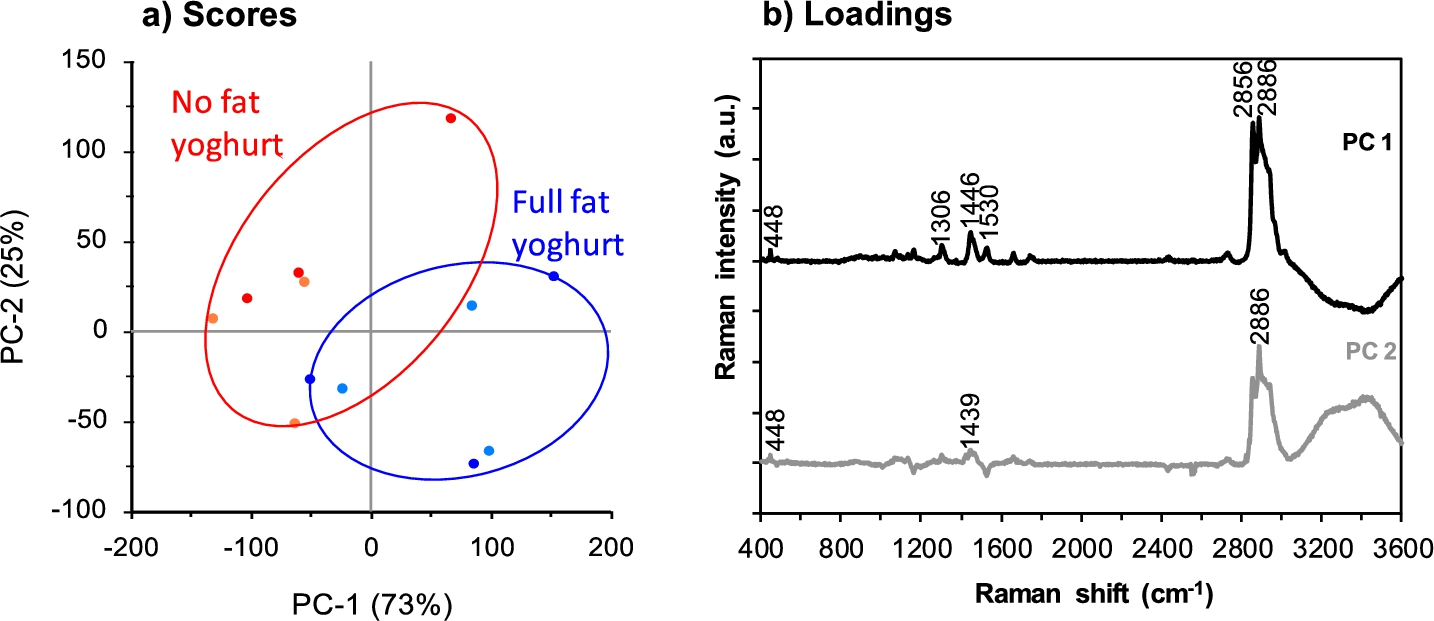
PCA scores (a) and loadings plots (b) for spectra in the collected region 3600–400 cm−1 for Greek style full fat (two replicate samples shown in dark blue and light blue) and non-fat (two replicate samples shown in red and orange) yoghurt samples. Three spectra were used to represent each sample based on the True Component Analysis. The colour version is available online.
The combination of the spectral data and exploratory data analysis like PCA provides a way to explore similar samples that does not rely on visual observation of micrographs. Here, the Greek style full fat yoghurt observed by confocal Raman microscopy had a microstructure similar to that observed by other microscopy techniques. Despite some difficulties imaging non-fat yoghurt, likely due to the optical tweezers effect, at a similar length scale the features, such as some high protein areas, are largely similar to that observed by other microscopy techniques. The reduction in protein network detail and a coarser protein network with lower fat content in yoghurts is in agreement with previous studies [17,19]. The protein network structure is likely important in determining the final yoghurt viscosity and consistency, with a less coarse protein network leading to a smoother product.
Confocal Raman microscopy allowed the chemical structure of Camembert cheese to be spatially mapped. Both the centre and the surface of the Camembert cheese had some common features, with key differences in some peaks as would be expected due to the differing compositions in these sections of the cheese [1,27]. In particular, the presence of the lactose and monosaccharides peak at 1162 cm−1 in the centre of Camembert cheese only is consistent with prior observations that lactose and related monosaccharides are consumed faster on the surface due to microbial activity and remain present in the centre of the cheese with up to four weeks of ripening [11,22]. Key peaks present in both sections of Camembert were associated with lipids and proteins at 1662 cm−1, carotenoids at 1525 cm−1 and proteins at 1457 cm−1 [24,30,38]. The peak at 479 cm−1 associated with the pyranose ring of carbohydrates [38] was also present in both Camembert sections. The peak at 2432 cm−1 is likely associated with atmospheric CO2 [15].
The structure of the surface of the Camembert cheese (Fig. 4a–c) displayed the characteristic fungal strands or hyphae that would be expected for a white mould cheese that contains the filamentous
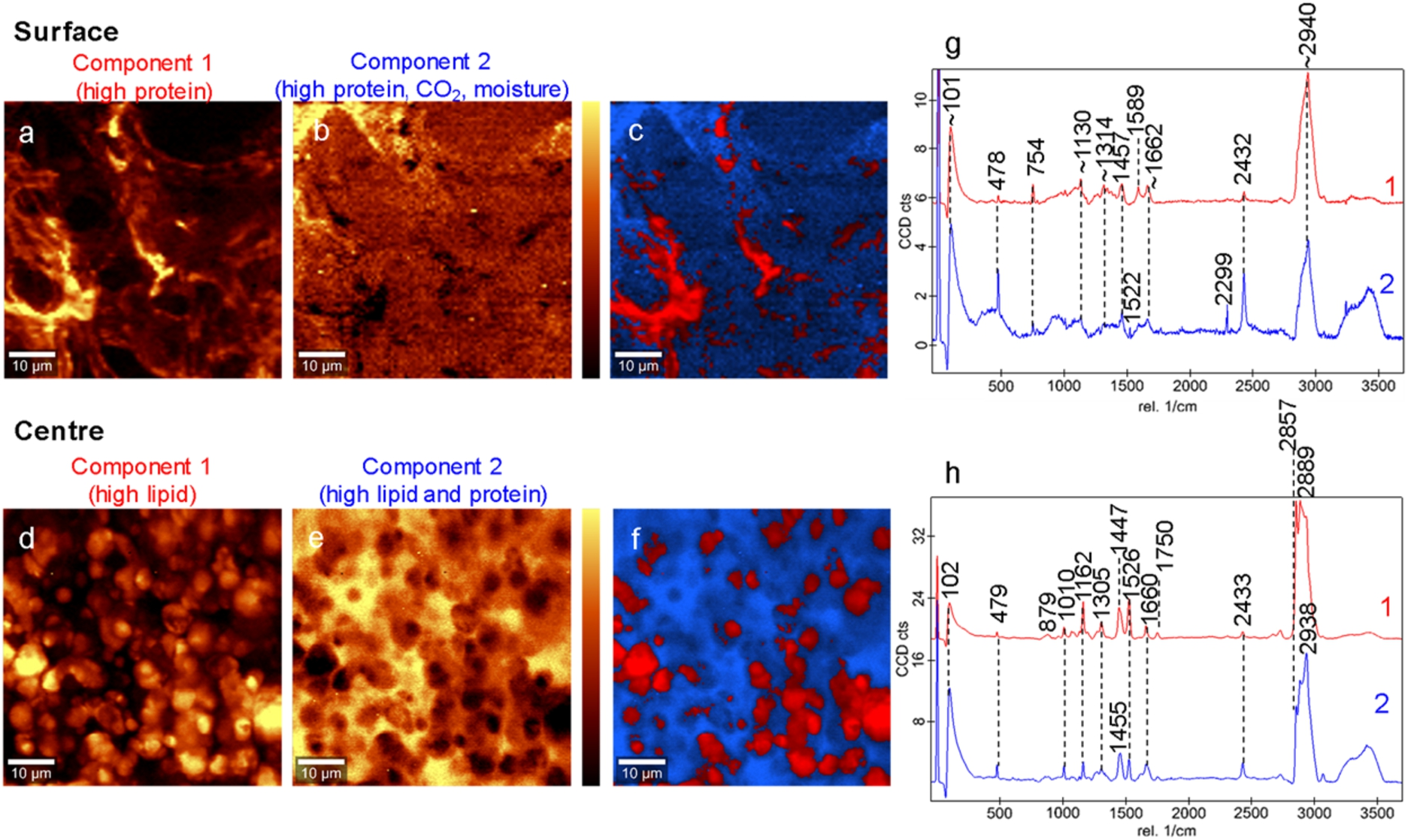
Confocal Raman images showing the distribution of two components for the Camembert cheese surface (a–b) and centre (d–e). Light areas indicate high component intensity and dark areas indicate low component intensity. The combined overlay images are also shown for the surface (c) and centre (f) of Camembert cheese. The average spectra for each of the identified components are also shown for the Camembert surface (g) and centre (h). The scale bars are 10
The two components that described the centre of the Camembert cheese also contained similar peaks, with some variation in concentrations. The first component contained a number of peaks associated with lipids (Fig. 4d). The second component contained similar peaks to the first component, but with a proportion of peaks also associated with proteins, such as at 1455 cm−1 and ∼2940 cm−1 (Fig. 4e). The centre of the Camembert cheese also had several peaks identified only in the centre portion and not on the surface of the cheese. These include the peak at 2857 cm−1 associated with lipids and proteins, the lipid only peak at 1750 cm−1, two peaks associated with proteins attributed to the amino acid tryptophan at 1010 cm−1 and 879 cm−1 [30] and the peak at 1161 cm−1 attributed to lactose and monosaccharides [27].
The centre of Camembert cheese also displays a characteristic microstructure that is porous and includes large fat globules within a heterogeneous protein network [5]. It is known that ammonia and sulphur compounds are released with the growth of these fungal species, however Raman studies of ammonia and water mixtures have found limited bands without overlap with other compounds, which is also an issue here [3]. The quantification of microbial growth in cheese remains limited to bulk values, as there is no technique as yet able to quantify each species individually [3,23].
It is well known that microorganisms such as the
PCA highlighted differences between the averaged component spectra of Camembert centre and surface samples within the range 3600–400 cm−1 (Fig. 5). The different cheese sections were clearly separated in PC1, which accounted for 97% of the variation between points (Fig. 5a). PC1 also described the variation between the average component spectra of the centre portions of Camembert cheese. Separation in PC1 was associated with absorbance peaks relating to lipids and proteins at 2857 cm−1, 1657 cm−1, 1525 cm−1 and 1446 cm−1, as well as lipids only at 1306 cm−1 and lactose or other monosaccharides at 1163 cm−1 (Fig. 5b).
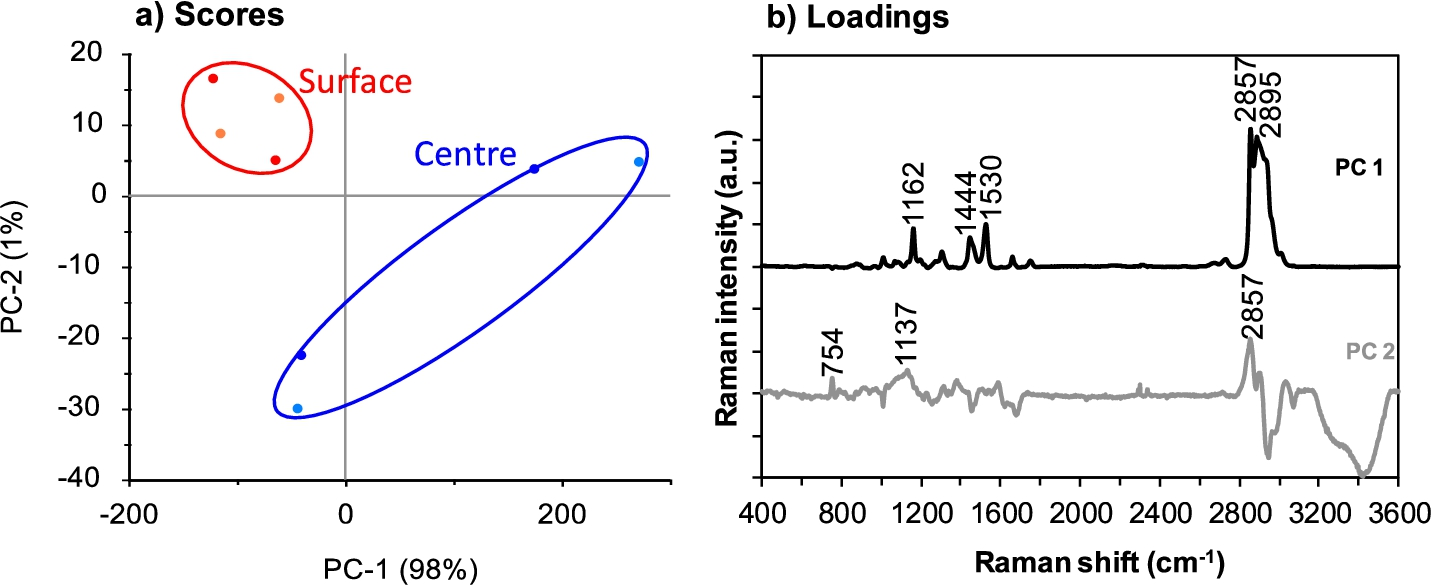
PCA scores (a) and loadings plots (b) for spectra in the collected region 3600–400 cm−1 for Camembert cheese surface (two replicate samples shown in red and orange) and centre (two replicate samples shown in dark blue and light blue) sections. Two spectra were used to represent each sample based on the True Component Analysis. The colour version is available online.
The surface and centre sections of Camembert cheese were also separated in PC2, which described 2% of the variation. The spread between data points of both sample sections was also accounted for in PC2. Variation in PC2 was mostly due to the peak at 2857 cm−1 associated with lipids and proteins.
These results highlight the potential of the technique to differentiate between different sections of the sample that may be used to develop a tool to monitor and predict ripening stages. Prior studies have also demonstrated other spectroscopic methods to validate mould-ripened cheese ripening [27]. The advantage of the techniques described here are that the spatial distribution of components is also possible at a resolution where individual features may be visualised. The structural alterations that were observed between cheese sections may also be used to develop structure-function relationships.
Confocal Raman microscopy was successfully applied to observe the chemical microstructure of Cheddar cheese, a harder and lower moisture cheese than Camembert. Absorbance peaks were identified that are associated with lipids and proteins at 2856 cm−1, 1659 cm−1, 1445 cm−1, lipids only at 1748 cm−1, carotenoids at 1525 cm−1, lactose and monosaccharides at 1161 cm−1 and the pyranose ring of carbohydrates at 478 cm−1 [27,30,38].
Employing True Component Analysis software (WITec) indicated that the cheese sample could be explained by three components (Fig. 6). The first component tended to describe mostly lipids with a little influence from the proteins in the sample (Fig. 6a). The second component explained moisture and protein in the sample whilst the third component tended to cover lipid-related absorbance in the sample (Fig. 6b–c). All components contained peaks associated with lipids, proteins and carbohydrates due to the both the small size of structural features and the sampling depth of the method.
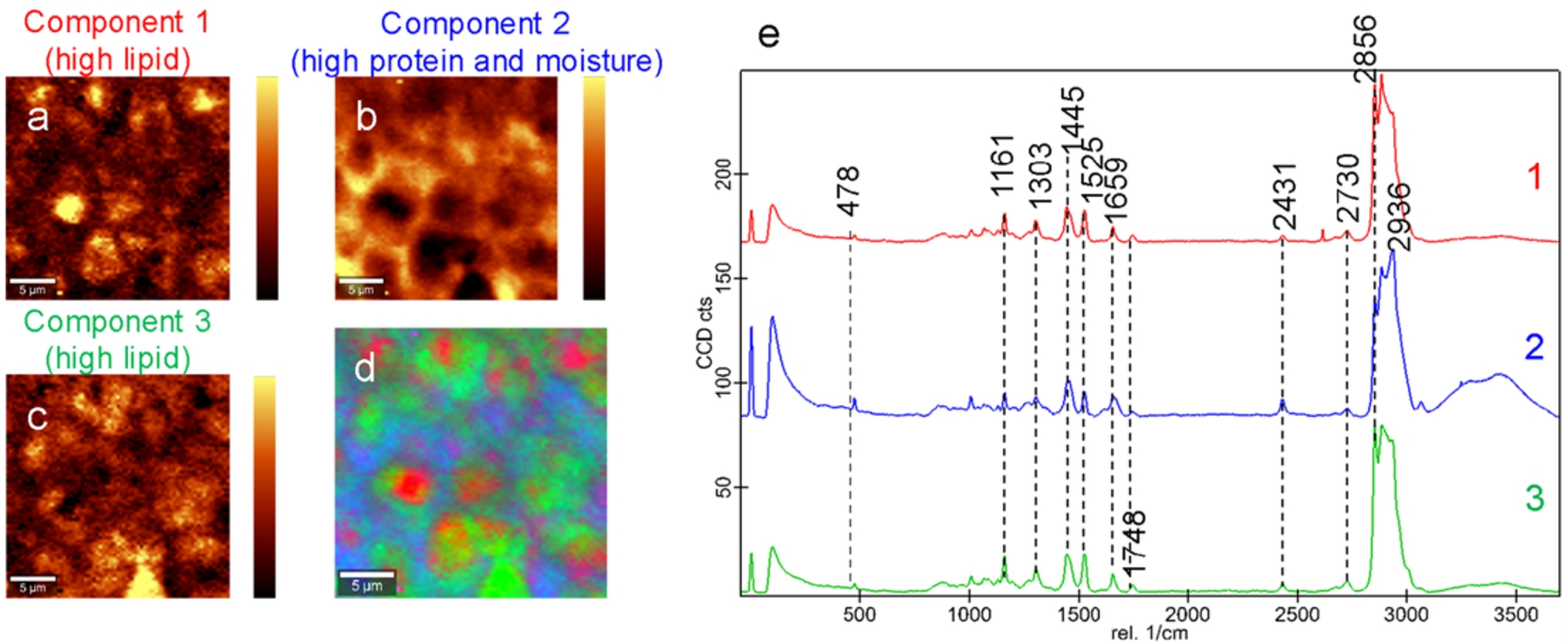
Confocal Raman images showing the distribution of three components for mature Cheddar cheese (a–c). Light areas indicate high component intensity and dark areas indicate low component intensity. The combined overlay image is also shown (d). The average spectra for each of the identified components is shown in (e). The scale bars are 5
In comparison to the yoghurt and camembert cheese discussed in Sections 3.1 and 3.2, the Cheddar cheese had a distinctly smaller water related band, consistent with the lower water content of this cheese (Fig. 6e).
Molecular information of proteins and lipids that form the Cheddar cheese microstructure is not possible through techniques such as SEM or CLSM. Prior studies have highlighted the potential of spectroscopic techniques as a quality tool to assess cheese ripening, texture and sensory properties [9,13,20]. The Raman technique employed here offers the advantage of minimal sample preparation and imaging time. Detailed information on the product microstructure is also possible with confocal Raman microscopy at a resolution where individual features such as fat globules may be observed.
Confocal Raman microscopy was applied to identify the protein, lipid and carbohydrate structures and their spatial distribution in a number of fermented dairy products. As expected, lipid related absorbance dominated spectral features across all samples, however components of high protein or high moisture were also able to be identified. Peaks associated with carbohydrates such as lactose were also clearly identified in yoghurt samples, suggesting future potential of the technique to map carbohydrate based ingredients. The spatial resolution allowed the identification of small features in yoghurt and Cheddar cheese samples, such as individual fat globules, and clearly revealed the structural heterogeneity that occurs between the centre and surface of mould-ripened cheese such as Camembert. Examining the surface of Camembert cheese with its high mould concentration has revealed some spectral features that may be associated with microbial growth. Future refinement of the technique may enable identification of growth markers or metabolites for microbes. The diverse products surveyed here highlight the applicability of confocal Raman microscopy to fermented dairy structure analysis. The current method may be applied as a tool to assist with correlating product structure and function in dairy and other food products.
Footnotes
Acknowledgements
This project has received funding from the European Union’s Horizon 2020 research and innovation programme under the Marie Skłodowska-Curie grant agreement No [713654].
Conflict of interest
The authors have no conflict of interest to report.
